1423:
1279:
1370:
1362:
1347:
33:
287:
178:
24:
1243:
1432:
1403:
1457:
1269:
1305:
1323:
389:
671:
Adam MP, Feldman J, Mirzaa GM, Pagon RA, Wallace SE, Bean LJ, Gripp KW, Amemiya A, Ah Mew N, Simpson KL, Gropman AL, Lanpher BC, Chapman KA, Summar ML (1993). "Urea Cycle
Disorders Overview". In Adam MP, Feldman J, Mirzaa GM, et al. (eds.).
830:
402:
823:
929:
935:
816:
326:
947:
917:
798:
715:
1199:
911:
32:
301:
448:, NAD dependent protein deacetylases, and ATP to form carbamoyl phosphate. CP then enters the urea cycle in which it reacts with
521:
23:
1080:
409:
95:
893:
244:
1422:
1278:
905:
628:
A defect in the CPS I enzyme, and a subsequent deficiency in the production of carbamoyl phosphate has been linked to
265:
1369:
1361:
1158:
641:
453:
444:. Its enzymatic counterpart, carbamoyl phosphate synthetase I (CPS I), interacts with a class of molecules called
211:
1498:
923:
1508:
1192:
996:
205:
1346:
1383:
1060:
987:
964:
173:
1354:
1148:
1125:
1070:
842:
517:
469:
429:
135:
1413:
1085:
1043:
45:
282:
1513:
1503:
1185:
941:
899:
61:
1242:
887:
870:
1211:
1014:
850:
794:
762:
711:
698:
Bhagavan NV, Ha CE (2015). "Protein and Amino Acid
Metabolism". In Bhagavan NV, Ha CE (eds.).
677:
473:
973:
752:
744:
703:
349:
253:
155:
71:
286:
177:
115:
1431:
1153:
787:
757:
732:
707:
629:
380:
1402:
1492:
1120:
876:
808:
166:
1019:
485:
1115:
733:"SIRT5 Deacetylates carbamoyl phosphate synthetase 1 and regulates the urea cycle"
1075:
509:
501:
233:
1177:
748:
1456:
1261:
1208:
839:
651:
646:
489:
481:
457:
441:
437:
368:
146:
1339:
1297:
1024:
582:
513:
449:
766:
681:
1268:
1463:
1442:
1029:
477:
433:
1304:
1137:
505:
445:
220:
1322:
1104:
126:
379:
Except where otherwise noted, data are given for materials in their
194:
425:
106:
94:
84:
1315:
702:(Second ed.). San Diego: Academic Press. pp. 227–268.
428:
of biochemical significance. In land-dwelling animals, it is an
185:
1181:
812:
270:
731:
Nakagawa T, Lomb DJ, Haigis MC, Guarente L (May 2009).
397:
310:
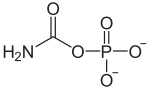
InChI=1S/CH4NO5P/c2-1(3)7-8(4,5)6/h(H2,2,3)(H2,4,5,6)
1134:
1101:
1094:
1053:
1042:
1007:
980:
957:
867:
860:
849:
789:
Lehninger
Principles of Biochemistry fourth edition
786:
232:
70:
936:Phosphoribosylaminoimidazolesuccinocarboxamide
1193:
824:
8:
930:5′-Phosphoribosyl-4-carboxy-5-aminoimidazole
520:). The synthesis is catalyzed by the enzyme
948:5-Formamidoimidazole-4-carboxamide ribotide
1224:
1200:
1186:
1178:
1098:
1050:
864:
857:
831:
817:
809:
285:
176:
154:
15:
756:
252:
1220:
524:. This uses three reactions as follows:
793:. New York: W. H. Freeman and company.
663:
331:
306:
281:
210:
167:
676:. University of Washington, Seattle.
313:Key: FFQKYPRQEYGKAF-UHFFFAOYSA-N
134:
114:
7:
918:5′-Phosphoribosylformylglycinamidine
693:
691:
912:Phosphoribosyl-N-formylglycineamide
452:(a process catalyzed by the enzyme
223:
193:
708:10.1016/b978-0-12-416687-5.00015-4
700:Essentials of Medical Biochemistry
14:
1455:
1430:
1421:
1401:
1368:
1360:
1345:
1321:
1303:
1277:
1267:
1241:
387:
31:
22:
383:(at 25 °C , 100 kPa).
522:carbamoyl phosphate synthetase
1:
49:(Carbamoyloxy)phosphonic acid
894:Phosphoribosyl pyrophosphate
906:Glycineamide ribonucleotide
373:141.020 g/mol
1530:
1477:
1081:Orotidine 5'-monophosphate
785:Nelson DL, Cox MM (2005).
749:10.1016/j.cell.2009.02.026
642:Ornithine transcarbamylase
454:ornithine transcarbamylase
1427:
1398:
1396:
1387:
1366:
1359:
1311:
1275:
1218:
924:5-Aminoimidazole ribotide
468:Carbamoyl phosphate is a
377:
342:
322:
297:
54:
44:
39:
30:
21:
997:Xanthosine monophosphate
1159:β-Ureidoisobutyric acid
1061:Carbamoyl aspartic acid
843:metabolic intermediates
430:intermediary metabolite
1149:3-Aminoisobutyric acid
1126:3-Ureidopropionic acid
1071:4,5-Dihydroorotic acid
470:metabolic intermediate
1086:Uridine monophosphate
624:Clinical significance
480:disposal through the
440:and the synthesis of
436:disposal through the
551:(carboxyl phosphate)
500:It is produced from
17:Carbamoyl phosphate
1236:Carbamoyl phosphate
1066:Carbamoyl phosphate
942:AICA ribonucleotide
900:Phosphoribosylamine
422:Carbamoyl phosphate
212:Carbamoyl+phosphate
18:
888:Ribose 5-phosphate
599:+ ATP → ADP +
410:Infobox references
334:C(=O)(N)OP(=O)(O)O
16:
1486:
1485:
1481:
1480:
1473:
1472:
1440:
1414:argininosuccinate
1411:
1337:
1295:
1259:
1212:Metabolic Pathway
1175:
1174:
1171:
1170:
1167:
1166:
1038:
1037:
1015:5-Hydroxyisourate
1003:
1002:
800:978-0-7167-4339-2
717:978-0-12-416687-5
418:Chemical compound
416:
415:
266:CompTox Dashboard
96:Interactive image
1521:
1499:Organophosphates
1459:
1438:
1434:
1425:
1409:
1405:
1372:
1364:
1357:
1349:
1335:
1325:
1307:
1293:
1281:
1271:
1257:
1245:
1225:
1221:
1202:
1195:
1188:
1179:
1141:
1108:
1099:
1051:
991:
974:Adenylosuccinate
968:
880:
865:
858:
833:
826:
819:
810:
804:
792:
771:
770:
760:
728:
722:
721:
695:
686:
685:
668:
619:
618:
617:
609:
608:
580:
579:
578:
564:
563:
562:
550:
549:
548:
538:
537:
536:
400:
394:
391:
390:
350:Chemical formula
290:
289:
274:
272:
256:
236:
225:
214:
197:
180:
169:
158:
138:
118:
98:
74:
35:
26:
19:
1529:
1528:
1524:
1523:
1522:
1520:
1519:
1518:
1509:Acid anhydrides
1489:
1488:
1487:
1482:
1469:
1468:
1467:
1460:
1447:
1446:
1435:
1418:
1417:
1406:
1391:
1381:
1373:
1352:
1351:
1350:
1343:
1327:
1326:
1319:
1309:
1308:
1301:
1287:
1286:
1282:
1273:
1272:
1265:
1247:
1246:
1239:
1214:
1206:
1176:
1163:
1136:
1130:
1103:
1090:
1045:
1034:
999:
982:
976:
959:
953:
869:
852:
845:
837:
807:
801:
784:
780:
778:Further reading
775:
774:
730:
729:
725:
718:
697:
696:
689:
670:
669:
665:
660:
638:
626:
616:
613:
612:
611:
607:
604:
603:
602:
600:
598:
591:
586:
577:
574:
573:
572:
570:
568:
561:
558:
557:
556:
554:
547:
544:
543:
542:
540:
535:
532:
531:
530:
528:
498:
466:
419:
412:
407:
406:
405: ?)
396:
392:
388:
384:
362:
358:
352:
338:
335:
330:
329:
318:
315:
314:
311:
305:
304:
293:
275:
268:
259:
239:
226:
200:
161:
141:
121:
101:
88:
77:
64:
50:
12:
11:
5:
1527:
1525:
1517:
1516:
1511:
1506:
1501:
1491:
1490:
1484:
1483:
1479:
1478:
1475:
1474:
1471:
1470:
1461:
1454:
1453:
1452:
1449:
1448:
1436:
1429:
1428:
1426:
1419:
1407:
1400:
1399:
1397:
1394:
1393:
1389:
1386:
1379:
1375:
1374:
1367:
1365:
1358:
1344:
1333:
1332:
1329:
1328:
1320:
1313:
1312:
1310:
1302:
1291:
1290:
1288:
1284:
1276:
1274:
1266:
1255:
1254:
1252:
1249:
1248:
1240:
1233:
1232:
1230:
1228:
1219:
1216:
1215:
1207:
1205:
1204:
1197:
1190:
1182:
1173:
1172:
1169:
1168:
1165:
1164:
1162:
1161:
1156:
1154:Dihydrothymine
1151:
1145:
1143:
1132:
1131:
1129:
1128:
1123:
1118:
1112:
1110:
1096:
1092:
1091:
1089:
1088:
1083:
1078:
1073:
1068:
1063:
1057:
1055:
1048:
1040:
1039:
1036:
1035:
1033:
1032:
1027:
1022:
1017:
1011:
1009:
1005:
1004:
1001:
1000:
995:
993:
978:
977:
972:
970:
955:
954:
952:
951:
945:
939:
933:
927:
921:
915:
909:
903:
897:
891:
884:
882:
862:
855:
847:
846:
838:
836:
835:
828:
821:
813:
806:
805:
799:
781:
779:
776:
773:
772:
743:(3): 560–570.
723:
716:
687:
662:
661:
659:
656:
655:
654:
649:
644:
637:
634:
630:hyperammonemia
625:
622:
621:
620:
614:
605:
596:
593:
589:
584:
575:
566:
559:
552:
545:
539:+ ATP → ADP +
533:
508:(derived from
497:
494:
476:that involves
465:
464:Classification
462:
417:
414:
413:
408:
386:
385:
381:standard state
378:
375:
374:
371:
365:
364:
360:
356:
353:
348:
345:
344:
340:
339:
337:
336:
333:
325:
324:
323:
320:
319:
317:
316:
312:
309:
308:
300:
299:
298:
295:
294:
292:
291:
283:DTXSID70862253
278:
276:
264:
261:
260:
258:
257:
249:
247:
241:
240:
238:
237:
229:
227:
219:
216:
215:
208:
202:
201:
199:
198:
190:
188:
182:
181:
171:
163:
162:
160:
159:
151:
149:
143:
142:
140:
139:
131:
129:
123:
122:
120:
119:
111:
109:
103:
102:
100:
99:
91:
89:
82:
79:
78:
76:
75:
67:
65:
60:
57:
56:
52:
51:
48:
42:
41:
37:
36:
28:
27:
13:
10:
9:
6:
4:
3:
2:
1526:
1515:
1512:
1510:
1507:
1505:
1502:
1500:
1497:
1496:
1494:
1476:
1466:
1465:
1458:
1451:
1450:
1445:
1444:
1433:
1424:
1420:
1416:
1415:
1404:
1395:
1385:
1377:
1376:
1371:
1363:
1356:
1348:
1342:
1341:
1331:
1330:
1324:
1318:
1317:
1306:
1300:
1299:
1289:
1280:
1270:
1264:
1263:
1253:
1251:
1250:
1244:
1238:
1237:
1231:
1229:
1227:
1226:
1223:
1222:
1217:
1213:
1210:
1203:
1198:
1196:
1191:
1189:
1184:
1183:
1180:
1160:
1157:
1155:
1152:
1150:
1147:
1146:
1144:
1142:
1139:
1133:
1127:
1124:
1122:
1121:Dihydrouracil
1119:
1117:
1114:
1113:
1111:
1109:
1106:
1100:
1097:
1093:
1087:
1084:
1082:
1079:
1077:
1074:
1072:
1069:
1067:
1064:
1062:
1059:
1058:
1056:
1052:
1049:
1047:
1041:
1031:
1028:
1026:
1023:
1021:
1018:
1016:
1013:
1012:
1010:
1006:
998:
994:
992:
989:
986:
979:
975:
971:
969:
966:
963:
956:
949:
946:
943:
940:
937:
934:
931:
928:
925:
922:
919:
916:
913:
910:
907:
904:
901:
898:
895:
892:
889:
886:
885:
883:
881:
878:
875:
872:
866:
863:
859:
856:
854:
848:
844:
841:
834:
829:
827:
822:
820:
815:
814:
811:
802:
796:
791:
790:
783:
782:
777:
768:
764:
759:
754:
750:
746:
742:
738:
734:
727:
724:
719:
713:
709:
705:
701:
694:
692:
688:
683:
679:
675:
667:
664:
657:
653:
650:
648:
645:
643:
640:
639:
635:
633:
631:
623:
594:
587:
553:
527:
526:
525:
523:
519:
515:
511:
507:
503:
495:
493:
491:
487:
483:
479:
475:
471:
463:
461:
459:
455:
451:
447:
443:
439:
435:
431:
427:
423:
411:
404:
399:
382:
376:
372:
370:
367:
366:
354:
351:
347:
346:
341:
332:
328:
321:
307:
303:
296:
288:
284:
280:
279:
277:
267:
263:
262:
255:
251:
250:
248:
246:
243:
242:
235:
231:
230:
228:
222:
218:
217:
213:
209:
207:
204:
203:
196:
192:
191:
189:
187:
184:
183:
179:
175:
172:
170:
168:ECHA InfoCard
165:
164:
157:
153:
152:
150:
148:
145:
144:
137:
133:
132:
130:
128:
125:
124:
117:
113:
112:
110:
108:
105:
104:
97:
93:
92:
90:
86:
81:
80:
73:
69:
68:
66:
63:
59:
58:
53:
47:
43:
38:
34:
29:
25:
20:
1462:
1437:
1408:
1334:
1314:
1292:
1256:
1235:
1234:
1135:
1102:
1065:
1020:Hypoxanthine
984:
981:
961:
958:
873:
868:
788:
740:
736:
726:
699:
674:Gene Reviews
673:
666:
627:
499:
486:biosynthesis
467:
421:
420:
136:ChEMBL369105
55:Identifiers
1076:Orotic acid
632:in humans.
555:HO–C(O)–OPO
541:HO–C(O)–OPO
510:amino acids
502:bicarbonate
490:pyrimidines
442:pyrimidines
343:Properties
174:100.230.975
116:CHEBI:17672
1514:Urea cycle
1504:Carbamates
1493:Categories
1262:citrulline
1209:Urea cycle
1095:catabolism
1046:metabolism
1044:pyrimidine
1008:catabolism
853:metabolism
840:Nucleotide
658:References
652:Urea Cycle
647:Citrulline
569:+ OH →
496:Production
482:urea cycle
458:citrulline
456:) to form
438:urea cycle
369:Molar mass
254:GC5KZW01Q3
147:ChemSpider
83:3D model (
62:CAS Number
46:IUPAC name
1340:aspartate
1298:ornithine
1116:β-Alanine
1054:anabolism
1025:Uric acid
861:anabolism
514:phosphate
450:ornithine
1464:Fumarate
1443:arginine
1030:Xanthine
950:(FAICAR)
938:(SAICAR)
767:19410549
682:20301396
636:See also
610:NC(O)OPO
595:O–C(O)NH
583:O–C(O)NH
484:and the
478:nitrogen
446:sirtuins
434:nitrogen
363:P
72:590-55-6
1138:thymine
944:(AICAR)
758:2698666
512:), and
506:ammonia
474:pathway
403:what is
401: (
221:PubChem
1105:uracil
932:(CAIR)
920:(FGAM)
914:(FGAR)
896:(PRPP)
851:purine
797:
765:
755:
714:
680:
516:(from
424:is an
398:verify
395:
327:SMILES
195:C00169
127:ChEMBL
40:Names
926:(AIR)
908:(GAR)
902:(PRA)
890:(R5P)
565:+ NH
472:in a
426:anion
302:InChI
107:ChEBI
85:JSmol
1316:Urea
795:ISBN
763:PMID
737:Cell
712:ISBN
678:PMID
588:+ H
245:UNII
206:MeSH
186:KEGG
1384:AMP
1355:ATP
988:GMP
983:IMP
965:AMP
960:IMP
877:IMP
871:R5P
753:PMC
745:doi
741:137
704:doi
581:+
571:HPO
529:HCO
518:ATP
488:of
432:in
271:EPA
234:278
224:CID
156:272
1495::
1392:O
1382:+
1378:PP
1353:+
761:.
751:.
739:.
735:.
710:.
690:^
504:,
492:.
460:.
359:NO
355:CH
1441:-
1439:L
1412:-
1410:L
1390:2
1388:H
1380:i
1338:-
1336:L
1296:-
1294:L
1285:i
1283:P
1260:-
1258:L
1201:e
1194:t
1187:v
1140::
1107::
990::
985:→
967::
962:→
879::
874:→
832:e
825:t
818:v
803:.
769:.
747::
720:.
706::
684:.
615:3
606:2
601:H
597:2
592:O
590:2
585:2
576:4
567:3
560:3
546:3
534:3
393:N
361:5
357:2
273:)
269:(
87:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.