270:, respectively. Many transamination reactions occur in tissues, catalysed by transaminases specific for a particular amino/keto acid pair. The reactions are readily reversible, the direction being determined by which of the reactants are in excess. This reversibility can be exploited for synthetic chemistry applications to achieve the synthesis of valuable chiral amines. The specific enzymes are named from one of the reactant pairs, for example; the reaction between glutamic acid and pyruvic acid to make alpha ketoglutaric acid and alanine is called
40:
1221:
290:
281:
with various amino/keto acid pairs. Transamination is demonstrated if the corresponding new amino acid and keto acid are formed, as revealed by paper chromatography. Reversibility is demonstrated by using the complementary keto/amino acid pair as starting reactants. After chromatogram has been taken
400:(ALT), also called alanine aminotransferase (ALAT) or serum glutamate-pyruvate transaminase (SGPT). These transaminases were discovered in 1954 and their clinical importance was described in 1955.
777:
140:
857:
720:
Ghany M, Hoofnagle JH (2005). "Approach to the
Patient With Liver Disease". In Kasper DL, Fauci AS, Longo DL, Braunwald E, Hauser SL, Jameson JL (eds.).
231:
group on one molecule is exchanged with the =O group on the other molecule. The amino acid becomes a keto acid, and the keto acid becomes an amino acid.
574:
Ladue JS, Wroblewski F, Karmen A (September 1954). "Serum glutamic oxaloacetic transaminase activity in human acute transmural myocardial infarction".
847:
770:
690:
862:
907:
940:
763:
641:
837:
413:
1096:
160:
1211:
852:
1081:
1197:
1184:
1171:
1158:
1145:
1132:
1119:
898:
879:
842:
803:
1091:
1045:
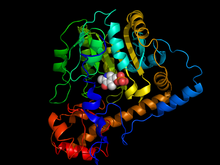
988:
794:
749:
148:
993:
665:
815:
393:
389:
44:
1241:
1014:
933:
1086:
144:
832:
583:
397:
369:
271:
1251:
1050:
380:
The transaminase enzymes are important in the production of various amino acids, and measuring the
331:
255:
235:
96:
983:
888:
1246:
599:
556:
507:
468:
135:
1029:
1024:
998:
926:
591:
546:
538:
527:"A note on the spectrometric assay of glutamic-oxalacetic transaminase in human blood serum"
499:
458:
450:
755:
618:"Biblioteca Nazionale di Napoli. News: Serata in onore di Mario Coltorti e Giuseppe Giusti"
346:
for conversion to urea for excretion of nitrogen. In similar manner, in muscles the use of
242:
in the first half-reaction, when an amino acid is converted into a keto acid. Enzyme-bound
127:
1076:
1060:
973:
361:
298:
745:
360:). Here other transaminases regenerate pyruvate, which provides a valuable precursor for
587:
392:
can be an indicator of liver and cardiac damage. Two important transaminase enzymes are
1225:
1114:
1055:
384:
of various transaminases in the blood is important in the diagnosing and tracking many
224:
188:
39:
551:
526:
463:
438:
354:, which is carried by the bloodstream to the liver (the overall reaction being termed
1235:
1019:
978:
409:
381:
356:
267:
263:
77:
968:
691:"Il Resto Del Carlino - Macerata - E' morto Mario Coltorti: scoprì la transaminasi"
327:
251:
243:
239:
123:
617:
101:
595:
89:
17:
1192:
1127:
963:
786:
319:
278:
1220:
365:
335:
294:
289:
200:
192:
511:
488:"Application of Paper Chromatography to the Study of the Transaminase System"
1166:
1140:
343:
339:
283:
196:
603:
560:
487:
472:
274:
and was originally called glutamic-pyruvic transaminase or GPT for short.
790:
733:(3rd ed.). New York: Worth Publishers. pp. 628–31, 634, 828–30.
396:(AST), also known as serum glutamic oxaloacetic transaminase (SGOT); and
347:
315:
247:
184:
84:
385:
351:
259:
49:
542:
454:
1179:
949:
503:
220:
180:
155:
277:
Tissue transaminase activities can be investigated by incubating a
1153:
323:
288:
212:
867:
825:
820:
117:
72:
922:
759:
918:
282:
out of the solvent the chromatogram is then treated with
724:(16th ed.). New York: McGraw-Hill. pp. 1814–5.
318:
to amino acids, at the expense of muscle tissue, when
1209:
642:"E' morto il prof. Coltorti: scoprì le transaminasi"
1105:
1069:
1038:
1007:
956:
897:
878:
802:
154:
134:
116:
111:
95:
83:
71:
63:
58:
32:
437:Karmen A, Wroblewski F, Ladue JS (January 1955).
486:Tulpule, P. G.; Patwardhan, V. N. (April 1952).
432:
430:
305:) group and the keto (=O) group are exchanged.
934:
771:
8:
334:plays a key role in funneling nitrogen from
941:
927:
919:
858:Branched-chain amino acid aminotransferase
778:
764:
756:
722:Harrison's Principles of Internal Medicine
108:
38:
748:at the U.S. National Library of Medicine
646:notizie-segreteria-liver-pool.blogspot.it
550:
462:
364:. This alanine cycle is analogous to the
199:. They are important in the synthesis of
1216:
426:
439:"Transaminase activity in human blood"
29:
531:The Journal of Clinical Investigation
443:The Journal of Clinical Investigation
7:
848:Kynurenine—oxoglutarate transaminase
731:Lehninger Principles of Biochemistry
53:with Pyridoxal 5' Phosphate cofactor
234:Transaminases require the coenzyme
293:Aminotransfer reaction between an
25:
1219:
908:Pyridoxine 5'-phosphate synthase
310:Amino acid metabolism in animals
219:) group. A keto acid contains a
863:Alanine—glyoxylate transaminase
388:. For example, the presence of
1:
112:Available protein structures:
689:MonrifNet (3 January 2009).
596:10.1126/science.120.3117.497
1268:
853:Ornithine aminotransferase
729:Nelson DL, Cox MM (2000).
322:is low. The preference of
238:, which is converted into
211:An amino acid contains an
1097:Michaelis–Menten kinetics
843:Tyrosine aminotransferase
525:Karmen A (January 1955).
350:for transamination gives
107:
37:
989:Diffusion-limited enzyme
750:Medical Subject Headings
695:www.ilrestodelcarlino.it
670:www.istitutobioetica.org
622:vecchiosito.bnnonline.it
314:Animals must metabolize
203:, which form proteins.
816:Aspartate transaminase
666:"Campania su Coltorti"
394:aspartate transaminase
390:elevated transaminases
306:
207:Function and mechanism
45:Aspartate transaminase
1082:Eadie–Hofstee diagram
1015:Allosteric regulation
882:: Oximinotransferases
357:glucose-alanine cycle
292:
286:to locate the spots..
1092:Lineweaver–Burk plot
833:Alanine transaminase
398:alanine transaminase
370:anaerobic metabolism
272:alanine transaminase
246:in turn reacts with
191:reaction between an
588:1954Sci...120..497L
332:alpha-ketoglutarate
256:alpha-ketoglutarate
236:pyridoxal phosphate
1051:Enzyme superfamily
984:Enzyme promiscuity
889:Oximinotransferase
326:transaminases for
307:
1207:
1206:
916:
915:
838:GABA transaminase
543:10.1172/JCI103055
498:(4303): 671–671.
455:10.1172/jci103055
414:GABA transaminase
177:aminotransferases
170:
169:
166:
165:
161:structure summary
18:Aminotransferases
16:(Redirected from
1259:
1224:
1223:
1215:
1087:Hanes–Woolf plot
1030:Enzyme activator
1025:Enzyme inhibitor
999:Enzyme catalysis
943:
936:
929:
920:
780:
773:
766:
757:
734:
725:
706:
705:
703:
702:
686:
680:
679:
677:
676:
662:
656:
655:
653:
652:
638:
632:
631:
629:
628:
614:
608:
607:
571:
565:
564:
554:
522:
516:
515:
504:10.1038/169671a0
483:
477:
476:
466:
434:
109:
67:Aminotransferase
42:
33:Aminotransferase
30:
27:Class of enzymes
21:
1267:
1266:
1262:
1261:
1260:
1258:
1257:
1256:
1232:
1231:
1230:
1218:
1210:
1208:
1203:
1115:Oxidoreductases
1101:
1077:Enzyme kinetics
1065:
1061:List of enzymes
1034:
1003:
974:Catalytic triad
952:
947:
917:
912:
893:
874:
798:
784:
742:
737:
728:
719:
715:
713:Further reading
710:
709:
700:
698:
688:
687:
683:
674:
672:
664:
663:
659:
650:
648:
640:
639:
635:
626:
624:
616:
615:
611:
582:(3117): 497–9.
573:
572:
568:
524:
523:
519:
485:
484:
480:
436:
435:
428:
423:
406:
378:
376:Diagnostic uses
368:, which allows
362:gluconeogenesis
312:
304:
301:. The amino (NH
299:alpha-keto acid
230:
223:(=O) group. In
218:
209:
54:
28:
23:
22:
15:
12:
11:
5:
1265:
1263:
1255:
1254:
1249:
1244:
1234:
1233:
1229:
1228:
1205:
1204:
1202:
1201:
1188:
1175:
1162:
1149:
1136:
1123:
1109:
1107:
1103:
1102:
1100:
1099:
1094:
1089:
1084:
1079:
1073:
1071:
1067:
1066:
1064:
1063:
1058:
1053:
1048:
1042:
1040:
1039:Classification
1036:
1035:
1033:
1032:
1027:
1022:
1017:
1011:
1009:
1005:
1004:
1002:
1001:
996:
991:
986:
981:
976:
971:
966:
960:
958:
954:
953:
948:
946:
945:
938:
931:
923:
914:
913:
911:
910:
904:
902:
895:
894:
892:
891:
885:
883:
876:
875:
873:
872:
871:
870:
860:
855:
850:
845:
840:
835:
830:
829:
828:
823:
812:
810:
800:
799:
785:
783:
782:
775:
768:
760:
754:
753:
741:
740:External links
738:
736:
735:
726:
716:
714:
711:
708:
707:
681:
657:
633:
609:
566:
517:
478:
425:
424:
422:
419:
418:
417:
405:
402:
382:concentrations
377:
374:
338:metabolism to
311:
308:
302:
228:
225:transamination
216:
208:
205:
189:transamination
168:
167:
164:
163:
158:
152:
151:
138:
132:
131:
121:
114:
113:
105:
104:
99:
93:
92:
87:
81:
80:
75:
69:
68:
65:
61:
60:
56:
55:
43:
35:
34:
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
1264:
1253:
1250:
1248:
1245:
1243:
1240:
1239:
1237:
1227:
1222:
1217:
1213:
1199:
1195:
1194:
1189:
1186:
1182:
1181:
1176:
1173:
1169:
1168:
1163:
1160:
1156:
1155:
1150:
1147:
1143:
1142:
1137:
1134:
1130:
1129:
1124:
1121:
1117:
1116:
1111:
1110:
1108:
1104:
1098:
1095:
1093:
1090:
1088:
1085:
1083:
1080:
1078:
1075:
1074:
1072:
1068:
1062:
1059:
1057:
1056:Enzyme family
1054:
1052:
1049:
1047:
1044:
1043:
1041:
1037:
1031:
1028:
1026:
1023:
1021:
1020:Cooperativity
1018:
1016:
1013:
1012:
1010:
1006:
1000:
997:
995:
992:
990:
987:
985:
982:
980:
979:Oxyanion hole
977:
975:
972:
970:
967:
965:
962:
961:
959:
955:
951:
944:
939:
937:
932:
930:
925:
924:
921:
909:
906:
905:
903:
900:
896:
890:
887:
886:
884:
881:
877:
869:
866:
865:
864:
861:
859:
856:
854:
851:
849:
846:
844:
841:
839:
836:
834:
831:
827:
824:
822:
819:
818:
817:
814:
813:
811:
809:
808:Transaminases
805:
801:
796:
792:
788:
781:
776:
774:
769:
767:
762:
761:
758:
751:
747:
746:Transaminases
744:
743:
739:
732:
727:
723:
718:
717:
712:
696:
692:
685:
682:
671:
667:
661:
658:
647:
643:
637:
634:
623:
619:
613:
610:
605:
601:
597:
593:
589:
585:
581:
577:
570:
567:
562:
558:
553:
548:
544:
540:
536:
532:
528:
521:
518:
513:
509:
505:
501:
497:
493:
489:
482:
479:
474:
470:
465:
460:
456:
452:
449:(1): 126–31.
448:
444:
440:
433:
431:
427:
420:
415:
411:
410:Valproic acid
408:
407:
403:
401:
399:
395:
391:
387:
383:
375:
373:
371:
367:
363:
359:
358:
353:
349:
345:
341:
337:
333:
329:
325:
321:
317:
309:
300:
296:
291:
287:
285:
280:
275:
273:
269:
268:glutamic acid
265:
264:aspartic acid
261:
257:
253:
249:
245:
241:
237:
232:
226:
222:
214:
206:
204:
202:
198:
194:
190:
186:
182:
178:
174:
173:Transaminases
162:
159:
157:
153:
150:
146:
142:
139:
137:
133:
129:
125:
122:
119:
115:
110:
106:
103:
100:
98:
94:
91:
88:
86:
82:
79:
76:
74:
70:
66:
62:
57:
52:
51:
46:
41:
36:
31:
19:
1242:Transferases
1193:Translocases
1190:
1177:
1164:
1151:
1138:
1128:Transferases
1125:
1112:
969:Binding site
807:
730:
721:
699:. Retrieved
697:(in Italian)
694:
684:
673:. Retrieved
669:
660:
649:. Retrieved
645:
636:
625:. Retrieved
621:
612:
579:
575:
569:
537:(1): 131–3.
534:
530:
520:
495:
491:
481:
446:
442:
379:
372:by muscles.
355:
328:oxaloacetate
313:
276:
252:oxaloacetate
244:pyridoxamine
240:pyridoxamine
233:
210:
176:
172:
171:
48:
964:Active site
791:nitrogenous
787:Transferase
320:blood sugar
201:amino acids
59:Identifiers
1252:Hepatology
1236:Categories
1167:Isomerases
1141:Hydrolases
1008:Regulation
701:2017-09-10
675:2017-09-10
651:2017-09-10
627:2017-09-10
421:References
366:Cori cycle
336:amino acid
295:amino acid
279:homogenate
193:amino acid
124:structures
97:Membranome
1046:EC number
512:1476-4687
416:inhibitor
344:glutamate
340:aspartate
284:ninhydrin
258:, giving
197:keto acid
195:and an α-
90:IPR004839
1247:EC 2.6.1
1070:Kinetics
994:Cofactor
957:Activity
793:groups (
604:13195683
561:13221664
473:13221663
404:See also
386:diseases
348:pyruvate
316:proteins
248:pyruvate
227:, the NH
185:catalyze
141:RCSB PDB
85:InterPro
1226:Biology
1180:Ligases
950:Enzymes
901:: Other
584:Bibcode
576:Science
352:alanine
297:and an
260:alanine
181:enzymes
78:PF00155
50:E. coli
1212:Portal
1154:Lyases
899:2.6.99
752:(MeSH)
602:
559:
552:438594
549:
510:
492:Nature
471:
464:438594
461:
156:PDBsum
130:
120:
64:Symbol
1106:Types
880:2.6.3
804:2.6.1
324:liver
266:, or
254:, or
213:amino
183:that
47:from
1198:list
1191:EC7
1185:list
1178:EC6
1172:list
1165:EC5
1159:list
1152:EC4
1146:list
1139:EC3
1133:list
1126:EC2
1120:list
1113:EC1
868:AGXT
826:GOT2
821:GOT1
797:2.6)
600:PMID
557:PMID
508:ISSN
469:PMID
412:- a
342:and
221:keto
179:are
149:PDBj
145:PDBe
128:ECOD
118:Pfam
73:Pfam
592:doi
580:120
547:PMC
539:doi
500:doi
496:169
459:PMC
451:doi
330:or
215:(NH
175:or
136:PDB
102:273
1238::
806::
795:EC
789::
693:.
668:.
644:.
620:.
598:.
590:.
578:.
555:.
545:.
535:34
533:.
529:.
506:.
494:.
490:.
467:.
457:.
447:34
445:.
441:.
429:^
262:,
250:,
187:a
147:;
143:;
126:/
1214::
1200:)
1196:(
1187:)
1183:(
1174:)
1170:(
1161:)
1157:(
1148:)
1144:(
1135:)
1131:(
1122:)
1118:(
942:e
935:t
928:v
779:e
772:t
765:v
704:.
678:.
654:.
630:.
606:.
594::
586::
563:.
541::
514:.
502::
475:.
453::
303:2
229:2
217:2
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.