31:
600:
61:
595:
369:"Genome-based phylogeny and taxonomy of the 'Enterobacteriales': proposal for Enterobacterales ord. nov. divided into the families Enterobacteriaceae, Erwiniaceae fam. nov., Pectobacteriaceae fam. nov., Yersiniaceae fam. nov., Hafniaceae fam. nov., Morganellaceae fam. nov., and Budviciaceae fam. nov"
328:
was largely based on 16S rRNA genome sequence analyses, which is known to have low discriminatory power and the results of which changes depends on the algorithm and organism information used. Despite this, the analyses still exhibited polyphyletic branching, indicating the presence of distinct
336:
into 7 novel families based on comparative genomic analyses and the branching pattern of various phylogenetic trees constructed from conserved genome sequences, 16S rRNA sequences and multilocus sequence analyses. Molecular markers, specifically
373:
554:
324:, a large phylogenetically unrelated group of species with distinct biochemical characteristics and different ecological niches. The original assignment of species into the family
641:
541:
567:
30:
528:
634:
458:
299:, and stationary phase inducible protein CsiE. These CSIs provide a reliable molecular method of identification and differentiation of
257:
These bacteria are catalase-positive, oxidase-negative, and do not produce indole or hydrogen disulfide. Most species are positive for
675:
627:
660:
276:
292:
341:, specific to this family were also identified as evidence supporting the division independent of phylogenetic trees.
60:
572:
318:, as of 2021, contains eight validly published genera. Members of this family were originally members of the family
670:
338:
272:
665:
283:(subunit B), LPS assembly protein LptD, thiol:disulfide interchange protein DsbA precursor, two-component sensor
258:
238:, referring the type genus of the family and the suffix "-aceae", an ending used to denote a family. Together,
481:
202:
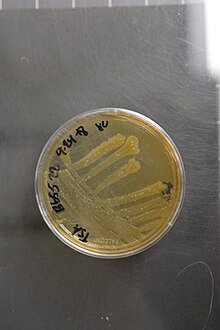
39:
296:
131:
439:"Phylogenetic Relationships of Bacteria with Special Reference to Endosymbionts and Enteric Species"
159:
607:
594:
214:
92:
320:
55:
180:
45:
546:
166:
454:
392:
611:
519:
275:(CSIs) were identified through genomic analyses as exclusive for this family in the proteins
599:
446:
382:
284:
210:
198:
102:
206:
218:
173:
82:
654:
288:
504:
513:
280:
438:
450:
396:
387:
368:
559:
413:
498:
72:
533:
223:
152:
138:
475:
145:
374:
International
Journal of Systematic and Evolutionary Microbiology
479:
367:
Adeolu, M; Alnajar, S; Naushad, S; S Gupta, R (December 2016).
209:
and insect endosymbionts. This family is a member of the order
437:
Francino, M. Pilar; Santos, Scott R.; Ochman, Howard (2006),
295:
TruD, glycine/betaine ABC transporter ATP-binding protein,
242:
refers to a family whose nomenclatural type is the genus
615:
250:
Biochemical characteristics and molecular signatures
488:
445:, New York, NY: Springer New York, pp. 41–59,
332:In 2016, Adeolu et al. proposed the division of
635:
362:
360:
358:
356:
354:
8:
311:Historical systematics and current taxonomy
642:
628:
476:
29:
20:
386:
303:species from other families in the order
350:
7:
591:
589:
408:
406:
263:Erwinia toletana, Erwinia ypographi
221:. The type genus of this family is
614:. You can help Knowledge (XXG) by
14:
598:
593:
59:
293:tRNA pseudouridine(13) synthase
234:is derived from the Latin term
329:subgroups within the family.
1:
205:which includes a number of
692:
588:
339:conserved signature indels
273:conserved signature indels
676:Gammaproteobacteria stubs
277:glutamate–cysteine ligase
127:
122:
56:Scientific classification
54:
37:
28:
23:
261:, with the exception of
451:10.1007/0-387-30746-x_2
661:Gram-negative bacteria
610:-related article is a
388:10.1099/ijsem.0.001485
203:Gram-negative bacteria
414:"Family: Erwiniaceae"
307:and other bacteria.
297:superoxide dismutase
265:and some strains of
259:Voges-Proskauer test
608:Gammaproteobacteria
215:Gammaproteobacteria
117:Adeolu et al., 2016
93:Gammaproteobacteria
334:Enterobacteriaceae
326:Enterobacteriaceae
321:Enterobacteriaceae
16:Family of bacteria
671:Bacteria families
623:
622:
583:
582:
482:Taxon identifiers
460:978-0-387-25496-8
381:(12): 5575–5599.
189:
188:
118:
40:Pantoea piersonii
683:
666:Enterobacterales
644:
637:
630:
602:
597:
590:
576:
575:
563:
562:
550:
549:
537:
536:
524:
523:
522:
509:
508:
507:
477:
470:
469:
468:
467:
434:
428:
427:
425:
424:
410:
401:
400:
390:
364:
305:Enterobacterales
285:histidine kinase
211:Enterobacterales
116:
103:Enterobacterales
64:
63:
33:
21:
691:
690:
686:
685:
684:
682:
681:
680:
651:
650:
649:
648:
586:
584:
579:
571:
566:
558:
553:
545:
540:
532:
527:
518:
517:
512:
503:
502:
497:
484:
474:
473:
465:
463:
461:
443:The Prokaryotes
436:
435:
431:
422:
420:
412:
411:
404:
366:
365:
352:
347:
313:
252:
207:plant pathogens
115:
58:
17:
12:
11:
5:
689:
687:
679:
678:
673:
668:
663:
653:
652:
647:
646:
639:
632:
624:
621:
620:
603:
581:
580:
578:
577:
564:
551:
538:
525:
510:
494:
492:
486:
485:
480:
472:
471:
459:
429:
402:
349:
348:
346:
343:
312:
309:
267:Erwinia oleae.
251:
248:
219:Pseudomonadota
217:of the phylum
187:
186:
185:
184:
177:
174:Wigglesworthia
170:
163:
160:Phaseolibacter
156:
149:
142:
135:
125:
124:
120:
119:
110:
106:
105:
100:
96:
95:
90:
86:
85:
83:Pseudomonadota
80:
76:
75:
70:
66:
65:
52:
51:
50:on agar plate
35:
34:
26:
25:
15:
13:
10:
9:
6:
4:
3:
2:
688:
677:
674:
672:
669:
667:
664:
662:
659:
658:
656:
645:
640:
638:
633:
631:
626:
625:
619:
617:
613:
609:
604:
601:
596:
592:
587:
574:
569:
565:
561:
556:
552:
548:
543:
539:
535:
530:
526:
521:
515:
511:
506:
500:
496:
495:
493:
491:
487:
483:
478:
462:
456:
452:
448:
444:
440:
433:
430:
419:
415:
409:
407:
403:
398:
394:
389:
384:
380:
376:
375:
370:
363:
361:
359:
357:
355:
351:
344:
342:
340:
335:
330:
327:
323:
322:
317:
310:
308:
306:
302:
298:
294:
290:
286:
282:
278:
274:
269:
268:
264:
260:
255:
249:
247:
245:
241:
237:
233:
228:
227:
225:
220:
216:
213:in the class
212:
208:
204:
200:
196:
195:
183:
182:
178:
176:
175:
171:
169:
168:
164:
162:
161:
157:
155:
154:
150:
148:
147:
143:
141:
140:
136:
134:
133:
129:
128:
126:
121:
114:
111:
108:
107:
104:
101:
98:
97:
94:
91:
88:
87:
84:
81:
78:
77:
74:
71:
68:
67:
62:
57:
53:
49:
47:
42:
41:
36:
32:
27:
22:
19:
616:expanding it
605:
585:
489:
464:, retrieved
442:
432:
421:. Retrieved
418:lpsn.dsmz.de
417:
378:
372:
333:
331:
325:
319:
315:
314:
304:
300:
289:RNA helicase
270:
266:
262:
256:
253:
243:
239:
235:
231:
229:
222:
193:
192:
190:
179:
172:
165:
158:
151:
144:
137:
130:
112:
44:
38:
24:Erwiniaceae
18:
560:erwiniaceae
520:Erwiniaceae
514:Wikispecies
490:Erwiniaceae
316:Erwiniaceae
301:Erwiniaceae
240:Erwiniaceae
232:Erwiniaceae
194:Erwiniaceae
113:Erwiniaceae
43:, formerly
655:Categories
466:2021-06-02
423:2021-06-05
345:References
281:DNA gyrase
181:Kalamiella
46:Kalamiella
505:Q28735287
230:The name
167:Tatumella
48:piersonii
547:11909049
534:10161084
499:Wikidata
397:27620848
254:Source:
132:Buchnera
109:Family:
79:Phylum:
73:Bacteria
69:Domain:
573:1903409
244:Erwinia
236:Erwinia
224:Erwinia
153:Pantoea
139:Erwinia
123:Genera
99:Order:
89:Class:
457:
395:
199:family
197:are a
606:This
542:IRMNG
146:Mixta
612:stub
568:NCBI
555:LPSN
529:GBIF
455:ISBN
393:PMID
191:The
447:doi
383:doi
271:12
201:of
657::
570::
557::
544::
531::
516::
501::
453:,
441:,
416:.
405:^
391:.
379:66
377:.
371:.
353:^
291:,
287:,
279:,
246:.
643:e
636:t
629:v
618:.
449::
426:.
399:.
385::
226:.
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.