542:
538:", blocking the replicase start. The start of the maturation protein gene is accessible in RNA being replicated but hidden within RNA secondary structure in the completed MS2 RNA; this ensures translation of only a very few copies of maturation protein per RNA. Finally, the lysis protein gene can only be initiated by ribosomes that have completed translation of the coat protein gene and "slip back" to the start of the lysis protein gene, at about a 5% frequency.
70:
253:
452:
44:
635:
601:
and his team, building upon their earlier milestone in 1972 of the first gene to be completely sequenced, the MS2 coat protein. These sequences were determined at the RNA level. The first effort at a statistical analysis of the MS2 genome was a search for patterns in the nucleotide sequence. Several
234:
The virus was isolated in 1961 and its genome was the first to be fully sequenced, in 1976, providing a crucial understanding of genetic codes. In practical applications, MS2's structural components have been used to detect RNA in living cells. The virus is also under research for potential uses in
230:
The MS2 lifecycle involves infecting bacteria with the fertility factor, enabling the virus to attach to the pilus, though the mechanism by which the virus's RNA enters the bacterium remains unknown. Once inside, the viral RNA starts functioning as a messenger RNA to produce viral proteins. MS2
770:
Fiers W, Contreras R, Duerinck F, Haegeman G, Iserentant D, Merregaert J, Min Jou W, Molemans F, Raeymaekers A, Van den Berghe A, Volckaert G, Ysebaert M (April 1976). "Complete nucleotide sequence of bacteriophage MS2 RNA: primary and secondary structure of the replicase gene".
549:
Replication of the plus-strand MS2 genome requires synthesis of the complementary minus strand RNA, which can then be used as a template for synthesis of a new plus strand RNA. MS2 replication has been much less well studied than replication of the highly related
529:
for the production of phage proteins. The gene for the most abundant protein, the coat protein, can be immediately translated. The translation start of the replicase gene is normally hidden within RNA secondary structure, but can be transiently opened as
562:
from coat proteins can occur in the absence of RNA; however, capsid assembly is nucleated by coat protein dimer binding to the operator hairpin, and assembly occurs at much lower concentrations of coat protein when MS2 RNA is present.
467:(viral particle) is about 27 nm in diameter, as determined by electron microscopy. It consists of one copy of the maturation protein and 180 copies of the coat protein (organized as 90 dimers) arranged into an
557:
The formation of the virion is thought to be initiated by binding of maturation protein to the MS2 RNA; in fact, the complex of maturation protein and RNA is infectious. The assembly of the icosahedral shell or
566:
Bacterial lysis and release of newly formed virions occurs when sufficient lysis protein has accumulated. Lysis (L) protein forms pores in the cytoplasmic membrane, which leads to loss of
534:
pass through the coat protein gene. Replicase translation is also shut down once large amounts of coat protein have been made; coat protein dimers bind and stabilize the RNA "operator
1197:
Sanger F, Air GM, Barrell BG, Brown NL, Coulson AR, Fiddes CA, Hutchison CA, Slocombe PM, Smith M (February 1977). "Nucleotide sequence of bacteriophage phi X174 DNA".
1444:
522:, which includes the F-pilin protein that serves as the viral receptor. MS2 attaches to the F-pilin on the side of the pilus using its single maturation protein.
623:, its similar optimum proliferation conditions, and non-pathogenicity to humans, has been used as substitute for noroviruses in studies of disease transmission.
1103:"National Academy of Sciences: Abstracts of Papers Presented at the Autumn Meeting, 29 October, La Jolla, California, 30 October-1 November 1961, Los Angeles".
231:
replicates its plus-strand genome by creating a minus strand RNA as a template. The virus then assembles, and the bacterial cell lyses, releasing new viruses.
239:, its preferred proliferation conditions, and its lack of pathogenicity to humans, MS2 serves as a substitute in studies of norovirus disease transmission.
439:, and is translated upon viral uncoating within the host cell. Although the four proteins are encoded by the same messenger/viral RNA, they are not all
227:
It is small and contains a maturation protein, coat protein, and genomic RNA. It also has one of the smallest known genomes, encoding four proteins.
914:
602:
non-coding sequences were identified, however at the time of this investigation (1979), the functions of the non-coding patterns were unknown.
395:
is one of the smallest known, consisting of 3569 nucleotides of single-stranded RNA. It encodes just four proteins: the maturation protein (
1146:
Min Jou W, Haegeman G, Ysebaert M, Fiers W (May 1972). "Nucleotide sequence of the gene coding for the bacteriophage MS2 coat protein".
702:
968:
Golmohammadi R, Valegård K, Fridborg K, Liljas L (December 1993). "The refined structure of bacteriophage MS2 at 2.8 A resolution".
582:
and appears to be essential to the lysis activity, although their different locations suggest that they have evolved independently.
910:"Delineating the Specific Influence of Virus Isoelectric Point and Size on Virus Adsorption and Transport Through Sandy Soils"
361:
336:
311:
281:
822:
Strauss JH, Sinsheimer RL (July 1963). "Purification and properties of bacteriophage MS2 and of its ribonucleic acid".
605:
Since 1998, the MS2 operator hairpin and coat protein have found utility in the detection of RNA in living cells (see
507:
857:
Valegård K, Liljas L, Fridborg K, Unge T (May 1990). "The three-dimensional structure of the bacterial virus MS2".
69:
200:
498:
is assembled, the helices and hairpin face the exterior of the particle, while the β-sheet faces the interior.
1513:
1248:
Erickson JW, Altman GG (April 1979). "A search for patterns in the nucleotide sequence of the MS2 genome".
235:
drug delivery, tumor imaging, and light harvesting. Furthermore, because of its structural similarities to
1412:
1518:
590:
In 1961, MS2 was isolated by Alvin John Clark and recognized as an RNA-containing phage very similar to
515:
1368:
1206:
1155:
1112:
922:
866:
780:
64:
1324:
Glasgow J, Tullman-Ercek D (July 2014). "Production and applications of engineered viral capsids".
658:
597:
In 1976, the MS2 genome was the first genome to be completely sequenced. This was accomplished by
578:
via an important P330 residue. A LS dipeptide motif on the L protein is found throughout the genus
551:
221:
609:). MS2 and other viral capsids are also currently under investigation as agents in drug delivery,
541:
1349:
1265:
1230:
1179:
890:
804:
720:"Crystal structure of the coat protein from the GA bacteriophage: model of the unassembled dimer"
567:
213:
392:
1449:
718:
Ni CZ, White CA, Mitchell RS, Wickersham J, Kodandapani R, Peabody DS, Ely KR (December 1996).
554:, partly because the MS2 replicase has been difficult to isolate, but is likely to be similar.
1485:
1341:
1306:
1222:
1171:
1128:
1085:
1034:
985:
950:
882:
839:
796:
749:
698:
693:
van Duin J, Tsareva N (2006). "Single-stranded RNA phages. Chapter 15". In
Calendar RL (ed.).
476:
1333:
1296:
1257:
1214:
1163:
1120:
1075:
1065:
1024:
1016:
977:
940:
930:
874:
831:
788:
739:
731:
653:
591:
432:
217:
208:
613:
440:
1210:
1159:
1116:
926:
870:
784:
1080:
1053:
1029:
1004:
744:
719:
663:
640:
1301:
1284:
945:
909:
835:
1507:
648:
526:
436:
130:
118:
106:
1269:
1490:
1476:
1353:
1283:
Bertrand E, Chartrand P, Schaefer M, Shenoy SM, Singer RH, Long RM (October 1998).
1234:
1183:
894:
808:
598:
491:
154:
142:
52:
1400:
1396:
1392:
1388:
1124:
935:
451:
252:
216:. MS2 is a member of a family of closely related bacterial viruses that includes
17:
606:
487:
468:
236:
1435:
379:
374:
349:
324:
298:
294:
1337:
630:
483:
272:
56:
620:
571:
535:
412:
166:
94:
1345:
1132:
1089:
1038:
981:
843:
735:
1310:
1175:
989:
954:
886:
800:
753:
1470:
1429:
1226:
1070:
531:
1020:
1261:
511:
634:
43:
1218:
1167:
878:
792:
559:
495:
472:
464:
1406:
525:
Once the viral RNA has entered the cell, it begins to function as a
59:
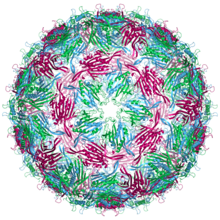
are labelled blue (chain a), green (chain b) and magenta (chain c)
610:
540:
519:
450:
400:
251:
81:
575:
1410:
1005:"MS2 Lysis of Escherichia coli Depends on Host Chaperone DnaJ"
697:(Second ed.). Oxford University Press. pp. 175–196.
203:
518:. Genes on the F plasmid specifies the proteins of the F
1369:"Viruses spread 'like crazy' in an office, study finds"
475:, protecting the genomic RNA inside. The virion has an
1285:"Localization of ASH1 mRNA particles in living yeast"
482:
The structure of the coat protein is a five-stranded
1460:
1419:
908:Dowd SE, Pillai SD, Wang S, Corapcioglu MY (1998).
1054:"Mutational analysis of the MS2 lysis protein L"
1052:Chamakura KR, Edwards GB, Young R (July 2017).
435:. The positive-stranded RNA genome serves as a
423:overlaps both the 3'-end of the upstream gene (
431:), and was one of the first known examples of
8:
1003:Chamakura KR, Tran JS, Young R (June 2017).
514:that allows cells to serve as DNA donors in
619:MS2, due to its structural similarities to
199:), commonly called MS2, is an icosahedral,
1407:
688:
686:
684:
682:
680:
678:
506:MS2 infects enteric bacteria carrying the
259:
42:
31:
1300:
1079:
1069:
1028:
944:
934:
743:
427:) and the 5'-end of the downstream gene (
574:. The lysis protein is known to bind to
765:
763:
674:
1326:Applied Microbiology and Biotechnology
616:, and light harvesting applications.
7:
459:virion (cross section and side view)
25:
586:MS2 in History of Science and Use
206:virus that infects the bacterium
1111:(3488): 1425–37. November 1961.
633:
471:shell with triangulation number
68:
1250:Journal of Mathematical Biology
419:) protein. The gene encoding
201:positive-sense single-stranded
1:
1302:10.1016/S1097-2765(00)80143-4
1125:10.1126/science.134.3488.1425
936:10.1128/aem.64.2.405-410.1998
836:10.1016/S0022-2836(63)80017-0
407:) protein, the coat protein (
55:. The three quasi-equivalent
970:Journal of Molecular Biology
824:Journal of Molecular Biology
545:Bacteriophage MS2 life cycle
1535:
1367:Fox M (8 September 2014).
1338:10.1007/s00253-014-5787-3
915:Appl. Environ. Microbiol.
256:Bacteriophage MS2 genome
212:and other members of the
63:
50:
41:
34:
1421:Enterobacteria phage MS2
1009:Journal of Bacteriology
455:Schematic drawing of a
982:10.1006/jmbi.1993.1616
736:10.1002/pro.5560051211
546:
460:
257:
570:and breakdown of the
544:
516:bacterial conjugation
454:
255:
1071:10.1099/mic.0.000485
508:fertility (F) factor
443:at the same levels.
65:Virus classification
1211:1977Natur.265..687S
1160:1972Natur.237...82J
1117:1961Sci...134.1425.
1021:10.1128/JB.00058-17
927:1998ApEnM..64..405D
871:1990Natur.345...36V
785:1976Natur.260..500F
1262:10.1007/BF00275725
695:The Bacteriophages
568:membrane potential
547:
461:
258:
214:Enterobacteriaceae
51:Bacteriophage MS2
1501:
1500:
1462:Emesvirus zinderi
1413:Taxon identifiers
477:isoelectric point
433:overlapping genes
389:
388:
196:Emesvirus zinderi
191:Bacteriophage MS2
188:
187:
181:Emesvirus zinderi
36:Emesvirus zinderi
18:Emesvirus zinderi
16:(Redirected from
1526:
1494:
1493:
1481:
1480:
1479:
1453:
1452:
1440:
1439:
1438:
1408:
1377:
1376:
1364:
1358:
1357:
1321:
1315:
1314:
1304:
1280:
1274:
1273:
1245:
1239:
1238:
1219:10.1038/265687a0
1205:(5596): 687–95.
1194:
1188:
1187:
1168:10.1038/237082a0
1143:
1137:
1136:
1100:
1094:
1093:
1083:
1073:
1049:
1043:
1042:
1032:
1000:
994:
993:
965:
959:
958:
948:
938:
905:
899:
898:
879:10.1038/345036a0
854:
848:
847:
819:
813:
812:
793:10.1038/260500a0
767:
758:
757:
747:
715:
709:
708:
690:
659:bacteriophage Qβ
654:bacteriophage f2
643:
638:
637:
592:bacteriophage f2
552:bacteriophage Qβ
260:
222:bacteriophage Qβ
218:bacteriophage f2
209:Escherichia coli
73:
72:
53:capsid structure
46:
32:
27:Species of virus
21:
1534:
1533:
1529:
1528:
1527:
1525:
1524:
1523:
1504:
1503:
1502:
1497:
1489:
1484:
1475:
1474:
1469:
1456:
1448:
1443:
1434:
1433:
1428:
1415:
1391:(also isolates
1389:Complete genome
1385:
1380:
1366:
1365:
1361:
1332:(13): 5847–58.
1323:
1322:
1318:
1282:
1281:
1277:
1247:
1246:
1242:
1196:
1195:
1191:
1145:
1144:
1140:
1102:
1101:
1097:
1051:
1050:
1046:
1002:
1001:
997:
967:
966:
962:
907:
906:
902:
865:(6270): 36–41.
856:
855:
851:
821:
820:
816:
779:(5551): 500–7.
769:
768:
761:
730:(12): 2485–93.
724:Protein Science
717:
716:
712:
705:
692:
691:
676:
672:
639:
632:
629:
588:
504:
449:
399:-protein), the
250:
245:
224:, R17, and GA.
184:
67:
28:
23:
22:
15:
12:
11:
5:
1532:
1530:
1522:
1521:
1516:
1514:Bacteriophages
1506:
1505:
1499:
1498:
1496:
1495:
1482:
1466:
1464:
1458:
1457:
1455:
1454:
1441:
1425:
1423:
1417:
1416:
1411:
1405:
1404:
1384:
1383:External links
1381:
1379:
1378:
1373:The Today Show
1359:
1316:
1289:Molecular Cell
1275:
1240:
1189:
1154:(5350): 82–8.
1138:
1095:
1064:(7): 961–969.
1044:
995:
960:
921:(2): 405–410.
900:
849:
814:
759:
710:
704:978-0195148503
703:
673:
671:
668:
667:
666:
664:phi-X174 phage
661:
656:
651:
645:
644:
641:Viruses portal
628:
625:
587:
584:
503:
500:
448:
445:
387:
386:
383:
372:
369:
356:
355:
352:
347:
344:
331:
330:
327:
322:
319:
306:
305:
302:
292:
289:
276:
275:
270:
267:
264:
249:
246:
244:
241:
186:
185:
178:
176:
172:
171:
164:
160:
159:
152:
148:
147:
140:
136:
135:
128:
124:
123:
116:
112:
111:
104:
100:
99:
92:
85:
84:
79:
75:
74:
61:
60:
48:
47:
39:
38:
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
1531:
1520:
1517:
1515:
1512:
1511:
1509:
1492:
1487:
1483:
1478:
1472:
1468:
1467:
1465:
1463:
1459:
1451:
1446:
1442:
1437:
1431:
1427:
1426:
1424:
1422:
1418:
1414:
1409:
1402:
1398:
1394:
1390:
1387:
1386:
1382:
1374:
1370:
1363:
1360:
1355:
1351:
1347:
1343:
1339:
1335:
1331:
1327:
1320:
1317:
1312:
1308:
1303:
1298:
1295:(4): 437–45.
1294:
1290:
1286:
1279:
1276:
1271:
1267:
1263:
1259:
1256:(3): 219–30.
1255:
1251:
1244:
1241:
1236:
1232:
1228:
1224:
1220:
1216:
1212:
1208:
1204:
1200:
1193:
1190:
1185:
1181:
1177:
1173:
1169:
1165:
1161:
1157:
1153:
1149:
1142:
1139:
1134:
1130:
1126:
1122:
1118:
1114:
1110:
1106:
1099:
1096:
1091:
1087:
1082:
1077:
1072:
1067:
1063:
1059:
1055:
1048:
1045:
1040:
1036:
1031:
1026:
1022:
1018:
1014:
1010:
1006:
999:
996:
991:
987:
983:
979:
976:(3): 620–39.
975:
971:
964:
961:
956:
952:
947:
942:
937:
932:
928:
924:
920:
917:
916:
911:
904:
901:
896:
892:
888:
884:
880:
876:
872:
868:
864:
860:
853:
850:
845:
841:
837:
833:
829:
825:
818:
815:
810:
806:
802:
798:
794:
790:
786:
782:
778:
774:
766:
764:
760:
755:
751:
746:
741:
737:
733:
729:
725:
721:
714:
711:
706:
700:
696:
689:
687:
685:
683:
681:
679:
675:
669:
665:
662:
660:
657:
655:
652:
650:
649:bacteriophage
647:
646:
642:
636:
631:
626:
624:
622:
617:
615:
612:
608:
603:
600:
595:
593:
585:
583:
581:
577:
573:
569:
564:
561:
555:
553:
543:
539:
537:
533:
528:
527:messenger RNA
523:
521:
517:
513:
509:
501:
499:
497:
493:
489:
485:
480:
479:(pI) of 3.9.
478:
474:
470:
466:
458:
453:
446:
444:
442:
438:
437:messenger RNA
434:
430:
426:
422:
418:
414:
410:
406:
402:
398:
394:
384:
382:
381:
376:
375:RNA replicase
373:
370:
368:
365:
364:
363:
358:
357:
353:
351:
350:lysis protein
348:
345:
343:
340:
339:
338:
333:
332:
328:
326:
323:
320:
318:
315:
314:
313:
308:
307:
303:
301:
300:
296:
293:
290:
288:
285:
284:
283:
278:
277:
274:
271:
268:
265:
262:
261:
254:
247:
242:
240:
238:
232:
228:
225:
223:
219:
215:
211:
210:
205:
202:
198:
197:
192:
183:
182:
177:
174:
173:
170:
169:
165:
162:
161:
158:
157:
153:
150:
149:
146:
145:
141:
138:
137:
134:
133:
132:Leviviricetes
129:
126:
125:
122:
121:
120:Lenarviricota
117:
114:
113:
110:
109:
108:Orthornavirae
105:
102:
101:
98:
97:
93:
90:
87:
86:
83:
80:
77:
76:
71:
66:
62:
58:
54:
49:
45:
40:
37:
33:
30:
19:
1519:Fiersviridae
1461:
1420:
1372:
1362:
1329:
1325:
1319:
1292:
1288:
1278:
1253:
1249:
1243:
1202:
1198:
1192:
1151:
1147:
1141:
1108:
1104:
1098:
1061:
1058:Microbiology
1057:
1047:
1012:
1008:
998:
973:
969:
963:
918:
913:
903:
862:
858:
852:
827:
823:
817:
776:
772:
727:
723:
713:
694:
618:
604:
599:Walter Fiers
596:
589:
579:
565:
556:
548:
524:
505:
481:
462:
456:
428:
424:
420:
416:
408:
404:
396:
390:
380:beta subunit
378:
366:
360:
359:
341:
335:
334:
325:coat protein
316:
310:
309:
297:
286:
280:
279:
269:Gene product
233:
229:
226:
207:
195:
194:
190:
189:
180:
179:
167:
156:Fiersviridae
155:
144:Norzivirales
143:
131:
119:
107:
95:
88:
78:(unranked):
35:
29:
621:noroviruses
607:MS2 tagging
494:. When the
469:icosahedral
411:), and the
237:noroviruses
1508:Categories
1477:Q106960351
670:References
502:Life cycle
393:MS2 genome
295:maturation
57:conformers
830:: 43–54.
580:Levivirus
572:cell wall
532:ribosomes
488:α-helices
486:with two
457:Levivirus
447:Structure
441:expressed
413:replicase
175:Species:
168:Emesvirus
103:Kingdom:
96:Riboviria
1471:Wikidata
1450:11459711
1436:Q4840020
1430:Wikidata
1346:24816622
1270:85199492
1133:17795773
1090:28691656
1039:28396351
844:13978804
627:See also
367:(MS2g4)
342:(MS2g3)
317:(MS2g2)
287:(MS2g1)
243:Virology
151:Family:
115:Phylum:
1354:6212583
1311:9809065
1235:4206886
1207:Bibcode
1184:4153893
1176:4555447
1156:Bibcode
1113:Bibcode
1105:Science
1081:5775895
1030:5446614
990:8254664
955:9464373
923:Bibcode
895:2803228
887:2330049
867:Bibcode
809:4289674
801:1264203
781:Bibcode
754:8976557
745:2143325
614:imaging
536:hairpin
512:plasmid
492:hairpin
484:β-sheet
463:An MS2
371:2055 nt
299:protein
291:1487 nt
163:Genus:
139:Order:
127:Class:
1399:, and
1352:
1344:
1309:
1268:
1233:
1227:870828
1225:
1199:Nature
1182:
1174:
1148:Nature
1131:
1088:
1078:
1037:
1027:
1015:(12).
988:
953:
946:106058
943:
893:
885:
859:Nature
842:
807:
799:
773:Nature
752:
742:
701:
560:capsid
496:capsid
490:and a
465:virion
346:295 nt
321:510 nt
248:Genome
1491:8TT92
1445:IRMNG
1350:S2CID
1266:S2CID
1231:S2CID
1180:S2CID
891:S2CID
805:S2CID
611:tumor
520:pilus
401:lysis
89:Realm
82:Virus
1397:DL16
1342:PMID
1307:PMID
1223:PMID
1172:PMID
1129:PMID
1086:PMID
1035:PMID
986:PMID
951:PMID
883:PMID
840:PMID
797:PMID
750:PMID
699:ISBN
576:DnaJ
510:, a
391:The
385:545
329:130
304:393
266:Size
263:Gene
1486:CoL
1401:J20
1393:R17
1334:doi
1297:doi
1258:doi
1215:doi
1203:265
1164:doi
1152:237
1121:doi
1109:134
1076:PMC
1066:doi
1062:163
1025:PMC
1017:doi
1013:199
978:doi
974:234
941:PMC
931:doi
875:doi
863:345
832:doi
789:doi
777:260
740:PMC
732:doi
473:T=3
429:rep
421:lys
417:rep
405:lys
362:rep
354:75
337:lys
282:mat
204:RNA
1510::
1488::
1473::
1447::
1432::
1395:,
1371:.
1348:.
1340:.
1330:98
1328:.
1305:.
1291:.
1287:.
1264:.
1252:.
1229:.
1221:.
1213:.
1201:.
1178:.
1170:.
1162:.
1150:.
1127:.
1119:.
1107:.
1084:.
1074:.
1060:.
1056:.
1033:.
1023:.
1011:.
1007:.
984:.
972:.
949:.
939:.
929:.
919:64
912:.
889:.
881:.
873:.
861:.
838:.
826:.
803:.
795:.
787:.
775:.
762:^
748:.
738:.
726:.
722:.
677:^
594:.
425:cp
409:cp
377:,
312:cp
273:aa
220:,
91::
1403:)
1375:.
1356:.
1336::
1313:.
1299::
1293:2
1272:.
1260::
1254:7
1237:.
1217::
1209::
1186:.
1166::
1158::
1135:.
1123::
1115::
1092:.
1068::
1041:.
1019::
992:.
980::
957:.
933::
925::
897:.
877::
869::
846:.
834::
828:7
811:.
791::
783::
756:.
734::
728:5
707:.
415:(
403:(
397:A
193:(
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.