587:
390:
444:
110:
599:
degradation, but in DNA methylation and other epigenetic regulation, through small RNA (smRNA) pathway. AGO10 is involved in plant development. AGO7 has a function distinct from AGO 1 and 10, and is not found in gene silencing induced by transgenes. Instead, it is related to developmental timing in plants.
613:
coordination of cell proliferation and cell death during development and metabolism have been uncovered. It is trusted that the miRNAs can direct negative or positive regulation at different levels, which depends on the specific miRNAs and target base pair interaction and the cofactors that recognize them.
633:
fungus (in which it is known as quelling), plants (post-transcriptional gene silencing) and mammalian cells(RNAi). If there is a complete or near complete sequence complementarity between the small RNA and the target, the
Argonaute protein component of RISC mediates cleavage of the target transcript,
594:
In humans, there are eight AGO family members, some of which are investigated intensively. However, even though AGO1–4 are capable of loading miRNA, endonuclease activity and thus RNAi-dependent gene silencing exclusively belongs to AGO2. Considering the sequence conservation of PAZ and PIWI domains
505:
In animals, Argonaute associated with miRNA binds to the 3′-untranslated region of mRNA and prevents the production of proteins in various ways. The recruitment of
Argonaute proteins to targeted mRNA can induce mRNA degradation. The Argonaute-miRNA complex can also affect the formation of functional
577:
At the interface of PIWI and Mid domains sits the 5′ phosphate of a siRNA, miRNA or piRNA, which is found essential in the functionality. Within Mid lies a MC motif, a homologue structure proposed to mimic the cap-binding structure motif found in eIF4E. It was later found that the MC motif is not
573:
PIWI is named after the
Drosophila Piwi protein. Structurally resembling RNaseH, the PIWI domain is essential for the target cleavage. The active site with aspartate–aspartate–glutamate triad harbors a divalent metal ion, necessary for the catalysis. Family members of AGO that lost this conserved
620:
have RNA rather than DNA as their genetic material and go through at least one stage in their life cycle when they make double-stranded RNA, RNA interference has been considered to be a potentially evolutionarily ancient mechanism for protecting organisms from viruses. The small interfering RNAs
607:
Argonaute proteins were reported to be associated with cancers. For the diseases that are involved with selective or elevated expression of particular identified genes, such as pancreatic cancer, the high sequence specificity of RNA interference might make it suitable to be a suitable treatment,
598:
Several AGO family members in plants also attract study. AGO1 is involved in miRNA related RNA degradation, and plays a central role in morphogenesis. In some organisms, it is strictly required for epigenetic silencing. It is regulated by miRNA itself. AGO4 does not involve in RNAi directed RNA
501:
domain cleaves only the passenger strand of the small interfering RNA. RNA strand separation and incorporation into the
Argonaute protein are guided by the strength of the hydrogen bond interaction at the 5′-ends of the RNA duplex, known as the asymmetry rule. Also the degree of complementarity
612:
gene sequences. It has been reported several tiny non-coding RNAs(microRNAs) are related with human cancers, like miR-15a and miR-16a are frequently deleted and/or down-regulated in patients. Even though the biological functions of miRNAs are not fully understood, the roles for miRNAs in the
478:) molecules into short double stranded fragments of around 20 nucleotide siRNAs. The dsRNA is then separated into two single-stranded RNAs (ssRNA) – the passenger strand and the guide strand. Subsequently, the passenger strand is degraded, while the guide strand is incorporated into the
646:. However, evidence for application of Argonaute proteins as DNA-guided nucleases for genome editing have been questioned, with the retraction of the claim from the leading journal. In 2017, a group from University of Illinois reported using a prokaryotic Argonaute protein taken from
482:(RISC). The most well-studied outcome of the RNAi is post-transcriptional gene silencing, which occurs when the guide strand pairs with a complementary sequence in a messenger RNA molecule and induces cleavage by Argonaute, that lies in the core of RNA-induced silencing complex.
466:, via either destruction of specific mRNA molecules or suppressing translation. RNAi has a significant role in defending cells against parasitic nucleotide sequences . In eukaryotes, including animals, RNAi is initiated by the enzyme
525:
double-stranded (ds) RNA duplexes are generated with the target mRNA, an unknown RNase-III-like enzyme produces new siRNAs, which are then loaded onto the
Argonaute proteins containing PIWI domains, lacking the catalytic
494:. It is known as the guide strand, incorporated into the Argonaute protein and leads gene silencing. The other single-stranded named passenger strand is degraded during the RNA-induced silencing complex process.
489:
produces short double-stranded fragments so there should be also two functional single-stranded siRNA produced. But only one of the two single-stranded RNA here will be utilized to base pair with target
557:
Zwille (also known as pinhead, and later renamed argonaute-10), where the domain was first recognized to be conserved. The PAZ domain is an RNA binding module that recognizes single-stranded 3′ ends of
574:
feature during evolution lack the cleavage activity. In human AGO, the PIWI motif also mediates protein-protein interaction at the PIWI box, where it binds to Dicer at an RNase III domain.
1148:
Qiao D, Zeeman AM, Deng W, Looijenga LH, Lin H (June 2002). "Molecular characterization of hiwi, a human member of the piwi gene family whose overexpression is correlated to seminomas".
357:
228:
485:
Argonaute proteins are the active part of RNA-induced silencing complex, cleaving the target mRNA strand complementary to their bound siRNA. Theoretically the
51:
family, first discovered for its evolutionarily conserved stem cell function, plays a central role in RNA silencing processes as essential components of the
75:(piRNAs). Small RNAs guide Argonaute proteins to their specific targets through sequence complementarity (base pairing), which then leads to mRNA cleavage,
538:
The
Argonaute (AGO) gene family encodes six characteristic domains: N- terminal (N), Linker-1 (L1), PAZ, Linker-2 (L2), Mid, and a C-terminal
595:
across the family, the uniqueness of AGO2 is presumed to arise from either the N-terminus or the spacing region linking PAZ and PIWI motifs.
305:
642:
In 2016, a group from Hebei
University of Science and Technology reported genome editing using a prokaryotic Argonaute protein from
502:
between the two strands of the intermediate RNA duplex defines how the miRNA are sorted into different types of
Argonaute proteins.
660:. PfAgo based artificial restriction enzymes were also used for storing data on native DNA sequences via enzymatic nicking.
377:
248:
1460:
657:
626:
479:
52:
1450:
471:
1465:
514:
assembly. Also, the
Argonaute-miRNA complex can adjust protein production by recruiting cellular factors such as
184:
38:
365:
236:
510:
at the 5′-end of the mRNA. The complex here competes with the translation initiation factors and/or abrogate
772:
Jonas S, Izaurralde E (July 2015). "Towards a molecular understanding of microRNA-mediated gene silencing".
1445:
1414:
1113:
Meins F, Si-Ammour A, Blevins T (2005). "RNA silencing systems and their relevance to plant development".
678:"A novel class of evolutionarily conserved genes defined by piwi are essential for stem cell self-renewal"
318:
586:
475:
68:
31:
361:
232:
82:
The name of this protein family is derived from a mutant phenotype resulting from mutation of AGO1 in
1368:
1253:
934:
877:
84:
72:
189:
648:
422:
402:
116:
1355:
Tabatabaei SK, Wang B, Athreya NB, Enghiad B, Hernandez AG, Fields CJ, et al. (April 2020).
1337:
1173:
1051:
797:
389:
1455:
1394:
1329:
1271:
1222:
1165:
1130:
1095:
1043:
993:
952:
903:
846:
789:
754:
707:
425:
406:
352:
223:
120:
1126:
625:
cause sequence specific, post-transcriptional gene silencing by guiding an endonuclease, the
497:
Once the
Argonaute is associated with the small RNA, the enzymatic activity conferred by the
1384:
1376:
1321:
1294:
1261:
1212:
1204:
1157:
1122:
1085:
1033:
1025:
983:
942:
893:
885:
836:
828:
781:
744:
734:
697:
689:
459:
56:
1016:
Hutvagner G, Simard MJ (January 2008). "Argonaute proteins: key players in RNA silencing".
344:
215:
463:
310:
1372:
1312:
Enghiad B, Zhao H (May 2017). "Programmable DNA-Guided Artificial Restriction Enzymes".
1257:
938:
881:
431:. The base-stacking interaction between the 5′ base on the guide strand and a conserved
1389:
1356:
1217:
1192:
898:
865:
841:
816:
749:
726:
90:
60:
702:
677:
450:
delivery of designed shRNA's and the mechanism of RNA interference in mammalian cells.
1439:
801:
518:
or post translational modifying enzymes, which degrade the growing of polypeptides.
286:
153:
1341:
1177:
1055:
590:
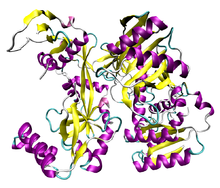
AGO2 (grey) in complex with a microRNA (light blue) and its target mRNA (dark blue)
340:
211:
1072:
Meister G, Landthaler M, Patkaniowska A, Dorsett Y, Teng G, Tuschl T (July 2004).
629:(RISC), to mRNA. This process has been seen in a wide range of organisms, such as
1090:
1073:
947:
922:
314:
298:
165:
1357:"DNA punch cards for storing data on native DNA sequences via enzymatic nicking"
418:
101:
76:
1380:
988:
972:"Human RISC couples microRNA biogenesis and posttranscriptional gene silencing"
971:
815:
Bohmert K, Camus I, Bellini C, Bouchez D, Caboche M, Benning C (January 1998).
443:
88:, which was likened by Bohmert et al. to the appearance of the pelagic octopus
1325:
1298:
1289:
Cyranoski D (2017). "Authors retract controversial NgAgo gene-editing study".
609:
527:
447:
832:
1425:
739:
436:
124:
109:
1398:
1333:
1275:
1226:
1169:
1161:
1134:
1099:
1047:
997:
956:
907:
793:
758:
693:
850:
711:
1429:
1421:
511:
507:
432:
293:
160:
64:
1208:
889:
817:"AGO1 defines a novel locus of Arabidopsis controlling leaf development"
1038:
608:
particularly appropriate for combating cancers associated with mutated
530:
residues, which might induce another level of specific gene silencing.
515:
397:
172:
55:(RISC). RISC is responsible for the gene silencing phenomenon known as
48:
1074:"Human Argonaute2 mediates RNA cleavage targeted by miRNAs and siRNAs"
866:"Mammalian microRNAs predominantly act to decrease target mRNA levels"
435:
residue (light blue) is highlighted; the stabilizing divalent cation (
725:
Mauro M, Berretta M, Palermo G, Cavalieri V, La Rocca G (June 2022).
372:
243:
177:
1266:
1241:
1029:
970:
Gregory RI, Chendrimada TP, Cooch N, Shiekhattar R (November 2005).
785:
17:
622:
617:
585:
567:
563:
559:
486:
467:
442:
388:
676:
Cox DN, Chao A, Baker J, Chang L, Qiao D, Lin H (December 1998).
634:
the mechanism involves repression of translation predominantly.
539:
498:
491:
334:
281:
205:
148:
1417:: a database for exploring microRNA–mRNA interaction maps from
638:
Biotechnological applications of prokaryotic Argonaute proteins
462:(RNAi) is a biological process in which RNA molecules inhibit
727:"The multiplicity of Argonaute complexes in mammalian cells"
428:
409:
864:
Guo H, Ingolia NT, Weissman JS, Bartel DP (August 2010).
128:. PIWI domain is on the right, PAZ domain to the left.
1193:"PIWI proteins and PIWI-interacting RNAs in the soma"
59:. Argonaute proteins bind different classes of small
371:
351:
333:
328:
304:
292:
280:
272:
267:
262:
242:
222:
204:
199:
183:
171:
159:
147:
139:
134:
99:
27:Protein that plays a role in RNA silencing process
470:. Dicer cleaves long double-stranded RNA (dsRNA,
79:inhibition, and/or the initiation of mRNA decay.
1011:
1009:
1007:
1115:Annual Review of Cell and Developmental Biology
8:
1067:
1065:
616:Because it has been widely known that many
1191:Ross RJ, Weiner MM, Lin H (January 2014).
325:
196:
108:
1388:
1265:
1216:
1089:
1037:
987:
946:
897:
840:
748:
738:
701:
652:(PfAgo) along with guide DNA to edit DNA
396:A full-length argonaute protein from the
1127:10.1146/annurev.cellbio.21.122303.114706
421:of an argonaute protein in complex with
668:
1018:Nature Reviews. Molecular Cell Biology
259:
96:
570:, in a sequence independent manner.
7:
25:
534:Functional domains and mechanism
921:Kupferschmidt K (August 2013).
658:artificial restriction enzymes
578:involved in mRNA cap binding
1:
627:RNA-induced silencing complex
603:Disease and therapeutic tools
480:RNA-induced silencing complex
329:Available protein structures:
200:Available protein structures:
53:RNA-induced silencing complex
1091:10.1016/j.molcel.2004.07.007
948:10.1126/science.341.6147.732
545:The PAZ domain is named for
439:) is shown as a gray sphere.
1482:
1381:10.1038/s41467-020-15588-z
989:10.1016/j.cell.2005.10.022
114:An argonaute protein from
36:
30:For the French ships, see
29:
1432:) and Degradome-Seq data.
1326:10.1021/acssynbio.6b00324
1299:10.1038/nature.2017.22412
644:Natronobacterium gregoryi
324:
195:
107:
39:Argonaut (disambiguation)
774:Nature Reviews. Genetics
37:Not to be confused with
1240:Hannon GJ (July 2002).
740:10.1124/jpet.122.001158
682:Genes & Development
57:RNA interference (RNAi)
1162:10.1038/sj.onc.1205505
923:"A lethal dose of RNA"
833:10.1093/emboj/17.1.170
694:10.1101/gad.12.23.3715
591:
472:often found in viruses
451:
440:
69:small interfering RNAs
1361:Nature Communications
1314:ACS Synthetic Biology
589:
476:small interfering RNA
446:
392:
73:Piwi-interacting RNAs
32:French ship Argonaute
1461:RNA-binding proteins
731:J Pharmacol Exp Ther
263:Argonaute Paz domain
85:Arabidopsis thaliana
1373:2020NatCo..11.1742T
1258:2002Natur.418..244H
1209:10.1038/nature12987
939:2013Sci...341..732K
890:10.1038/nature09267
882:2010Natur.466..835G
649:Pyrococcus furiosus
423:double-stranded RNA
403:Pyrococcus furiosus
117:Pyrococcus furiosus
1451:Molecular genetics
1242:"RNA interference"
592:
452:
441:
1415:starBase database
1252:(6894): 244–251.
1203:(7483): 353–359.
1156:(25): 3988–3999.
933:(6147): 732–733.
876:(7308): 835–840.
688:(23): 3715–3727.
553:Argonaute-1, and
387:
386:
383:
382:
378:structure summary
258:
257:
254:
253:
249:structure summary
16:(Redirected from
1473:
1466:RNA interference
1403:
1402:
1392:
1352:
1346:
1345:
1309:
1303:
1302:
1286:
1280:
1279:
1269:
1237:
1231:
1230:
1220:
1188:
1182:
1181:
1145:
1139:
1138:
1110:
1104:
1103:
1093:
1069:
1060:
1059:
1041:
1013:
1002:
1001:
991:
967:
961:
960:
950:
918:
912:
911:
901:
861:
855:
854:
844:
821:The EMBO Journal
812:
806:
805:
769:
763:
762:
752:
742:
722:
716:
715:
705:
673:
521:In plants, once
460:RNA interference
455:RNA interference
326:
260:
197:
127:
112:
97:
21:
1481:
1480:
1476:
1475:
1474:
1472:
1471:
1470:
1436:
1435:
1411:
1406:
1354:
1353:
1349:
1311:
1310:
1306:
1288:
1287:
1283:
1267:10.1038/418244a
1239:
1238:
1234:
1190:
1189:
1185:
1147:
1146:
1142:
1112:
1111:
1107:
1071:
1070:
1063:
1030:10.1038/nrm2321
1015:
1014:
1005:
969:
968:
964:
920:
919:
915:
863:
862:
858:
814:
813:
809:
786:10.1038/nrg3965
771:
770:
766:
724:
723:
719:
675:
674:
670:
666:
640:
605:
584:
536:
464:gene expression
457:
130:
123:
61:non-coding RNAs
42:
35:
28:
23:
22:
15:
12:
11:
5:
1479:
1477:
1469:
1468:
1463:
1458:
1453:
1448:
1438:
1437:
1434:
1433:
1410:
1409:External links
1407:
1405:
1404:
1347:
1320:(5): 752–757.
1304:
1281:
1232:
1183:
1140:
1121:(1): 297–318.
1105:
1084:(2): 185–197.
1078:Molecular Cell
1061:
1003:
982:(4): 631–640.
962:
913:
856:
827:(1): 170–180.
807:
780:(7): 421–433.
764:
717:
667:
665:
662:
639:
636:
604:
601:
583:
582:Family members
580:
535:
532:
456:
453:
385:
384:
381:
380:
375:
369:
368:
355:
349:
348:
338:
331:
330:
322:
321:
308:
302:
301:
296:
290:
289:
284:
278:
277:
274:
270:
269:
265:
264:
256:
255:
252:
251:
246:
240:
239:
226:
220:
219:
209:
202:
201:
193:
192:
187:
181:
180:
175:
169:
168:
163:
157:
156:
151:
145:
144:
141:
137:
136:
132:
131:
113:
105:
104:
91:Argonauta argo
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
1478:
1467:
1464:
1462:
1459:
1457:
1454:
1452:
1449:
1447:
1446:Ribonucleases
1444:
1443:
1441:
1431:
1427:
1423:
1420:
1416:
1413:
1412:
1408:
1400:
1396:
1391:
1386:
1382:
1378:
1374:
1370:
1366:
1362:
1358:
1351:
1348:
1343:
1339:
1335:
1331:
1327:
1323:
1319:
1315:
1308:
1305:
1300:
1296:
1292:
1285:
1282:
1277:
1273:
1268:
1263:
1259:
1255:
1251:
1247:
1243:
1236:
1233:
1228:
1224:
1219:
1214:
1210:
1206:
1202:
1198:
1194:
1187:
1184:
1179:
1175:
1171:
1167:
1163:
1159:
1155:
1151:
1144:
1141:
1136:
1132:
1128:
1124:
1120:
1116:
1109:
1106:
1101:
1097:
1092:
1087:
1083:
1079:
1075:
1068:
1066:
1062:
1057:
1053:
1049:
1045:
1040:
1035:
1031:
1027:
1023:
1019:
1012:
1010:
1008:
1004:
999:
995:
990:
985:
981:
977:
973:
966:
963:
958:
954:
949:
944:
940:
936:
932:
928:
924:
917:
914:
909:
905:
900:
895:
891:
887:
883:
879:
875:
871:
867:
860:
857:
852:
848:
843:
838:
834:
830:
826:
822:
818:
811:
808:
803:
799:
795:
791:
787:
783:
779:
775:
768:
765:
760:
756:
751:
746:
741:
736:
732:
728:
721:
718:
713:
709:
704:
699:
695:
691:
687:
683:
679:
672:
669:
663:
661:
659:
655:
651:
650:
645:
637:
635:
632:
628:
624:
619:
614:
611:
602:
600:
596:
588:
581:
579:
575:
571:
569:
565:
561:
556:
552:
548:
543:
541:
533:
531:
529:
524:
519:
517:
513:
509:
503:
500:
495:
493:
488:
483:
481:
477:
473:
469:
465:
461:
454:
449:
445:
438:
434:
430:
427:
424:
420:
416:
412:
411:
408:
404:
399:
395:
391:
379:
376:
374:
370:
367:
363:
359:
356:
354:
350:
346:
342:
339:
336:
332:
327:
323:
320:
316:
312:
309:
307:
303:
300:
297:
295:
291:
288:
285:
283:
279:
275:
271:
266:
261:
250:
247:
245:
241:
238:
234:
230:
227:
225:
221:
217:
213:
210:
207:
203:
198:
194:
191:
188:
186:
182:
179:
176:
174:
170:
167:
164:
162:
158:
155:
152:
150:
146:
142:
138:
133:
129:
126:
122:
118:
111:
106:
103:
98:
95:
93:
92:
87:
86:
80:
78:
74:
71:(siRNAs) and
70:
66:
62:
58:
54:
50:
47:
40:
33:
19:
1418:
1364:
1360:
1350:
1317:
1313:
1307:
1290:
1284:
1249:
1245:
1235:
1200:
1196:
1186:
1153:
1149:
1143:
1118:
1114:
1108:
1081:
1077:
1024:(1): 22–32.
1021:
1017:
979:
975:
965:
930:
926:
916:
873:
869:
859:
824:
820:
810:
777:
773:
767:
730:
720:
685:
681:
671:
653:
647:
643:
641:
630:
621:produced by
615:
606:
597:
593:
576:
572:
554:
550:
546:
544:
537:
522:
520:
504:
496:
484:
458:
414:
401:
393:
115:
89:
83:
81:
63:, including
45:
43:
1367:(1): 1742.
1039:10453/15429
555:Arabidopsis
551:Arabidopsis
419:PIWI domain
268:Identifiers
135:Identifiers
102:Piwi domain
77:translation
1440:Categories
664:References
631:Neurospora
610:endogenous
547:Drosophila
528:amino acid
448:Lentiviral
341:structures
212:structures
100:Argonaute
67:(miRNAs),
1426:HITS-CLIP
1419:Argonaute
508:ribosomes
437:magnesium
311:b.34.14.1
299:IPR021103
166:IPR003165
65:microRNAs
46:Argonaute
1456:MicroRNA
1430:PAR-CLIP
1422:CLIP-Seq
1399:32269230
1334:28165224
1276:12110901
1227:24429634
1170:12037681
1150:Oncogene
1135:16212497
1100:15260970
1048:18073770
998:16271387
957:23950525
908:20703300
802:24892348
794:26077373
759:35667689
654:in vitro
542:domain.
516:peptides
512:ribosome
433:tyrosine
400:species
358:RCSB PDB
294:InterPro
229:RCSB PDB
161:InterPro
1390:7142088
1369:Bibcode
1342:3833124
1254:Bibcode
1218:4265809
1178:6078065
1056:8822503
935:Bibcode
927:Science
899:2990499
878:Bibcode
851:9427751
842:1170368
750:9827513
712:9851978
618:viruses
523:de novo
398:archaea
287:PF12212
190:cd02826
178:PS50822
173:PROSITE
154:PF02171
49:protein
1397:
1387:
1340:
1332:
1291:Nature
1274:
1246:Nature
1225:
1215:
1197:Nature
1176:
1168:
1133:
1098:
1054:
1046:
996:
955:
906:
896:
870:Nature
849:
839:
800:
792:
757:
747:
710:
703:317255
700:
549:Piwi,
415:Right:
373:PDBsum
347:
337:
319:SUPFAM
273:Symbol
244:PDBsum
218:
208:
140:Symbol
1338:S2CID
1174:S2CID
1052:S2CID
798:S2CID
623:Dicer
568:piRNA
564:miRNA
560:siRNA
487:dicer
468:Dicer
394:Left:
315:SCOPe
306:SCOP2
1395:PMID
1330:PMID
1272:PMID
1223:PMID
1166:PMID
1131:PMID
1096:PMID
1044:PMID
994:PMID
976:Cell
953:PMID
904:PMID
847:PMID
790:PMID
755:PMID
708:PMID
566:and
540:PIWI
499:PIWI
492:mRNA
474:and
429:1YTU
417:The
410:1U04
366:PDBj
362:PDBe
345:ECOD
335:Pfam
282:Pfam
237:PDBj
233:PDBe
216:ECOD
206:Pfam
149:Pfam
143:Piwi
125:1U04
44:The
18:Ago2
1385:PMC
1377:doi
1322:doi
1295:doi
1262:doi
1250:418
1213:PMC
1205:doi
1201:505
1158:doi
1123:doi
1086:doi
1034:hdl
1026:doi
984:doi
980:123
943:doi
931:341
894:PMC
886:doi
874:466
837:PMC
829:doi
782:doi
745:PMC
735:doi
698:PMC
690:doi
656:as
426:PDB
407:PDB
353:PDB
276:Paz
224:PDB
185:CDD
121:PDB
1442::
1428:,
1393:.
1383:.
1375:.
1365:11
1363:.
1359:.
1336:.
1328:.
1316:.
1293:.
1270:.
1260:.
1248:.
1244:.
1221:.
1211:.
1199:.
1195:.
1172:.
1164:.
1154:21
1152:.
1129:.
1119:21
1117:.
1094:.
1082:15
1080:.
1076:.
1064:^
1050:.
1042:.
1032:.
1020:.
1006:^
992:.
978:.
974:.
951:.
941:.
929:.
925:.
902:.
892:.
884:.
872:.
868:.
845:.
835:.
825:17
823:.
819:.
796:.
788:.
778:16
776:.
753:.
743:.
733:.
729:.
706:.
696:.
686:12
684:.
680:.
562:,
413:.
364:;
360:;
343:/
317:/
313:/
235:;
231:;
214:/
119:.
94:.
1424:(
1401:.
1379::
1371::
1344:.
1324::
1318:6
1301:.
1297::
1278:.
1264::
1256::
1229:.
1207::
1180:.
1160::
1137:.
1125::
1102:.
1088::
1058:.
1036::
1028::
1022:9
1000:.
986::
959:.
945::
937::
910:.
888::
880::
853:.
831::
804:.
784::
761:.
737::
714:.
692::
405:.
41:.
34:.
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.