253:
230:
127:
152:
259:
158:
1186:
ability to have an inhibitory effect on the RYR2 channel activity. As more Ca binding sites on CASQ2 become occupied, there is an increasing probability of the RYR2 channel being able to open. Eventually, CASQ2 completely dissociates from
Triadin and the RYR2 channel becomes completely uninhibited, although Triadin remains bound to RYR2 at all luminal concentrations of Ca.
1198:
are found in RYR2 or CASQ2 genes, however a third of CPVT patients have no mutations in either of these proteins, making a mutation in
Triadin the most likely cause Because Triadin is necessary in the regulation of Ca release by the RyR channel during cardiac contraction, a mutation that prevents
1185:
via
Triadin. At low luminal Ca concentrations, Triadin is bound to both RYR2 and CASQ2, so that CSQ prevents RYR2 from opening. At high luminal Ca concentrations, Ca binding sites on CASQ2 become occupied with Ca, leading to a weakened interaction between CASQ2 and Triadin. This removes CASQ2's
1917:
Roux-Buisson, N.; Cacheux, M.; Fourest-Lieuvin, A.; Fauconnier, J.; Brocard, J.; Denjoy, I.; Durand, P.; Guicheney, P.; Kyndt, F.; Leenhardt, A.; Le Marec, H.; Lucet, V.; Mabo, P.; Probst, V.; Monnier, N.; Ray, P. F.; Santoni, E.; Tremeaux, P.; Lacampagne, A.; Faure, J.; Lunardi, J.; Marty, I.
1147:(inner compartment of the sarcoplasmic reticulum) section of Triadin has areas of highly charged amino acid residues that act as luminal Ca receptors. Triadin is also able to sense luminal Ca concentrations by mediating interactions between RYR2 and CASQ2. Triadin has several
1174:
and CASQ2 proteins, so that RYR2 channel activity can be regulated by CASQ2. The linkage of RYR2 with CASQ2 occurs via highly charged luminal sections of
Triadin that are characterized as alternating positively and negatively charged amino acids, known as the KEKE motif.
1230:
at position 59 of the TRDN gene (pT59R) causes instability of
Triadin, leading to degradation of the protein. Any of these naturally occurring mutations result in an absence of functional Triadin protein, resulting in CPVT in patients.
1340:
Thevenon, D.; Smida-Rezgui, S.; Chevessier, F.; Groh, S.; Henry-Berger, J.; Romero, N. B.; Villaz, M.; De Waard, M; Marty, I (2003). "Human skeletal muscle triadin: gene organization and cloning of the major isoform, Trisk 51".
1195:
1431:"Purification, primary structure, and immunological characterization of the 26-kDa calsequestrin binding protein (junctin) from cardiac junctional sarcoplasmic reticulum"
2396:
Kim E, Shin DW, Hong CS, et al. (2003). "Increased Ca storage capacity in the sarcoplasmic reticulum by overexpression of HRC (histidine-rich Ca binding protein)".
2135:
Sacchetto R, Turcato F, Damiani E, Margreth A (1999). "Interaction of triadin with histidine-rich Ca-binding protein at the triadic junction in skeletal muscle fibers".
1972:
Taske NL, Eyre HJ, O'Brien RO, et al. (1995). "Molecular cloning of the cDNA encoding human skeletal muscle triadin and its localisation to chromosome 6q22-6q23".
266:
165:
2174:"Localization and characterization of the calsequestrin-binding domain of triadin 1. Evidence for a charged beta-strand in mediating the protein-protein interaction"
1525:"Localization and characterization of the calsequestrin-binding domain of triadin 1. Evidence for a charged beta-strand in mediating the protein-protein interaction"
2425:
Thevenon D, Smida-Rezgui S, Chevessier F, et al. (2003). "Human skeletal muscle triadin: gene organization and cloning of the major isoform, Trisk 51".
2038:"Complex formation between junctin, triadin, calsequestrin, and the ryanodine receptor. Proteins of the cardiac junctional sarcoplasmic reticulum membrane"
1477:"Complex formation between junctin, triadin, calsequestrin, and the ryanodine receptor. Proteins of the cardiac junctional sarcoplasmic reticulum membrane"
1127:. It is a transmembrane protein on the sarcoplasmic reticulum due to a well defined hydrophobic section and it forms a quaternary complex with the cardiac
793:
774:
88:
1863:
Shin, D. W.; Ma, J.; Kim, D. H. (2000). "The asp-rich region at the carboxyl-terminus of calsequestrin binds to Ca and interacts with triadin".
1308:
1290:
1222:
from being translated into the
Triadin protein, or can result in a shortened, nonfunctional Triadin protein. A replacement of the amino acid
2532:
Arvanitis DA, Vafiadaki E, Fan GC, et al. (2007). "Histidine-rich Ca-binding protein interacts with sarcoplasmic reticulum Ca-ATPase".
2613:
252:
1045:
2242:
Shin DW, Ma J, Kim DH (2001). "The asp-rich region at the carboxyl-terminus of calsequestrin binds to Ca and interacts with triadin".
2576:
1052:
229:
1662:
Caswell, A H; Motoike H K; Fan H; Brandt N R (Jan 1999). "Location of ryanodine receptor binding site on skeletal muscle triadin".
1326:
1151:; Trisk 95 and Trisk 51, which are expressed in skeletal muscle, and Trisk 32 (CT1), which is mainly expressed in cardiac muscle.
2456:"Negatively charged amino acids within the intraluminal loop of ryanodine receptor are involved in the interaction with triadin"
1920:"Absence of triadin, a protein of the calcium release complex, is responsible for cardiac arrhythmia with sudden death in human"
1615:"Negatively charged amino acids within the intraluminal loop of ryanodine receptor are involved in the interaction with triadin"
1199:
Triadin from being formed will make CASQ2 unable to inhibit the RYR2 channel activity, allowing Ca leaks and the development of
1277:
1256:
1120:
1808:"The Role of Calsequestrin, Triadin, and Junctin in Conferring Cardiac Ryanodine Receptor Responsiveness to Luminal Calcium"
1273:
2071:
Caswell AH, Motoike HK, Fan H, Brandt NR (1999). "Location of ryanodine receptor binding site on skeletal muscle triadin".
1613:
Lee, Jae Man; Rho Seong-Hwan; Shin Dong Wook; Cho
Chunghee; Park Woo Jin; Eom Soo Hyun; Ma Jianjie; Kim Do Han (Feb 2004).
151:
126:
2588:
1252:
2314:
Hong CS, Ji JH, Kim JP, et al. (2002). "Molecular cloning and characterization of mouse cardiac triadin isoforms".
68:
1160:
265:
164:
2003:"Association of triadin with the ryanodine receptor and calsequestrin in the lumen of the sarcoplasmic reticulum"
1707:"Association of triadin with the ryanodine receptor and calsequestrin in the lumen of the sarcoplasmic reticulum"
258:
157:
838:
76:
819:
2281:"Interaction of HRC (histidine-rich Ca-binding protein) and triadin in the lumen of sarcoplasmic reticulum"
1116:
2356:
1819:
999:
140:
55:
2594:
2582:
1028:
1024:
985:
981:
977:
973:
953:
949:
920:
916:
912:
908:
2102:"Functional interaction of the cytoplasmic domain of triadin with the skeletal ryanodine receptor"
1756:"Functional interaction of the cytoplasmic domain of triadin with the skeletal ryanodine receptor"
1383:"Identification of triadin 1 as the predominant triadin isoform expressed in mammalian myocardium"
2557:
2520:
2267:
2160:
1888:
1128:
100:
2345:"Generation and initial analysis of more than 15,000 full-length human and mouse cDNA sequences"
1020:
989:
945:
924:
634:
positive regulation of cell communication by electrical coupling involved in cardiac conduction
2549:
2512:
2477:
2442:
2413:
2384:
2331:
2302:
2259:
2230:
2195:
2152:
2123:
2088:
2059:
2024:
1989:
1949:
1880:
1845:
1785:
1777:
1736:
1728:
1687:
1679:
1644:
1636:
1595:
1564:
Marty, I.; Fauré, J.; Fourest-Lieuvin, A.; Vassilopoulos, S.; Oddoux, S.; Brocard, J. (2009).
1546:
1498:
1452:
1404:
1358:
48:
2541:
2502:
2467:
2434:
2405:
2374:
2364:
2323:
2292:
2251:
2220:
2185:
2144:
2113:
2080:
2049:
2014:
1981:
1939:
1931:
1872:
1835:
1827:
1767:
1718:
1671:
1626:
1585:
1577:
1536:
1488:
1442:
1394:
1350:
345:
276:
220:
175:
1178:
1144:
1123:. Triadin is a multiprotein family, arising from different processing of the TRDN gene on
320:
96:
2209:"Cardiac hypertrophy and impaired relaxation in transgenic mice overexpressing triadin 1"
2360:
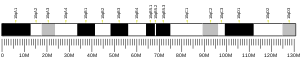
1823:
1944:
1840:
1807:
1590:
1565:
704:
regulation of release of sequestered calcium ion into cytosol by sarcoplasmic reticulum
2438:
2409:
2379:
2344:
2327:
2255:
1876:
1831:
1354:
708:
703:
698:
693:
688:
683:
678:
673:
668:
663:
658:
653:
648:
643:
638:
633:
617:
612:
607:
602:
597:
592:
587:
582:
577:
572:
567:
551:
546:
541:
536:
2607:
1985:
1181:
concentration levels of Ca are sensed by CSQ, and this information is transmitted to
523:
2561:
2164:
1919:
1754:
Groh, S; Marty I; Ottolia M; Prestipino G; Chapel A; Villaz M; Ronjat M (Apr 1999).
80:
2524:
2491:"Global, in vivo, and site-specific phosphorylation dynamics in signaling networks"
2271:
1892:
1124:
338:
117:
1581:
104:
2545:
1313:
National Center for
Biotechnology Information, U.S. National Library of Medicine
1295:
National Center for
Biotechnology Information, U.S. National Library of Medicine
1207:
2507:
2490:
421:
2148:
1475:
Zhang, L.; Kelley, J.; Schmeisser, G.; Kobayashi, Y. M.; Jones, L. R. (1997).
1215:
1211:
237:
134:
84:
2118:
2101:
2054:
2037:
1781:
1772:
1755:
1732:
1683:
1640:
1493:
1476:
1447:
1430:
1399:
1382:
2019:
2002:
1723:
1706:
1429:
Jones, L. R.; Zhang, L.; Sanborn, K.; Jorgensen, A. O.; Kelley., J. (1995).
1227:
1182:
738:
481:
359:
304:
291:
203:
190:
92:
2553:
2516:
2481:
2472:
2455:
2446:
2417:
2388:
2369:
2335:
2306:
2297:
2280:
2263:
2234:
2225:
2208:
2199:
2190:
2173:
2156:
2127:
1953:
1884:
1849:
1789:
1648:
1631:
1614:
1599:
1550:
1541:
1524:
1408:
1362:
674:
positive regulation of ryanodine-sensitive calcium-release channel activity
669:
negative regulation of ryanodine-sensitive calcium-release channel activity
2092:
2063:
2028:
1993:
1740:
1691:
1502:
1456:
1092:
1087:
1935:
1223:
1076:
883:
864:
649:
release of sequestered calcium ion into cytosol by sarcoplasmic reticulum
2084:
1675:
1148:
1140:
850:
805:
1060:
760:
17:
1136:
1190:
Relation to catecholaminergic polymorphic ventricular tachycardia
723:
719:
1219:
1200:
1171:
1164:
1132:
1112:
72:
654:
regulation of release of sequestered calcium ion into cytosol
2343:
Strausberg RL, Feingold EA, Grouse LH, et al. (2003).
1806:
Gyorke, I.; Hester, N.; Jones, L. R.; Gyorke, S. (2004).
1523:
Kobayashi, Y. M.; Alseikhan, B. A.; Jones, L. R. (2000).
328:
689:
regulation of cell communication by electrical coupling
2207:
Kirchhefer U, Neumann J, Baba HA, et al. (2001).
2036:
Zhang L, Kelley J, Schmeisser G, et al. (1997).
1115:
associated with the release of calcium ions from the
493:
699:
regulation of cardiac muscle cell membrane potential
1013:
972:
938:
901:
1269:
1267:
1265:
1248:
1246:
1244:
2489:Olsen JV, Blagoev B, Gnad F, et al. (2006).
1566:"Triadin: what possible function 20 years later?"
275:
174:
2100:Groh S, Marty I, Ottolia M, et al. (1999).
1274:GRCm38: Ensembl release 89: ENSMUSG00000019787
2454:Lee JM, Rho SH, Shin DW, et al. (2004).
2172:Kobayashi YM, Alseikhan BA, Jones LR (2000).
8:
1253:GRCh38: Ensembl release 89: ENSG00000186439
1170:Triadin is required to physically link the
664:endoplasmic reticulum membrane organization
1801:
1799:
969:
734:
583:junctional sarcoplasmic reticulum membrane
519:
370:Skeletal muscle tissue of rectus abdominis
316:
215:
112:
2506:
2471:
2378:
2368:
2296:
2224:
2189:
2117:
2053:
2018:
1943:
1839:
1771:
1722:
1630:
1589:
1540:
1492:
1446:
1398:
2279:Lee HG, Kang H, Kim DH, Park WJ (2001).
1912:
1910:
1908:
1906:
1904:
1902:
1518:
1516:
1514:
1512:
1470:
1468:
1466:
1376:
1374:
1372:
1210:in the TRDN gene can result in an early
1119:triggering muscular contraction through
382:Skeletal muscle tissue of biceps brachii
1424:
1422:
1420:
1418:
1381:Kobayashi, Y. M.; Jones, L. R. (1999).
1240:
537:protein-macromolecule adaptor activity
29:
568:voltage-gated calcium channel complex
280:
241:
236:
179:
138:
133:
7:
684:cytoplasmic microtubule organization
59:, CPVT5, TDN, TRISK, triadin, CARDAR
2534:Am. J. Physiol. Heart Circ. Physiol
1010:
935:
898:
874:
855:
829:
810:
784:
765:
498:
416:
354:
333:
25:
1705:Guo, W; Campbell K P (Apr 1995).
552:transmembrane transporter binding
1986:10.1111/j.1432-1033.1995.258_1.x
659:cellular calcium ion homeostasis
639:regulation of cardiac conduction
264:
257:
251:
228:
163:
156:
150:
125:
1766:(18). United States: 12278–83.
1625:(8). United States: 6994–7000.
1121:calcium-induced calcium release
598:sarcoplasmic reticulum membrane
1717:(16). United States: 9027–30.
1194:Most mutations that result in
573:integral component of membrane
482:More reference expression data
1:
2439:10.1016/S0006-291X(03)00406-6
2427:Biochem. Biophys. Res. Commun
2410:10.1016/S0006-291X(02)02829-2
2398:Biochem. Biophys. Res. Commun
2328:10.1016/S0378-1119(01)00718-1
2256:10.1016/S0014-5793(00)02246-8
1877:10.1016/S0014-5793(00)02246-8
1832:10.1016/S0006-3495(04)74271-X
1355:10.1016/s0006-291x(03)00406-6
1343:Biochem. Biophys. Res. Commun
249:
148:
27:Protein-coding gene in humans
2349:Proc. Natl. Acad. Sci. U.S.A
1582:10.1113/jphysiol.2009.171892
608:sarcoplasmic reticulum lumen
2614:Genes on human chromosome 6
2581:human gene location in the
2546:10.1152/ajpheart.00278.2007
2001:Guo W, Campbell KP (1995).
1327:"Entrez Gene: TRDN triadin"
679:ion transmembrane transport
603:junctional membrane complex
2630:
2593:human gene details in the
2508:10.1016/j.cell.2006.09.026
2137:J. Muscle Res. Cell. Motil
1670:(1). United States: 90–7.
547:signaling receptor binding
452:sternocleidomastoid muscle
1309:"Mouse PubMed Reference:"
1291:"Human PubMed Reference:"
1091:
1086:
1082:
1075:
1059:
1046:Chr 6: 123.22 – 123.64 Mb
1040:
1017:
996:
965:
942:
905:
894:
881:
877:
862:
858:
849:
836:
832:
817:
813:
804:
791:
787:
772:
768:
759:
744:
737:
733:
717:
522:
518:
506:
501:
492:
479:
428:
419:
366:
357:
327:
319:
315:
298:
285:
248:
227:
218:
214:
197:
184:
147:
124:
115:
111:
66:
63:
53:
46:
41:
37:
32:
2119:10.1074/jbc.274.18.12278
2055:10.1074/jbc.272.37.23389
1924:Human Molecular Genetics
1773:10.1074/jbc.274.18.12278
1494:10.1074/jbc.272.37.23389
1448:10.1074/jbc.270.51.30787
1400:10.1074/jbc.274.40.28660
1053:Chr 10: 32.96 – 33.35 Mb
464:tibialis anterior muscle
2149:10.1023/A:1005580609414
2020:10.1074/jbc.270.16.9027
1724:10.1074/jbc.270.16.9027
1218:can either prevent the
1159:TRDN has been shown to
460:vastus lateralis muscle
456:interventricular septum
390:vastus lateralis muscle
2473:10.1074/jbc.M312446200
2370:10.1073/pnas.242603899
2298:10.1074/jbc.M010664200
2226:10.1074/jbc.M006443200
2191:10.1074/jbc.M002091200
1632:10.1074/jbc.M312446200
1542:10.1074/jbc.M002091200
1117:sarcoplasmic reticulum
588:sarcoplasmic reticulum
436:triceps brachii muscle
406:triceps brachii muscle
709:response to bacterium
593:endoplasmic reticulum
243:Chromosome 10 (mouse)
378:gastrocnemius muscle
141:Chromosome 6 (human)
2595:UCSC Genome Browser
2583:UCSC Genome Browser
2361:2002PNAS...9916899M
1824:2004BpJ....86.2121G
1812:Biophysical Journal
1535:(23): 17639–17646.
1487:(37): 23389–23397.
1441:(51): 30787–30796.
1393:(40): 28660–28668.
1226:for the amino acid
1936:10.1093/hmg/dds104
1135:), calsequestrin (
1129:ryanodine receptor
839:ENSMUSG00000019787
644:muscle contraction
627:Biological process
561:Cellular component
530:Molecular function
444:extraocular muscle
2355:(26): 16899–903.
2085:10.1021/bi981306+
1930:(12): 2759–2767.
1676:10.1021/bi981306+
1576:(13): 3117–3121.
1102:
1101:
1098:
1097:
1071:
1070:
1036:
1035:
1007:
1006:
961:
960:
932:
931:
890:
889:
871:
870:
845:
844:
826:
825:
800:
799:
781:
780:
729:
728:
694:heart contraction
514:
513:
510:
509:
488:
487:
475:
474:
413:
412:
311:
310:
210:
209:
16:(Redirected from
2621:
2565:
2528:
2510:
2485:
2475:
2466:(8): 6994–7000.
2450:
2421:
2392:
2382:
2372:
2339:
2310:
2300:
2275:
2238:
2228:
2203:
2193:
2184:(23): 17639–46.
2168:
2131:
2121:
2112:(18): 12278–83.
2096:
2067:
2057:
2048:(37): 23389–97.
2032:
2022:
1997:
1958:
1957:
1947:
1914:
1897:
1896:
1860:
1854:
1853:
1843:
1818:(4): 2121–2128.
1803:
1794:
1793:
1775:
1751:
1745:
1744:
1726:
1702:
1696:
1695:
1659:
1653:
1652:
1634:
1610:
1604:
1603:
1593:
1561:
1555:
1554:
1544:
1520:
1507:
1506:
1496:
1472:
1461:
1460:
1450:
1426:
1413:
1412:
1402:
1378:
1367:
1366:
1337:
1331:
1330:
1323:
1317:
1316:
1305:
1299:
1298:
1287:
1281:
1271:
1260:
1250:
1107:, also known as
1084:
1083:
1055:
1048:
1031:
1011:
1002:
970:
966:RefSeq (protein)
956:
936:
927:
899:
875:
856:
830:
811:
785:
766:
735:
520:
499:
484:
468:digastric muscle
424:
422:Top expressed in
417:
362:
360:Top expressed in
355:
334:
317:
307:
294:
283:
268:
261:
255:
244:
232:
216:
206:
193:
182:
167:
160:
154:
143:
129:
113:
107:
105:TRDN - orthologs
58:
51:
30:
21:
2629:
2628:
2624:
2623:
2622:
2620:
2619:
2618:
2604:
2603:
2601:
2573:
2568:
2531:
2488:
2453:
2424:
2395:
2342:
2313:
2291:(43): 39533–8.
2278:
2241:
2206:
2171:
2134:
2099:
2070:
2035:
2013:(16): 9027–30.
2000:
1974:Eur. J. Biochem
1971:
1967:
1965:Further reading
1962:
1961:
1916:
1915:
1900:
1862:
1861:
1857:
1805:
1804:
1797:
1753:
1752:
1748:
1704:
1703:
1699:
1661:
1660:
1656:
1612:
1611:
1607:
1563:
1562:
1558:
1522:
1521:
1510:
1474:
1473:
1464:
1428:
1427:
1416:
1380:
1379:
1370:
1349:(12): 669–675.
1339:
1338:
1334:
1325:
1324:
1320:
1307:
1306:
1302:
1289:
1288:
1284:
1272:
1263:
1251:
1242:
1237:
1192:
1157:
1149:different forms
1093:View/Edit Mouse
1088:View/Edit Human
1051:
1044:
1041:Location (UCSC)
1027:
1023:
1019:
998:
992:
988:
984:
980:
976:
952:
948:
944:
923:
919:
915:
911:
907:
820:ENSG00000186439
713:
622:
618:plasma membrane
556:
542:protein binding
480:
471:
466:
462:
458:
454:
450:
448:temporal muscle
446:
442:
440:muscle of thigh
438:
434:
420:
409:
404:
400:
396:
392:
388:
386:muscle of thigh
384:
380:
376:
372:
358:
302:
289:
281:
271:
270:
269:
262:
242:
219:Gene location (
201:
188:
180:
170:
169:
168:
161:
139:
116:Gene location (
67:
54:
47:
28:
23:
22:
15:
12:
11:
5:
2627:
2625:
2617:
2616:
2606:
2605:
2599:
2598:
2586:
2572:
2571:External links
2569:
2567:
2566:
2540:(3): H1581–9.
2529:
2486:
2451:
2422:
2393:
2340:
2322:(1–2): 193–9.
2311:
2276:
2239:
2204:
2169:
2132:
2097:
2068:
2033:
1998:
1968:
1966:
1963:
1960:
1959:
1898:
1871:(2): 178–182.
1855:
1795:
1746:
1697:
1654:
1605:
1556:
1508:
1462:
1414:
1368:
1332:
1318:
1300:
1282:
1261:
1239:
1238:
1236:
1233:
1214:. A premature
1206:A deletion of
1191:
1188:
1156:
1153:
1143:proteins. The
1100:
1099:
1096:
1095:
1090:
1080:
1079:
1073:
1072:
1069:
1068:
1066:
1064:
1057:
1056:
1049:
1042:
1038:
1037:
1034:
1033:
1015:
1014:
1008:
1005:
1004:
1000:NP_001242950.1
994:
993:
967:
963:
962:
959:
958:
940:
939:
933:
930:
929:
903:
902:
896:
892:
891:
888:
887:
879:
878:
872:
869:
868:
860:
859:
853:
847:
846:
843:
842:
834:
833:
827:
824:
823:
815:
814:
808:
802:
801:
798:
797:
789:
788:
782:
779:
778:
770:
769:
763:
757:
756:
751:
746:
742:
741:
731:
730:
727:
726:
715:
714:
712:
711:
706:
701:
696:
691:
686:
681:
676:
671:
666:
661:
656:
651:
646:
641:
636:
630:
628:
624:
623:
621:
620:
615:
610:
605:
600:
595:
590:
585:
580:
575:
570:
564:
562:
558:
557:
555:
554:
549:
544:
539:
533:
531:
527:
526:
516:
515:
512:
511:
508:
507:
504:
503:
496:
490:
489:
486:
485:
477:
476:
473:
472:
470:
469:
465:
461:
457:
453:
449:
445:
441:
437:
433:
429:
426:
425:
414:
411:
410:
408:
407:
403:
402:body of tongue
399:
395:
391:
387:
383:
379:
375:
374:biceps brachii
371:
367:
364:
363:
351:
350:
342:
331:
325:
324:
321:RNA expression
313:
312:
309:
308:
300:
296:
295:
287:
284:
279:
273:
272:
263:
256:
250:
246:
245:
240:
234:
233:
225:
224:
212:
211:
208:
207:
199:
195:
194:
186:
183:
178:
172:
171:
162:
155:
149:
145:
144:
137:
131:
130:
122:
121:
109:
108:
65:
61:
60:
52:
44:
43:
39:
38:
35:
34:
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
2626:
2615:
2612:
2611:
2609:
2602:
2596:
2592:
2591:
2587:
2584:
2580:
2579:
2575:
2574:
2570:
2563:
2559:
2555:
2551:
2547:
2543:
2539:
2535:
2530:
2526:
2522:
2518:
2514:
2509:
2504:
2501:(3): 635–48.
2500:
2496:
2492:
2487:
2483:
2479:
2474:
2469:
2465:
2461:
2460:J. Biol. Chem
2457:
2452:
2448:
2444:
2440:
2436:
2433:(2): 669–75.
2432:
2428:
2423:
2419:
2415:
2411:
2407:
2403:
2399:
2394:
2390:
2386:
2381:
2376:
2371:
2366:
2362:
2358:
2354:
2350:
2346:
2341:
2337:
2333:
2329:
2325:
2321:
2317:
2312:
2308:
2304:
2299:
2294:
2290:
2286:
2285:J. Biol. Chem
2282:
2277:
2273:
2269:
2265:
2261:
2257:
2253:
2250:(2): 178–82.
2249:
2245:
2240:
2236:
2232:
2227:
2222:
2219:(6): 4142–9.
2218:
2214:
2213:J. Biol. Chem
2210:
2205:
2201:
2197:
2192:
2187:
2183:
2179:
2178:J. Biol. Chem
2175:
2170:
2166:
2162:
2158:
2154:
2150:
2146:
2143:(4): 403–15.
2142:
2138:
2133:
2129:
2125:
2120:
2115:
2111:
2107:
2106:J. Biol. Chem
2103:
2098:
2094:
2090:
2086:
2082:
2078:
2074:
2069:
2065:
2061:
2056:
2051:
2047:
2043:
2042:J. Biol. Chem
2039:
2034:
2030:
2026:
2021:
2016:
2012:
2008:
2007:J. Biol. Chem
2004:
1999:
1995:
1991:
1987:
1983:
1980:(1): 258–65.
1979:
1975:
1970:
1969:
1964:
1955:
1951:
1946:
1941:
1937:
1933:
1929:
1925:
1921:
1913:
1911:
1909:
1907:
1905:
1903:
1899:
1894:
1890:
1886:
1882:
1878:
1874:
1870:
1866:
1859:
1856:
1851:
1847:
1842:
1837:
1833:
1829:
1825:
1821:
1817:
1813:
1809:
1802:
1800:
1796:
1791:
1787:
1783:
1779:
1774:
1769:
1765:
1761:
1760:J. Biol. Chem
1757:
1750:
1747:
1742:
1738:
1734:
1730:
1725:
1720:
1716:
1712:
1711:J. Biol. Chem
1708:
1701:
1698:
1693:
1689:
1685:
1681:
1677:
1673:
1669:
1665:
1658:
1655:
1650:
1646:
1642:
1638:
1633:
1628:
1624:
1620:
1619:J. Biol. Chem
1616:
1609:
1606:
1601:
1597:
1592:
1587:
1583:
1579:
1575:
1571:
1567:
1560:
1557:
1552:
1548:
1543:
1538:
1534:
1530:
1529:J. Biol. Chem
1526:
1519:
1517:
1515:
1513:
1509:
1504:
1500:
1495:
1490:
1486:
1482:
1481:J. Biol. Chem
1478:
1471:
1469:
1467:
1463:
1458:
1454:
1449:
1444:
1440:
1436:
1435:J. Biol. Chem
1432:
1425:
1423:
1421:
1419:
1415:
1410:
1406:
1401:
1396:
1392:
1388:
1387:J. Biol. Chem
1384:
1377:
1375:
1373:
1369:
1364:
1360:
1356:
1352:
1348:
1344:
1336:
1333:
1328:
1322:
1319:
1314:
1310:
1304:
1301:
1296:
1292:
1286:
1283:
1279:
1275:
1270:
1268:
1266:
1262:
1258:
1254:
1249:
1247:
1245:
1241:
1234:
1232:
1229:
1225:
1221:
1217:
1213:
1209:
1204:
1202:
1197:
1189:
1187:
1184:
1180:
1176:
1173:
1168:
1166:
1162:
1154:
1152:
1150:
1146:
1142:
1138:
1134:
1130:
1126:
1122:
1118:
1114:
1111:, is a human
1110:
1106:
1094:
1089:
1085:
1081:
1078:
1074:
1067:
1065:
1062:
1058:
1054:
1050:
1047:
1043:
1039:
1032:
1030:
1026:
1022:
1016:
1012:
1009:
1003:
1001:
995:
991:
987:
983:
979:
975:
971:
968:
964:
957:
955:
951:
947:
941:
937:
934:
928:
926:
922:
918:
914:
910:
904:
900:
897:
895:RefSeq (mRNA)
893:
886:
885:
880:
876:
873:
867:
866:
861:
857:
854:
852:
848:
841:
840:
835:
831:
828:
822:
821:
816:
812:
809:
807:
803:
796:
795:
790:
786:
783:
777:
776:
771:
767:
764:
762:
758:
755:
752:
750:
747:
743:
740:
736:
732:
725:
721:
716:
710:
707:
705:
702:
700:
697:
695:
692:
690:
687:
685:
682:
680:
677:
675:
672:
670:
667:
665:
662:
660:
657:
655:
652:
650:
647:
645:
642:
640:
637:
635:
632:
631:
629:
626:
625:
619:
616:
614:
611:
609:
606:
604:
601:
599:
596:
594:
591:
589:
586:
584:
581:
579:
576:
574:
571:
569:
566:
565:
563:
560:
559:
553:
550:
548:
545:
543:
540:
538:
535:
534:
532:
529:
528:
525:
524:Gene ontology
521:
517:
505:
500:
497:
495:
491:
483:
478:
467:
463:
459:
455:
451:
447:
443:
439:
435:
431:
430:
427:
423:
418:
415:
405:
401:
398:apex of heart
397:
393:
389:
385:
381:
377:
373:
369:
368:
365:
361:
356:
353:
352:
349:
347:
343:
341:
340:
336:
335:
332:
330:
326:
322:
318:
314:
306:
301:
297:
293:
288:
282:10|10 A4
278:
274:
267:
260:
254:
247:
239:
235:
231:
226:
222:
217:
213:
205:
200:
196:
192:
187:
177:
173:
166:
159:
153:
146:
142:
136:
132:
128:
123:
119:
114:
110:
106:
102:
98:
94:
90:
86:
82:
78:
74:
70:
62:
57:
50:
45:
40:
36:
31:
19:
2600:
2589:
2577:
2537:
2533:
2498:
2494:
2463:
2459:
2430:
2426:
2404:(1): 192–6.
2401:
2397:
2352:
2348:
2319:
2315:
2288:
2284:
2247:
2243:
2216:
2212:
2181:
2177:
2140:
2136:
2109:
2105:
2076:
2073:Biochemistry
2072:
2045:
2041:
2010:
2006:
1977:
1973:
1927:
1923:
1868:
1864:
1858:
1815:
1811:
1763:
1759:
1749:
1714:
1710:
1700:
1667:
1664:Biochemistry
1663:
1657:
1622:
1618:
1608:
1573:
1569:
1559:
1532:
1528:
1484:
1480:
1438:
1434:
1390:
1386:
1346:
1342:
1335:
1321:
1312:
1303:
1294:
1285:
1205:
1193:
1177:
1169:
1158:
1155:Interactions
1125:chromosome 6
1108:
1104:
1103:
1029:NP_001351626
1025:NP_001351625
1018:
997:
986:NP_001242951
982:NP_001242950
978:NP_001242949
974:NP_001238916
954:NM_001364697
950:NM_001364696
943:
921:NM_001256022
917:NM_001256021
913:NM_001256020
909:NM_001251987
906:
882:
863:
837:
818:
792:
773:
753:
748:
344:
337:
202:123,637,093
189:123,216,339
64:External IDs
2079:(1): 90–7.
1208:amino acids
303:33,352,705
290:32,956,550
42:Identifiers
1570:J. Physiol
1280:, May 2017
1259:, May 2017
1235:References
1216:stop codon
1212:stop codon
348:(ortholog)
85:HomoloGene
2244:FEBS Lett
1865:FEBS Lett
1782:0021-9258
1733:0021-9258
1684:0006-2960
1641:0021-9258
1228:Threonine
1021:NP_084002
990:NP_006064
946:NM_029726
925:NM_006073
739:Orthologs
93:GeneCards
2608:Category
2562:12820507
2554:17526652
2517:17081983
2482:14638677
2447:12659871
2418:12480542
2389:12477932
2336:11707337
2307:11504710
2264:11113462
2235:11069905
2200:10748065
2165:21796512
2157:10531621
2128:10212196
1954:22422768
1918:(2012).
1885:11113462
1850:15041652
1790:10212196
1649:14638677
1600:19403623
1551:10748065
1409:10497235
1363:12659871
1276:–
1255:–
1224:Arginine
1161:interact
1077:Wikidata
718:Sources:
578:membrane
2525:7827573
2357:Bibcode
2272:3135618
2093:9890886
2064:9287354
2029:7721813
1994:7588753
1945:3363337
1893:3135618
1841:1304063
1820:Bibcode
1741:7721813
1692:9890886
1591:2727022
1503:9287354
1457:8530521
1278:Ensembl
1257:Ensembl
1179:Luminal
1145:luminal
1141:junctin
1105:Triadin
851:UniProt
806:Ensembl
745:Species
724:QuickGO
613:cytosol
323:pattern
181:6q22.31
81:1924007
49:Aliases
2560:
2552:
2523:
2515:
2480:
2445:
2416:
2387:
2380:139241
2377:
2334:
2305:
2270:
2262:
2233:
2198:
2163:
2155:
2126:
2091:
2062:
2027:
1992:
1952:
1942:
1891:
1883:
1848:
1838:
1788:
1780:
1739:
1731:
1690:
1682:
1647:
1639:
1598:
1588:
1549:
1501:
1455:
1407:
1361:
1139:) and
1063:search
1061:PubMed
884:E9Q9K5
865:Q13061
761:Entrez
494:BioGPS
394:glutes
73:603283
2558:S2CID
2521:S2CID
2268:S2CID
2161:S2CID
1889:S2CID
1163:with
1137:CASQ2
794:76757
775:10345
754:Mouse
749:Human
720:Amigo
432:ankle
346:Mouse
339:Human
286:Start
221:Mouse
185:Start
118:Human
89:38137
2590:TRDN
2578:TRDN
2550:PMID
2513:PMID
2495:Cell
2478:PMID
2443:PMID
2414:PMID
2385:PMID
2332:PMID
2316:Gene
2303:PMID
2260:PMID
2231:PMID
2196:PMID
2153:PMID
2124:PMID
2089:PMID
2060:PMID
2025:PMID
1990:PMID
1950:PMID
1881:PMID
1846:PMID
1786:PMID
1778:ISSN
1737:PMID
1729:ISSN
1688:PMID
1680:ISSN
1645:PMID
1637:ISSN
1596:PMID
1547:PMID
1499:PMID
1453:PMID
1405:PMID
1359:PMID
1220:gene
1201:CPVT
1196:CPVT
1172:RYR2
1165:RYR1
1133:RYR2
1113:gene
1109:TRDN
329:Bgee
277:Band
238:Chr.
176:Band
135:Chr.
97:TRDN
69:OMIM
56:TRDN
33:TRDN
18:TRDN
2542:doi
2538:293
2503:doi
2499:127
2468:doi
2464:279
2435:doi
2431:303
2406:doi
2402:300
2375:PMC
2365:doi
2324:doi
2320:278
2293:doi
2289:276
2252:doi
2248:486
2221:doi
2217:276
2186:doi
2182:275
2145:doi
2114:doi
2110:274
2081:doi
2050:doi
2046:272
2015:doi
2011:270
1982:doi
1978:233
1940:PMC
1932:doi
1873:doi
1869:486
1836:PMC
1828:doi
1768:doi
1764:274
1719:doi
1715:270
1672:doi
1627:doi
1623:279
1586:PMC
1578:doi
1574:587
1537:doi
1533:275
1489:doi
1485:272
1443:doi
1439:270
1395:doi
1391:274
1351:doi
1347:303
1183:RyR
502:n/a
299:End
198:End
101:OMA
77:MGI
2610::
2556:.
2548:.
2536:.
2519:.
2511:.
2497:.
2493:.
2476:.
2462:.
2458:.
2441:.
2429:.
2412:.
2400:.
2383:.
2373:.
2363:.
2353:99
2351:.
2347:.
2330:.
2318:.
2301:.
2287:.
2283:.
2266:.
2258:.
2246:.
2229:.
2215:.
2211:.
2194:.
2180:.
2176:.
2159:.
2151:.
2141:20
2139:.
2122:.
2108:.
2104:.
2087:.
2077:38
2075:.
2058:.
2044:.
2040:.
2023:.
2009:.
2005:.
1988:.
1976:.
1948:.
1938:.
1928:21
1926:.
1922:.
1901:^
1887:.
1879:.
1867:.
1844:.
1834:.
1826:.
1816:86
1814:.
1810:.
1798:^
1784:.
1776:.
1762:.
1758:.
1735:.
1727:.
1713:.
1709:.
1686:.
1678:.
1668:38
1666:.
1643:.
1635:.
1621:.
1617:.
1594:.
1584:.
1572:.
1568:.
1545:.
1531:.
1527:.
1511:^
1497:.
1483:.
1479:.
1465:^
1451:.
1437:.
1433:.
1417:^
1403:.
1389:.
1385:.
1371:^
1357:.
1345:.
1311:.
1293:.
1264:^
1243:^
1203:.
1167:.
722:/
305:bp
292:bp
204:bp
191:bp
99:;
95::
91:;
87::
83:;
79::
75:;
71::
2597:.
2585:.
2564:.
2544::
2527:.
2505::
2484:.
2470::
2449:.
2437::
2420:.
2408::
2391:.
2367::
2359::
2338:.
2326::
2309:.
2295::
2274:.
2254::
2237:.
2223::
2202:.
2188::
2167:.
2147::
2130:.
2116::
2095:.
2083::
2066:.
2052::
2031:.
2017::
1996:.
1984::
1956:.
1934::
1895:.
1875::
1852:.
1830::
1822::
1792:.
1770::
1743:.
1721::
1694:.
1674::
1651:.
1629::
1602:.
1580::
1553:.
1539::
1505:.
1491::
1459:.
1445::
1411:.
1397::
1365:.
1353::
1329:.
1315:.
1297:.
1131:(
223:)
120:)
103::
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.