230:
643:
551:
622:
24:
538:(TC) motif Cys-Cys-Xxx-Xxx-Cys-Cys and becoming fluorescent when bound. It displays non-specific binding to endogenous cysteine-rich proteins, meaning it binds to sites other than the one of interest (CCXXCC). Further optimization of the TC motif has revealed improved FlAsH binding affinity for a CCPGCC motif, and higher
635:. Such strong sulfur-arsenic bond can be, again, regulated by designing a peptide domain that exhibits higher affinity toward the arsenic, such as tetracysteine motif. By modulating the distance between the two pairs of cysteine residues and the space between the arsenic centers of FlAsH-EDT
825:
Adams, Stephen R.; Campbell, Robert E.; Gross, Larry A.; Martin, Brent R.; Walkup, Grant K.; Yao, Yong; Llopis, Juan; Tsien, Roger Y. (2002). "New biarsenical ligands and tetracysteine motifs for protein labeling in vitro and in vivo: synthesis and biological applications".
630:
Many studies show that trivalent arsenic compounds bind to pairs of cysteine residues. This binding is responsible for the toxicity of many arsenic compounds. Binding is reversed by 1,2-ethanedithiol, which binds tightly to arsenic compounds, as shown by the stability of
698:
enables less toxic and more specific fluorescent labeling that is membrane permeable. The modification of the fluorescein moiety also allows multicolor analysis. It has been proven to be a good alternative to green fluorescent proteins (GFP) with the advantage that
91:
677:
has 0.1-0.6 fluorescence quantum efficiencies with several μM detection limits for diffuse cytosolic tag and 30 - 80 extinction coefficients L mmol cm. The FlAsH-peptide complex also has demonstrated
861:
Martin, Brent R.; Giepmans, Ben N.G.; Adams, Stephen R.; Tsien, Roger Y. (2005). "Mammalian cell–based optimization of the biarsenical-binding tetracysteine motif for improved fluorescence and affinity".
84:
253:
InChI=1S/C24H18As2O5S4/c27-17-7-5-15-19(13-3-1-2-4-14(13)24(29)30)16-6-8-18(28)21(26-34-11-12-35-26)23(16)31-22(15)20(17)25-32-9-10-33-25/h1-8,27H,9-12H2,(H,29,30)
1149:
727:
cellular events and subcellular structures in animal cells, Ebola virus matrix protein, and protein misfolding. With the electron microscopic imaging, FlAsH-EDT
666:
FlAsH becomes fluorescent upon the binding of tetracysteine motif. It is excited at 508 nm and emits 528 nm, a green-yellow, of free fluorescein. The
905:
Kalef, Edna; Gitler, Carlos (1994). "Purification of
Vicinal Dithiol-containing Proteins by Arsenical-Based Affinity Chromatography". In Sies, Helmut (ed.).
269:
1332:
1101:
Hoffmann, Carsten; Gaietta, Guido; Zürn, Alexander; Adams, Stephen R.; Terrillon, Sonia; Ellisman, Mark H.; Tsien, Roger Y.; Lohse, Martin J. (2010).
679:
1065:(2002). "New Biarsenical Ligands and Tetracysteine Motifs for Protein Labeling in Vitro and in Vivo: Synthesis and Biological Applications".
1357:
1198:
930:
244:
654:
is thus subject to equilibration. The FlAsH-peptide adduct formation can be favored in low concentration of EDT (below 10
670:
is 0.49 for 250 nM FlAsH is bound to a model tetracysteine-containing peptide in a phosphate-buffered saline at pH 7.4.
1362:
376:
1352:
711:) as compared to GFPs (~30 kDa), therefore minimizing the perturbation of activity of the protein under the study.
187:
1222:
Gaietta, Guido; Deerinck, Thomas J.; Adams, Stephen R.; Bouwer, James; Tour, Oded; Laird, Dale W.; Sosinsky, Gina E.;
1061:
Adams, Stephen R.; Campbell, Robert E.; Gross, Larry A.; Martin, Brent R.; Walkup, Grant K.; Yao, Yong; Llopis, Juan;
208:
517:
1337:
683:
584:
451:
1347:
395:
427:
131:
1178:
910:
1165:(2000). "Fluorescent labeling of recombinant proteins in living cells with FlAsH". In Thorner, Jeremy;
1237:
1027:
562:
225:
1226:; Ellisman, Mark H. (2002). "Multicolor and Electron Microscopic Imaging of Connexin Trafficking".
954:
655:
570:
50:
1261:
887:
473:
1150:
TC-FlAsH™ II In-Cell
Tetracysteine Tag Detection Kit (Green Fluorescence), for live-cell imaging
642:
542:
when the tetracysteine motif is flanked with specific residues (HRWCCPGCCKTF or FLNCCPGCCMEP).
1308:
1253:
1228:
1204:
1194:
1132:
1083:
1067:
1043:
1018:
989:
936:
926:
879:
843:
807:
592:
577:
558:
458:
399:
151:
1342:
1298:
1245:
1186:
1122:
1114:
1075:
1035:
979:
971:
918:
871:
835:
797:
789:
297:
60:
682:(FRET) from fluorescent proteins, such as from enhanced cyan fluorescent protein (ECFP) of
430:
substituents. It is used in bioanalytical research as a fluorescent label for visualising
196:
229:
1016:(1998). "Specific Covalent Labeling of Recombinant Protein Molecules Inside Live Cells".
534:
is used for site-specific labelling, selectively binding to proteins containing the tetra
1241:
1031:
111:
1223:
1182:
1175:
Applications of
Chimeric Genes and Hybrid Proteins, Part B: Cell Biology and Physiology
1162:
1127:
1102:
1062:
1013:
984:
958:
914:
802:
768:
764:
370:
1190:
922:
1326:
1289:
708:
667:
539:
358:
1265:
891:
176:
1170:
1166:
781:
524:
1039:
639:, a cooperative and entropically favored dithiol arsenic bond could be achieved.
963:
737:
621:
608:
550:
423:
1281:"FlAsH-based live-cell fluorescent imaging of synthetic peptides expressed in
959:"An Experimental Investigation of the 'Ring Hypothesis' of Arsenical Toxicity"
704:
337:
142:
1249:
1312:
1257:
1208:
1136:
1118:
1087:
993:
883:
847:
811:
793:
1047:
940:
535:
513:
1103:"Fluorescent labeling of tetracysteine-tagged proteins in intact cells"
735:. More recently, it was used in an extended study of plant cells like
431:
163:
23:
1303:
1280:
1079:
975:
839:
512:
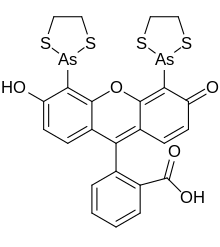
COOH, representing the dithiarsolane substituents bound to the hydroxy
122:
875:
472:, and is a pale yellow or pinkish fluorogenic solid. It has a semi-
369:
Except where otherwise noted, data are given for materials in their
282:
OC(=O)C1=C(C=CC=C1)C1=C2C=CC(=O)C(3SCCS3)=C2OC2=C(3SCCS3)C(O)=CC=C12
658:) and be reversed in high concentration of EDT (above 1 mM).
641:
549:
102:
90:
83:
73:
277:
c1ccc(c(c1)c2c3ccc(c(c3oc-4c(c(=O)ccc24)5SCCS5)6SCCS6)O)C(=O)O
769:"Preparation of the membrane-permeant biarsenicals FlAsH-EDT
620:
213:
731:
is also used to study the processes of protein trafficking
363:
169 to 172 °C (336 to 342 °F; 442 to 445 K)
777:
for fluorescent labeling of tetracysteine-tagged proteins"
1161:Griffin, B. Albert; Adams, Stephen R.; Jones, Jay;
38:Fluorescein Arsenical Hairpin Binder; Lumio green
175:
59:
907:Oxygen Radicals in Biological Systems, Part C
8:
1279:Estévez, José M.; Somerville, Chris (2006).
758:
756:
754:
723:has been widely used to study a number of
228:
150:
15:
1302:
1126:
983:
801:
195:
828:Journal of the American Chemical Society
1012:Griffin, B. Albert; Adams, Stephen R.;
1007:
1005:
1003:
750:
274:
249:
224:
680:fluorescence resonance energy transfer
256:Key: KCPRYVGBEBFLIG-UHFFFAOYSA-N
130:
110:
7:
607:can be prepared in three steps from
422:. Its structure is based around a
166:
14:
1333:Biochemistry detection reactions
309:
22:
373:(at 25 °C , 100 kPa).
315:
327:
321:
303:
1:
1191:10.1016/S0076-6879(00)27302-3
923:10.1016/S0076-6879(94)33046-8
1040:10.1126/science.281.5374.269
646:Formation of FlAsH-TC adduct
615:Formation of FlAsH-TC adduct
434:in living cells. FlAsH-EDT
1379:
1358:Organoarsenic dithiolates
684:Green Fluorescent Protein
367:
290:
265:
240:
43:
35:
30:
21:
650:The binding of FlAsH-EDT
1250:10.1126/science.1068793
707: < 1
438:is an abbreviation for
1119:10.1038/nprot.2010.129
794:10.1038/nprot.2008.144
719:In the past, FlAsH-EDT
647:
625:
600:
396:organoarsenic compound
1179:Methods in Enzymology
911:Methods in Enzymology
645:
624:
553:
516:core, attached to an
1363:Arsenic heterocycles
1185:. pp. 565–578.
955:Whittaker, Victor P.
917:. pp. 395–403.
864:Nature Biotechnology
673:Generally, FlAsH-EDT
1353:Triarylmethane dyes
1242:2002Sci...296..503G
1032:1998Sci...281..269G
763:Adams, Stephen R.;
599:in aqueous acetone.
428:1,3,2-dithiarsolane
345: g·mol
18:
648:
626:
601:
474:structural formula
377:Infobox references
16:
1304:10.2144/000112264
1236:(5567): 503–507.
1181:. Vol. 327.
1113:(10): 1666–1677.
1080:10.1021/ja017687n
1074:(21): 6063–6076.
1068:J. Am. Chem. Soc.
1026:(5374): 269–272.
976:10.1042/bj0410056
913:. Vol. 233.
870:(10): 1308–1314.
840:10.1021/ja017687n
834:(21): 6063–6076.
703:is much smaller (
400:molecular formula
385:Chemical compound
383:
382:
209:CompTox Dashboard
92:Interactive image
85:Interactive image
1370:
1338:Fluorescent dyes
1317:
1316:
1306:
1276:
1270:
1269:
1219:
1213:
1212:
1171:Abelson, John N.
1158:
1152:
1147:
1141:
1140:
1130:
1107:Nature Protocols
1098:
1092:
1091:
1058:
1052:
1051:
1009:
998:
997:
987:
951:
945:
944:
902:
896:
895:
858:
852:
851:
822:
816:
815:
805:
788:(9): 1527–1534.
760:
344:
329:
323:
317:
311:
305:
298:Chemical formula
233:
232:
217:
215:
199:
179:
168:
154:
134:
114:
94:
87:
63:
26:
19:
1378:
1377:
1373:
1372:
1371:
1369:
1368:
1367:
1323:
1322:
1321:
1320:
1278:
1277:
1273:
1224:Tsien, Roger Y.
1221:
1220:
1216:
1201:
1163:Tsien, Roger Y.
1160:
1159:
1155:
1148:
1144:
1100:
1099:
1095:
1063:Tsien, Roger Y.
1060:
1059:
1055:
1014:Tsien, Roger Y.
1011:
1010:
1001:
953:
952:
948:
933:
904:
903:
899:
876:10.1038/nbt1136
860:
859:
855:
824:
823:
819:
776:
772:
765:Tsien, Roger Y.
762:
761:
752:
747:
730:
722:
717:
702:
697:
692:
676:
664:
653:
638:
634:
617:
606:
596:
581:
574:
548:
533:
511:
507:
503:
499:
495:
491:
487:
483:
479:
437:
421:
417:
413:
409:
405:
392:
386:
379:
374:
342:
332:
326:
320:
314:
308:
300:
286:
283:
278:
273:
272:
261:
258:
257:
254:
248:
247:
236:
218:
211:
202:
182:
169:
157:
137:
117:
97:
77:
66:
53:
39:
12:
11:
5:
1376:
1374:
1366:
1365:
1360:
1355:
1350:
1345:
1340:
1335:
1325:
1324:
1319:
1318:
1297:(5): 569–574.
1271:
1214:
1199:
1183:Academic Press
1153:
1142:
1093:
1053:
999:
946:
931:
915:Academic Press
897:
853:
817:
774:
770:
749:
748:
746:
743:
728:
720:
716:
713:
700:
695:
691:
688:
674:
663:
660:
651:
636:
632:
628:
627:
616:
613:
611:(see figure).
604:
594:
579:
576:, followed by
572:
547:
544:
531:
509:
505:
501:
497:
493:
489:
485:
481:
477:
435:
426:core with two
419:
415:
411:
407:
403:
390:
384:
381:
380:
375:
371:standard state
368:
365:
364:
361:
355:
354:
351:
347:
346:
340:
334:
333:
330:
324:
318:
312:
306:
301:
296:
293:
292:
288:
287:
285:
284:
281:
279:
276:
268:
267:
266:
263:
262:
260:
259:
255:
252:
251:
243:
242:
241:
238:
237:
235:
234:
226:DTXSID90431400
221:
219:
207:
204:
203:
201:
200:
192:
190:
184:
183:
181:
180:
172:
170:
162:
159:
158:
156:
155:
147:
145:
139:
138:
136:
135:
127:
125:
119:
118:
116:
115:
107:
105:
99:
98:
96:
95:
88:
80:
78:
71:
68:
67:
65:
64:
56:
54:
49:
46:
45:
41:
40:
37:
33:
32:
28:
27:
13:
10:
9:
6:
4:
3:
2:
1375:
1364:
1361:
1359:
1356:
1354:
1351:
1349:
1348:Benzoic acids
1346:
1344:
1341:
1339:
1336:
1334:
1331:
1330:
1328:
1314:
1310:
1305:
1300:
1296:
1292:
1291:
1290:BioTechniques
1286:
1284:
1275:
1272:
1267:
1263:
1259:
1255:
1251:
1247:
1243:
1239:
1235:
1231:
1230:
1225:
1218:
1215:
1210:
1206:
1202:
1200:9780080496825
1196:
1192:
1188:
1184:
1180:
1176:
1172:
1168:
1167:Emr, Scott D.
1164:
1157:
1154:
1151:
1146:
1143:
1138:
1134:
1129:
1124:
1120:
1116:
1112:
1108:
1104:
1097:
1094:
1089:
1085:
1081:
1077:
1073:
1070:
1069:
1064:
1057:
1054:
1049:
1045:
1041:
1037:
1033:
1029:
1025:
1021:
1020:
1015:
1008:
1006:
1004:
1000:
995:
991:
986:
981:
977:
973:
969:
966:
965:
960:
956:
950:
947:
942:
938:
934:
932:9780080883465
928:
924:
920:
916:
912:
908:
901:
898:
893:
889:
885:
881:
877:
873:
869:
865:
857:
854:
849:
845:
841:
837:
833:
829:
821:
818:
813:
809:
804:
799:
795:
791:
787:
784:
783:
778:
773:and ReAsH-EDT
766:
759:
757:
755:
751:
744:
742:
741:and tobacco.
740:
739:
734:
726:
714:
712:
710:
706:
689:
687:
685:
681:
671:
669:
668:quantum yield
661:
659:
657:
644:
640:
623:
619:
618:
614:
612:
610:
598:
590:
586:
582:
575:
568:
564:
560:
556:
552:
545:
543:
541:
540:quantum yield
537:
528:
526:
522:
520:
515:
475:
471:
469:
465:
461:
456:
455:airpin binder
454:
449:
445:
441:
433:
429:
425:
401:
397:
393:
378:
372:
366:
362:
360:
359:Melting point
357:
356:
352:
349:
348:
341:
339:
336:
335:
302:
299:
295:
294:
289:
280:
275:
271:
264:
250:
246:
239:
231:
227:
223:
222:
220:
210:
206:
205:
198:
194:
193:
191:
189:
186:
185:
178:
174:
173:
171:
165:
161:
160:
153:
149:
148:
146:
144:
141:
140:
133:
129:
128:
126:
124:
121:
120:
113:
109:
108:
106:
104:
101:
100:
93:
89:
86:
82:
81:
79:
75:
70:
69:
62:
58:
57:
55:
52:
48:
47:
42:
34:
29:
25:
20:
1294:
1288:
1285:and tobacco"
1282:
1274:
1233:
1227:
1217:
1174:
1156:
1145:
1110:
1106:
1096:
1071:
1066:
1056:
1023:
1017:
970:(1): 56–62.
967:
962:
949:
906:
900:
867:
863:
856:
831:
827:
820:
785:
782:Nat. Protoc.
780:
736:
732:
724:
718:
693:
672:
665:
649:
629:
602:
588:
566:
554:
529:
525:benzoic acid
523:molecule of
521:-substituted
518:
467:
463:
459:
452:
447:
443:
439:
388:
387:
132:ChEMBL454668
44:Identifiers
36:Other names
1283:Arabidopsis
964:Biochem. J.
738:Arabidopsis
690:Application
609:fluorescein
546:Preparation
424:fluorescein
350:Appearance
291:Properties
112:CHEBI:52107
61:212118-77-9
17:FlAsH-EDT2
1327:Categories
745:References
705:molar mass
662:Properties
338:Molar mass
197:Z8CH38Y9L3
143:ChemSpider
72:3D model (
51:CAS Number
699:FlAsH-EDT
694:FlAsH-EDT
631:FlAsH-EDT
603:FlAsH-EDT
530:FlAsH-EDT
442:uorescin
389:FlAsH-EDT
1313:17140113
1266:16397816
1258:11964472
1209:11045009
1173:(eds.).
1137:20885379
1088:12022841
994:16748119
957:(1947).
892:16456334
884:16155565
848:12022841
812:18772880
767:(2008).
536:cysteine
514:xanthone
432:proteins
1343:Phenols
1238:Bibcode
1229:Science
1128:3086663
1048:9657724
1028:Bibcode
1019:Science
985:1258423
941:8015475
803:2843588
733:in situ
725:in vivo
686:(GFP).
578:Pd(OAc)
450:enical
177:2763100
164:PubChem
152:2043798
1311:
1264:
1256:
1207:
1197:
1135:
1125:
1086:
1046:
992:
982:
939:
929:
890:
882:
846:
810:
800:
589:Step 3
567:Step 2
555:Step 1
394:is an
353:Solid
343:664.49
270:SMILES
123:ChEMBL
31:Names
1262:S2CID
888:S2CID
462:thane
398:with
245:InChI
103:ChEBI
74:JSmol
1309:PMID
1254:PMID
1205:PMID
1195:ISBN
1133:PMID
1084:PMID
1044:PMID
990:PMID
937:PMID
927:ISBN
880:PMID
844:PMID
808:PMID
585:DIEA
583:and
571:AsCl
470:hiol
188:UNII
1299:doi
1246:doi
1234:296
1187:doi
1123:PMC
1115:doi
1076:doi
1072:124
1036:doi
1024:281
980:PMC
972:doi
919:doi
872:doi
836:doi
832:124
798:PMC
790:doi
715:Use
709:kDa
597:EDT
563:TFA
561:in
559:HgO
504:)-C
492:-(C
484:AsS
214:EPA
167:CID
1329::
1307:.
1295:41
1293:.
1287:.
1260:.
1252:.
1244:.
1232:.
1203:.
1193:.
1177:.
1169:;
1131:.
1121:.
1109:.
1105:.
1082:.
1042:.
1034:.
1022:.
1002:^
988:.
978:.
968:41
961:.
935:.
925:.
909:.
886:.
878:.
868:23
866:.
842:.
830:.
806:.
796:.
779:.
753:^
656:μM
591::
587:;
569::
565:;
557::
527:.
494:13
476:(C
440:fl
410:As
408:18
404:24
316:As
313:18
307:24
1315:.
1301::
1268:.
1248::
1240::
1211:.
1189::
1139:.
1117::
1111:5
1090:.
1078::
1050:.
1038::
1030::
996:.
974::
943:.
921::
894:.
874::
850:.
838::
814:.
792::
786:3
775:2
771:2
729:2
721:2
701:2
696:2
675:2
652:2
637:2
633:2
605:2
595:2
593:H
580:2
573:3
532:2
519:o
510:4
508:H
506:6
502:3
500:O
498:5
496:H
490:2
488:)
486:2
482:4
480:H
478:2
468:t
466:i
464:d
460:e
457:-
453:h
448:s
446:r
444:a
436:2
420:4
418:S
416:5
414:O
412:2
406:H
402:C
391:2
331:4
328:S
325:5
322:O
319:2
310:H
304:C
216:)
212:(
76:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.