122:
33:
344:
Takeshima, Hirohiko; Iguchi, Kei-ichiro & Nishida, Mutsumi (2005): Unexpected
Ceiling of Genetic Differentiation in the Control Region of the Mitochondrial DNA between Different Subspecies of the Ayu
141:
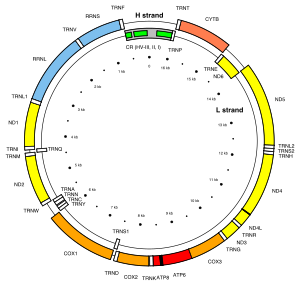
of the human mitogenome, the most variable sites of HVR1 are numbered 16024-16383 (this subsequence is called HVR-I), and the most variable sites of HVR2 are numbered 57-372 (
408:
Antibodies are remarkably specific, thanks to hypervariable regions in both light and heavy chains. The hyperbariable regions for the antigen-binding site.
133:. HVR1 is considered a "low resolution" region and HVR2 is considered a "high resolution" region. Getting HVR1 and HVR2 DNA tests can help determine one's
109:
repeat (in the case of nuclear DNA) or have substitutions (in the case of mitochondrial DNA). Changes or repeats in the hypervariable region are highly
380:
263:
238:
134:
394:
172:
accumulate mutations faster and more freely. It is not known whether such hypovariable control regions are more widespread. In the
125:
Human mitochondrial genome showing hypervariable regions I to III (green boxes) located in the control region (CR; grey box).
48:
453:
253:
138:
54:
448:
219:
215:
59:
427:
192:
are far lower in the control region than elsewhere. This phenomenon completely defies explanation at present.
121:
110:
42:
258:
205:
130:
386:
268:
165:
458:
201:
157:
351:
423:
433:
390:
311:
98:
359:
301:
242:
17:
223:
332:
442:
290:"Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation"
173:
333:"Annotated mtDNA reference sequences: revised Cambridge Reference Sequence (rCRS)"
90:
231:
211:
189:
129:
There are two mitochondrial hypervariable regions used in human mitochondrial
106:
214:, hypervariable regions form the antigen-binding site and are found on both
181:
153:
102:
315:
434:
DNA: Forensic and Legal
Applications, Explanation of Hypervariable Regions
185:
363:
306:
289:
161:
94:
222:. They also contribute to the specificity of each antibody. In a
169:
379:
Michael Stein; Paul Zei; Gloria Hwang; Radhika
Breaden (2000).
188:
rate is not markedly lowered, but sequence differences between
26:
230:
segments of each heavy or light chain fold together at the
168:
evolves remarkably slowly. Even functional mitochondrial
78:
A highly polymorphic nuclear or mitochondrial DNA region
40:It has been suggested that this article should be
8:
374:
372:
426:at the U.S. National Library of Medicine
305:
120:
280:
288:van Oven M, Kayser M (February 2009).
184:protacanthopterygian, control region
7:
234:to form an antigen binding pocket.
49:Hypervariable region (nucleic acid)
264:Human mitochondrial DNA haplogroup
239:Complementarity-determining region
25:
382:Cracking The Boards: USMLE Step 1
31:
335:. Retrieved on 4 February 2016.
55:Hypervariable region (antibody)
1:
254:Cambridge Reference Sequence
166:mitochondrial control region
139:Cambridge Reference Sequence
18:Hypervariable control region
475:
236:
199:
428:Medical Subject Headings
131:genealogical DNA testing
89:) is a location within
347:Plecoglossus altivelis
178:Plecoglossus altivelis
156:, for example certain
126:
259:Genealogical DNA test
206:Somatic hypermutation
145:HVR-II) and 438-574 (
124:
46:into articles titled
424:Hypervariable+region
387:The Princeton Review
269:mtDNA control region
83:hypervariable region
454:Genetic engineering
202:V(D)J recombination
158:Protacanthopterygii
364:10.2108/zsj.22.401
307:10.1002/humu.20921
127:
449:Genetic genealogy
137:. In the revised
99:mitochondrial DNA
76:
75:
16:(Redirected from
466:
411:
410:
405:
403:
376:
367:
342:
336:
326:
320:
319:
309:
285:
243:Framework region
71:
68:
35:
34:
27:
21:
474:
473:
469:
468:
467:
465:
464:
463:
439:
438:
420:
415:
414:
401:
399:
397:
378:
377:
370:
366:(HTML abstract)
343:
339:
327:
323:
287:
286:
282:
277:
250:
245:
229:
224:variable domain
208:
200:Main articles:
198:
119:
79:
72:
66:
63:
36:
32:
23:
22:
15:
12:
11:
5:
472:
470:
462:
461:
456:
451:
441:
440:
437:
436:
431:
419:
418:External links
416:
413:
412:
395:
368:
358:(4): 401–410.
337:
321:
300:(2): E386–94.
294:Human Mutation
279:
278:
276:
273:
272:
271:
266:
261:
256:
249:
246:
227:
197:
194:
118:
115:
77:
74:
73:
39:
37:
30:
24:
14:
13:
10:
9:
6:
4:
3:
2:
471:
460:
457:
455:
452:
450:
447:
446:
444:
435:
432:
429:
425:
422:
421:
417:
409:
398:
396:9780375761638
392:
388:
384:
383:
375:
373:
369:
365:
361:
357:
354:
353:
348:
341:
338:
334:
330:
325:
322:
317:
313:
308:
303:
299:
295:
291:
284:
281:
274:
270:
267:
265:
262:
260:
257:
255:
252:
251:
247:
244:
240:
235:
233:
225:
221:
217:
213:
207:
203:
195:
193:
191:
187:
183:
179:
175:
171:
167:
163:
159:
155:
150:
148:
144:
140:
136:
132:
123:
117:Mitochondrial
116:
114:
112:
108:
104:
100:
96:
92:
88:
84:
70:
61:
57:
56:
51:
50:
45:
44:
38:
29:
28:
19:
407:
400:. Retrieved
381:
355:
350:
346:
340:
329:PhyloTree mt
328:
324:
297:
293:
283:
220:heavy chains
209:
177:
151:
146:
142:
128:
86:
82:
80:
64:
53:
47:
41:
402:5 September
154:bony fishes
111:polymorphic
107:nucleotides
91:nuclear DNA
459:Antibodies
443:Categories
352:Zool. Sci.
275:References
237:See also:
232:N-terminus
212:antibodies
196:Antibodies
190:subspecies
182:East Asian
149:HVR-III).
135:haplogroup
103:base pairs
226:, the 3 H
101:in which
67:June 2022
316:18853457
248:See also
186:mutation
152:In some
162:Gadidae
93:or the
60:discuss
430:(MeSH)
393:
314:
180:), an
164:, the
95:D-loop
216:light
170:genes
147:i.e.,
143:i.e.,
43:split
404:2011
391:ISBN
312:PMID
241:and
218:and
204:and
160:and
52:and
360:doi
302:doi
210:In
174:Ayu
105:of
97:of
87:HVR
62:)
58:. (
445::
406:.
389:.
385:.
371:^
356:22
349:.
331:.
310:.
298:30
296:.
292:.
113:.
81:A
362::
318:.
304::
228:V
176:(
85:(
69:)
65:(
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.