665:) revealed that a majority of m6A locates in the last exon of mRNAs in multiple tissues/cultured cells of mouse and human, and the m6A enrichment around stop codons is a coincidence that many stop codons locate round the start of last exons where m6A is truly enriched. The major presence of m6A in last exon (>=70%) allows the potential for 3'UTR regulation, including alternative polyadenylation. The study combining m6A-CLIP with rigorous cell fractionation biochemistry reveals that m6A mRNA modifications are deposited in nascent pre-mRNA and are not required for splicing but do specify cytoplasmic turnover.
261:
868:(PCGs) have all been shown to undergo mA modifications during growth and differentiation. Depending on the stage of development, modifications to HSCs can either promote or inhibit stem cell differentiation by affecting the epithelial-to-hemopoietic transition via METTL3 inhibition or depletion. mA modifications to NSCs can causes changes in brain size, neuron formation, long-term memory, and learning ability. These changes are often caused by inhibition of either
773:. The impacts of mA on cancer cell proliferation might be much more profound with more data emerging. The depletion of METTL3 is known to cause apoptosis of cancer cells and reduce invasiveness of cancer cells, while the activation of ALKBH5 by hypoxia was shown to cause cancer stem cell enrichment. mA has also been indicated in the regulation of energy homeostasis and obesity, as FTO is a key regulatory gene for energy metabolism and obesity. SNPs of
41:
547:. Exon junction complexes suppress mA methylation near exon-exon junctions by packaging nearby RNA and protecting it from methylation by the mA methyltransferase complex. mA regions in long internal and terminal exons, away from exon-exon junctions and exon junction complexes, escape suppression and can be methylated by the methyltransferase complex.
794:
govern the efficiency of infection, replication, translation and transport of RNA viruses such as human immunodeficiency virus (HIV), hepatitis B virus (HBV), hepatitis C virus (HCV), and Zika virus (ZIKV). These results suggest mA and its cognate factors play crucial roles in regulating virus life cycles and host-viral interactions.
723:, composed of a transcription feedback loop tightly regulated to oscillate with a period of about 24 hours, is therefore extremely sensitive to perturbations in mA-dependent RNA processing, likely due to the presence of mA sites within clock gene transcripts. The effects of global methylation inhibition on the
672:
RNA reveals that mA levels are low during embryonic development and increase dramatically by adulthood. In mESCs and during mouse development, FTO has been shown to mediated LINE1 RNA mA demethylation and consequently affect local chromatin state and nearby gene transcription. Additionally, silencing
781:
differentiation has been suggested. The connection between mA and neuronal disorders has also been studied. For instance, neurodegenerative diseases may be affected by mA as the cognate dopamine signalling was shown to be dependent on FTO and correct mA methylation on key signalling transcripts. The
740:
Considering the versatile functions of mA in various physiological processes, it is thus not surprising to find links between mA and numerous human diseases; many originated from mutations or single nucleotide polymorphisms (SNPs) of cognate factors of mA. The linkages between mA and numerous cancer
539:
to selectively recognize m6A-containing RNAs and promote their translation and stability. These mA readers, together with mA methyltransferases (writers) and demethylases (erasers), establish a complex mechanism of mA regulation in which writers and erasers determine the distributions of mA on RNA,
793:
Additionally, mA has been reported to impact viral infections. Many RNA viruses including SV40, adenovirus, herpes virus, Rous sarcoma virus, and influenza virus have been known to contain internal mA methylation on virus genomic RNA. Several more recent studies have revealed that mA regulators
657:
binding sites; roughly 2/3 of the mRNAs which contain an mA site within their 3' UTR also have at least one microRNA binding site. By integrating all mA sequencing data, a novel database called RMBase has identified and provided ~200,000 sites in the human and mouse genomes corresponding to
786:, a potential reader of mA, have been known to cause neurodegeneration. The IGF2BP1–3, a novel class of mA reader, has oncogenic functions. IGF2BP1–3 knockdown or knockout decreased MYC protein expression, cell proliferation and colony formation in human cancer cell lines. The
727:
period in mouse cells can be prevented by ectopic expression of an enzyme from the bacterial methyl metabolism. Mouse cells expressing this bacterial protein were resistant to pharmacological inhibition of methyl metabolism, showing no decrease in mRNA mA methylation or protein
4892:
Blow, Matthew J.; Clark, Tyson A.; Daum, Chris G.; Deutschbauer, Adam M.; Fomenkov, Alexey; Fries, Roxanne; Froula, Jeff; Kang, Dongwan D.; Malmstrom, Rex R.; Morgan, Richard D.; Posfai, Janos; Singh, Kanwar; Visel, Axel; Wetmore, Kelly; Zhao, Zhiying (2016-02-12).
572:, IME4, is induced in diploid cells in response to nitrogen and fermentable carbon source starvation and is required for mRNA methylation and the initiation of correct meiosis and sporulation. mRNAs of IME1 and IME2, key early regulators of
3981:
Kalnina I, Zaharenko L, Vaivade I, Rovite V, Nikitina-Zake L, Peculis R, et al. (September 2013). "Polymorphisms in FTO and near TMEM18 associate with type 2 diabetes and predispose to younger age at diagnosis of diabetes".
852:. One such role is introducing a methyltransferase which recognizes the same target site that restriction enzymes (Type 1 restriction enzymes) attack and modifying it in order to stop such enzymes from attacking bacteria DNA.
3134:
Akilzhanova A, Nurkina Z, Momynaliev K, Ramanculov E, Zhumadilov Z, Rakhypbekov T, et al. (September 2013). "Genetic profile and determinants of homocysteine levels in
Kazakhstan patients with breast cancer".
5083:
1395:
Kennedy TD, Lane BG (June 1979). "Wheat embryo ribonucleates. XIII. Methyl-substituted nucleoside constituents and 5'-terminal dinucleotide sequences in bulk poly(AR)-rich RNA from imbibing wheat embryos".
797:
Aside from affecting viruses themselves, mA modifications can also disrupt the innate immune response. For example, in HBV, mA modifications were shown to disrupt the recognition of viruses by RIG-1, a
4780:
Balzarolo, Melania; Engels, Sander; de Jong, Anja J.; Franke, Katka; van den Berg, Timo K.; Gulen, Muhammet F.; Ablasser, Andrea; Janssen, Edith M.; van
Steensel, Bas; Wolkers, Monika C. (March 2021).
1333:
Levis R, Penman S (April 1978). "5'-terminal structures of poly(A)+ cytoplasmic messenger RNA and of poly(A)+ and poly(A)- heterogeneous nuclear RNA of cells of the dipteran
Drosophila melanogaster".
703:
The importance of mA methylation for physiological processes was recently demonstrated. Inhibition of mA methylation via pharmacological inhibition of cellular methylations or more specifically by
4231:
Hess ME, Hess S, Meyer KD, Verhagen LA, Koch L, Brönneke HS, et al. (August 2013). "The fat mass and obesity associated gene (Fto) regulates activity of the dopaminergic midbrain circuitry".
503:
The biological functions of mA are mediated through a group of RNA binding proteins that specifically recognize the methylated adenosine on RNA. These binding proteins are named mA readers. The
5450:
1655:
Dominissini D, Moshitch-Moshkovitz S, Schwartz S, Salmon-Divon M, Ungar L, Osenberg S, et al. (April 2012). "Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq".
5076:
142:
802:
in the immune system. Modifications can also disrupt downstream signaling pathways via mechanisms including ubiquitination and changes in the levels of protein expression.
5069:
880:
and negatively affect both gamete formation and fertility. Similar to NSCs, inhibition of the METTL and YTHDF families of proteins is often a catalyst for these changes.
3946:
Wang L, Yu Q, Xiong Y, Liu L, Zhang X, Zhang Z, et al. (2013). "Variant rs1421085 in the FTO gene contribute childhood obesity in
Chinese children aged 3-6 years".
1485:"Induction of sporulation in Saccharomyces cerevisiae leads to the formation of N6-methyladenosine in mRNA: a potential mechanism for the activity of the IME4 gene"
829:
In replication, M6A modifications mark DNA regions where the initiation stage takes place as well as regulates precise timing via the Dam methyltransferase in
492:. Following a 2010 speculation of mA in mRNA being dynamic and reversible, the discovery of the first mA demethylase, fat mass and obesity-associated protein (
2391:
Xu C, Wang X, Liu K, Roundtree IA, Tempel W, Li Y, et al. (November 2014). "Structural basis for selective binding of m6A RNA by the YTHDC1 YTH domain".
300:
840:
In some cases of DNA protection, M6A methylations (along with M4C modifications) play a role in the protection of bacterial DNA by influencing certain
3493:"Association of type 2 diabetes susceptibility variants with advanced prostate cancer risk in the Breast and Prostate Cancer Cohort Consortium"
4741:
860:
mA modifications, along with other epigenetic changes, have been shown to play important roles during eukaryotic development. Hematopoietic
3405:
Casalegno-Garduño R, Schmitt A, Wang X, Xu X, Schmitt M (October 2010). "Wilms' tumor 1 as a novel target for immunotherapy of leukemia".
523:) have been characterized as direct mA readers and have a conserved mA-binding pocket. Insulin-like growth factor-2 mRNA-binding proteins
284:
InChI=1S/C11H15N5O4/c1-12-9-6-10(14-3-13-9)16(4-15-6)11-8(19)7(18)5(2-17)20-11/h3-5,7-8,11,17-19H,2H2,1H3,(H,12,13,14)/t5-,7-,8-,11-/m1/s1
625:
of mAC consistent with the previously identified motif. The localization of individual mA sites in many mRNAs is highly similar between
777:
have been shown to associate with body mass index in human populations and occurrence of obesity and diabetes. The influence of FTO on
3356:"Association between variations in the fat mass and obesity-associated gene and pancreatic cancer risk: a case-control study in Japan"
613:. A >90% reduction of mA levels in mature plants leads to dramatically altered growth patterns and floral homeotic abnormalities.
4389:
3716:
662:
275:
3652:"Association study of type 2 diabetes genetic susceptibility variants and risk of pancreatic cancer: an analysis of PanScan-I data"
3699:
Bokar JA (2005-01-01). "The biosynthesis and functional roles of methylated nucleosides in eukaryotic mRNA". In
Grosjean H (ed.).
621:
Mapping of mA in human and mouse RNA has identified over 18,000 mA sites in the transcripts of more than 7,000 human genes with a
5666:
1798:"Precise localization of m6A in Rous sarcoma virus RNA reveals clustering of methylation sites: implications for RNA processing"
404:
and DNA. It is found within some viruses, and most eukaryotes including mammals, insects, plants and yeast. It is also found in
2799:"mA mRNA modifications are deposited in nascent pre-mRNA and are not required for splicing but do specify cytoplasmic turnover"
845:
497:
5430:
5425:
5009:
Jiang, Xiulin; Liu, Baiyang; Nie, Zhi; Duan, Lincan; Xiong, Qiuxia; Jin, Zhixian; Yang, Cuiping; Chen, Yongbin (2021-02-21).
1110:"N6-Methyladenosine in RNA and DNA: An Epitranscriptomic and Epigenetic Player Implicated in Determination of Stem Cell Fate"
3783:"Hypoxia induces the breast cancer stem cell phenotype by HIF-dependent and ALKBH5-mediated m⁶A-demethylation of NANOG mRNA"
476:
mRNA. More recent studies have characterized other key components of the mA methyltransferase complex in mammals, including
3254:"Biochemical function of female-lethal (2)D/Wilms' tumor suppressor-1-associated proteins in alternative pre-mRNA splicing"
1257:
Wei CM, Gershowitz A, Moss B (January 1976). "5'-Terminal and internal methylated nucleotide sequences in HeLa cell mRNA".
668:
mA is susceptible to dynamic regulation both throughout development and in response to cellular stimuli. Analysis of mA in
540:
whereas readers mediate mA-dependent functions. mA has also been shown to mediate a structural switch termed mA switch.
378:
5568:
5382:
5377:
4174:"The Demethylase Activity of FTO (Fat Mass and Obesity Associated Protein) Is Required for Preadipocyte Differentiation"
2640:"Adenosine Methylation in Arabidopsis mRNA is Associated with the 3' End and Reduced Levels Cause Developmental Defects"
1433:"MTA is an Arabidopsis messenger RNA adenosine methylase and interacts with a homolog of a sex-specific splicing factor"
799:
218:
5606:
5601:
5397:
790:, a member of the m6A methyltransferase complex, markedly inhibited colorectal cancer cells growth when knocked down.
610:
239:
3891:"A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity"
5621:
5517:
5512:
5407:
5402:
5392:
5339:
5631:
5532:
5484:
638:
1708:"Purification and cDNA cloning of the AdoMet-binding subunit of the human mRNA (N6-adenosine)-methyltransferase"
5626:
5616:
5359:
3442:"Identification of an MSI-H tumor-specific cytotoxic T cell epitope generated by the (-1) frame of U79260(FTO)"
581:
5527:
496:) in 2011 confirmed this hypothesis and revitalized the interests in the study of mA. A second mA demethylase
4837:
Raghunathan, Nalini; Goswami, Sayantan; Leela, Jakku K.; Pandiyan, Apuratha; Gowrishankar, Jayaraman (2019).
5588:
5546:
5499:
5387:
5334:
5329:
4068:"FTO-dependent demethylation of N6-methyladenosine regulates mRNA splicing and is required for adipogenesis"
877:
446:
3542:"Evaluating genome-wide association study-identified breast cancer risk variants in African-American women"
5611:
5563:
5558:
5349:
2694:"RMBase: a resource for decoding the landscape of RNA modifications from high-throughput sequencing data"
1294:"The methylated constituents of L cell messenger RNA: evidence for an unusual cluster at the 5' terminus"
400:) was originally identified and partially characterised in the 1970s, and is an abundant modification in
5578:
5522:
5479:
5474:
5344:
544:
66:
53:
4665:"The Significance of N6-Methyladenosine RNA Methylation in Regulating the Hepatitis B Virus Life Cycle"
2056:"Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5' sites"
1073:
Beemon K, Keith J (June 1977). "Localization of N6-methyladenosine in the Rous sarcoma virus genome".
5656:
5583:
5573:
5494:
5296:
4839:"A new role for Escherichia coli Dam DNA methylase in prevention of aberrant chromosomal replication"
4611:
Gokhale NS, McIntyre AB, McFadden MJ, Roder AE, Kennedy EM, Gandara JA, et al. (November 2016).
4287:
4185:
4128:
3902:
3794:
3553:
3491:
Machiela MJ, Lindström S, Allen NE, Haiman CA, Albanes D, Barricarte A, et al. (December 2012).
2908:
2591:
2534:
2298:
1858:
1664:
1208:
Gokhale NS, McIntyre AB, McFadden MJ, Roder AE, Kennedy EM, Gandara JA, et al. (November 2016).
972:
912:
601:
4720:
O’Brown, Zach
Klapholz; Greer, Eric Lieberman (2016), Jeltsch, Albert; Jurkowska, Renata Z. (eds.),
5489:
5369:
5258:
5161:
4376:. Advances in Enzymology - and Related Areas of Molecular Biology. Vol. 65. pp. 255–285.
3540:
Long J, Zhang B, Signorello LB, Cai Q, Deming-Halverson S, Shrubsole MJ, et al. (2013-04-08).
834:
417:
256:
108:
5291:
4276:"Mutations in prion-like domains in hnRNPA2B1 and hnRNPA1 cause multisystem proteinopathy and ALS"
3211:
Heiliger KJ, Hess J, Vitagliano D, Salerno P, Braselmann H, Salvatore G, et al. (June 2012).
837:
using M6A modifications which influence other repair proteins by recognizing specific mismatches.
4256:
4017:
Karra E, O'Daly OG, Choudhury AI, Yousseif A, Millership S, Neary MT, et al. (August 2013).
2474:"Recognition of RNA N-methyladenosine by IGF2BP proteins enhances mRNA stability and translation"
1688:
938:
876:
readers and writers. In the reproductive system, mA modifications have been shown to disrupt the
873:
622:
504:
469:
3889:
Frayling TM, Timpson NJ, Weedon MN, Zeggini E, Freathy RM, Lindgren CM, et al. (May 2007).
2750:"A majority of m6A residues are in the last exons, allowing the potential for 3' UTR regulation"
1958:"N6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells"
1599:"Comprehensive analysis of mRNA methylation reveals enrichment in 3' UTRs and near stop codons"
5445:
5417:
5048:
5030:
4991:
4973:
4934:
4916:
4868:
4819:
4801:
4757:
4737:
4702:
4684:
4642:
4593:
4544:
4495:
4464:"N(6)-methyladenosine of HIV-1 RNA regulates viral infection and HIV-1 Gag protein expression"
4444:
4395:
4385:
4354:
4313:
4248:
4213:
4154:
4115:
Merkestein M, Laber S, McMurray F, Andrew D, Sachse G, Sanderson J, et al. (April 2015).
4097:
4048:
3999:
3963:
3928:
3871:
3822:
3763:
3712:
3681:
3632:
3581:
3522:
3473:
3422:
3387:
3336:
3285:
3234:
3193:
3144:
3116:
3065:
3024:
2983:
2934:
2877:
2828:
2779:
2723:
2671:
2617:
2560:
2503:
2449:
2408:
2373:
2324:
2267:
2218:
2169:
2134:
2103:
van Tran N, Ernst FG, Hawley BR, Zorbas C, Ulryck N, Hackert P, et al. (September 2019).
2085:
2036:
1987:
1938:
1886:
1827:
1778:
1729:
1680:
1628:
1568:
1514:
1462:
1413:
1350:
1315:
1274:
1239:
1190:
1141:
1090:
1052:
1000:
930:
869:
754:
674:
565:
438:
413:
4413:
Kennedy EM, Bogerd HP, Kornepati AV, Kang D, Ghoshal D, Marshall JB, et al. (May 2016).
2897:"FTO mediates LINE1 mA demethylation and chromatin regulation in mESCs and mouse development"
5661:
5215:
5038:
5022:
4981:
4965:
4924:
4906:
4858:
4850:
4809:
4793:
4747:
4729:
4692:
4676:
4632:
4624:
4583:
4575:
4534:
4526:
4513:
Lichinchi G, Gao S, Saletore Y, Gonzalez GM, Bansal V, Wang Y, et al. (February 2016).
4485:
4475:
4434:
4426:
4377:
4344:
4303:
4295:
4240:
4203:
4193:
4144:
4136:
4087:
4079:
4038:
4030:
3991:
3955:
3918:
3910:
3861:
3853:
3812:
3802:
3753:
3745:
3704:
3671:
3663:
3622:
3612:
3571:
3561:
3512:
3504:
3463:
3453:
3414:
3377:
3367:
3326:
3316:
3275:
3265:
3224:
3183:
3175:
3106:
3096:
3055:
3014:
3001:
Fustin JM, Doi M, Yamaguchi Y, Hida H, Nishimura S, Yoshida M, et al. (November 2013).
2973:
2965:
2924:
2916:
2867:
2859:
2848:"Settling the mA debate: methylation of mature mRNA is not dynamic but accelerates turnover"
2818:
2810:
2769:
2761:
2713:
2705:
2661:
2651:
2607:
2599:
2550:
2542:
2493:
2485:
2439:
2400:
2363:
2355:
2314:
2306:
2257:
2249:
2208:
2200:
2161:
2124:
2116:
2075:
2067:
2054:
Schwartz S, Mumbach MR, Jovanovic M, Wang T, Maciag K, Bushkin GG, et al. (July 2014).
2026:
2018:
1977:
1969:
1928:
1920:
1876:
1866:
1817:
1809:
1768:
1760:
1719:
1672:
1618:
1610:
1558:
1550:
1504:
1496:
1452:
1444:
1405:
1377:
1342:
1305:
1266:
1229:
1221:
1180:
1172:
1159:
Courtney DG, Kennedy EM, Dumm RE, Bogerd HP, Tsai K, Heaton NS, Cullen BR (September 2017).
1131:
1121:
1082:
1042:
1034:
990:
980:
920:
830:
489:
461:
323:
485:
227:
182:
5205:
5134:
4838:
3703:. Topics in Current Genetics. Vol. 12. Springer Berlin Heidelberg. pp. 141–177.
2952:
Abakir A, Giles TC, Cristini A, Foster JM, Dai N, Starczak M, et al. (January 2020).
2523:"N(6)-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions"
823:
815:
746:
724:
720:
712:
592:
In plants, the majority of the mA is found within 150 nucleotides before the start of the
118:
3164:"Clinical and genetic predictors of weight gain in patients diagnosed with breast cancer"
260:
4697:
4664:
4515:"Dynamics of the human and viral m(6)A RNA methylomes during HIV-1 infection of T cells"
4291:
4189:
4132:
3906:
3798:
3557:
2912:
2595:
2538:
2302:
2007:"Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase"
1862:
1668:
976:
961:"Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells"
916:
162:
5321:
5271:
5266:
5187:
5043:
5010:
4986:
4953:
4929:
4894:
4863:
4814:
4782:"m6A methylation potentiates cytosolic dsDNA recognition in a sequence-specific manner"
4781:
4752:
4637:
4612:
4588:
4563:
4539:
4514:
4490:
4463:
4439:
4414:
4308:
4275:
4208:
4173:
4149:
4116:
4092:
4067:
4043:
4018:
3923:
3890:
3866:
3841:
3817:
3782:
3758:
3733:
3676:
3651:
3627:
3600:
3576:
3541:
3517:
3492:
3468:
3441:
3418:
3382:
3355:
3331:
3304:
3188:
3163:
3111:
3084:
2978:
2953:
2929:
2896:
2872:
2847:
2823:
2798:
2774:
2749:
2718:
2693:
2666:
2639:
2612:
2579:
2555:
2522:
2498:
2473:
2368:
2343:
2319:
2286:
2262:
2238:"ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility"
2237:
2213:
2188:
2129:
2104:
2080:
2055:
2031:
2006:
1982:
1957:
1933:
1908:
1724:
1707:
1623:
1598:
1563:
1538:
1457:
1432:
1234:
1209:
1185:
1160:
1136:
1109:
742:
372:
4663:
Moon, Jae-Su; Lee, Wooseong; Cho, Yong-Hee; Kim, Yonghyo; Kim, Geon-Woo (2024-02-28).
3252:
Ortega A, Niksic M, Bachi A, Wilm M, Sánchez L, Hastie N, Valcárcel J (January 2003).
3213:"Novel candidate genes of thyroid tumourigenesis identified in Trk-T1 transgenic mice"
2189:"N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO"
1881:
1846:
1822:
1797:
1773:
1748:
1509:
1484:
1047:
1022:
995:
960:
5650:
5440:
5435:
5354:
5286:
5225:
4333:"Comprehensive Genomic Characterization of RNA-Binding Proteins across Human Cancers"
1381:
1346:
1310:
1293:
1086:
849:
778:
758:
750:
634:
4274:
Kim HJ, Kim NC, Wang YD, Scarborough EA, Moore J, Diaz Z, et al. (March 2013).
4260:
3162:
Reddy SM, Sadim M, Li J, Yi N, Agarwal S, Mantzoros CS, Kaklamani VG (August 2013).
3003:"RNA-methylation-dependent RNA processing controls the speed of the circadian clock"
1161:"Epitranscriptomic Enhancement of Influenza A Virus Gene Expression and Replication"
700:
in human cells, where it is involved in regulation of stability of RNA:DNA hybrids.
5281:
5124:
5092:
3599:
Kaklamani V, Yi N, Sadim M, Siziopikou K, Zhang K, Xu Y, et al. (April 2011).
3083:
Fustin JM, Ye S, Rakers C, Kaneko K, Fukumoto K, Yamano M, et al. (May 2020).
2748:
Ke S, Alemu EA, Mertens C, Gantman EC, Fak JJ, Mele A, et al. (October 2015).
2426:
Xiao W, Adhikari S, Dahal U, Chen YS, Hao YJ, Sun BF, et al. (February 2016).
2236:
Zheng G, Dahl JA, Niu Y, Fedorcsak P, Huang CM, Li CJ, et al. (January 2013).
1692:
942:
841:
762:
678:
593:
207:
3601:"The role of the fat mass and obesity associated gene (FTO) in breast cancer risk"
2797:
Ke S, Pandya-Jones A, Saito Y, Fak JJ, Vågbø CB, Geula S, et al. (May 2017).
1909:"A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation"
543:
The specificity of mA installation on mRNA is controlled by exon architecture and
40:
17:
4911:
4733:
4415:"Posttranscriptional m(6)A Editing of HIV-1 mRNAs Enhances Viral Gene Expression"
4349:
4332:
4198:
3749:
3566:
2444:
2427:
2253:
2071:
4952:
Loenen, W. A. M.; Dryden, D. T. F.; Raleigh, E. A.; Wilson, G. G. (2014-01-01).
3781:
Zhang C, Samanta D, Lu H, Bullen JW, Zhang H, Chen I, et al. (April 2016).
1038:
729:
669:
650:
577:
481:
430:
5026:
4721:
4628:
4579:
4530:
4430:
3995:
3959:
3787:
Proceedings of the
National Academy of Sciences of the United States of America
3734:"The m(6)A Methyltransferase METTL3 Promotes Translation in Human Cancer Cells"
3060:
3043:
3019:
3002:
2359:
2005:
Ping XL, Sun BF, Wang L, Xiao W, Yang X, Wang WJ, et al. (February 2014).
1851:
Proceedings of the
National Academy of Sciences of the United States of America
1614:
1225:
1176:
965:
Proceedings of the
National Academy of Sciences of the United States of America
5310:
5248:
5243:
5178:
5141:
5101:
4381:
3667:
3101:
2969:
2489:
819:
357:
173:
5034:
4977:
4920:
4805:
4688:
3372:
3303:
Jin DI, Lee SW, Han ME, Kim HJ, Seo SA, Hur GY, et al. (December 2012).
2954:"N-methyladenosine regulates the stability of RNA:DNA hybrids in human cells"
1764:
1597:
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR (June 2012).
5276:
5200:
5195:
5168:
5061:
4728:, vol. 945, Cham: Springer International Publishing, pp. 213–246,
4331:
Wang ZL, Li B, Luo YX, Lin Q, Liu SR, Zhang XQ, et al. (January 2018).
4066:
Zhao X, Yang Y, Sun BF, Shi Y, Yang X, Xiao W, et al. (December 2014).
3914:
3857:
3807:
3617:
3305:"Expression and roles of Wilms' tumor 1-associating protein in glioblastoma"
3085:"Methylation deficiency disrupts biological rhythms from bacteria to humans"
2920:
2656:
2603:
2578:
He PC, Wei J, Dou X, Harada BT, Zhang Z, Ge R, et al. (February 2023).
1871:
985:
865:
861:
783:
770:
690:
536:
473:
434:
5052:
4995:
4938:
4872:
4823:
4761:
4706:
4680:
4646:
4597:
4548:
4499:
4448:
4358:
4317:
4252:
4217:
4158:
4101:
4052:
4003:
3967:
3932:
3875:
3826:
3767:
3685:
3636:
3585:
3526:
3477:
3426:
3391:
3354:
Lin Y, Ueda J, Yagyu K, Ishii H, Ueno M, Egawa N, et al. (July 2013).
3340:
3289:
3270:
3253:
3238:
3197:
3148:
3120:
3069:
3028:
2987:
2938:
2881:
2863:
2832:
2814:
2783:
2765:
2727:
2675:
2621:
2564:
2507:
2453:
2412:
2377:
2342:
Wang X, Zhao BS, Roundtree IA, Lu Z, Han D, Ma H, et al. (June 2015).
2328:
2271:
2222:
2173:
2138:
2089:
2040:
1991:
1942:
1684:
1632:
1572:
1518:
1466:
1448:
1243:
1194:
1145:
1126:
646:
4613:"N6-Methyladenosine in Flaviviridae Viral RNA Genomes Regulates Infection"
4399:
3458:
2709:
2404:
2105:"The human 18S rRNA m6A methyltransferase METTL5 is stabilized by TRMT112"
1924:
1890:
1845:
Horowitz S, Horowitz A, Nilsen TW, Munns TW, Rottman FM (September 1984).
1831:
1782:
1733:
1706:
Bokar JA, Shambaugh ME, Polayes D, Matera AG, Rottman FM (November 1997).
1319:
1210:"N6-Methyladenosine in Flaviviridae Viral RNA Genomes Regulates Infection"
1004:
934:
5220:
5156:
5129:
4969:
4854:
4564:"Dynamics of Human and Viral RNA Methylation during Zika Virus Infection"
4172:
Zhang M, Zhang Y, Ma J, Guo F, Cao Q, Zhang Y, et al. (2015-07-28).
3508:
3179:
2285:
Wang X, Lu Z, Gomez A, Hon GC, Yue Y, Han D, et al. (January 2014).
2204:
2187:
Jia G, Fu Y, Zhao X, Dai Q, Zheng G, Yang Y, et al. (October 2011).
2165:
2120:
1907:
Liu J, Yue Y, Han D, Wang X, Fu Y, Zhang L, et al. (February 2014).
1813:
1554:
1500:
1417:
1354:
1278:
1094:
1056:
811:
654:
493:
4797:
4562:
Lichinchi G, Zhao BS, Wu Y, Lu Z, Qin Y, He C, Rana TM (November 2016).
4480:
4299:
4083:
3280:
3229:
3212:
2546:
2310:
1749:"Sequence specificity of the human mRNA N6-adenosine methylase in vitro"
1676:
1431:
Zhong S, Li H, Bodi Z, Button J, Vespa L, Herzog M, Fray RG (May 2008).
1270:
5230:
5210:
5151:
5119:
5114:
5011:"The role of m6A modification in the biological functions and diseases"
4372:
Narayan P, Rottman FM (1992). "Methylation of mRNA". In Nord FF (ed.).
4140:
4019:"A link between FTO, ghrelin, and impaired brain food-cue responsivity"
766:
573:
532:
528:
524:
477:
194:
3321:
2638:
Bodi Z, Zhong S, Mehra S, Song J, Graham N, Li H, et al. (2012).
2472:
Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, et al. (March 2018).
2152:
He C (December 2010). "Grand challenge commentary: RNA epigenetics?".
2022:
1956:
Wang Y, Li Y, Toth JI, Petroski MD, Zhang Z, Zhao JC (February 2014).
5146:
5109:
4034:
2344:"N(6)-methyladenosine Modulates Messenger RNA Translation Efficiency"
925:
900:
787:
697:
606:
569:
535:(IGF2BP1–3) are reported as a novel class of mA readers. IGF2BPs use
520:
516:
512:
508:
442:
4244:
4117:"FTO influences adipogenesis by regulating mitotic clonal expansion"
3708:
2895:
Wei J, Yu X, Yang L, Liu X, Gao B, Huang B, et al. (May 2022).
2580:"Exon architecture controls mRNA mA suppression and gene expression"
2287:"N6-methyladenosine-dependent regulation of messenger RNA stability"
1973:
1409:
371:
Except where otherwise noted, data are given for materials in their
3842:"The bigger picture of FTO: the first GWAS-identified obesity gene"
2692:
Sun WJ, Li JH, Liu S, Wu J, Zhou H, Qu LH, Yang JH (January 2016).
901:"Modified nucleosides and bizarre 5'-termini in mouse myeloma mRNA"
704:
642:
630:
626:
153:
141:
131:
1847:"Mapping of N6-methyladenosine residues in bovine prolactin mRNA"
2521:
Liu N, Dai Q, Zheng G, He C, Parisien M, Pan T (February 2015).
686:
422:
409:
405:
401:
5065:
460:, this methyltransferase complex preferentially methylates RNA
682:
4374:
Advances in
Enzymology and Related Areas of Molecular Biology
1483:
Clancy MJ, Shambaugh ME, Timpte CS, Bokar JA (October 2002).
1747:
Harper JE, Miceli SM, Roberts RJ, Manley JL (October 1990).
653:. mA within 3' UTRs is also associated with the presence of
244:
637:-wide analysis reveals that mA is found in regions of high
1292:
Perry RP, Kelley DE, Friderici K, Rottman F (April 1975).
4462:
Tirumuru N, Zhao BS, Lu W, Lu Z, He C, Wu L (July 2016).
3732:
Lin S, Choe J, Du P, Triboulet R, Gregory RI (May 2016).
1537:
Bodi Z, Button JD, Grierson D, Fray RG (September 2010).
464:
containing GGACU and a similar preference was identified
3701:
Fine-Tuning of RNA Functions by Modification and Editing
3440:
Linnebacher M, Wienck A, Boeck I, Klar E (2010-03-18).
1368:
Nichols JL (1979). "In maize poly(A)-containing RNA".
677:
significantly affects gene expression and alternative
2428:"Nuclear m(6)A Reader YTHDC1 Regulates mRNA Splicing"
959:
Desrosiers R, Friderici K, Rottman F (October 1974).
4722:"N6-Methyladenine: A Conserved and Dynamic DNA Mark"
3044:"m(6)A mRNA methylation: a new circadian pacesetter"
5545:
5463:
5416:
5368:
5320:
5309:
5257:
5186:
5177:
5100:
2846:Rosa-Mercado NA, Withers JB, Steitz JA (May 2017).
1650:
1648:
1646:
1644:
1642:
864:(HSCs), Neuronal Stem Cells (NSCs) and Primordial
741:types have been indicated in reports that include
661:Precise m6A mapping by m6A-CLIP/IP (briefly m6A-
4954:"Type I restriction enzymes and their relatives"
206:
117:
1478:
1476:
833:. Another enzyme, Dam DNA methylase regulates
5077:
2467:
2465:
2463:
1592:
1590:
1588:
1586:
1584:
1582:
1023:"Methylation of nuclear simian virus 40 RNAs"
954:
952:
894:
892:
8:
2633:
2631:
1532:
1530:
1528:
1068:
1066:
1016:
1014:
3650:Pierce BL, Austin MA, Ahsan H (June 2011).
2687:
2685:
5551:
5467:
5317:
5183:
5084:
5070:
5062:
4726:DNA Methyltransferases - Role and Function
3446:Journal of Biomedicine & Biotechnology
1021:Aloni Y, Dhar R, Khoury G (October 1979).
696:mA is also found on the RNA components of
259:
181:
29:
5042:
4985:
4928:
4910:
4895:"The Epigenomic Landscape of Prokaryotes"
4862:
4813:
4751:
4696:
4669:Journal of Microbiology and Biotechnology
4636:
4587:
4538:
4489:
4479:
4438:
4348:
4307:
4207:
4197:
4148:
4091:
4042:
3922:
3865:
3816:
3806:
3757:
3675:
3626:
3616:
3575:
3565:
3516:
3467:
3457:
3381:
3371:
3330:
3320:
3279:
3269:
3228:
3187:
3110:
3100:
3059:
3018:
2977:
2928:
2871:
2822:
2773:
2743:
2741:
2739:
2737:
2717:
2665:
2655:
2611:
2554:
2497:
2443:
2367:
2318:
2261:
2212:
2128:
2079:
2030:
1981:
1932:
1880:
1870:
1821:
1772:
1723:
1622:
1562:
1508:
1456:
1309:
1233:
1184:
1135:
1125:
1046:
994:
984:
924:
681:patterns, resulting in modulation of the
226:
5015:Signal Transduction and Targeted Therapy
3948:Obesity Research & Clinical Practice
707:-mediated silencing of the mA methylase
888:
719:led to a shorter period. The mammalian
715:period. In contrast, overexpression of
500:(ALKBH5) was later discovered as well.
305:
280:
255:
810:M6A methylation is also widespread in
645:and is preferentially enriched within
87:)-2-(Hydroxymethyl)-5-oxolane-2,3-diol
4658:
4656:
4023:The Journal of Clinical Investigation
1902:
1900:
287:Key: VQAYFKKCNSOZKM-IOSLPCCCSA-N
161:
7:
1796:Kane SE, Beemon K (September 1985).
1539:"Yeast targets for mRNA methylation"
480:, Wilms tumor 1 associated protein (
3258:The Journal of Biological Chemistry
878:maternal-to-zygotic mRNA transition
641:. mA is found within long internal
197:
3419:10.1016/j.transproceed.2010.07.034
1108:Ji P, Wang X, Xie N, Li Y (2018).
445:, which is the subunit that binds
25:
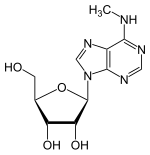
658:N6-Methyladenosines (mA) in RNA.
3840:Loos RJ, Yeo GS (January 2014).
3497:American Journal of Epidemiology
1398:Canadian Journal of Biochemistry
814:, influencing functions such as
605:homologue of METTL3, results in
341:
335:
39:
846:restriction-modification system
375:(at 25 °C , 100 kPa).
1802:Molecular and Cellular Biology
848:, decreasing the influence of
576:, are known to be targets for
347:
329:
308:n2c1c(ncnc1NC)n(c2)3O((O)3O)CO
1:
3846:Nature Reviews. Endocrinology
3042:Hastings MH (November 2013).
899:Adams JM, Cory S (May 1975).
711:led to the elongation of the
505:YT521-B homology (YTH) domain
4912:10.1371/journal.pgen.1005854
4734:10.1007/978-3-319-43624-1_10
4350:10.1016/j.celrep.2017.12.035
4199:10.1371/journal.pone.0133788
3750:10.1016/j.molcel.2016.03.021
3567:10.1371/journal.pone.0058350
2445:10.1016/j.molcel.2016.01.012
2254:10.1016/j.molcel.2012.10.015
2072:10.1016/j.celrep.2014.05.048
1382:10.1016/0304-4211(79)90141-X
1347:10.1016/0022-2836(78)90350-9
1335:Journal of Molecular Biology
1311:10.1016/0092-8674(75)90159-2
1087:10.1016/0022-2836(77)90047-X
1075:Journal of Molecular Biology
800:pattern recognition receptor
3656:Cancer Causes & Control
3407:Transplantation Proceedings
1039:10.1128/JVI.32.1.52-60.1979
826:, and prokaryotic defense.
416:(snRNA) as well as several
5683:
5314:(Nucleoside monophosphate)
5027:10.1038/s41392-020-00450-x
4629:10.1016/j.chom.2016.09.015
4580:10.1016/j.chom.2016.10.002
4531:10.1038/nmicrobiol.2016.11
4431:10.1016/j.chom.2016.04.002
3996:10.1016/j.gene.2013.06.079
3960:10.1016/j.orcp.2011.12.007
3061:10.1016/j.cell.2013.10.028
3020:10.1016/j.cell.2013.10.026
2644:Frontiers in Plant Science
2360:10.1016/j.cell.2015.05.014
1615:10.1016/j.cell.2012.05.003
1226:10.1016/j.chom.2016.09.015
1177:10.1016/j.chom.2017.08.004
472:genomic RNA and in bovine
437:is directed by a large mA
5597:
5554:
5508:
5470:
5169:Unnatural base pair (UBP)
4382:10.1002/9780470123119.ch7
3668:10.1007/s10552-011-9760-5
3168:British Journal of Cancer
3102:10.1038/s42003-020-0942-0
2970:10.1038/s41588-019-0549-x
2490:10.1038/s41556-018-0045-z
689:) signalling pathway and
639:evolutionary conservation
564:), the expression of the
369:
316:
296:
271:
101:
93:
65:
52:
47:
38:
27:Modification in mRNA, DNA
3373:10.1186/1471-2407-13-337
3217:Endocrine-Related Cancer
1114:Stem Cells International
562:Saccharomyces cerevisiae
5667:Hydroxymethyl compounds
5547:Nucleoside triphosphate
4617:Cell Host & Microbe
4568:Cell Host & Microbe
4419:Cell Host & Microbe
3915:10.1126/science.1141634
3858:10.1038/nrendo.2013.227
3808:10.1073/pnas.1602883113
3618:10.1186/1471-2350-12-52
2921:10.1126/science.abe9582
2852:Genes & Development
2803:Genes & Development
2754:Genes & Development
2657:10.3389/fpls.2012.00048
2604:10.1126/science.abj9090
2393:Nature Chemical Biology
2193:Nature Chemical Biology
2154:Nature Chemical Biology
1913:Nature Chemical Biology
1872:10.1073/pnas.81.18.5667
1214:Cell Host & Microbe
1165:Cell Host & Microbe
986:10.1073/pnas.71.10.3971
545:exon junction complexes
537:K homology (KH) domains
5500:Xanthosine diphosphate
5464:Nucleoside diphosphate
4958:Nucleic Acids Research
4843:Nucleic Acids Research
4681:10.4014/jmb.2309.09013
3271:10.1074/jbc.M210737200
3089:Communications Biology
2864:10.1101/gad.302695.117
2815:10.1101/gad.301036.117
2766:10.1101/gad.269415.115
2698:Nucleic Acids Research
2109:Nucleic Acids Research
1765:10.1093/nar/18.19.5735
1753:Nucleic Acids Research
1543:Nucleic Acids Research
1489:Nucleic Acids Research
1449:10.1105/tpc.108.058883
599:Mutations of MTA, the
468:in mapped mA sites in
4121:Nature Communications
2405:10.1038/nchembio.1654
1925:10.1038/nchembio.1432
1370:Plant Science Letters
736:Clinical significance
67:Systematic IUPAC name
3605:BMC Medical Genetics
3180:10.1038/bjc.2013.441
2205:10.1038/nchembio.687
2166:10.1038/nchembio.482
1814:10.1128/mcb.5.9.2298
1127:10.1155/2018/3256524
602:Arabidopsis thaliana
551:Species distribution
507:family of proteins (
5370:Deoxyribonucleotide
5259:Deoxyribonucleoside
5162:Pyrimidine analogue
4798:10.1098/rsob.210030
4519:Nature Microbiology
4481:10.7554/eLife.15528
4300:10.1038/nature11922
4292:2013Natur.495..467K
4233:Nature Neuroscience
4190:2015PLoSO..1033788Z
4133:2015NatCo...6.6792M
4084:10.1038/cr.2014.151
3907:2007Sci...316..889F
3799:2016PNAS..113E2047Z
3793:(14): E2047–E2056.
3558:2013PLoSO...858350L
3459:10.1155/2010/841451
3230:10.1530/ERC-11-0387
3137:Anticancer Research
2913:2022Sci...376..968W
2710:10.1093/nar/gkv1036
2596:2023Sci...379..677H
2547:10.1038/nature14234
2539:2015Natur.518..560L
2478:Nature Cell Biology
2311:10.1038/nature12730
2303:2014Natur.505..117W
1962:Nature Cell Biology
1863:1984PNAS...81.5667H
1677:10.1038/nature11112
1669:2012Natur.485..201D
1271:10.1021/bi00647a024
1027:Journal of Virology
977:1974PNAS...71.3971D
917:1975Natur.255...28A
441:complex containing
418:long non-coding RNA
365: g·mol
35:
4970:10.1093/nar/gkt847
4855:10.1093/nar/gkz242
4141:10.1038/ncomms7792
3509:10.1093/aje/kws191
2121:10.1093/nar/gkz619
1555:10.1093/nar/gkq266
1501:10.1093/nar/gkf573
623:consensus sequence
560:In budding yeast (
470:Rous sarcoma virus
379:Infobox references
30:
18:N6-methyladenosine
5644:
5643:
5640:
5639:
5541:
5540:
5459:
5458:
5418:Cyclic nucleotide
5305:
5304:
4849:(11): 5698–5711.
4743:978-3-319-43624-1
4286:(7442): 467–473.
4078:(12): 1403–1419.
3901:(5826): 889–894.
3503:(12): 1121–1129.
3322:10.1111/cas.12022
3315:(12): 2102–2109.
2907:(6596): 968–973.
2760:(19): 2037–2053.
2704:(D1): D259–D265.
2590:(6633): 677–682.
2533:(7540): 560–564.
2297:(7481): 117–120.
2115:(15): 7719–7733.
2023:10.1038/cr.2014.3
1857:(18): 5667–5671.
1759:(19): 5735–5741.
1718:(11): 1233–1247.
1663:(7397): 201–206.
1549:(16): 5327–5335.
1495:(20): 4509–4518.
1171:(3): 377–386.e5.
971:(10): 3971–3975.
755:pancreatic cancer
675:methyltransferase
453:
439:methyltransferase
414:small nuclear RNA
387:Chemical compound
385:
384:
240:CompTox Dashboard
143:Interactive image
34:-Methyladenosine
16:(Redirected from
5674:
5552:
5468:
5318:
5239:-Methyladenosine
5216:5-Methylcytidine
5184:
5086:
5079:
5072:
5063:
5057:
5056:
5046:
5006:
5000:
4999:
4989:
4949:
4943:
4942:
4932:
4914:
4889:
4883:
4882:
4880:
4879:
4866:
4834:
4828:
4827:
4817:
4777:
4771:
4770:
4769:
4768:
4755:
4717:
4711:
4710:
4700:
4660:
4651:
4650:
4640:
4608:
4602:
4601:
4591:
4559:
4553:
4552:
4542:
4510:
4504:
4503:
4493:
4483:
4459:
4453:
4452:
4442:
4410:
4404:
4403:
4369:
4363:
4362:
4352:
4328:
4322:
4321:
4311:
4271:
4265:
4264:
4239:(8): 1042–1048.
4228:
4222:
4221:
4211:
4201:
4169:
4163:
4162:
4152:
4112:
4106:
4105:
4095:
4063:
4057:
4056:
4046:
4035:10.1172/jci44403
4029:(8): 3539–3551.
4014:
4008:
4007:
3978:
3972:
3971:
3943:
3937:
3936:
3926:
3886:
3880:
3879:
3869:
3837:
3831:
3830:
3820:
3810:
3778:
3772:
3771:
3761:
3729:
3723:
3722:
3696:
3690:
3689:
3679:
3647:
3641:
3640:
3630:
3620:
3596:
3590:
3589:
3579:
3569:
3537:
3531:
3530:
3520:
3488:
3482:
3481:
3471:
3461:
3437:
3431:
3430:
3413:(8): 3309–3311.
3402:
3396:
3395:
3385:
3375:
3351:
3345:
3344:
3334:
3324:
3300:
3294:
3293:
3283:
3273:
3264:(5): 3040–3047.
3249:
3243:
3242:
3232:
3208:
3202:
3201:
3191:
3159:
3153:
3152:
3143:(9): 4049–4059.
3131:
3125:
3124:
3114:
3104:
3080:
3074:
3073:
3063:
3039:
3033:
3032:
3022:
2998:
2992:
2991:
2981:
2949:
2943:
2942:
2932:
2892:
2886:
2885:
2875:
2843:
2837:
2836:
2826:
2809:(10): 990–1006.
2794:
2788:
2787:
2777:
2745:
2732:
2731:
2721:
2689:
2680:
2679:
2669:
2659:
2635:
2626:
2625:
2615:
2575:
2569:
2568:
2558:
2518:
2512:
2511:
2501:
2469:
2458:
2457:
2447:
2423:
2417:
2416:
2388:
2382:
2381:
2371:
2354:(6): 1388–1399.
2339:
2333:
2332:
2322:
2282:
2276:
2275:
2265:
2233:
2227:
2226:
2216:
2184:
2178:
2177:
2149:
2143:
2142:
2132:
2100:
2094:
2093:
2083:
2051:
2045:
2044:
2034:
2002:
1996:
1995:
1985:
1953:
1947:
1946:
1936:
1904:
1895:
1894:
1884:
1874:
1842:
1836:
1835:
1825:
1808:(9): 2298–2306.
1793:
1787:
1786:
1776:
1744:
1738:
1737:
1727:
1703:
1697:
1696:
1652:
1637:
1636:
1626:
1609:(7): 1635–1646.
1594:
1577:
1576:
1566:
1534:
1523:
1522:
1512:
1480:
1471:
1470:
1460:
1443:(5): 1278–1288.
1428:
1422:
1421:
1392:
1386:
1385:
1365:
1359:
1358:
1330:
1324:
1323:
1313:
1289:
1283:
1282:
1254:
1248:
1247:
1237:
1205:
1199:
1198:
1188:
1156:
1150:
1149:
1139:
1129:
1105:
1099:
1098:
1070:
1061:
1060:
1050:
1018:
1009:
1008:
998:
988:
956:
947:
946:
928:
926:10.1038/255028a0
896:
584:of IME4 itself.
462:oligonucleotides
451:
394:-Methyladenosine
364:
349:
343:
337:
331:
324:Chemical formula
264:
263:
248:
246:
230:
210:
199:
185:
165:
145:
121:
60:-Methyladenosine
43:
36:
21:
5682:
5681:
5677:
5676:
5675:
5673:
5672:
5671:
5647:
5646:
5645:
5636:
5593:
5537:
5504:
5455:
5412:
5364:
5313:
5301:
5297:Deoxyxanthosine
5253:
5206:5-Methyluridine
5173:
5135:Purine analogue
5096:
5090:
5060:
5008:
5007:
5003:
4951:
4950:
4946:
4905:(2): e1005854.
4891:
4890:
4886:
4877:
4875:
4836:
4835:
4831:
4779:
4778:
4774:
4766:
4764:
4744:
4719:
4718:
4714:
4662:
4661:
4654:
4610:
4609:
4605:
4561:
4560:
4556:
4512:
4511:
4507:
4461:
4460:
4456:
4412:
4411:
4407:
4392:
4371:
4370:
4366:
4330:
4329:
4325:
4273:
4272:
4268:
4245:10.1038/nn.3449
4230:
4229:
4225:
4184:(7): e0133788.
4171:
4170:
4166:
4114:
4113:
4109:
4065:
4064:
4060:
4016:
4015:
4011:
3980:
3979:
3975:
3945:
3944:
3940:
3888:
3887:
3883:
3839:
3838:
3834:
3780:
3779:
3775:
3731:
3730:
3726:
3719:
3709:10.1007/b106365
3698:
3697:
3693:
3649:
3648:
3644:
3598:
3597:
3593:
3539:
3538:
3534:
3490:
3489:
3485:
3439:
3438:
3434:
3404:
3403:
3399:
3353:
3352:
3348:
3302:
3301:
3297:
3251:
3250:
3246:
3210:
3209:
3205:
3161:
3160:
3156:
3133:
3132:
3128:
3082:
3081:
3077:
3041:
3040:
3036:
3000:
2999:
2995:
2958:Nature Genetics
2951:
2950:
2946:
2894:
2893:
2889:
2858:(10): 957–958.
2845:
2844:
2840:
2796:
2795:
2791:
2747:
2746:
2735:
2691:
2690:
2683:
2637:
2636:
2629:
2577:
2576:
2572:
2520:
2519:
2515:
2471:
2470:
2461:
2425:
2424:
2420:
2399:(11): 927–929.
2390:
2389:
2385:
2341:
2340:
2336:
2284:
2283:
2279:
2235:
2234:
2230:
2199:(12): 885–887.
2186:
2185:
2181:
2160:(12): 863–865.
2151:
2150:
2146:
2102:
2101:
2097:
2053:
2052:
2048:
2004:
2003:
1999:
1974:10.1038/ncb2902
1955:
1954:
1950:
1906:
1905:
1898:
1844:
1843:
1839:
1795:
1794:
1790:
1746:
1745:
1741:
1705:
1704:
1700:
1654:
1653:
1640:
1596:
1595:
1580:
1536:
1535:
1526:
1482:
1481:
1474:
1430:
1429:
1425:
1410:10.1139/o79-112
1394:
1393:
1389:
1367:
1366:
1362:
1332:
1331:
1327:
1291:
1290:
1286:
1256:
1255:
1251:
1207:
1206:
1202:
1158:
1157:
1153:
1107:
1106:
1102:
1072:
1071:
1064:
1020:
1019:
1012:
958:
957:
950:
911:(5503): 28–33.
898:
897:
890:
886:
858:
835:mismatch repair
824:gene expression
816:DNA replication
808:
747:prostate cancer
738:
721:circadian clock
685:(also known as
619:
590:
558:
553:
388:
381:
376:
362:
352:
346:
340:
334:
326:
312:
309:
304:
303:
292:
289:
288:
285:
279:
278:
267:
249:
242:
233:
213:
200:
188:
168:
148:
135:
124:
111:
97:
89:
88:
61:
28:
23:
22:
15:
12:
11:
5:
5680:
5678:
5670:
5669:
5664:
5659:
5649:
5648:
5642:
5641:
5638:
5637:
5635:
5634:
5629:
5624:
5619:
5614:
5609:
5604:
5598:
5595:
5594:
5592:
5591:
5586:
5581:
5576:
5571:
5566:
5561:
5555:
5549:
5543:
5542:
5539:
5538:
5536:
5535:
5530:
5525:
5520:
5515:
5509:
5506:
5505:
5503:
5502:
5497:
5492:
5487:
5482:
5477:
5471:
5465:
5461:
5460:
5457:
5456:
5454:
5453:
5448:
5443:
5438:
5433:
5428:
5422:
5420:
5414:
5413:
5411:
5410:
5405:
5400:
5395:
5390:
5385:
5380:
5374:
5372:
5366:
5365:
5363:
5362:
5357:
5352:
5347:
5342:
5337:
5332:
5326:
5324:
5322:Ribonucleotide
5315:
5307:
5306:
5303:
5302:
5300:
5299:
5294:
5289:
5284:
5279:
5274:
5272:Deoxyguanosine
5269:
5267:Deoxyadenosine
5263:
5261:
5255:
5254:
5252:
5251:
5246:
5241:
5233:
5228:
5223:
5218:
5213:
5208:
5203:
5198:
5192:
5190:
5188:Ribonucleoside
5181:
5175:
5174:
5172:
5171:
5166:
5165:
5164:
5159:
5154:
5149:
5139:
5138:
5137:
5132:
5127:
5122:
5117:
5106:
5104:
5098:
5097:
5091:
5089:
5088:
5081:
5074:
5066:
5059:
5058:
5001:
4944:
4884:
4829:
4772:
4742:
4712:
4675:(2): 233–239.
4652:
4623:(5): 654–665.
4603:
4574:(5): 666–673.
4554:
4505:
4454:
4425:(5): 675–685.
4405:
4390:
4364:
4343:(1): 286–298.
4323:
4266:
4223:
4164:
4107:
4058:
4009:
3990:(2): 462–468.
3973:
3954:(1): e14–e22.
3938:
3881:
3832:
3773:
3744:(3): 335–345.
3738:Molecular Cell
3724:
3717:
3691:
3662:(6): 877–883.
3642:
3591:
3532:
3483:
3432:
3397:
3346:
3309:Cancer Science
3295:
3244:
3223:(3): 409–421.
3203:
3174:(4): 872–881.
3154:
3126:
3075:
3054:(4): 740–741.
3034:
3013:(4): 793–806.
2993:
2944:
2887:
2838:
2789:
2733:
2681:
2627:
2570:
2513:
2484:(3): 285–295.
2459:
2438:(4): 507–519.
2432:Molecular Cell
2418:
2383:
2334:
2277:
2242:Molecular Cell
2228:
2179:
2144:
2095:
2066:(1): 284–296.
2046:
2017:(2): 177–189.
1997:
1968:(2): 191–198.
1948:
1896:
1837:
1788:
1739:
1698:
1638:
1578:
1524:
1472:
1437:The Plant Cell
1423:
1404:(6): 927–931.
1387:
1376:(4): 357–361.
1360:
1341:(4): 487–515.
1325:
1304:(4): 387–394.
1284:
1265:(2): 397–401.
1249:
1220:(5): 654–665.
1200:
1151:
1100:
1081:(1): 165–179.
1062:
1010:
948:
887:
885:
882:
857:
856:In Development
854:
850:bacteriophages
807:
804:
743:stomach cancer
737:
734:
618:
615:
611:globular stage
609:arrest at the
589:
586:
557:
554:
552:
549:
498:alkB homolog 5
386:
383:
382:
377:
373:standard state
370:
367:
366:
360:
354:
353:
350:
344:
338:
332:
327:
322:
319:
318:
314:
313:
311:
310:
307:
299:
298:
297:
294:
293:
291:
290:
286:
283:
282:
274:
273:
272:
269:
268:
266:
265:
252:
250:
238:
235:
234:
232:
231:
223:
221:
215:
214:
212:
211:
203:
201:
193:
190:
189:
187:
186:
178:
176:
170:
169:
167:
166:
158:
156:
150:
149:
147:
146:
138:
136:
129:
126:
125:
123:
122:
114:
112:
107:
104:
103:
99:
98:
95:
91:
90:
70:
69:
63:
62:
56:
50:
49:
45:
44:
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
5679:
5668:
5665:
5663:
5660:
5658:
5655:
5654:
5652:
5633:
5630:
5628:
5625:
5623:
5620:
5618:
5615:
5613:
5610:
5608:
5605:
5603:
5600:
5599:
5596:
5590:
5587:
5585:
5582:
5580:
5577:
5575:
5572:
5570:
5567:
5565:
5562:
5560:
5557:
5556:
5553:
5550:
5548:
5544:
5534:
5531:
5529:
5526:
5524:
5521:
5519:
5516:
5514:
5511:
5510:
5507:
5501:
5498:
5496:
5493:
5491:
5488:
5486:
5483:
5481:
5478:
5476:
5473:
5472:
5469:
5466:
5462:
5452:
5449:
5447:
5444:
5442:
5439:
5437:
5434:
5432:
5429:
5427:
5424:
5423:
5421:
5419:
5415:
5409:
5406:
5404:
5401:
5399:
5396:
5394:
5391:
5389:
5386:
5384:
5381:
5379:
5376:
5375:
5373:
5371:
5367:
5361:
5358:
5356:
5353:
5351:
5348:
5346:
5343:
5341:
5338:
5336:
5333:
5331:
5328:
5327:
5325:
5323:
5319:
5316:
5312:
5308:
5298:
5295:
5293:
5290:
5288:
5287:Deoxycytidine
5285:
5283:
5280:
5278:
5275:
5273:
5270:
5268:
5265:
5264:
5262:
5260:
5256:
5250:
5247:
5245:
5242:
5240:
5238:
5234:
5232:
5229:
5227:
5226:Pseudouridine
5224:
5222:
5219:
5217:
5214:
5212:
5209:
5207:
5204:
5202:
5199:
5197:
5194:
5193:
5191:
5189:
5185:
5182:
5180:
5176:
5170:
5167:
5163:
5160:
5158:
5155:
5153:
5150:
5148:
5145:
5144:
5143:
5140:
5136:
5133:
5131:
5128:
5126:
5123:
5121:
5118:
5116:
5113:
5112:
5111:
5108:
5107:
5105:
5103:
5099:
5094:
5087:
5082:
5080:
5075:
5073:
5068:
5067:
5064:
5054:
5050:
5045:
5040:
5036:
5032:
5028:
5024:
5020:
5016:
5012:
5005:
5002:
4997:
4993:
4988:
4983:
4979:
4975:
4971:
4967:
4963:
4959:
4955:
4948:
4945:
4940:
4936:
4931:
4926:
4922:
4918:
4913:
4908:
4904:
4900:
4899:PLOS Genetics
4896:
4888:
4885:
4874:
4870:
4865:
4860:
4856:
4852:
4848:
4844:
4840:
4833:
4830:
4825:
4821:
4816:
4811:
4807:
4803:
4799:
4795:
4791:
4787:
4783:
4776:
4773:
4763:
4759:
4754:
4749:
4745:
4739:
4735:
4731:
4727:
4723:
4716:
4713:
4708:
4704:
4699:
4694:
4690:
4686:
4682:
4678:
4674:
4670:
4666:
4659:
4657:
4653:
4648:
4644:
4639:
4634:
4630:
4626:
4622:
4618:
4614:
4607:
4604:
4599:
4595:
4590:
4585:
4581:
4577:
4573:
4569:
4565:
4558:
4555:
4550:
4546:
4541:
4536:
4532:
4528:
4524:
4520:
4516:
4509:
4506:
4501:
4497:
4492:
4487:
4482:
4477:
4473:
4469:
4465:
4458:
4455:
4450:
4446:
4441:
4436:
4432:
4428:
4424:
4420:
4416:
4409:
4406:
4401:
4397:
4393:
4391:9780470123119
4387:
4383:
4379:
4375:
4368:
4365:
4360:
4356:
4351:
4346:
4342:
4338:
4334:
4327:
4324:
4319:
4315:
4310:
4305:
4301:
4297:
4293:
4289:
4285:
4281:
4277:
4270:
4267:
4262:
4258:
4254:
4250:
4246:
4242:
4238:
4234:
4227:
4224:
4219:
4215:
4210:
4205:
4200:
4195:
4191:
4187:
4183:
4179:
4175:
4168:
4165:
4160:
4156:
4151:
4146:
4142:
4138:
4134:
4130:
4126:
4122:
4118:
4111:
4108:
4103:
4099:
4094:
4089:
4085:
4081:
4077:
4073:
4072:Cell Research
4069:
4062:
4059:
4054:
4050:
4045:
4040:
4036:
4032:
4028:
4024:
4020:
4013:
4010:
4005:
4001:
3997:
3993:
3989:
3985:
3977:
3974:
3969:
3965:
3961:
3957:
3953:
3949:
3942:
3939:
3934:
3930:
3925:
3920:
3916:
3912:
3908:
3904:
3900:
3896:
3892:
3885:
3882:
3877:
3873:
3868:
3863:
3859:
3855:
3851:
3847:
3843:
3836:
3833:
3828:
3824:
3819:
3814:
3809:
3804:
3800:
3796:
3792:
3788:
3784:
3777:
3774:
3769:
3765:
3760:
3755:
3751:
3747:
3743:
3739:
3735:
3728:
3725:
3720:
3718:9783540244950
3714:
3710:
3706:
3702:
3695:
3692:
3687:
3683:
3678:
3673:
3669:
3665:
3661:
3657:
3653:
3646:
3643:
3638:
3634:
3629:
3624:
3619:
3614:
3610:
3606:
3602:
3595:
3592:
3587:
3583:
3578:
3573:
3568:
3563:
3559:
3555:
3552:(4): e58350.
3551:
3547:
3543:
3536:
3533:
3528:
3524:
3519:
3514:
3510:
3506:
3502:
3498:
3494:
3487:
3484:
3479:
3475:
3470:
3465:
3460:
3455:
3451:
3447:
3443:
3436:
3433:
3428:
3424:
3420:
3416:
3412:
3408:
3401:
3398:
3393:
3389:
3384:
3379:
3374:
3369:
3365:
3361:
3357:
3350:
3347:
3342:
3338:
3333:
3328:
3323:
3318:
3314:
3310:
3306:
3299:
3296:
3291:
3287:
3282:
3277:
3272:
3267:
3263:
3259:
3255:
3248:
3245:
3240:
3236:
3231:
3226:
3222:
3218:
3214:
3207:
3204:
3199:
3195:
3190:
3185:
3181:
3177:
3173:
3169:
3165:
3158:
3155:
3150:
3146:
3142:
3138:
3130:
3127:
3122:
3118:
3113:
3108:
3103:
3098:
3094:
3090:
3086:
3079:
3076:
3071:
3067:
3062:
3057:
3053:
3049:
3045:
3038:
3035:
3030:
3026:
3021:
3016:
3012:
3008:
3004:
2997:
2994:
2989:
2985:
2980:
2975:
2971:
2967:
2963:
2959:
2955:
2948:
2945:
2940:
2936:
2931:
2926:
2922:
2918:
2914:
2910:
2906:
2902:
2898:
2891:
2888:
2883:
2879:
2874:
2869:
2865:
2861:
2857:
2853:
2849:
2842:
2839:
2834:
2830:
2825:
2820:
2816:
2812:
2808:
2804:
2800:
2793:
2790:
2785:
2781:
2776:
2771:
2767:
2763:
2759:
2755:
2751:
2744:
2742:
2740:
2738:
2734:
2729:
2725:
2720:
2715:
2711:
2707:
2703:
2699:
2695:
2688:
2686:
2682:
2677:
2673:
2668:
2663:
2658:
2653:
2649:
2645:
2641:
2634:
2632:
2628:
2623:
2619:
2614:
2609:
2605:
2601:
2597:
2593:
2589:
2585:
2581:
2574:
2571:
2566:
2562:
2557:
2552:
2548:
2544:
2540:
2536:
2532:
2528:
2524:
2517:
2514:
2509:
2505:
2500:
2495:
2491:
2487:
2483:
2479:
2475:
2468:
2466:
2464:
2460:
2455:
2451:
2446:
2441:
2437:
2433:
2429:
2422:
2419:
2414:
2410:
2406:
2402:
2398:
2394:
2387:
2384:
2379:
2375:
2370:
2365:
2361:
2357:
2353:
2349:
2345:
2338:
2335:
2330:
2326:
2321:
2316:
2312:
2308:
2304:
2300:
2296:
2292:
2288:
2281:
2278:
2273:
2269:
2264:
2259:
2255:
2251:
2247:
2243:
2239:
2232:
2229:
2224:
2220:
2215:
2210:
2206:
2202:
2198:
2194:
2190:
2183:
2180:
2175:
2171:
2167:
2163:
2159:
2155:
2148:
2145:
2140:
2136:
2131:
2126:
2122:
2118:
2114:
2110:
2106:
2099:
2096:
2091:
2087:
2082:
2077:
2073:
2069:
2065:
2061:
2057:
2050:
2047:
2042:
2038:
2033:
2028:
2024:
2020:
2016:
2012:
2011:Cell Research
2008:
2001:
1998:
1993:
1989:
1984:
1979:
1975:
1971:
1967:
1963:
1959:
1952:
1949:
1944:
1940:
1935:
1930:
1926:
1922:
1918:
1914:
1910:
1903:
1901:
1897:
1892:
1888:
1883:
1878:
1873:
1868:
1864:
1860:
1856:
1852:
1848:
1841:
1838:
1833:
1829:
1824:
1819:
1815:
1811:
1807:
1803:
1799:
1792:
1789:
1784:
1780:
1775:
1770:
1766:
1762:
1758:
1754:
1750:
1743:
1740:
1735:
1731:
1726:
1721:
1717:
1713:
1709:
1702:
1699:
1694:
1690:
1686:
1682:
1678:
1674:
1670:
1666:
1662:
1658:
1651:
1649:
1647:
1645:
1643:
1639:
1634:
1630:
1625:
1620:
1616:
1612:
1608:
1604:
1600:
1593:
1591:
1589:
1587:
1585:
1583:
1579:
1574:
1570:
1565:
1560:
1556:
1552:
1548:
1544:
1540:
1533:
1531:
1529:
1525:
1520:
1516:
1511:
1506:
1502:
1498:
1494:
1490:
1486:
1479:
1477:
1473:
1468:
1464:
1459:
1454:
1450:
1446:
1442:
1438:
1434:
1427:
1424:
1419:
1415:
1411:
1407:
1403:
1399:
1391:
1388:
1383:
1379:
1375:
1371:
1364:
1361:
1356:
1352:
1348:
1344:
1340:
1336:
1329:
1326:
1321:
1317:
1312:
1307:
1303:
1299:
1295:
1288:
1285:
1280:
1276:
1272:
1268:
1264:
1260:
1253:
1250:
1245:
1241:
1236:
1231:
1227:
1223:
1219:
1215:
1211:
1204:
1201:
1196:
1192:
1187:
1182:
1178:
1174:
1170:
1166:
1162:
1155:
1152:
1147:
1143:
1138:
1133:
1128:
1123:
1119:
1115:
1111:
1104:
1101:
1096:
1092:
1088:
1084:
1080:
1076:
1069:
1067:
1063:
1058:
1054:
1049:
1044:
1040:
1036:
1032:
1028:
1024:
1017:
1015:
1011:
1006:
1002:
997:
992:
987:
982:
978:
974:
970:
966:
962:
955:
953:
949:
944:
940:
936:
932:
927:
922:
918:
914:
910:
906:
902:
895:
893:
889:
883:
881:
879:
875:
871:
867:
863:
855:
853:
851:
847:
843:
842:endonucleases
838:
836:
832:
827:
825:
821:
817:
813:
805:
803:
801:
795:
791:
789:
785:
782:mutations in
780:
779:pre-adipocyte
776:
772:
768:
764:
760:
759:kidney cancer
756:
752:
751:breast cancer
748:
744:
735:
733:
731:
726:
722:
718:
714:
710:
706:
701:
699:
694:
692:
688:
684:
680:
676:
671:
666:
664:
659:
656:
652:
648:
644:
640:
636:
635:transcriptome
632:
628:
624:
616:
614:
612:
608:
604:
603:
597:
595:
587:
585:
583:
579:
575:
571:
567:
563:
555:
550:
548:
546:
541:
538:
534:
530:
526:
522:
518:
514:
510:
506:
501:
499:
495:
491:
487:
483:
479:
475:
471:
467:
463:
459:
455:
449:
444:
440:
436:
432:
427:
425:
424:
419:
415:
411:
407:
403:
399:
395:
393:
380:
374:
368:
361:
359:
356:
355:
328:
325:
321:
320:
315:
306:
302:
295:
281:
277:
270:
262:
258:
257:DTXSID6020858
254:
253:
251:
241:
237:
236:
229:
225:
224:
222:
220:
217:
216:
209:
205:
204:
202:
196:
192:
191:
184:
180:
179:
177:
175:
172:
171:
164:
160:
159:
157:
155:
152:
151:
144:
140:
139:
137:
133:
128:
127:
120:
116:
115:
113:
110:
106:
105:
100:
92:
86:
82:
78:
74:
68:
64:
59:
55:
51:
46:
42:
37:
33:
19:
5292:Deoxyinosine
5282:Deoxyuridine
5236:
5235:
5125:Hypoxanthine
5095:constituents
5093:Nucleic acid
5018:
5014:
5004:
4964:(1): 20–44.
4961:
4957:
4947:
4902:
4898:
4887:
4876:. Retrieved
4846:
4842:
4832:
4789:
4786:Open Biology
4785:
4775:
4765:, retrieved
4725:
4715:
4672:
4668:
4620:
4616:
4606:
4571:
4567:
4557:
4525:(4): 16011.
4522:
4518:
4508:
4471:
4467:
4457:
4422:
4418:
4408:
4373:
4367:
4340:
4337:Cell Reports
4336:
4326:
4283:
4279:
4269:
4236:
4232:
4226:
4181:
4177:
4167:
4124:
4120:
4110:
4075:
4071:
4061:
4026:
4022:
4012:
3987:
3983:
3976:
3951:
3947:
3941:
3898:
3894:
3884:
3852:(1): 51–61.
3849:
3845:
3835:
3790:
3786:
3776:
3741:
3737:
3727:
3700:
3694:
3659:
3655:
3645:
3608:
3604:
3594:
3549:
3545:
3535:
3500:
3496:
3486:
3449:
3445:
3435:
3410:
3406:
3400:
3363:
3359:
3349:
3312:
3308:
3298:
3281:10261/162976
3261:
3257:
3247:
3220:
3216:
3206:
3171:
3167:
3157:
3140:
3136:
3129:
3092:
3088:
3078:
3051:
3047:
3037:
3010:
3006:
2996:
2964:(1): 48–55.
2961:
2957:
2947:
2904:
2900:
2890:
2855:
2851:
2841:
2806:
2802:
2792:
2757:
2753:
2701:
2697:
2647:
2643:
2587:
2583:
2573:
2530:
2526:
2516:
2481:
2477:
2435:
2431:
2421:
2396:
2392:
2386:
2351:
2347:
2337:
2294:
2290:
2280:
2248:(1): 18–29.
2245:
2241:
2231:
2196:
2192:
2182:
2157:
2153:
2147:
2112:
2108:
2098:
2063:
2060:Cell Reports
2059:
2049:
2014:
2010:
2000:
1965:
1961:
1951:
1919:(2): 93–95.
1916:
1912:
1854:
1850:
1840:
1805:
1801:
1791:
1756:
1752:
1742:
1715:
1711:
1701:
1660:
1656:
1606:
1602:
1546:
1542:
1492:
1488:
1440:
1436:
1426:
1401:
1397:
1390:
1373:
1369:
1363:
1338:
1334:
1328:
1301:
1297:
1287:
1262:
1259:Biochemistry
1258:
1252:
1217:
1213:
1203:
1168:
1164:
1154:
1117:
1113:
1103:
1078:
1074:
1033:(1): 52–60.
1030:
1026:
968:
964:
908:
904:
859:
839:
828:
809:
796:
792:
774:
763:mesothelioma
739:
716:
708:
702:
695:
679:RNA splicing
667:
660:
620:
600:
598:
594:poly(A) tail
591:
561:
559:
542:
502:
465:
457:
447:
428:
421:
397:
391:
390:
389:
102:Identifiers
94:Other names
84:
80:
76:
72:
57:
31:
5657:Nucleosides
1120:: 3256524.
806:In bacteria
730:methylation
670:mouse brain
651:stop codons
649:and around
582:transcripts
578:methylation
454:-methionine
431:methylation
317:Properties
163:CHEBI:21891
5651:Categories
5311:Nucleotide
5249:Wybutosine
5244:Xanthosine
5179:Nucleoside
5142:Pyrimidine
5102:Nucleobase
4878:2024-04-07
4767:2024-04-07
3452:: 841451.
3360:BMC Cancer
3095:(1): 211.
884:References
866:Germ Cells
862:Stem Cells
450:-adenosyl-
420:, such as
358:Molar mass
228:CLE6G00625
174:ChemSpider
130:3D model (
109:CAS Number
54:IUPAC name
5277:Thymidine
5201:Guanosine
5196:Adenosine
5035:2059-3635
5021:(1): 74.
4978:0305-1048
4921:1553-7404
4806:2046-2441
4689:1738-8872
784:HNRNPA2B1
771:leukaemia
725:circadian
713:circadian
691:apoptosis
580:, as are
566:homologue
474:prolactin
435:adenosine
119:1867-73-8
5441:c-di-AMP
5436:c-di-GMP
5221:Cytidine
5157:Cytosine
5130:Xanthine
5053:33611339
4996:24068554
4939:26870957
4873:30957852
4824:33715389
4762:27826841
4707:37942519
4698:10940779
4647:27773535
4598:27773536
4549:27572442
4500:27371828
4449:27117054
4359:29298429
4318:23455423
4261:11452560
4253:23817550
4218:26218273
4178:PLOS ONE
4159:25881961
4127:: 6792.
4102:25412662
4053:23867619
4004:23860325
3968:24331679
3933:17434869
3876:24247219
3827:27001847
3768:27117702
3686:21445555
3637:21489227
3586:23593120
3546:PLOS ONE
3527:23193118
3478:20339516
3427:20970678
3392:23835106
3341:22957919
3290:12444081
3239:22454401
3198:23922112
3149:24023349
3121:32376902
3070:24209613
3029:24209618
2988:31844323
2939:35511947
2882:28637691
2833:28637692
2784:26404942
2728:26464443
2676:22639649
2622:36705538
2565:25719671
2508:29476152
2454:26876937
2413:25242552
2378:26046440
2329:24284625
2272:23177736
2223:22002720
2174:21079590
2139:31328227
2090:24981863
2041:24407421
1992:24394384
1943:24316715
1685:22575960
1633:22608085
1573:20421205
1519:12384598
1467:18505803
1244:27773535
1195:28910636
1146:30405719
844:via the
812:bacteria
655:microRNA
458:In vitro
5662:Purines
5231:Inosine
5211:Uridine
5152:Thymine
5120:Guanine
5115:Adenine
5044:7897327
4987:3874165
4930:4752239
4864:6582345
4815:8101014
4753:5291743
4638:5123813
4589:5155635
4540:6053355
4491:4961459
4440:4867121
4400:1315118
4309:3756911
4288:Bibcode
4209:4517749
4186:Bibcode
4150:4410642
4129:Bibcode
4093:4260349
4044:3726147
3924:2646098
3903:Bibcode
3895:Science
3867:4188449
3818:4833258
3795:Bibcode
3759:4860043
3677:7043136
3628:3089782
3577:3620157
3554:Bibcode
3518:3571230
3469:2842904
3383:3716552
3366:: 337.
3332:7659328
3189:3749587
3112:7203018
2979:6974403
2930:9746489
2909:Bibcode
2901:Science
2873:5495124
2824:5495127
2775:4604345
2719:4702777
2667:3355605
2613:9990141
2592:Bibcode
2584:Science
2556:4355918
2535:Bibcode
2499:5826585
2369:4825696
2320:3877715
2299:Bibcode
2263:3646334
2214:3218240
2130:6735865
2081:4142486
2032:3915904
1983:4640932
1934:3911877
1891:6592581
1859:Bibcode
1832:3016525
1783:2216767
1734:9409616
1725:1369564
1693:3517716
1665:Bibcode
1624:3383396
1564:2938207
1458:2438467
1320:1168101
1235:5123813
1186:5615858
1137:6199872
1005:4372599
973:Bibcode
943:4199864
935:1128665
913:Bibcode
831:E. coli
767:sarcoma
698:R-loops
673:the mA
647:3' UTRs
617:Mammals
574:meiosis
478:METTL14
466:in vivo
456:(SAM).
363:281.272
195:PubChem
5147:Uracil
5110:Purine
5051:
5041:
5033:
4994:
4984:
4976:
4937:
4927:
4919:
4871:
4861:
4822:
4812:
4804:
4760:
4750:
4740:
4705:
4695:
4687:
4645:
4635:
4596:
4586:
4547:
4537:
4498:
4488:
4447:
4437:
4398:
4388:
4357:
4316:
4306:
4280:Nature
4259:
4251:
4216:
4206:
4157:
4147:
4100:
4090:
4051:
4041:
4002:
3966:
3931:
3921:
3874:
3864:
3825:
3815:
3766:
3756:
3715:
3684:
3674:
3635:
3625:
3611:: 52.
3584:
3574:
3525:
3515:
3476:
3466:
3425:
3390:
3380:
3339:
3329:
3288:
3237:
3196:
3186:
3147:
3119:
3109:
3068:
3027:
2986:
2976:
2937:
2927:
2880:
2870:
2831:
2821:
2782:
2772:
2726:
2716:
2674:
2664:
2650:: 48.
2620:
2610:
2563:
2553:
2527:Nature
2506:
2496:
2452:
2411:
2376:
2366:
2327:
2317:
2291:Nature
2270:
2260:
2221:
2211:
2172:
2137:
2127:
2088:
2078:
2039:
2029:
1990:
1980:
1941:
1931:
1889:
1882:391771
1879:
1830:
1823:366956
1820:
1781:
1774:332308
1771:
1732:
1722:
1691:
1683:
1657:Nature
1631:
1621:
1571:
1561:
1517:
1510:137137
1507:
1465:
1455:
1418:476526
1416:
1355:418182
1353:
1318:
1279:174715
1277:
1242:
1232:
1193:
1183:
1144:
1134:
1095:196091
1093:
1057:232187
1055:
1048:353526
1045:
1003:
996:434308
993:
941:
933:
905:Nature
822:, and
820:repair
788:ZC3H13
769:, and
717:Mettl3
709:Mettl3
633:, and
607:embryo
588:Plants
570:METTL3
531:, and
521:YTHDC1
517:YTHDF3
513:YTHDF2
509:YTHDF1
490:METTL5
443:METTL3
412:, and
301:SMILES
208:102175
48:Names
5451:cGAMP
5446:cADPR
4792:(3).
4468:eLife
4257:S2CID
1689:S2CID
939:S2CID
874:YTHDF
870:METTL
705:siRNA
643:exons
631:mouse
627:human
556:Yeast
486:VIRMA
276:InChI
183:92307
154:ChEBI
132:JSmol
5632:dXTP
5627:dITP
5622:dCTP
5617:dUTP
5612:dTTP
5607:dGTP
5602:dATP
5569:mUTP
5533:dCDP
5528:dUDP
5523:dTDP
5518:dGDP
5513:dADP
5485:mUDP
5431:cGMP
5426:cAMP
5408:dXMP
5403:dIMP
5398:dCMP
5393:dUMP
5388:dTMP
5383:dGMP
5378:dAMP
5340:mUMP
5049:PMID
5031:ISSN
4992:PMID
4974:ISSN
4935:PMID
4917:ISSN
4869:PMID
4820:PMID
4802:ISSN
4758:PMID
4738:ISBN
4703:PMID
4685:ISSN
4643:PMID
4594:PMID
4545:PMID
4496:PMID
4445:PMID
4396:PMID
4386:ISBN
4355:PMID
4314:PMID
4249:PMID
4214:PMID
4155:PMID
4098:PMID
4049:PMID
4000:PMID
3984:Gene
3964:PMID
3929:PMID
3872:PMID
3823:PMID
3764:PMID
3713:ISBN
3682:PMID
3633:PMID
3582:PMID
3523:PMID
3474:PMID
3450:2010
3423:PMID
3388:PMID
3337:PMID
3286:PMID
3235:PMID
3194:PMID
3145:PMID
3117:PMID
3066:PMID
3048:Cell
3025:PMID
3007:Cell
2984:PMID
2935:PMID
2878:PMID
2829:PMID
2780:PMID
2724:PMID
2672:PMID
2618:PMID
2561:PMID
2504:PMID
2450:PMID
2409:PMID
2374:PMID
2348:Cell
2325:PMID
2268:PMID
2219:PMID
2170:PMID
2135:PMID
2086:PMID
2037:PMID
1988:PMID
1939:PMID
1887:PMID
1828:PMID
1779:PMID
1730:PMID
1681:PMID
1629:PMID
1603:Cell
1569:PMID
1515:PMID
1463:PMID
1414:PMID
1351:PMID
1316:PMID
1298:Cell
1275:PMID
1240:PMID
1191:PMID
1142:PMID
1118:2018
1091:PMID
1053:PMID
1001:PMID
931:PMID
687:TP53
663:CLIP
629:and
519:and
488:and
482:WTAP
429:The
423:Xist
410:rRNA
406:tRNA
402:mRNA
219:UNII
5589:XTP
5584:ITP
5579:CTP
5574:UTP
5564:GTP
5559:ATP
5495:CDP
5490:UDP
5480:GDP
5475:ADP
5360:XMP
5355:IMP
5350:CMP
5345:UMP
5335:GMP
5330:AMP
5039:PMC
5023:doi
4982:PMC
4966:doi
4925:PMC
4907:doi
4859:PMC
4851:doi
4810:PMC
4794:doi
4748:PMC
4730:doi
4693:PMC
4677:doi
4633:PMC
4625:doi
4584:PMC
4576:doi
4535:PMC
4527:doi
4486:PMC
4476:doi
4435:PMC
4427:doi
4378:doi
4345:doi
4304:PMC
4296:doi
4284:495
4241:doi
4204:PMC
4194:doi
4145:PMC
4137:doi
4088:PMC
4080:doi
4039:PMC
4031:doi
4027:123
3992:doi
3988:527
3956:doi
3919:PMC
3911:doi
3899:316
3862:PMC
3854:doi
3813:PMC
3803:doi
3791:113
3754:PMC
3746:doi
3705:doi
3672:PMC
3664:doi
3623:PMC
3613:doi
3572:PMC
3562:doi
3513:PMC
3505:doi
3501:176
3464:PMC
3454:doi
3415:doi
3378:PMC
3368:doi
3327:PMC
3317:doi
3313:103
3276:hdl
3266:doi
3262:278
3225:doi
3184:PMC
3176:doi
3172:109
3107:PMC
3097:doi
3056:doi
3052:155
3015:doi
3011:155
2974:PMC
2966:doi
2925:PMC
2917:doi
2905:376
2868:PMC
2860:doi
2819:PMC
2811:doi
2770:PMC
2762:doi
2714:PMC
2706:doi
2662:PMC
2652:doi
2608:PMC
2600:doi
2588:379
2551:PMC
2543:doi
2531:518
2494:PMC
2486:doi
2440:doi
2401:doi
2364:PMC
2356:doi
2352:161
2315:PMC
2307:doi
2295:505
2258:PMC
2250:doi
2209:PMC
2201:doi
2162:doi
2125:PMC
2117:doi
2076:PMC
2068:doi
2027:PMC
2019:doi
1978:PMC
1970:doi
1929:PMC
1921:doi
1877:PMC
1867:doi
1818:PMC
1810:doi
1769:PMC
1761:doi
1720:PMC
1712:RNA
1673:doi
1661:485
1619:PMC
1611:doi
1607:149
1559:PMC
1551:doi
1505:PMC
1497:doi
1453:PMC
1445:doi
1406:doi
1378:doi
1343:doi
1339:120
1306:doi
1267:doi
1230:PMC
1222:doi
1181:PMC
1173:doi
1132:PMC
1122:doi
1083:doi
1079:113
1043:PMC
1035:doi
991:PMC
981:doi
921:doi
909:255
872:or
775:FTO
683:p53
568:of
494:FTO
484:),
433:of
245:EPA
198:CID
5653::
5047:.
5037:.
5029:.
5017:.
5013:.
4990:.
4980:.
4972:.
4962:42
4960:.
4956:.
4933:.
4923:.
4915:.
4903:12
4901:.
4897:.
4867:.
4857:.
4847:47
4845:.
4841:.
4818:.
4808:.
4800:.
4790:11
4788:.
4784:.
4756:,
4746:,
4736:,
4724:,
4701:.
4691:.
4683:.
4673:34
4671:.
4667:.
4655:^
4641:.
4631:.
4621:20
4619:.
4615:.
4592:.
4582:.
4572:20
4570:.
4566:.
4543:.
4533:.
4521:.
4517:.
4494:.
4484:.
4474:.
4470:.
4466:.
4443:.
4433:.
4423:19
4421:.
4417:.
4394:.
4384:.
4353:.
4341:22
4339:.
4335:.
4312:.
4302:.
4294:.
4282:.
4278:.
4255:.
4247:.
4237:16
4235:.
4212:.
4202:.
4192:.
4182:10
4180:.
4176:.
4153:.
4143:.
4135:.
4123:.
4119:.
4096:.
4086:.
4076:24
4074:.
4070:.
4047:.
4037:.
4025:.
4021:.
3998:.
3986:.
3962:.
3950:.
3927:.
3917:.
3909:.
3897:.
3893:.
3870:.
3860:.
3850:10
3848:.
3844:.
3821:.
3811:.
3801:.
3789:.
3785:.
3762:.
3752:.
3742:62
3740:.
3736:.
3711:.
3680:.
3670:.
3660:22
3658:.
3654:.
3631:.
3621:.
3609:12
3607:.
3603:.
3580:.
3570:.
3560:.
3548:.
3544:.
3521:.
3511:.
3499:.
3495:.
3472:.
3462:.
3448:.
3444:.
3421:.
3411:42
3409:.
3386:.
3376:.
3364:13
3362:.
3358:.
3335:.
3325:.
3311:.
3307:.
3284:.
3274:.
3260:.
3256:.
3233:.
3221:19
3219:.
3215:.
3192:.
3182:.
3170:.
3166:.
3141:33
3139:.
3115:.
3105:.
3091:.
3087:.
3064:.
3050:.
3046:.
3023:.
3009:.
3005:.
2982:.
2972:.
2962:52
2960:.
2956:.
2933:.
2923:.
2915:.
2903:.
2899:.
2876:.
2866:.
2856:31
2854:.
2850:.
2827:.
2817:.
2807:31
2805:.
2801:.
2778:.
2768:.
2758:29
2756:.
2752:.
2736:^
2722:.
2712:.
2702:44
2700:.
2696:.
2684:^
2670:.
2660:.
2646:.
2642:.
2630:^
2616:.
2606:.
2598:.
2586:.
2582:.
2559:.
2549:.
2541:.
2529:.
2525:.
2502:.
2492:.
2482:20
2480:.
2476:.
2462:^
2448:.
2436:61
2434:.
2430:.
2407:.
2397:10
2395:.
2372:.
2362:.
2350:.
2346:.
2323:.
2313:.
2305:.
2293:.
2289:.
2266:.
2256:.
2246:49
2244:.
2240:.
2217:.
2207:.
2195:.
2191:.
2168:.
2156:.
2133:.
2123:.
2113:47
2111:.
2107:.
2084:.
2074:.
2062:.
2058:.
2035:.
2025:.
2015:24
2013:.
2009:.
1986:.
1976:.
1966:16
1964:.
1960:.
1937:.
1927:.
1917:10
1915:.
1911:.
1899:^
1885:.
1875:.
1865:.
1855:81
1853:.
1849:.
1826:.
1816:.
1804:.
1800:.
1777:.
1767:.
1757:18
1755:.
1751:.
1728:.
1714:.
1710:.
1687:.
1679:.
1671:.
1659:.
1641:^
1627:.
1617:.
1605:.
1601:.
1581:^
1567:.
1557:.
1547:38
1545:.
1541:.
1527:^
1513:.
1503:.
1493:30
1491:.
1487:.
1475:^
1461:.
1451:.
1441:20
1439:.
1435:.
1412:.
1402:57
1400:.
1374:15
1372:.
1349:.
1337:.
1314:.
1300:.
1296:.
1273:.
1263:15
1261:.
1238:.
1228:.
1218:20
1216:.
1212:.
1189:.
1179:.
1169:22
1167:.
1163:.
1140:.
1130:.
1116:.
1112:.
1089:.
1077:.
1065:^
1051:.
1041:.
1031:32
1029:.
1025:.
1013:^
999:.
989:.
979:.
969:71
967:.
963:.
951:^
937:.
929:.
919:.
907:.
903:.
891:^
818:,
765:,
761:,
757:,
753:,
749:,
745:,
732:.
693:.
596:.
527:,
515:,
511:,
426:.
408:,
398:mA
339:15
333:11
96:mA
83:,5
79:,4
75:,3
71:(2
5237:N
5085:e
5078:t
5071:v
5055:.
5025::
5019:6
4998:.
4968::
4941:.
4909::
4881:.
4853::
4826:.
4796::
4732::
4709:.
4679::
4649:.
4627::
4600:.
4578::
4551:.
4529::
4523:1
4502:.
4478::
4472:5
4451:.
4429::
4402:.
4380::
4361:.
4347::
4320:.
4298::
4290::
4263:.
4243::
4220:.
4196::
4188::
4161:.
4139::
4131::
4125:6
4104:.
4082::
4055:.
4033::
4006:.
3994::
3970:.
3958::
3952:7
3935:.
3913::
3905::
3878:.
3856::
3829:.
3805::
3797::
3770:.
3748::
3721:.
3707::
3688:.
3666::
3639:.
3615::
3588:.
3564::
3556::
3550:8
3529:.
3507::
3480:.
3456::
3429:.
3417::
3394:.
3370::
3343:.
3319::
3292:.
3278::
3268::
3241:.
3227::
3200:.
3178::
3151:.
3123:.
3099::
3093:3
3072:.
3058::
3031:.
3017::
2990:.
2968::
2941:.
2919::
2911::
2884:.
2862::
2835:.
2813::
2786:.
2764::
2730:.
2708::
2678:.
2654::
2648:3
2624:.
2602::
2594::
2567:.
2545::
2537::
2510:.
2488::
2456:.
2442::
2415:.
2403::
2380:.
2358::
2331:.
2309::
2301::
2274:.
2252::
2225:.
2203::
2197:7
2176:.
2164::
2158:6
2141:.
2119::
2092:.
2070::
2064:8
2043:.
2021::
1994:.
1972::
1945:.
1923::
1893:.
1869::
1861::
1834:.
1812::
1806:5
1785:.
1763::
1736:.
1716:3
1695:.
1675::
1667::
1635:.
1613::
1575:.
1553::
1521:.
1499::
1469:.
1447::
1420:.
1408::
1384:.
1380::
1357:.
1345::
1322:.
1308::
1302:4
1281:.
1269::
1246:.
1224::
1197:.
1175::
1148:.
1124::
1097:.
1085::
1059:.
1037::
1007:.
983::
975::
945:.
923::
915::
533:3
529:2
525:1
452:L
448:S
396:(
392:N
351:4
348:O
345:5
342:N
336:H
330:C
247:)
243:(
134:)
85:R
81:R
77:S
73:R
58:N
32:N
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.