852:
637:, a common thiol reductant. In a model system, it was found that the 5,9-bicyclic core of kedarcidin chromophore exists in equilibrium with the corresponding 5,5,6-tricyclic cycloaromatized biradical. The rate of pseudo-first-order decay of this model enediyne is highly dependent on the solvent hydrogen-donor ability, indicating that the hydrogen abstraction step following biradical formation is kinetically significant in the cycloaromatization of the enediyne, as opposed to acyclic systems, where formation of the biradical itself is known to be the rate-limiting step. It is noteworthy that of the solvents examined,
506:
888:). Comparative studies of these biosynthetic apparatus have shown that the enediyne core of these molecules is initiated by a common enzyme, enediyne polyketide synthase (PKS). The polyene product of this enzyme is then divergently elaborated into the 9- or 10-membered cores of the enediynes depending on the specific PKS-associated enzymes present. A convergent biosynthetic strategy is then employed by the producing organisms, whereby the varying peripheral appendages of the enediynes are attached to the core structure to furnish the final product.
773:
489:
824:
24:
978:
923:
746:
677:
421:, chemical degradation, and derivatization experiments enabled the isolation team to identify the key structural features of kedarcidin chromophore, including the enediyne bicyclic core, the ansa-bridging chloropyridyl ring, the mycarose and kedarosamine sugars, and the naphthoamide appendage. However, due to the challenges posed by the complex structure, the initial report had several errors. The bicyclic core proved particularly difficult to deconvolute, as the interpretation of
571:
613:
97:
714:
33:
519:
188:
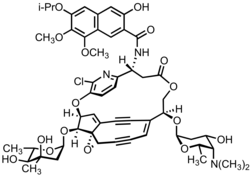
InChI=1S/C53H60ClN3O16/c1-25(2)67-38-18-29-17-35(58)32(20-31(29)46(64-8)47(38)65-9)51(62)56-34-21-42(60)66-24-40(71-43-22-36(59)45(57(6)7)26(3)68-43)28-11-10-12-41-53(73-41)30(14-13-28)19-39(70-37-16-15-33(34)55-50(37)54)49(53)72-44-23-52(5,63)48(61)27(4)69-44/h11,15-20,25-27,34,36,39-41,43-45,48-49,58-59,61,63H,21-24H2,1-9H3,(H,56,62)/b28-11+/t26-,27-,34+,36-,39-,40+,41-,43-,44-,45+,48-,49-,52+,53+/m0/s1
1055:, a calicheamicin-based antibody-drug conjugate for the treatment of non-Hodgkin lymphoma, reinforces the potential of enediynes to find critical use in the treatment of human disease. Thus, the biological potential and complex molecular architecture of kedarcidin may likely inspire further scientific inquiry into this substance, and possibly deliver new ordnance in the war against cancer.
851:
1037:
Owing to its non-specific cytotoxicity, instability under ambient conditions, and relative expense of isolation and manufacture, kedarcidin chromophore has not been investigated rigorously as a therapeutic candidate. However, the recent scientific advances discussed above have served to diminish this
595:
to C12 and consequent opening of the core epoxide has been hypothesized to trigger
Bergman cyclization in kedarcidin chromophore. Nucleophilic activation is thought to diminish the ring strain incurred by formation of the cycloaromatized product, and thus activate kedarcidin chromophore toward DNA
1005:
substructures, respectively. Five genes, KedN1–N5, bear high sequence homology with the enzymes responsible for naphthonate synthesis in neocarzinostatin—consequently, the intermediacy of 3,6,8-trihydroxy-2-naphthoic acid is proposed in kedarcidin biosynthesis. This compound is believed to be
632:
note that the 5,9-fused enediyne core is susceptible to cycloaromatization–reduction in the absence of both thiol "activating agents" and (non-solvent) hydrogen donors. The kedarcidin chromophore aglycone similarly undergoes reductive cycloaromatization at comparable rates irrespective of the
604:
reduction of kedarcidin chromophore induced rapid cycloaromatization and so facilitated studies of the otherwise unstable natural product. Consequently, C12-nucleophilic activation is proposed extensively in review literature as a possible means for triggering the cycloaromatization event
501:
Like other enediynes, kedarcidin chromophore comprises a core structure that forms destructive free radicals, as well as appendages that deliver this "warhead" to its DNA target. Thus, the general mechanism by which kedarcidin chromophore damages DNA is known; however, the details of this
477:. to invert the other aglycone stereocenters as well, affording a revised structure of kedarcidin chromophore that differed only in the relative stereochemistry of the mycarose-bearing carbon, C10. Finally, in 2007, Myers and co-workers synthesized the structure proposed by Hirama
737:
of the C10 hydroxyl would lead to the desired α-face epoxidation product by steric occlusion of the β face of the olefin; however, without a directing group to accelerate the oxidation of a proximal alkene, this hypothetical reaction would likely suffer from poor
855:
Proposed component-based biosynthesis of kedarcidin chromophore. Inversion at C11 of the common enediyne core is invoked to explain the relative stereochemistry of the final product, which differs from most other enediynes, including neocarzinostatin
672:
that focused on the convergent coupling of components with roughly equal chemical complexity. Several of the major challenges of C10-epi-kedarcidin chromophore, as well as the strategies used in addressing these difficulties are discussed below.
546:
With considerable sequence selectivity, kedarcidin chromophore binds and cleaves DNA preferentially at TCCTn-mer sites, producing single-strand breaks. Puzzlingly, while the structure of kedarcidin chromophore is most closely related to that of
701:
installation of the olefin. Without this unsaturation linking the two alkynyl bridges, synthetic intermediates are not disposed toward
Bergman-type decomposition, and risk of decomposition is mitigated. In this case, dehydration of a
542:
sugars of DNA. This generates a carbon-centered free radical on DNA, which undergoes oxidation by molecular oxygen. The resulting peroxide decomposes to form single- or double-stranded breaks in DNA, ultimately leading to cell death.
1050:
therapies, toxicity liabilities may be mitigated through targeted delivery of this potent cytotoxin, potentially enabling efficient therapies that use minimal quantities of this complex material. The recent development of
847:
substructure has not been identified in any other known natural product; and despite its seeming simplicity, little literature precedence exists for the biosynthesis of the isopropoxy substituent of the naphthonate group.
425:
correlations led the researchers to misassign the relative stereochemistry of the core stereotetrad. Moreover, as global absolute chemistry was assigned on the basis of NOE correlations between the stereodefined
835:
The means by which bacteria construct enediynes like kedarcidin continues to motivate research. Kedarcidin chromophore, beyond the carbocyclic core it shares with other enediynes, presents additional
761:. In the first incarnation, hydride delivery to a cyclic tetrayne was guided by aluminum coordination to a proximal alkoxide, thus generating the desired enediyne core in one step via two successive
780:
The bicyclic core of C10-epi-kedarcidin chromophore was prepared by the sequential application of three carbon-carbon bond forming reactions, as shown in the retrosynthetic schematic above. First, a
729:. sought to install the epoxide functionality syn to the adjacent C10 hydroxyl group. This was accomplished by vanadium-catalyzed epoxidation directed by the C10 hydroxyl group. Had the natural C10-(
628:
calculations show that the C1–C12 double bond in the bicyclic core imparts a considerable amount of ring strain (ca. 14 kcal·mol) to the tricycle formed upon
Bergman cyclization–reduction, Hirama
574:
Borohydride-induced tandem epoxide opening–Bergman cycloaromatization of kedarcidin chromophore. Analogous nucleophilic "bioactivation" was initially implicated in the mechanism of action as well.
505:
772:
481:.; the corresponding NMR spectroscopic data were distinct from that of the natural product, leading the Myers group to revise the stereochemistry of the mycarose-bearing carbon to 10-(
453:. In 1997, en route to the originally reported structure, researchers under the direction of Masahiro Hirama discovered that the spectroscopic data of the proposed chloroazatyrosyl (
375:
damage, however, kedarcidin is capable of harming tumor cells, as well. Kedarcidin is thus the subject of scientific research, both for its structural complexity as well as its
757:
Myers and co-workers have pioneered the application of transannular anionic cyclization reactions in the synthesis of the 5,9-fused bicyclic core of kedarcidin chromophore and
664:
In 2007, Myers and co-workers at
Harvard University reported the synthesis of C10-epi-kedarcidin chromophore, corresponding to the 1997 revised structure advanced by Hirama
612:
1468:
Ahlert, J.; Shepard, E.; Lomovskaya, N.; Zazopoulos, E.; Staffa, A.; Bachmann, B. O.; Huang, K, Fonstein, L.; Czisny, A.; Whitwam, R. E.; Farnet, C. M.; Thorson, T. S.
620:
Recent evidence suggests that spontaneous cycloaromatization of kedarcidin chromophore is competitive with nucleophilic bioactivation, if not the predominant mechanism
860:
The biosynthetic gene clusters encoding the biological machinery responsible for producing enediynes have been cloned and characterized for five 9-membered enediynes (
839:
puzzles: The relative stereochemistry of the groups appended to the carbocyclic core of kedarcidin chomophore differs from that of closely related enediynes; the (
399:
bound chromophore from its apoprotein host. This isolate—kedarcidin chromophore—decomposed readily under ambient conditions and was shown to possess cytotoxicity (
922:
49:
N-oxy-14-oxy-11-oxo-4,12,20-trioxa-7-azapentacyclohexacosa-1,5,7,15,25-pentaen-17,22-diyn-9-yl]-3-hydroxy-7,8-dimethoxy-6-propan-2-yloxynaphthalene-2-carboxamide
977:
930:
The 2-aza-β-tyrosine subunit of kedarcidin chromophore is altogether unknown in any other natural product; this lack of precedence frustrates any attempt at
204:
1076:
Leet, J. E.; Schroeder, D. R.; Langley, D. R.; Colson, K. L.; Huang, S.; Klohr, S. E.; Lee, M. S.; Golik, J.; Hofstead, S. J.; Doyle, T. W.; Matson, J. A.
934:
identification of the genes responsible for synthesizing this structure. However, six genes are conserved among the biosynthetic clusters of kedarcidin,
391:
indicated the presence of a DNA-damaging chromoprotein in the fermentation broth of an
Actinomycete strain. The involvement of a non-peptidic
212:(chromophore): C1((C(O1)O\2COC(=O)C(c3ccc(c(n3)Cl)O4C=C5C#C/C2=C\C#C65(4O7C(((O7)C)O)(C)O)O6)NC(=O)c8cc9c(cc8O)cc(c(c9OC)OC)OC(C)C)O)N(C)C
1379:
Liu, W.; Nonaka, K.; Nie, L.; Zhang, J.; Christenson, S. D.; Bae, J.; Van Lanen, S. G.; Zazopoulos, E.; Farnet, C. M.; Yang, C. F.; Shen, B.
434:, the errors of the stereotetrad propagated to the other two stereocenters of the aglycone. Connectivity of the naphthoamide group to the
784:
was carried out between a bromovinyl electrophile and alkynyl nucleophile; ring closure to give a cyclic triyne was then accomplished by
1570:
371:
appendages—kedarcidin was likely evolved to kill bacteria that compete with the producing organism. Because it achieves this by causing
559:. To this end, kedarcidin chromophore–induced DNA cleavage is diminished by the addition of divalent cations such as Ca and Mg, which
1488:(a) Zazopoulos, E.; Huang, K.; Staffa, A.; Liu, W.; Bachmann, B. O.; Nonaka, K.; Ahlert, J.; Thorson, J. S.; Shen, B.; Farnet, C. M.
809:
179:
900:
656:
independently remark that deoxyribose 4'-hydrogen abstraction is most likely operative in kedarcidin chromophore bioactivity.
295:
985:
Insight into the biosynthesis of the isopropoxy-2-naphthonate appendage was similarly gained by comparative analysis of the
563:
bind the naphthoic acid group of kedarcidin chromophore and thus lessen its affinity for DNA. Competition experiments with
1665:
322:
1705:
1700:
1690:
971:
896:
555:
enediyne antitumor antibiotic. The naphthoic acid substructure has been implicated in DNA binding, likely through
1544:
Kedarcidin, a new chromoprotein antitumor antibiotic. II. Isolation, purification and physico-chemical properties
805:
556:
422:
1448:
Lohman, J. R.; Huang, S.-X.; Horsman, G. P.; Dilfer, P. E.; Huang, T.; Chen, Y.; Wendt-Pienkowski, E.; Shen, B.
1675:
488:
958:-tyrosine. This α-amino acid is thus believed to be converted to the corresponding β-amino acid by KedY4, an
1563:
1047:
813:
680:
Retrosynthetic approach to 10-epi-kedarcidin chromophore employed by Ren, Hogan, Anderson, and Myers (2007).
669:
625:
1046:
routes toward scalable kedarcidin production are now within reach. Moreover, with the rising popularity of
23:
919:
led to the formation of a signature heptaene product previously implicated in enediyne core biosynthesis.
765:–type cyclizations. Later-generation syntheses of the core intercept this cascade cyclization, relying on
567:, a known binder of the DNA minor groove, indicate that kedarcidin likely binds the minor groove as well.
1685:
1052:
766:
45:
745:
676:
942:—while these later two enediynes do not contain a 2-aza-β-tyrosine subunit, they do feature similar (
781:
698:
388:
954:
to propose a pathway for the synthesis of the corresponding kedarcidin subunit beginning with 2-aza-
570:
804:
The ansa-bridging macrolactone was constructed following the first
Sonogashira coupling, using the
742:, as oxidation of other C–C unsaturations in the molecule would compete with the desired reaction.
690:
527:
396:
61:
1639:
1217:
Zein, N.; Colson, K. L.; Leet, J. E.; Schroeder, D. R.; Solomon, W.; Doyle, T. W.; Casazza, A. M.
1680:
1556:
1039:
601:
538:
biradical activated toward homolytic abstraction of hydrogen from suitable donors, including the
450:
823:
762:
907:, a gene in the cluster that encodes the previously isolated kedarcidin apoprotein, as well as
1695:
1670:
1026:
891:
In 2013, the successful cloning and characterization of the kedarcidin biosynthetic cluster ("
634:
446:
442:
418:
349:
118:
1608:
1499:, 187–190. (b) Liu, W.; Ahlert, J.; Gao, Q.; Wendt-Pienkowski, E.; Shen, B.; Thorson, J. S.
990:
865:
758:
739:
734:
713:
707:
548:
458:
227:
72:
461:
derivative were not consistent with those of the degradation product characterized by Leet
395:
was deduced by UV spectroscopy, and reverse-phase chromatography was used to separate this
1644:
967:
785:
638:
470:
776:
Transannular cyclization in the synthesis of the bicyclic core of kedarcidin chromophore.
769:
on a cyclic vinyl bromide to generate the vinyl anion precursor to the bicyclic product.
808:. This protocol was performed on the gram-scale without diminishing its yield employing
32:
998:
596:
scission. In the isolation and structural characterization studies carried out by Leet
376:
356:
345:
316:
530:, wherein the enediyne portion undergoes spontaneous cycloaromatization to generate a
1659:
1624:
877:
817:
552:
522:
Proposed mechanism of DNA scission by kedarcidin chromophore–induced 4'-H abstraction
338:
693:–reduction decomposition pathways poses a major threat to any proposed synthesis of
518:
473:
derivative was proposed and validated by the Hirama group. This revision led Hirama
165:
1603:
1043:
994:
939:
869:
836:
793:
407:
341:
616:
Ring strain associated with the C1-C12 double bond in kedarcidin chromophore core.
502:
process—particularly the necessity of nucleophilic activation—have been disputed.
359:
that serves to stabilize the toxin in the
Actinomycete. Like other members of the
509:
Equilibrium of kedarcidin chromophore core and
Bergman-cycloaromatized biradical.
1634:
1629:
1023:
885:
881:
642:
588:
539:
392:
363:
class of drugs—so named for the nine-or-ten-membered core structure bearing an
1543:
873:
264:
108:
645:—led to comparatively fast decomposition of the 5,9-fused enediyne scaffold (
703:
564:
560:
966:
cluster. The resulting product is believed to be loaded onto the peptidyl
1579:
1002:
947:
844:
694:
592:
526:
The unifying mechanism of bioactivity in all enediyne antibiotics is the
431:
360:
352:
788:
of two terminal alkynes. The 5,9-fused bicyclic core was established by
551:, the former shares sequence-specificity with the structurally distinct
1014:-methyltransferase. To furnish the unique isopropoxy substituent, Shen
584:
535:
151:
822:
1593:
1399:
Van Lanen, S. G.; Oh, T.-J.; Liu, W.; Wendt-Pienkowski, E.; Shen, B.
959:
935:
861:
368:
364:
387:
Kedarcidin was first discovered in 1992 when bioassays conducted at
315:
Except where otherwise noted, data are given for materials in their
139:
675:
569:
517:
487:
96:
85:
400:
129:
1552:
1006:
oxygenated to the 3,6,7,8-tetrahydroxy derivative, then triply
903:. The identity of this cloned gene cluster was corroborated by
926:
Proposed biosynthesis of the activated aza-β-tyrosine subunit.
697:. Myers and co-workers addressed this liability by late-stage
414:
372:
492:
Evolution of the proposed structure of kedarcidin chromophore
1548:
976:
921:
850:
771:
744:
712:
611:
504:
796:
species that underwent transannular 5-exo-dig cyclization.
981:
Proposed biosynthesis of activated 2-naphthonate subunit.
1165:
1163:
1022:-methylation of the corresponding methoxy group by the
1119:
Ren, F.; Hogan, P. C.; Anderson, A. J.; Myers, A. G.
441:
These errors were later corrected by the independent
1359:
Liu, W.; Christenson, S. D.; Standage, S.; Shen, B.
1277:
Rossiter, B. E.; Verhoeven, T. R.; Sharpless, K. B.
876:, and kedarcidin), and three 10-membered enediynes (
820:
as a base to promote intramolecular esterification.
1617:
1586:
1338:Shiina, I.; Kubota, M.; Oshiumi, H.; Hashizume, M.
725:In targeting 10-epi-kedarcidin chromophore, Myers
438:bridge was also misjudged in the initial report.
970:KedY2 and subsequently chlorinated by KedY3, an
164:
668:. Critical to the success of this endeavor was
71:
733:)-epimer been desired, it is conceivable that
1564:
1142:
1140:
1138:
1136:
8:
1444:
1442:
1440:
1438:
1436:
1115:
1113:
408:HCT-116 human colorectal carcinoma cell line
1072:
1070:
1068:
1571:
1557:
1549:
1419:McGlinchey, R. P.; Nett, M.; Moore, B. S.
117:
15:
685:Inherent instability of the enediyne core
1237:Myers, A. G.; Hurd, A. R.; Hogan, P. C.
1064:
660:Synthesis of epi-kedarcidin chromophore
209:
184:
1213:
1211:
1209:
1191:, 553. (c) Zein, N.; Schroeder, D. R.
895:") was reported by researchers at the
706:alcohol was induced by treatment with
1096:Kawata, S.; Ashizawa, S.; Hirama, M.
191:Key: RSXFZXJOBQZOOM-WXIIGEIKSA-N
7:
1180:, 2103. (b) Xi, Z.; Goldberg, I. H.
383:Discovery and structure elucidation
154:
138:
1169:(a) Smith, A. L.; Nicolaou, K. C.
950:-derived components, leading Shen
355:chromophore (shown) as well as an
14:
1193:Adv. DNA Sequence-Specific Agents
810:2-methyl-6-nitrobenzoic anhydride
753:Construction of the bicyclic core
915:, the co-expression of which in
248:
245:
239:
31:
22:
1501:Proc. Natl. Acad. Sci. U. S. A.
901:University of Wisconsin-Madison
319:(at 25 °C , 100 kPa).
1318:Myers, A. G.; Goldberg, S. D.
1297:Myers, A. G.; Goldberg, S. D.
641:—structurally homologous with
296:Occupational safety and health
254:
233:
1:
1257:Jones, R. R.; Bergman, R. G.
284:Buff-colored amorphous solid
759:neocarzinostatin chromophore
549:neocarzinostatin chromophore
972:flavin adenine dinucleotide
1722:
1219:Proc. Natl. Acad. Sci. USA
897:Scripps Research Institute
800:Ansa-bridging macrolactone
1010:-methylated by KedN1, an
806:Shiina macrolactonization
367:directly attached to two
313:
293:
288:
220:
200:
175:
54:
44:
39:
30:
21:
1519:Gao, Q.; Thorson, J. S.
1146:Iida, K.-I.; Hirama, M.
767:lithium-halogen exchange
735:trialkylsilyl protection
430:-mycarose sugar and the
1048:antibody-drug conjugate
974:-dependent halogenase.
814:4-dimethylaminopyridine
721:Epoxide stereochemistry
689:The instability toward
670:retrosynthetic analysis
579:Nucleophilic activation
514:Free-radical DNA damage
348:in 1992, comprising an
344:first isolated from an
17:Kedarcidin chromophore
1182:Comp. Nat. Prod. Chem.
982:
927:
857:
827:
777:
749:
717:
681:
617:
575:
523:
510:
493:
1521:FEMS Microbiol. Lett.
1320:Angew. Chem. Int. Ed.
1053:inotuzumab ozogamicin
980:
925:
854:
826:
775:
748:
716:
679:
615:
573:
521:
508:
491:
1666:Antineoplastic drugs
989:cluster to those of
782:Sonogashira coupling
389:Bristol-Myers Squibb
342:antitumor antibiotic
163:(chromophore):
137:(chromophore):
116:(chromophore):
95:(chromophore):
70:(chromophore):
1706:Nine-membered rings
1701:Isopropyl compounds
843:)-2-aza-3-chloro-β-
691:Bergman cyclization
528:Bergman cyclization
497:Mechanism of action
309:Cytotoxic, mutagen
276: g·mol
18:
1691:Experimental drugs
983:
928:
858:
828:
778:
750:
718:
682:
618:
602:sodium borohydride
576:
524:
511:
494:
451:Harvard University
445:of researchers at
323:Infobox references
16:
1653:
1652:
1618:10 membered rings
1421:J. Am. Chem. Soc.
1401:J. Am. Chem. Soc.
1279:Tetrahedron Lett.
1259:J. Am. Chem. Soc.
1239:J. Am. Chem. Soc.
1148:J. Am. Chem. Soc.
1121:J. Am. Chem. Soc.
1098:J. Am. Chem. Soc.
1078:J. Am. Chem. Soc.
1027:methyltransferase
997:, enediynes with
957:
945:
635:β-mercaptoethanol
447:Tohoku University
443:synthetic efforts
429:
419:mass spectrometry
331:Chemical compound
329:
328:
98:Interactive image
1713:
1609:Neocarzinostatin
1587:9 membered rings
1573:
1566:
1559:
1550:
1531:
1517:
1511:
1490:Nat. Biotechnol.
1486:
1480:
1466:
1460:
1446:
1431:
1417:
1411:
1397:
1391:
1377:
1371:
1357:
1351:
1336:
1330:
1316:
1310:
1299:Tetrahedron Lett
1295:
1289:
1275:
1269:
1255:
1249:
1235:
1229:
1215:
1204:
1167:
1158:
1144:
1131:
1117:
1108:
1094:
1088:
1074:
1038:last hurdle, as
991:neocarzinostatin
955:
943:
866:neocarzinostatin
792:generation of a
740:regioselectivity
708:Martin sulfurane
652:= 68 min); Zein
427:
406:0.4 ng/ml,
275:
273:
256:
250:
247:
241:
235:
228:Chemical formula
168:
156:
142:
121:
100:
75:
35:
26:
19:
1721:
1720:
1716:
1715:
1714:
1712:
1711:
1710:
1676:Cancer research
1656:
1655:
1654:
1649:
1645:Shishijimicin A
1613:
1582:
1577:
1540:
1535:
1534:
1518:
1514:
1487:
1483:
1467:
1463:
1447:
1434:
1418:
1414:
1398:
1394:
1378:
1374:
1358:
1354:
1337:
1333:
1317:
1313:
1296:
1292:
1276:
1272:
1256:
1252:
1236:
1232:
1216:
1207:
1168:
1161:
1145:
1134:
1118:
1111:
1095:
1091:
1075:
1066:
1061:
1040:fully synthetic
1035:
968:carrier protein
962:encoded in the
833:
802:
786:Glaser coupling
755:
723:
687:
662:
651:
639:tetrahydrofuran
581:
516:
499:
465:. Instead, an (
404:
385:
332:
325:
320:
306:
271:
269:
259:
253:
244:
238:
230:
216:
213:
208:
207:
196:
193:
192:
189:
183:
182:
171:
157:
145:
124:
103:
89:
78:
64:
50:
12:
11:
5:
1719:
1717:
1709:
1708:
1703:
1698:
1693:
1688:
1683:
1678:
1673:
1668:
1658:
1657:
1651:
1650:
1648:
1647:
1642:
1637:
1632:
1627:
1621:
1619:
1615:
1614:
1612:
1611:
1606:
1601:
1596:
1590:
1588:
1584:
1583:
1578:
1576:
1575:
1568:
1561:
1553:
1547:
1546:
1539:
1538:External links
1536:
1533:
1532:
1512:
1510:, 11959–11963.
1481:
1461:
1432:
1412:
1410:, 13082–13094.
1392:
1372:
1352:
1331:
1311:
1290:
1270:
1250:
1230:
1205:
1159:
1132:
1109:
1107:, 12012–12013.
1089:
1063:
1062:
1060:
1057:
1034:
1031:
1018:invoke double
832:
829:
801:
798:
754:
751:
722:
719:
686:
683:
661:
658:
649:
580:
577:
515:
512:
498:
495:
402:
384:
381:
330:
327:
326:
321:
317:standard state
314:
311:
310:
307:
304:
301:
300:
291:
290:
286:
285:
282:
278:
277:
267:
261:
260:
257:
251:
242:
236:
231:
226:
223:
222:
218:
217:
215:
214:
211:
203:
202:
201:
198:
197:
195:
194:
190:
187:
186:
178:
177:
176:
173:
172:
170:
169:
160:
158:
150:
147:
146:
144:
143:
134:
132:
126:
125:
123:
122:
113:
111:
105:
104:
102:
101:
92:
90:
83:
80:
79:
77:
76:
67:
65:
60:
57:
56:
52:
51:
48:
42:
41:
37:
36:
28:
27:
13:
10:
9:
6:
4:
3:
2:
1718:
1707:
1704:
1702:
1699:
1697:
1694:
1692:
1689:
1687:
1684:
1682:
1679:
1677:
1674:
1672:
1669:
1667:
1664:
1663:
1661:
1646:
1643:
1641:
1638:
1636:
1633:
1631:
1628:
1626:
1625:Calicheamicin
1623:
1622:
1620:
1616:
1610:
1607:
1605:
1602:
1600:
1597:
1595:
1592:
1591:
1589:
1585:
1581:
1574:
1569:
1567:
1562:
1560:
1555:
1554:
1551:
1545:
1542:
1541:
1537:
1529:
1525:
1522:
1516:
1513:
1509:
1505:
1502:
1498:
1494:
1491:
1485:
1482:
1478:
1474:
1471:
1465:
1462:
1458:
1454:
1451:
1450:Mol. BioSyst.
1445:
1443:
1441:
1439:
1437:
1433:
1429:
1425:
1422:
1416:
1413:
1409:
1405:
1402:
1396:
1393:
1389:
1385:
1382:
1376:
1373:
1369:
1365:
1362:
1356:
1353:
1349:
1345:
1341:
1340:J. Org. Chem.
1335:
1332:
1328:
1324:
1321:
1315:
1312:
1308:
1304:
1300:
1294:
1291:
1287:
1283:
1280:
1274:
1271:
1267:
1263:
1260:
1254:
1251:
1247:
1243:
1240:
1234:
1231:
1227:
1223:
1220:
1214:
1212:
1210:
1206:
1202:
1198:
1194:
1190:
1186:
1183:
1179:
1175:
1172:
1171:J. Med. Chem.
1166:
1164:
1160:
1156:
1152:
1149:
1143:
1141:
1139:
1137:
1133:
1129:
1125:
1122:
1116:
1114:
1110:
1106:
1102:
1099:
1093:
1090:
1086:
1082:
1079:
1073:
1071:
1069:
1065:
1058:
1056:
1054:
1049:
1045:
1041:
1032:
1030:
1028:
1025:
1021:
1017:
1013:
1009:
1004:
1000:
996:
992:
988:
979:
975:
973:
969:
965:
961:
953:
949:
941:
937:
933:
924:
920:
918:
914:
910:
906:
902:
898:
894:
889:
887:
883:
879:
878:calicheamicin
875:
871:
867:
863:
853:
849:
846:
842:
838:
830:
825:
821:
819:
818:triethylamine
815:
811:
807:
799:
797:
795:
791:
787:
783:
774:
770:
768:
764:
760:
752:
747:
743:
741:
736:
732:
728:
720:
715:
711:
709:
705:
700:
696:
692:
684:
678:
674:
671:
667:
659:
657:
655:
648:
644:
640:
636:
631:
627:
623:
614:
610:
608:
603:
599:
594:
590:
587:
586:
578:
572:
568:
566:
562:
558:
557:intercalation
554:
553:calicheamicin
550:
544:
541:
537:
533:
529:
520:
513:
507:
503:
496:
490:
486:
484:
480:
476:
472:
468:
464:
460:
456:
452:
448:
444:
439:
437:
433:
424:
420:
416:
411:
409:
405:
398:
397:noncovalently
394:
390:
382:
380:
378:
374:
370:
366:
362:
358:
354:
351:
347:
343:
340:
339:chromoprotein
336:
324:
318:
312:
308:
303:
302:
298:
297:
292:
287:
283:
280:
279:
268:
266:
263:
262:
232:
229:
225:
224:
219:
210:
206:
199:
185:
181:
174:
167:
162:
161:
159:
153:
149:
148:
141:
136:
135:
133:
131:
128:
127:
120:
115:
114:
112:
110:
107:
106:
99:
94:
93:
91:
87:
82:
81:
74:
69:
68:
66:
63:
59:
58:
53:
47:
43:
38:
34:
29:
25:
20:
1686:Naphthalenes
1640:Golfomycin A
1604:Maduropeptin
1598:
1527:
1523:
1520:
1515:
1507:
1503:
1500:
1496:
1492:
1489:
1484:
1479:, 1173–1176.
1476:
1472:
1469:
1464:
1456:
1452:
1449:
1430:, 2406–2407.
1427:
1423:
1420:
1415:
1407:
1403:
1400:
1395:
1387:
1383:
1380:
1375:
1370:, 1170–1173.
1367:
1363:
1360:
1355:
1347:
1343:
1339:
1334:
1329:, 2732–2735.
1326:
1322:
1319:
1314:
1309:, 9633–9636.
1306:
1302:
1298:
1293:
1285:
1281:
1278:
1273:
1265:
1261:
1258:
1253:
1248:, 4583–4585.
1245:
1241:
1238:
1233:
1228:, 2822–2826.
1225:
1221:
1218:
1200:
1196:
1192:
1188:
1184:
1181:
1177:
1173:
1170:
1157:, 8875–8876.
1154:
1150:
1147:
1130:, 5381–5383.
1127:
1123:
1120:
1104:
1100:
1097:
1092:
1087:, 8432–8443.
1084:
1080:
1077:
1044:biosynthetic
1036:
1019:
1015:
1011:
1007:
995:maduropeptin
986:
984:
963:
951:
940:maduropeptin
931:
929:
916:
912:
908:
904:
892:
890:
870:maduropeptin
859:
856:chromophore.
840:
837:biosynthetic
834:
831:Biosynthesis
803:
794:vinyllithium
789:
779:
756:
730:
726:
724:
688:
665:
663:
653:
646:
633:presence of
629:
621:
619:
606:
597:
591:addition of
589:nucelophilic
583:
582:
545:
531:
525:
500:
482:
478:
474:
471:β-amino acid
466:
462:
459:α-amino acid
454:
440:
435:
412:
386:
379:properties.
350:ansa-bridged
346:Actinomycete
334:
333:
305:Main hazards
294:
55:Identifiers
1635:Esperamicin
1630:Dynemicin A
1381:Chem. Biol.
1350:, 1822–1830
1024:radical SAM
999:naphthonate
960:aminomutase
882:esperamicin
704:propargylic
699:dehydrative
643:deoxyribose
561:chelatively
540:deoxyribose
413:Subsequent
393:chromophore
299:(OHS/OSH):
281:Appearance
221:Properties
73:143591-04-2
1660:Categories
1599:Kedarcidin
1530:, 105–114.
1459:, 478–491.
1390:, 293–302.
1268:, 660–661.
1059:References
1033:Conclusion
874:sporolides
377:anticancer
357:apoprotein
335:Kedarcidin
265:Molar mass
109:ChemSpider
84:3D model (
62:CAS Number
46:IUPAC name
1681:Enediynes
1580:Enediynes
886:dynemicin
763:5-exo-dig
695:enediynes
593:thiolates
565:netropsin
1696:Lactones
1671:Epoxides
1003:benzoate
948:tyrosine
932:a priori
899:and the
845:tyrosine
624:. While
432:aglycone
361:enediyne
353:enediyne
289:Hazards
119:26286043
1470:Science
1361:Science
1288:, 4733.
1029:KedN5.
917:E. coli
790:in situ
622:in vivo
607:in vivo
600:., C12-
585:In vivo
536:benzyne
369:alkynyl
166:6444256
152:PubChem
1594:C-1027
1203:, 201.
1016:et al.
952:et al.
938:, and
936:C-1027
913:kedE10
884:, and
862:C-1027
816:, and
654:et al.
630:et al.
365:alkene
205:SMILES
140:C21301
40:Names
727:et al
666:et al
598:et al
479:et al
475:et al
463:et al
337:is a
180:InChI
86:JSmol
1524:2008
1504:2003
1493:2003
1473:2002
1453:2013
1424:2008
1404:2007
1384:2005
1364:2002
1344:2004
1323:2000
1303:1998
1282:1979
1262:1972
1242:2002
1222:1993
1197:1998
1185:1999
1174:1996
1151:1995
1124:2007
1101:1997
1081:1993
1042:and
993:and
911:and
909:kedE
905:kedA
532:para
449:and
436:ansa
130:KEGG
1528:282
1508:100
1477:297
1428:130
1408:129
1368:297
1246:124
1155:117
1128:129
1105:119
1085:115
1001:or
987:ked
964:ked
893:ked
626:MM2
485:).
423:NOE
415:NMR
410:).
373:DNA
274:.52
272:030
155:CID
1662::
1526:,
1506:,
1497:21
1495:,
1475:,
1455:,
1435:^
1426:,
1406:,
1388:12
1386:,
1366:,
1348:69
1346:,
1342:,
1327:39
1325:,
1307:39
1305:,
1301:.
1286:20
1284:,
1266:94
1264:,
1244:,
1226:90
1224:,
1208:^
1199:,
1195:,
1187:,
1178:39
1176:,
1162:^
1153:,
1135:^
1126:,
1112:^
1103:,
1083:,
1067:^
946:)-
880:,
872:,
868:,
864:,
812:,
710:.
609:.
469:)-
457:)-
417:,
403:50
401:IC
258:16
246:Cl
243:60
237:53
1572:e
1565:t
1558:v
1457:9
1201:3
1189:7
1020:C
1012:O
1008:O
956:L
944:L
841:R
731:S
650:½
647:t
534:-
483:S
467:R
455:S
428:L
270:1
255:O
252:3
249:N
240:H
234:C
88:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.