22:
1259:
1086:
166:
355:(SDSA) pathway. In the case of double strand breakage, the 3' end is degraded and the longer 5' end invades the contiguous sister chromatid, forming a replication bubble. As this bubble nears the broken DNA, the longer 5' antisense strand again invades the sense strand of this portion of DNA, transcribing a second copy. When replication ends, both tails are reconnected to form two Holliday Junctions, which are then cleaved in a variety of patterns by proteins. An animation of this process can be seen
135:
225:
conformer. In particular, junctions containing the sequence A-CC bridging the junction point appear to strongly prefer the conformer that allows a hydrogen bond to form between the second cytosine and one of the phosphates at the junction point. While most studies have focused on the identities of the four bases nearest to the junction on each arm, it is evident that bases farther out can also affect the observed stacking conformations.
571:
253:
533:
122:
653:
in the
Holliday junction. As each crossover strand reanneals to its original partner strand, it displaces the original complementary strand ahead of it. This causes the Holliday junction to migrate, creating the heteroduplex segments. Depending on which strand was used as a template to repair the other, the four cells resulting from
21:
584:
with the double-helical domains directly side by side, in contrast to their preferred angle of about 60°. The complex can be designed to force the junctions into either a parallel or antiparallel orientation, but in practice the antiparallel variety are more well-behaved, and the parallel version is rarely used.
755:
and the mobile
Holliday junction, but Seeman's insight was that immobile nucleic acid junctions could be created by properly designing the strand sequences to remove symmetry in the assembled molecule, and that these immobile junctions could in principle be combined into rigid crystalline lattices.
691:
DNA. Later in the 1980s, enzymes responsible for initiating the formation of, and binding to, Holliday junctions were identified, although as of 2004 the identification of mammalian
Holliday junction resolvases remained elusive (however, see section "Resolution of Holliday junctions," above for more
558:
rather than as the carriers of genetic information in living cells. The field uses branched DNA structures as fundamental components to create more complex, rationally designed structures. Holliday junctions are thus components of many such DNA structures. As isolated
Holliday junction complexes are
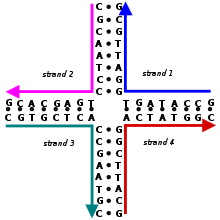
209:
Mg, the electrostatic repulsion is counteracted and the stacked structures predominate. As of 2000, it was not known with certainty whether the electrostatic shielding was the result of site-specific binding of cations to the junction, or the presence of a diffuse collection of the ions in solution.
652:
In the original
Holliday model for homologous recombination, single-strand breaks occur at the same point on one strand of each parental DNA. Free ends of each broken strand then migrate across to the other DNA helix. There, the invading strands are joined to the free ends they encounter, resulting
772:
technique for easily and robustly creating folded DNA structures of arbitrary shape. This method allowed the creation of much larger structures than were previously possible, and which are less technically demanding to design and synthesize. The synthesis of a three-dimensional lattice was finally
398:
hydrolysis to move the junction. The junction must then be resolved into two separate duplexes, restoring either the parental configuration or a crossed-over configuration. Resolution can occur in either a horizontal or vertical fashion during homologous recombination, giving patch products (if in
735:
conformation, because that would place the homologous duplexes in closer alignment to each other. Chemical analysis in the 1980s showed that the junction actually preferred the antiparallel conformation, a finding that was considered controversial, and Robin
Holliday himself initially doubted the
583:
The most common such motif is the double crossover (DX) complex, which contains two
Holliday junctions in close proximity to each other, resulting in a rigid structure that can self-assemble into larger arrays. The structure of the DX molecule forces the Holliday junctions to adopt a conformation
339:
that cleave the junctions, sometimes in a sequence-specific fashion. Such proteins distort the structure of the junction in various ways, often pulling the junction into an unstacked conformation, breaking the central base pairs, and/or changing the angles between the four arms. Other classes are
591:
method, which is used to make larger two- and three-dimensional structures of arbitrary shape. Instead of using individual DX tiles, a single long scaffold strand is folded into the desired shape by a number of short staple strands. When assembled, the scaffold strand is continuous through the
224:
The two possible stacked forms differ in which pairs of the arms are stacked with each other; which of the two dominates is highly dependent on the base sequences nearest to the junction. Some sequences result in an equilibrium between the two conformers, while others strongly prefer a single
243:
RNA Holliday junctions assume an antiparallel stacked conformation at high magnesium concentrations, a perpendicular stacked conformation at moderate concentrations, and rotate into a parallel stacked conformation at low concentrations, while even small calcium ion concentrations favor the
99:
Immobile
Holliday junctions, with asymmetrical sequences that lock the strands in a specific position, were artificially created by scientists to study their structure as a model for natural Holliday junctions. These junctions also later found use as basic structural building blocks in
514:
MSH4 and MSH5 act specifically to facilitate crossovers between homologous chromosomes during meiosis. The MSH4/MSH5 complex binds and stabilizes double
Holliday junctions and promotes their resolution into crossover products. An MSH4 hypomorphic (partially functional) mutant of
595:
Some tile types that retain the
Holliday junction's native 60° angle have been demonstrated. One such array uses tiles containing four Holliday junctions in a parallelogram arrangement. This structure had the benefit of allowing the junction angle to be directly visualized via
232:
process. The rate of branch migration varies dramatically with ion concentration, with single-step times increasing from 0.3 to 0.4 ms with no ions to 270−300 ms with 10 mM Mg. The change in rate is correlated with the formation of the stacked versus the unstacked structures.
519:
showed a 30% genome wide reduction in crossover numbers, and a large number of meioses with non exchange chromosomes. Nevertheless, this mutant gave rise to spore viability patterns suggesting that segregation of non-exchange chromosomes occurred efficiently. Thus in
487:
Double mutants deleted for both MLH3 (major pathway) and MMS4 (minor pathway) showed dramatically reduced crossing over compared to wild-type (6- to 17-fold); however spore viability was reasonably high (62%) and chromosomal disjunction appeared mostly functional.
1620:
Bocker T, Barusevicius A, Snowden T, Rasio D, Guerrette S, Robbins D, Schmidt C, Burczak J, Croce CM, Copeland T, Kovatich AJ, Fishel R (1999). "hMSH5: a human MutS homologue that forms a novel heterodimer with hMSH4 and is expressed during spermatogenesis".
127:
Molecular structure of a stacked Holliday junction, in which the four arms stack into two double-helical domains. Note how the blue and red strands remain roughly helical, while the green and yellow strands cross over between the two
756:
The first theoretical paper proposing this scheme was published in 1982, and the first experimental demonstration of an immobile DNA junction was published the following year. Seeman developed the more rigid double-crossover (DX)
399:
same orientation during double strand break repair) or splice products (if in different orientations during double strand break repair). RuvA and RuvB are branch migration proteins, while RuvC is a junction-resolving enzyme.
644:
with base mismatches between different versions of a single gene. He predicted that the cell would have a mechanism for mismatch repair, which was later discovered. Prior to Holliday's model, the accepted model involved a
476:). The MLH1-MLH3 heterodimer binds preferentially to Holliday junctions. It is an endonuclease that makes single-strand breaks in supercoiled double-stranded DNA. The MLH1-MLH3 heterodimer promotes the formation of
319:
process to occur where the strands move through the junction point. Cleavage, or resolution, of the Holliday junction can occur in two ways. Cleavage of the original set of strands leads to two molecules that may show
578:
triangle complex containing three Holliday junctions, both in isolation (a) and as part of a crystal (b, c). In addition to the two-dimensional array shown, this structure is capable of forming three-dimensional
240:, or break in one of the strands, at the junction point adopt a perpendicular orientation, and always prefer the stacking conformer that places the nick on a crossover strand rather than a helical strand.
193:
to bind to each other, by interactions between the exposed bases. There are three possible conformers: an unstacked (or open-X) form and two stacked forms. The unstacked form dominates in the absence of
315:
The Holliday junctions in homologous recombination are between identical or nearly identical sequences, leading to a symmetric arrangement of sequences around the central junction. This allows a
213:
The unstacked form is a nearly square planar, extended conformation. On the other hand, the stacked conformers have two continuous double-helical domains separated by an angle of about 60° in a
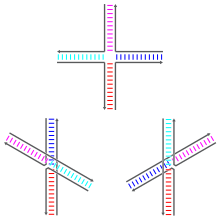
177:: at left, the stacks are red–blue and cyan–magenta, while at right the stacks are red–cyan and blue–magenta. The bases nearest to the junction point determine which stacked isomer dominates.
370:. Both the RecBCD and RecF pathways include a series of reactions known as branch migration, in which single DNA strands are exchanged between two intercrossed molecules of duplex DNA, and
668:
in 1975 introduced the idea of branch migration. Further observations in the 1980s led to the proposal of alternate mechanisms for recombination such as the double-strand break model (by
721:
495:, MUS81 appears to be part of an essential, if not the predominant crossover pathway. The MUS81 pathway also appears to be the predominant crossover pathway in the fission yeast
484:-MMS4, SLX1 and YEN1, respectively, can promote Holliday junction resolution in vivo, absence of all three nucleases has only a modest impact on formation of crossover products.
2176:
Rothemund, Paul W. K. (2006). "Scaffolded DNA origami: from generalized multicrossovers to polygonal networks". In Chen, Junghuei; Jonoska, Natasha; Rozenberg, Grzegorz (eds.).
344:. In prokaryotes, Holliday junction resolvases fall into two families, integrases and nucleases, that are each structurally similar although their sequences are not conserved.
347:
In eukaryotes, two primary models for how homologous recombination repairs double-strand breaks in DNA are the double-strand break repair (DSBR) pathway (sometimes called the
217:
direction. Two of the four strands stay roughly helical, remaining within each of the two double-helical domains, while the other two cross between the two domains in an
374:, in which those two intercrossed molecules of DNA are cut apart and restored to their normal double-stranded state. Homologous recombination occurs in several
660:
Holliday's original model assumed that heteroduplex DNA would be present on both chromosomes, but experimental data on yeast refuted this. An updated model by
2273:
366:
pathway of homologous recombination. Breaks that occur on only one of the two DNA strands, known as single-strand gaps, are thought to be repaired by the
619:
proposed the junction structure that now bears his name as part of his model of homologous recombination in 1964, based on his research on the organisms
328:, while cleavage of the other set of two strands causes the resulting recombinant molecules to show crossover. All products, regardless of cleavage, are
336:
676:, and others) and the single-strand annealing model. A third, the synthesis-dependent strand annealing model, did not involve Holliday junctions.
312:, these usually contain unpaired nucleotides in between the paired double-helical domains, and thus do not strictly adopt the Holliday structure.
386:, recombination occurs through a break-and-rejoin mechanism like in bacteria and eukaryotes. In bacteria, branch migration is facilitated by the
657:
might end up with three copies of one allele and only one of the other, instead of the normal two of each, a property known as gene conversion.
2197:
2103:
1912:
1650:"Variation in crossover frequencies perturb crossover assurance without affecting meiotic chromosome segregation in Saccharomyces cerevisiae"
700:, allowing for more direct study of their physical properties. Much of the early analysis of Holliday junction structure was inferred from
705:
37:. The sequence shown is only one of many possibilities. This is an immobile Holliday junction because the sequences are not symmetrical.
491:
Although MUS81 is a component of a minor crossover pathway in the meiosis of budding yeast, plants and vertebrates, in the protozoan
205:, because of electrostatic repulsion between the negatively charged backbones of the strands. In the presence of at least about 0.1 m
2037:
1112:
1006:
104:, where multiple Holliday junctions can be combined into specific designed geometries that provide molecules with a high degree of
950:
Sung, P; Klein, H (October 2006). "Mechanism of homologous recombination: mediators and helicases take on regulatory functions".
30:
600:. Tiles of three Holliday junctions in a triangular fashion have been used to make periodic three-dimensional arrays for use in
34:
444:, Holliday junctions can be resolved by four different pathways that account for essentially all Holliday junction resolution
1434:"Genetic analysis of mlh3 mutations reveals interactions between crossover promoting factors during meiosis in baker's yeast"
876:
1581:"Cloning and characterization of the human and Caenorhabditis elegans homologs of the Saccharomyces cerevisiae MSH5 gene"
1329:"The Saccharomyces cerevisiae Mlh1-Mlh3 heterodimer is an endonuclease that preferentially binds to Holliday junctions"
2290:
1488:"Mus81 nuclease and Sgs1 helicase are essential for meiotic recombination in a protist lacking a synaptonemal complex"
732:
341:
277:
218:
84:. These junctions usually have a symmetrical sequence and are thus mobile, meaning that the four individual arms may
554:
DNA nanotechnology is the design and manufacture of artificial nucleic acid structures as engineering materials for
423:
407:
744:
structure remained unclear, especially the structure of the junctions is often altered by proteins bound to it.
2262:
627:
604:
of biomolecules. These structures are named for their similarity to structural units based on the principle of
440:
367:
269:
257:
592:
double-helical domains, while the staple strands participate in the Holliday junctions as crossover strands.
597:
1258:
1206:
Boni, MF; de Jong, MD; van Doorn, HR; Holmes, EC; Martin, Darren P. (3 May 2010). Martin, Darren P. (ed.).
1085:
751:
in the early 1980s. A number of natural branched DNA structures were known at the time, including the DNA
692:
recent information). In 1983, artificial Holliday junction molecules were first constructed from synthetic
646:
1100:
537:
473:
395:
309:
1171:
Kowalczykowski SC (2000). "Initiation of genetic recombination and recombination-dependent replication".
544:
domains, on the top and the bottom in this image. This tile is capable of forming two-dimensional arrays.
1277:"Delineation of joint molecule resolution pathways in meiosis identifies a crossover-specific resolvase"
637:
601:
477:
449:
335:
Many proteins are able to recognize or distort the Holliday junction structure. One such class contains
325:
190:
182:
170:
144:
between the double-helical domains, and is stable only in solutions lacking divalent metal ions such as
77:
58:
50:
26:
1380:"Mlh1-Mlh3, a meiotic crossover and DNA mismatch repair factor, is a Msh2-Msh3-stimulated endonuclease"
736:
findings. The antiparallel structure later became widely accepted due to X-ray crystallography data on
165:
1854:
Pan, Keyao; Kim, Do-Nyun; Zhang, Fei; Adendorff, Matthew R.; Yan, Hao; Bathe, Mark (3 December 2014).
2300:
2222:
2132:
1867:
1764:
1219:
665:
375:
868:
725:
701:
680:
352:
301:
206:
173:
of the Holliday junction. The two stacked conformers differ in which sets of two arms are bound by
105:
70:
1943:
1153:
975:
932:
841:
549:
101:
2274:
Analysis of branch migration activities of proteins using synthetic DNA substrates (a protocol)
524:
proper segregation apparently does not entirely depend on crossovers between homologous pairs.
2258:
2238:
2193:
2158:
2099:
2076:
2033:
2009:
1935:
1893:
1836:
1780:
1730:
1679:
1630:
1602:
1561:
1517:
1463:
1411:
1360:
1306:
1247:
1188:
1145:
1108:
1074:
1002:
994:
967:
924:
872:
833:
150:
49:
structure that contains four double-stranded arms joined. These arms may adopt one of several
2027:
510:
proteins form a hetero-oligomeric structure (heterodimer) in yeast and humans. In the yeast
292:
involving Holliday junctions can arise to relieve helical strain in symmetrical sequences in
2230:
2185:
2177:
2148:
2140:
2066:
1999:
1991:
1927:
1883:
1875:
1826:
1818:
1772:
1720:
1710:
1669:
1661:
1592:
1551:
1507:
1499:
1453:
1445:
1401:
1391:
1350:
1340:
1296:
1288:
1237:
1227:
1180:
1137:
1064:
1054:
959:
916:
860:
825:
757:
752:
709:
661:
560:
316:
305:
186:
174:
134:
85:
1822:
717:
693:
633:
621:
391:
321:
214:
54:
2178:
1043:"Comparative and evolutionary analysis of the bacterial homologous recombination systems"
861:
683:
studies in the late 1970s, where the four-arm structure was clearly visible in images of
649:
where the new strand is synthesized directly from parts of the different parent strands.
2226:
2136:
1871:
1768:
1223:
1023:
570:
356:
228:
In junctions with symmetrical sequences, the branchpoint is mobile and can migrate in a
140:
Molecular structure of an unstacked (open-X) Holliday junction. This conformation lacks
2153:
2120:
2004:
1888:
1855:
1831:
1806:
1674:
1649:
1512:
1487:
1458:
1433:
1406:
1379:
1355:
1328:
1301:
1276:
1242:
1207:
1069:
1042:
765:
748:
697:
616:
555:
297:
189:
between the four double-helical arms. Coaxial stacking is the tendency of nucleic acid
93:
66:
1776:
1725:
1698:
1184:
340:
branch migration proteins that increase the exchange rate by orders of magnitude, and
272:, a biological process that increases genetic diversity by shifting genes between two
2284:
1856:"Lattice-free prediction of three-dimensional structure of programmed DNA assemblies"
773:
published by Seeman in 2009, nearly thirty years after he had set out to achieve it.
688:
679:
The first experimental evidence for the structure of the Holliday junction came from
457:
293:
289:
141:
1157:
979:
936:
907:
Liu Y, West S (2004). "Happy Hollidays: 40th anniversary of the Holliday junction".
845:
731:
Initially, geneticists assumed that the junction would adopt a parallel rather than
252:
2295:
1947:
761:
669:
641:
564:
541:
415:
329:
46:
2234:
532:
1995:
1715:
1648:
Krishnaprasad GN, Anand MT, Lin G, Tekkedil MM, Steinmetz LM, Nishant KT (2015).
1232:
1208:"Guidelines for identifying homologous recombination events in influenza a virus"
1059:
760:, suitable for forming two-dimensional lattices, demonstrated in 1998 by him and
1972:
1665:
816:
Lilley, David M. J. (2000). "Structures of helical junctions in nucleic acids".
769:
673:
588:
419:
383:
229:
121:
92:. Additionally, four-arm junctions similar to Holliday junctions appear in some
1292:
1128:
West SC (2003). "Molecular views of recombination proteins and their control".
2268:
829:
605:
575:
411:
285:
273:
237:
81:
62:
1556:
1539:
2189:
1396:
1345:
427:
403:
379:
281:
261:
202:
145:
89:
2242:
2162:
2144:
2080:
2071:
2054:
1939:
1931:
1897:
1840:
1784:
1734:
1683:
1634:
1597:
1580:
1540:"Conserved properties between functionally distinct MutS homologs in yeast"
1521:
1467:
1415:
1364:
1310:
1251:
1192:
1149:
1078:
971:
928:
837:
2013:
1606:
1565:
1449:
1024:"Double-Strand Break Repair via Double Holliday Junctions (Szostak Model)"
1503:
859:
Bloomfield, Victor A.; Crothers, Donald M.; Tinoco, Jr., Ignacio (2000).
713:
195:
1879:
1755:
Seeman, Nadrian C. (June 2004). "Nanotechnology and the double helix".
1378:
Rogacheva MV, Manhart CM, Chen C, Guarne A, Surtees J, Alani E (2014).
747:
The conceptual foundation for DNA nanotechnology was first laid out by
684:
654:
445:
422:. There is controversy over whether homologous recombination occurs in
76:
In biology, Holliday junctions are a key intermediate in many types of
154:
387:
363:
198:
1141:
963:
920:
995:"Chapter 6: Molecular Biology of DNA Replication and Recombination"
569:
531:
481:
251:
164:
88:
through the junction in a specific pattern that largely preserves
20:
2213:
Service, Robert F. (3 June 2011). "DNA nanotechnology grows up".
2096:
Self-assembly: the science of things that put themselves together
2055:"Caution! DNA Crossing: Crystal Structures of Holliday Junctions"
1107:(4th ed.). University of Texas Medical Branch at Galveston.
587:
The DX structural motif is the fundamental building block of the
2121:"Challenges and opportunities for structural DNA nanotechnology"
2119:
Pinheiro, A. V.; Han, D.; Shih, W. M.; Yan, H. (December 2011).
507:
503:
469:
465:
461:
2184:. Natural Computing Series. New York: Springer. pp. 3–21.
1977:
632:
The model provided a molecular mechanism that explained both
2098:. New York: Chapman & Hall/CRC. pp. 201, 242, 259.
1432:
Sonntag Brown M, Lim E, Chen C, Nishant KT, Alani E (2013).
264:, showing the formation and resolution of Holliday junctions
867:. Sausalito, California: University Science Books. p.
640:. Holliday realized that the proposed pathway would create
563:
with multiple Holliday junctions are used to create rigid "
608:, which utilizes members both in tension and compression.
456:
budding yeast, and possibly in mammals, involves proteins
1275:
Zakharyevich, K; Tang, S; Ma, Y; Hunter, N (April 2012).
362:
Double-strand DNA breaks in bacteria are repaired by the
740:
molecules, although as of 2004 the implications for the
480:. While the other three pathways, involving proteins
65:
closest to the junction. The structure is named after
1973:"The Holliday junction on its thirtieth anniversary"
863:
Nucleic acids: structures, properties, and functions
559:
too flexible to assemble into large ordered arrays,
724:methods became available, as well as computational
1538:Pochart P, Woltering D, Hollingsworth NM (1997).
394:protein, molecular motors that use the energy of
1041:Rocha, EPC; Cornet, E; Michel, B (August 2005).
540:contains two Holliday junctions between the two
1750:
1748:
1746:
1744:
1699:"The emergence of complexity: lessons from DNA"
1486:Lukaszewicz A, Howard-Till RA, Loidl J (2013).
993:Hartel, Daniel L.; Jones, Elizabeth W. (2009).
567:" that can then assemble into larger "arrays".
268:The Holliday junction is a key intermediate in
1911:Saccà, Barbara; Niemeyer, Christof M. (2012).
332:in the region of Holliday junction migration.
169:Schematic diagrams of the three base-stacking
1800:
1798:
1796:
1794:
811:
809:
807:
805:
448:. The pathway that produces the majority of
181:Holliday junctions may exist in a variety of
25:Schematic of a Holliday junction showing the
8:
803:
801:
799:
797:
795:
793:
791:
789:
787:
785:
402:There is evidence for recombination in some
2269:Conformational Change of Holliday Junction
1579:Winand NJ, Panzer JA, Kolodner RD (1998).
1533:
1531:
999:Genetics: Analysis of Genetics and Genomes
296:. While four-arm junctions also appear in
2261:at the U.S. National Library of Medicine
2152:
2070:
2003:
1887:
1830:
1724:
1714:
1673:
1596:
1555:
1511:
1457:
1405:
1395:
1354:
1344:
1322:
1320:
1300:
1241:
1231:
1068:
1058:
902:
900:
898:
896:
894:
892:
890:
888:
1966:
1964:
1481:
1479:
1477:
1427:
1425:
2180:Nanotechnology: science and computation
1920:Angewandte Chemie International Edition
1270:
1268:
781:
1327:Ranjha, L; Anand, R; Cejka, P (2014).
1913:"DNA Origami: The Art of Folding DNA"
1823:10.1146/annurev-biochem-060308-102244
1130:Nature Reviews Molecular Cell Biology
952:Nature Reviews Molecular Cell Biology
909:Nature Reviews Molecular Cell Biology
351:) and the synthesis-dependent strand
7:
1001:. Burlington: Jones & Bartlett.
716:footprinting studies. In the 1990s,
468:heterodimer (called MutL gamma) and
284:. They are additionally involved in
73:who proposed its existence in 1964.
14:
2053:Hays FA, Watson J, Ho PS (2003).
1777:10.1038/scientificamerican0604-64
1022:Helleday, T. (20 November 2018).
1257:
1084:
133:
120:
818:Quarterly Reviews of Biophysics
16:Branched nucleic acid structure
1697:Mao, Chengde (December 2004).
1173:Trends in Biochemical Sciences
349:double Holliday junction model
286:repair of double-strand breaks
1:
2235:10.1126/science.332.6034.1140
1811:Annual Review of Biochemistry
1185:10.1016/S0968-0004(00)01569-3
1807:"Nanomaterials based on DNA"
1716:10.1371/journal.pbio.0020431
1438:G3: Genes, Genomes, Genetics
1233:10.1371/journal.pone.0010434
1060:10.1371/journal.pgen.0010015
424:negative-sense ssRNA viruses
408:positive-sense ssRNA viruses
57:salt concentrations and the
1971:Stahl FW (1 October 1994).
1805:Seeman, Nadrian C. (2010).
1666:10.1534/genetics.114.172320
1099:Fleischmann Jr, WR (1996).
536:This double-crossover (DX)
278:site-specific recombination
185:with different patterns of
2317:
1996:10.1093/genetics/138.2.241
1293:10.1016/j.cell.2012.03.023
547:
342:site-specific recombinases
337:junction-resolving enzymes
236:Holliday junctions with a
82:double-strand break repair
2094:Pelesko, John A. (2007).
830:10.1017/S0033583500003590
642:heteroduplex DNA segments
528:Use in DNA nanotechnology
497:Schizosaccharomyces pombe
2263:Medical Subject Headings
2032:. Academic Press. 1971.
1557:10.1074/jbc.272.48.30345
628:Saccharomyces cerevisiae
512:Saccharomyces cerevisiae
441:Saccharomyces cerevisiae
270:homologous recombination
258:homologous recombination
244:antiparallel conformer.
2190:10.1007/3-540-30296-4_1
1397:10.1074/jbc.M113.534644
1346:10.1074/jbc.M113.533810
768:first demonstrated the
598:atomic force microscopy
493:Tetrahymena thermophila
474:Bloom syndrome helicase
2145:10.1038/nnano.2011.187
2072:10.1074/jbc.R300033200
1932:10.1002/anie.201105846
1598:10.1006/geno.1998.5447
580:
545:
538:supramolecular complex
478:crossover recombinants
310:tobacco ringspot virus
265:
183:conformational isomers
178:
171:conformational isomers
38:
2125:Nature Nanotechnology
1860:Nature Communications
1450:10.1534/g3.112.004622
647:copy-choice mechanism
638:chromosomal crossover
602:X-ray crystallography
573:
535:
326:chromosomal crossover
256:The two pathways for
255:
168:
78:genetic recombination
24:
2029:Advances in genetics
1105:Medical Microbiology
290:cruciform structures
2227:2011Sci...332.1140S
2221:(6034): 1140–1143.
2137:2011NatNa...6..763P
2065:(50): 49663–49666.
1872:2014NatCo...5.5578P
1769:2004SciAm.290f..64S
1757:Scientific American
1224:2010PLoSO...510434B
726:molecular modelling
702:gel electrophoresis
681:electron microscopy
302:U1 spliceosomal RNA
300:molecules, such as
248:Biological function
106:structural rigidity
71:molecular biologist
31:secondary structure
2291:Molecular genetics
2259:Holliday+junctions
1880:10.1038/ncomms6578
1504:10.1093/nar/gkt703
581:
550:DNA nanotechnology
546:
266:
179:
102:DNA nanotechnology
39:
35:tertiary structure
2199:978-3-540-30295-7
2105:978-1-58488-687-7
1709:(12): 2036–2038.
1492:Nucleic Acids Res
561:structural motifs
438:In budding yeast
280:events involving
43:Holliday junction
2308:
2247:
2246:
2210:
2204:
2203:
2183:
2173:
2167:
2166:
2156:
2116:
2110:
2109:
2091:
2085:
2084:
2074:
2050:
2044:
2043:
2024:
2018:
2017:
2007:
1981:
1968:
1959:
1958:
1956:
1954:
1917:
1908:
1902:
1901:
1891:
1851:
1845:
1844:
1834:
1802:
1789:
1788:
1752:
1739:
1738:
1728:
1718:
1694:
1688:
1687:
1677:
1645:
1639:
1638:
1617:
1611:
1610:
1600:
1576:
1570:
1569:
1559:
1535:
1526:
1525:
1515:
1498:(20): 9296–309.
1483:
1472:
1471:
1461:
1429:
1420:
1419:
1409:
1399:
1375:
1369:
1368:
1358:
1348:
1324:
1315:
1314:
1304:
1272:
1263:
1262:
1261:
1255:
1245:
1235:
1203:
1197:
1196:
1168:
1162:
1161:
1125:
1119:
1118:
1096:
1090:
1089:
1088:
1082:
1072:
1062:
1038:
1032:
1031:
1019:
1013:
1012:
990:
984:
983:
947:
941:
940:
904:
883:
882:
866:
856:
850:
849:
813:
753:replication fork
722:nucleic acid NMR
710:hydroxyl radical
694:oligonucleotides
317:branch migration
306:hairpin ribozyme
187:coaxial stacking
175:coaxial stacking
157:
137:
124:
80:, as well as in
2316:
2315:
2311:
2310:
2309:
2307:
2306:
2305:
2281:
2280:
2255:
2250:
2212:
2211:
2207:
2200:
2175:
2174:
2170:
2131:(12): 763–772.
2118:
2117:
2113:
2106:
2093:
2092:
2088:
2052:
2051:
2047:
2040:
2026:
2025:
2021:
1975:
1970:
1969:
1962:
1952:
1950:
1915:
1910:
1909:
1905:
1853:
1852:
1848:
1804:
1803:
1792:
1754:
1753:
1742:
1696:
1695:
1691:
1647:
1646:
1642:
1619:
1618:
1614:
1578:
1577:
1573:
1550:(48): 30345–9.
1537:
1536:
1529:
1485:
1484:
1475:
1431:
1430:
1423:
1377:
1376:
1372:
1326:
1325:
1318:
1274:
1273:
1266:
1256:
1205:
1204:
1200:
1170:
1169:
1165:
1142:10.1038/nrm1127
1127:
1126:
1122:
1115:
1098:
1097:
1093:
1083:
1040:
1039:
1035:
1021:
1020:
1016:
1009:
992:
991:
987:
964:10.1038/nrm2008
958:(10): 739–750.
949:
948:
944:
921:10.1038/nrm1502
906:
905:
886:
879:
858:
857:
853:
815:
814:
783:
779:
718:crystallography
666:Charley Radding
634:gene conversion
622:Ustilago maydis
614:
552:
530:
436:
406:, specifically
378:of viruses. In
322:gene conversion
288:. In addition,
250:
163:
162:
161:
160:
159:
149:
138:
130:
129:
125:
114:
17:
12:
11:
5:
2314:
2312:
2304:
2303:
2298:
2293:
2283:
2282:
2277:
2276:
2271:
2266:
2254:
2253:External links
2251:
2249:
2248:
2205:
2198:
2168:
2111:
2104:
2086:
2045:
2038:
2019:
1990:(2): 241–246.
1960:
1903:
1846:
1790:
1740:
1689:
1660:(2): 399–412.
1640:
1612:
1571:
1527:
1473:
1421:
1390:(9): 5664–73.
1370:
1339:(9): 5674–86.
1316:
1264:
1198:
1163:
1120:
1113:
1091:
1033:
1014:
1007:
985:
942:
915:(11): 937–44.
884:
877:
851:
824:(2): 109–159.
780:
778:
775:
766:Paul Rothemund
749:Nadrian Seeman
698:Nadrian Seeman
617:Robin Holliday
613:
610:
574:Diagrams of a
556:nanotechnology
548:Main article:
542:double-helical
529:
526:
435:
432:
416:picornaviruses
330:heteroduplexes
298:functional RNA
294:DNA supercoils
249:
246:
139:
132:
131:
126:
119:
118:
117:
116:
115:
113:
110:
94:functional RNA
67:Robin Holliday
45:is a branched
15:
13:
10:
9:
6:
4:
3:
2:
2313:
2302:
2299:
2297:
2294:
2292:
2289:
2288:
2286:
2279:
2275:
2272:
2270:
2267:
2264:
2260:
2257:
2256:
2252:
2244:
2240:
2236:
2232:
2228:
2224:
2220:
2216:
2209:
2206:
2201:
2195:
2191:
2187:
2182:
2181:
2172:
2169:
2164:
2160:
2155:
2150:
2146:
2142:
2138:
2134:
2130:
2126:
2122:
2115:
2112:
2107:
2101:
2097:
2090:
2087:
2082:
2078:
2073:
2068:
2064:
2060:
2056:
2049:
2046:
2041:
2039:9780080568027
2035:
2031:
2030:
2023:
2020:
2015:
2011:
2006:
2001:
1997:
1993:
1989:
1985:
1979:
1974:
1967:
1965:
1961:
1949:
1945:
1941:
1937:
1933:
1929:
1925:
1921:
1914:
1907:
1904:
1899:
1895:
1890:
1885:
1881:
1877:
1873:
1869:
1865:
1861:
1857:
1850:
1847:
1842:
1838:
1833:
1828:
1824:
1820:
1816:
1812:
1808:
1801:
1799:
1797:
1795:
1791:
1786:
1782:
1778:
1774:
1770:
1766:
1762:
1758:
1751:
1749:
1747:
1745:
1741:
1736:
1732:
1727:
1722:
1717:
1712:
1708:
1704:
1700:
1693:
1690:
1685:
1681:
1676:
1671:
1667:
1663:
1659:
1655:
1651:
1644:
1641:
1636:
1632:
1629:(4): 816–22.
1628:
1624:
1616:
1613:
1608:
1604:
1599:
1594:
1590:
1586:
1582:
1575:
1572:
1567:
1563:
1558:
1553:
1549:
1545:
1544:J. Biol. Chem
1541:
1534:
1532:
1528:
1523:
1519:
1514:
1509:
1505:
1501:
1497:
1493:
1489:
1482:
1480:
1478:
1474:
1469:
1465:
1460:
1455:
1451:
1447:
1443:
1439:
1435:
1428:
1426:
1422:
1417:
1413:
1408:
1403:
1398:
1393:
1389:
1385:
1384:J. Biol. Chem
1381:
1374:
1371:
1366:
1362:
1357:
1352:
1347:
1342:
1338:
1334:
1333:J. Biol. Chem
1330:
1323:
1321:
1317:
1312:
1308:
1303:
1298:
1294:
1290:
1287:(2): 334–47.
1286:
1282:
1278:
1271:
1269:
1265:
1260:
1253:
1249:
1244:
1239:
1234:
1229:
1225:
1221:
1218:(5): e10434.
1217:
1213:
1209:
1202:
1199:
1194:
1190:
1186:
1182:
1179:(4): 156–65.
1178:
1174:
1167:
1164:
1159:
1155:
1151:
1147:
1143:
1139:
1136:(6): 435–45.
1135:
1131:
1124:
1121:
1116:
1114:0-9631172-1-1
1110:
1106:
1102:
1095:
1092:
1087:
1080:
1076:
1071:
1066:
1061:
1056:
1052:
1048:
1047:PLOS Genetics
1044:
1037:
1034:
1029:
1025:
1018:
1015:
1010:
1008:9780763758684
1004:
1000:
996:
989:
986:
981:
977:
973:
969:
965:
961:
957:
953:
946:
943:
938:
934:
930:
926:
922:
918:
914:
910:
903:
901:
899:
897:
895:
893:
891:
889:
885:
880:
874:
870:
865:
864:
855:
852:
847:
843:
839:
835:
831:
827:
823:
819:
812:
810:
808:
806:
804:
802:
800:
798:
796:
794:
792:
790:
788:
786:
782:
776:
774:
771:
767:
763:
759:
754:
750:
745:
743:
739:
734:
729:
727:
723:
719:
715:
711:
707:
703:
699:
695:
690:
689:bacteriophage
686:
682:
677:
675:
671:
667:
663:
662:Matt Meselson
658:
656:
650:
648:
643:
639:
635:
631:
629:
624:
623:
618:
611:
609:
607:
603:
599:
593:
590:
585:
577:
572:
568:
566:
562:
557:
551:
543:
539:
534:
527:
525:
523:
522:S. cerevisiae
518:
517:S. cerevisiae
513:
509:
505:
500:
498:
494:
489:
485:
483:
479:
475:
472:(ortholog of
471:
467:
463:
459:
455:
454:S. cerevisiae
451:
447:
443:
442:
433:
431:
429:
425:
421:
420:coronaviruses
417:
413:
409:
405:
400:
397:
393:
389:
385:
381:
377:
373:
369:
365:
360:
358:
354:
350:
345:
343:
338:
333:
331:
327:
323:
318:
313:
311:
307:
303:
299:
295:
291:
287:
283:
279:
276:, as well as
275:
271:
263:
259:
254:
247:
245:
241:
239:
234:
231:
226:
222:
220:
216:
211:
208:
204:
200:
197:
192:
188:
184:
176:
172:
167:
156:
152:
147:
143:
142:base stacking
136:
123:
111:
109:
107:
103:
97:
95:
91:
87:
83:
79:
74:
72:
68:
64:
60:
56:
53:depending on
52:
51:conformations
48:
44:
36:
32:
28:
27:base sequence
23:
19:
2278:
2218:
2214:
2208:
2179:
2171:
2128:
2124:
2114:
2095:
2089:
2062:
2058:
2048:
2028:
2022:
1987:
1983:
1951:. Retrieved
1926:(1): 58–66.
1923:
1919:
1906:
1863:
1859:
1849:
1814:
1810:
1763:(6): 64–75.
1760:
1756:
1706:
1703:PLOS Biology
1702:
1692:
1657:
1653:
1643:
1626:
1622:
1615:
1591:(1): 69–80.
1588:
1584:
1574:
1547:
1543:
1495:
1491:
1441:
1437:
1387:
1383:
1373:
1336:
1332:
1284:
1280:
1215:
1211:
1201:
1176:
1172:
1166:
1133:
1129:
1123:
1104:
1101:"Chapter 43"
1094:
1050:
1046:
1036:
1027:
1017:
998:
988:
955:
951:
945:
912:
908:
862:
854:
821:
817:
762:Erik Winfree
746:
741:
737:
733:antiparallel
730:
678:
670:Jack Szostak
659:
651:
626:
620:
615:
594:
586:
582:
553:
521:
516:
511:
501:
496:
492:
490:
486:
453:
439:
437:
412:retroviruses
401:
371:
368:RecF pathway
361:
348:
346:
334:
314:
267:
242:
235:
227:
223:
219:antiparallel
215:right-handed
212:
180:
98:
90:base pairing
75:
47:nucleic acid
42:
40:
33:but not the
18:
2301:Chromosomes
2059:J Biol Chem
1953:25 February
1444:(1): 9–22.
770:DNA origami
764:. In 2006,
674:Frank Stahl
589:DNA origami
404:RNA viruses
390:complex or
384:herpesvirus
380:DNA viruses
274:chromosomes
230:random walk
96:molecules.
63:nucleobases
2285:Categories
1623:Cancer Res
1053:(2): e15.
878:0935702490
777:References
606:tensegrity
576:tensegrity
450:crossovers
434:Resolution
372:resolution
282:integrases
262:eukaryotes
191:blunt ends
1817:: 65–87.
1028:Animation
579:crystals.
428:influenza
353:annealing
221:fashion.
112:Structure
2243:21636754
2163:22056726
2081:14563836
1984:Genetics
1940:22162047
1898:25470497
1866:: 5578.
1841:20222824
1785:15195395
1735:15597116
1684:25467183
1654:Genetics
1635:10029069
1585:Genomics
1522:23935123
1468:23316435
1416:24403070
1365:24443562
1311:22500800
1252:20454662
1212:PLOS ONE
1193:10754547
1158:28474965
1150:12778123
1079:16132081
980:30324005
972:16926856
937:24520723
929:15520813
846:40501795
838:11131562
738:in vitro
714:nuclease
382:such as
324:but not
304:and the
201:such as
196:divalent
158:.
128:domains.
59:sequence
2223:Bibcode
2215:Science
2154:3334823
2133:Bibcode
2014:7828807
2005:1206142
1948:8014597
1889:4268701
1868:Bibcode
1832:3454582
1765:Bibcode
1675:4317650
1607:9787078
1566:9374523
1513:3814389
1459:3538346
1407:3937641
1356:3937642
1302:3377385
1243:2862710
1220:Bibcode
1070:1193525
742:in vivo
728:tools.
685:plasmid
655:meiosis
612:History
446:in vivo
308:of the
199:cations
148:. From
2265:(MeSH)
2241:
2196:
2161:
2151:
2102:
2079:
2036:
2012:
2002:
1946:
1938:
1896:
1886:
1839:
1829:
1783:
1733:
1726:535573
1723:
1682:
1672:
1633:
1605:
1564:
1520:
1510:
1466:
1456:
1414:
1404:
1363:
1353:
1309:
1299:
1250:
1240:
1191:
1156:
1148:
1111:
1077:
1067:
1030:. MIT.
1005:
978:
970:
935:
927:
875:
844:
836:
708:, and
418:, and
388:RuvABC
376:groups
364:RecBCD
69:, the
55:buffer
1944:S2CID
1916:(PDF)
1154:S2CID
976:S2CID
933:S2CID
842:S2CID
758:motif
565:tiles
482:MUS81
426:like
410:like
86:slide
2239:PMID
2194:ISBN
2159:PMID
2100:ISBN
2077:PMID
2034:ISBN
2010:PMID
1955:2015
1936:PMID
1894:PMID
1837:PMID
1781:PMID
1731:PMID
1680:PMID
1631:PMID
1603:PMID
1562:PMID
1518:PMID
1464:PMID
1412:PMID
1361:PMID
1307:PMID
1281:Cell
1248:PMID
1189:PMID
1146:PMID
1109:ISBN
1075:PMID
1003:ISBN
968:PMID
925:PMID
873:ISBN
834:PMID
720:and
712:and
706:FRET
687:and
664:and
636:and
625:and
508:MSH5
506:and
504:MSH4
502:The
470:SGS1
466:MLH3
462:MLH1
458:EXO1
392:RecG
357:here
238:nick
155:3CRX
29:and
2296:DNA
2231:doi
2219:332
2186:doi
2149:PMC
2141:doi
2067:doi
2063:278
2000:PMC
1992:doi
1988:138
1978:PDF
1928:doi
1884:PMC
1876:doi
1827:PMC
1819:doi
1773:doi
1761:290
1721:PMC
1711:doi
1670:PMC
1662:doi
1658:199
1593:doi
1552:doi
1548:272
1508:PMC
1500:doi
1454:PMC
1446:doi
1402:PMC
1392:doi
1388:289
1351:PMC
1341:doi
1337:289
1297:PMC
1289:doi
1285:149
1238:PMC
1228:doi
1181:doi
1138:doi
1065:PMC
1055:doi
960:doi
917:doi
869:468
826:doi
696:by
452:in
396:ATP
260:in
151:PDB
61:of
2287::
2237:.
2229:.
2217:.
2192:.
2157:.
2147:.
2139:.
2127:.
2123:.
2075:.
2061:.
2057:.
2008:.
1998:.
1986:.
1982:.
1963:^
1942:.
1934:.
1924:51
1922:.
1918:.
1892:.
1882:.
1874:.
1862:.
1858:.
1835:.
1825:.
1815:79
1813:.
1809:.
1793:^
1779:.
1771:.
1759:.
1743:^
1729:.
1719:.
1705:.
1701:.
1678:.
1668:.
1656:.
1652:.
1627:59
1625:.
1601:.
1589:53
1587:.
1583:.
1560:.
1546:.
1542:.
1530:^
1516:.
1506:.
1496:41
1494:.
1490:.
1476:^
1462:.
1452:.
1440:.
1436:.
1424:^
1410:.
1400:.
1386:.
1382:.
1359:.
1349:.
1335:.
1331:.
1319:^
1305:.
1295:.
1283:.
1279:.
1267:^
1246:.
1236:.
1226:.
1214:.
1210:.
1187:.
1177:25
1175:.
1152:.
1144:.
1132:.
1103:.
1073:.
1063:.
1049:.
1045:.
1026:.
997:.
974:.
966:.
954:.
931:.
923:.
911:.
887:^
871:.
840:.
832:.
822:33
820:.
784:^
704:,
672:,
499:.
460:,
430:.
414:,
359:.
203:Mg
153::
146:Mg
108:.
41:A
2245:.
2233::
2225::
2202:.
2188::
2165:.
2143::
2135::
2129:6
2108:.
2083:.
2069::
2042:.
2016:.
1994::
1980:)
1976:(
1957:.
1930::
1900:.
1878::
1870::
1864:5
1843:.
1821::
1787:.
1775::
1767::
1737:.
1713::
1707:2
1686:.
1664::
1637:.
1609:.
1595::
1568:.
1554::
1524:.
1502::
1470:.
1448::
1442:3
1418:.
1394::
1367:.
1343::
1313:.
1291::
1254:.
1230::
1222::
1216:5
1195:.
1183::
1160:.
1140::
1134:4
1117:.
1081:.
1057::
1051:1
1011:.
982:.
962::
956:7
939:.
919::
913:5
881:.
848:.
828::
630:.
464:-
207:M
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.