433:
2159:) and later incorporated some of these into the machinery of genetic coding. Although much circumstantial evidence has been found to suggest that fewer amino acid types were used in the past, precise and detailed hypotheses about which amino acids entered the code in what order are controversial. However, several studies have suggested that Gly, Ala, Asp, Val, Ser, Pro, Glu, Leu, Thr may belong to a group of early-addition amino acids, whereas Cys, Met, Tyr, Trp, His, Phe may belong to a group of later-addition amino acids.
2024:, GUG and UUG are common start codons. In rare cases, certain proteins may use alternative start codons. Surprisingly, variations in the interpretation of the genetic code exist also in human nuclear-encoded genes: In 2016, researchers studying the translation of malate dehydrogenase found that in about 4% of the mRNAs encoding this enzyme the stop codon is naturally used to encode the amino acids tryptophan and arginine. This type of recoding is induced by a high-readthrough stop codon context and it is referred to as
729:
814:
the first position of the codons are more important than changes in the second position on a global scale. The reason may be that charge reversal (from a positive to a negative charge or vice versa) can only occur upon mutations in the first position of certain codons, but not upon changes in the second position of any codon. Such charge reversal may have dramatic consequences for the structure or function of a protein. This aspect may have been largely underestimated by previous studies.
148:
1971:
1992:
universality of the genetic code. However, in his seminal paper on the origins of the genetic code in 1968, Francis Crick still stated that the universality of the genetic code in all organisms was an unproven assumption, and was probably not true in some instances. He predicted that "The code is universal (the same in all organisms) or nearly so". The first variation was discovered in 1979, by researchers studying
201:
Templates and the
Adaptor Hypothesis: A Note for the RNA Tie Club" to the members of the club in January 1955, which "totally changed the way we thought about protein synthesis", as Watson recalled. The hypothesis states that the triplet code was not passed on to amino acids as Gamow thought, but carried by a different molecule, an adaptor, that interacts with amino acids. The adaptor was later identified as tRNA.
9431:
7762:
590:
9455:
8229:
745:
33:
9443:
2035:, ribosomes, single direction reading and translating single codons into single amino acids. The most extreme variations occur in certain ciliates where the meaning of stop codons depends on their position within mRNA. When close to the 3' end they act as terminators while in internal positions they either code for amino acids as in
2166:. A recent hypothesis suggests that the triplet code was derived from codes that used longer than triplet codons (such as quadruplet codons). Longer than triplet decoding would increase codon redundancy and would be more error resistant. This feature could allow accurate decoding absent complex translational machinery such as the
481:" (ORF). For example, the string 5'-AAATGAACG-3' (see figure), if read from the first position, contains the codons AAA, TGA, and ACG ; if read from the second position, it contains the codons AAT and GAA ; and if read from the third position, it contains the codons ATG and AAC. Every sequence can, thus, be read in its
2202:
Stop codons: Codons for translational stops are also an interesting aspect to the problem of the origin of the genetic code. As an example for addressing stop codon evolution, it has been suggested that the stop codons are such that they are most likely to terminate translation early in the case of a
2113:
Given the non-random genetic triplet coding scheme, a tenable hypothesis for the origin of genetic code could address multiple aspects of the codon table, such as absence of codons for D-amino acids, secondary codon patterns for some amino acids, confinement of synonymous positions to third position,
1986:
is in use. The logo shows the 64 codons from left to right, predicted alternatives in red (relative to the standard genetic code). Red line: stop codons. The height of each amino acid in the stack shows how often it is aligned to the codon in homologous protein domains. The stack height indicates the
813:
Note in the table, below, eight amino acids are not affected at all by mutations at the third position of the codon, whereas in the figure above, a mutation at the second position is likely to cause a radical change in the physicochemical properties of the encoded amino acid. Nevertheless, changes in
362:
In a broad academic audience, the concept of the evolution of the genetic code from the original and ambiguous genetic code to a well-defined ("frozen") code with the repertoire of 20 (+2) canonical amino acids is widely accepted. However, there are different opinions, concepts, approaches and ideas,
677:
protein-coding sequences. One reason inheritance of frameshift mutations is rare is that, if the protein being translated is essential for growth under the selective pressures the organism faces, absence of a functional protein may cause death before the organism becomes viable. Frameshift mutations
2198:
combine elements of game theory, natural selection and information channels. Such models have been used to suggest that the first polypeptides were likely short and had non-enzymatic function. Game theoretic models suggested that the organization of RNA strings into cells may have been necessary to
2132:
Stereochemical affinity: the genetic code is a result of a high affinity between each amino acid and its codon or anti-codon; the latter option implies that pre-tRNA molecules matched their corresponding amino acids by this affinity. Later during evolution, this matching was gradually replaced with
2105:
A hypothetical randomly evolved genetic code further motivates a biochemical or evolutionary model for its origin. If amino acids were randomly assigned to triplet codons, there would be 1.5 × 10 possible genetic codes. This number is found by calculating the number of ways that 21 items
2064:
Variant genetic codes used by an organism can be inferred by identifying highly conserved genes encoded in that genome, and comparing its codon usage to the amino acids in homologous proteins of other organisms. For example, the program FACIL infers a genetic code by searching which amino acids in
2177:
approaches model the process of translating the genetic code into corresponding amino acids as an error-prone information channel. The inherent noise (that is, the error) in the channel poses the organism with a fundamental question: how can a genetic code be constructed to withstand noise while
1991:
There was originally a simple and widely accepted argument that the genetic code should be universal: namely, that any variation in the genetic code would be lethal to the organism (although Crick had stated that viruses were an exception). This is known as the "frozen accident" argument for the
200:
The first scientific contribution of the club, later recorded as "one of the most important unpublished articles in the history of science" and "the most famous unpublished paper in the annals of molecular biology", was made by Crick. Crick presented a type-written paper titled "On
Degenerate
2109:
Amino acids that share the same biosynthetic pathway tend to have the same first base in their codons. This could be an evolutionary relic of an early, simpler genetic code with fewer amino acids that later evolved to code a larger set of amino acids. It could also reflect steric and chemical
2182:
models suggest that the genetic code originated as a result of the interplay of the three conflicting evolutionary forces: the needs for diverse amino acids, for error-tolerance and for minimal resource cost. The code emerges at a transition when the mapping of codons to amino acids becomes
366:
Since 2001, 40 non-natural amino acids have been added into proteins by creating a unique codon (recoding) and a corresponding transfer-RNA:aminoacyl – tRNA-synthetase pair to encode it with diverse physicochemical and biological properties in order to be used as a tool to exploring
341:
The three stop codons were named by discoverers
Richard Epstein and Charles Steinberg. "Amber" was named after their friend Harris Bernstein, whose last name means "amber" in German. The other two stop codons were named "ochre" and "opal" in order to keep the "color names" theme.
792:(UCA, UCG, UCC, UCU, AGU, or AGC) codons (difference in the first, second, or third position). A practical consequence of redundancy is that errors in the third position of the triplet codon cause only a silent mutation or an error that would not affect the protein because the
448:(in black: positions 8,525 to 8,580 in the sequence accession NC_012920). There are three possible reading frames in the 5' → 3' forward direction, starting on the first (+1), second (+2) and third position (+3). For each codon (square brackets), the amino acid is given by the
423:
genome that is refactored (all overlaps expanded), recoded (removing the use of three out of 64 codons completely), and further modified to remove the now unnecessary tRNAs and release factors. It is fully viable and grows 1.6× slower than its wild-type counterpart "MDS42".
337:
RNA into protein. Unique triplets promoted the binding of specific tRNAs to the ribosome. Leder and
Nirenberg were able to determine the sequences of 54 out of 64 codons in their experiments. Khorana, Holley and Nirenberg received the Nobel Prize (1968) for their work.
1983:
489:, each producing a possibly distinct amino acid sequence: in the given example, Lys (K)-Trp (W)-Thr (T), Asn (N)-Glu (E), or Met (M)-Asn (N), respectively (when translating with the vertebrate mitochondrial code). When DNA is double-stranded, six possible
2073:
database. Despite the NCBI already providing 27 translation tables, the authors were able to find new 5 genetic code variations (corroborated by tRNA mutations) and correct several misattributions. Codetta was later used to analyze genetic code change in
2106:(20 amino acids plus one stop) can be placed in 64 bins, wherein each item is used at least once. However, the distribution of codon assignments in the genetic code is nonrandom. In particular, the genetic code clusters certain amino acid assignments.
181:
was the first to give a workable scheme for protein synthesis from DNA. He postulated that sets of three bases (triplets) must be employed to encode the 20 standard amino acids used by living cells to build proteins, which would allow a maximum of
1950:. Selenocysteine came to be seen as the 21st amino acid, and pyrrolysine as the 22nd. Both selenocysteine and pyrrolysine may be present in the same organism. Although the genetic code is normally fixed in an organism, the achaeal prokaryote
2055:
The origins and variation of the genetic code, including the mechanisms behind the evolvability of the genetic code, have been widely studied, and some studies have been done experimentally evolving the genetic code of some organisms.
800:
is maintained by equivalent substitution of amino acids; for example, a codon of NUN (where N = any nucleotide) tends to code for hydrophobic amino acids. NCN yields amino acid residues that are small in size and moderate in
2110:
properties that had another effect on the codon during its evolution. Amino acids with similar physical properties also tend to have similar codons, reducing the problems caused by point mutations and mistranslations.
577:. Stop codons are also called "termination" or "nonsense" codons. They signal release of the nascent polypeptide from the ribosome because no cognate tRNA has anticodons complementary to these stop signals, allowing a
2068:
As of
January 2022, the most complete survey of genetic codes is done by Shulgina and Eddy, who screened 250,000 prokaryotic genomes using their Codetta tool. This tool uses a similar approach to FACIL with a larger
685:
Although most mutations that change protein sequences are harmful or neutral, some mutations have benefits. These mutations may enable the mutant organism to withstand particular environmental stresses better than
2979:
The Nobel Prize in
Physiology or Medicine 1959 was awarded jointly to Severo Ochoa and Arthur Kornberg 'for their discovery of the mechanisms in the biological synthesis of ribonucleic acid and deoxyribonucleic
3064:
The Nobel Prize in
Physiology or Medicine 1968 was awarded jointly to Robert W. Holley, Har Gobind Khorana and Marshall W. Nirenberg 'for their interpretation of the genetic code and its function in protein
643:
respectively. Clinically important missense mutations generally change the properties of the coded amino acid residue among basic, acidic, polar or non-polar states, whereas nonsense mutations result in a
3507:"Scientists Created Bacteria With a Synthetic Genome. Is This Artificial Life? - In a milestone for synthetic biology, colonies of E. coli thrive with DNA constructed from scratch by humans, not nature"
2155:
Biosynthetic expansion. The genetic code grew from a simpler earlier code through a process of "biosynthetic expansion". Primordial life "discovered" new amino acids (for example, as by-products of
809:) of 12 variables (4 nucleotides x 3 positions) yields a remarkable correlation (C = 0.95) for predicting the hydropathicity of the encoded amino acid directly from the triplet nucleotide sequence,
764:(redundancy), neither specifies another amino acid (no ambiguity). The codons encoding one amino acid may differ in any of their three positions. For example, the amino acid leucine is specified by
6728:
Ntountoumi, Chrysa; Vlastaridis, Panayotis; Mossialos, Dimitris; Stathopoulos, Constantinos; Iliopoulos, Ioannis; Promponas, Vasilios; Oliver, Stephen G; Amoutzias, Grigoris D (4 November 2019).
2012:). Because viruses must use the same genetic code as their hosts, modifications to the standard genetic code could interfere with viral protein synthesis or functioning. However, viruses such as
197:
from genes. However, the club could have only 20 permanent members to represent each of the 20 amino acids; and four additional honorary members to represent the four nucleotides of DNA.
477:
A reading frame is defined by the initial triplet of nucleotides from which translation starts. It sets the frame for a run of successive, non-overlapping codons, which is known as an "
4588:
Dutilh BE, Jurgelenaite R, Szklarczyk R, van Hijum SA, Harhangi HR, Schmid M, de Wild B, Françoijs KJ, Stunnenberg HG, Strous M, Jetten MS, Op den Camp HJ, Huynen MA (July 2011).
617:, especially if they occur within the protein coding sequence of a gene. Error rates are typically 1 error in every 10–100 million bases—due to the "proofreading" ability of
7600:
2152:
showed that some amino acids have a selective chemical affinity for their codons. Experiments showed that of 8 amino acids tested, 6 show some RNA triplet-amino acid association.
1942:
In some proteins, non-standard amino acids are substituted for standard stop codons, depending on associated signal sequences in the messenger RNA. For example, UGA can code for
1996:. Many slight variants were discovered thereafter, including various alternative mitochondrial codes. These minor variants for example involve translation of the codon UGA as
8185:
2065:
homologous protein domains are most often aligned to every codon. The resulting amino acid (or stop codon) probabilities for each codon are displayed in a genetic code logo.
6671:
Chaliotis, Anargyros; Vlastaridis, Panayotis; Mossialos, Dimitris; Ibba, Michael; Becker, Hubert D.; Stathopoulos, Constantinos; Amoutzias, Grigorios D. (17 February 2017).
404:
In 2016 the first stable semisynthetic organism was created. It was a (single cell) bacterium with two synthetic bases (called X and Y). The bases survived cell division.
2125:-like ribozymes may have had different affinities for amino acids, with codons emerging from another part of the ribozyme that exhibited random variability. Once enough
310:(tRNA), the adapter molecule that facilitates the process of translating RNA into protein. This work was based upon Ochoa's earlier studies, yielding the latter the
407:
In 2017, researchers in South Korea reported that they had engineered a mouse with an extended genetic code that can produce proteins with unnatural amino acids.
8542:
3614:
2098:(RNA enzymes) to proteins as the principal enzymes in cells. In line with the RNA world hypothesis, transfer RNA molecules appear to have evolved before modern
363:
which is the best way to change it experimentally. Even models are proposed that predict "entry points" for synthetic amino acid invasion of the genetic code.
5427:
Neumann H, Wang K, Davis L, Garcia-Alai M, Chin JW (March 2010). "Encoding multiple unnatural amino acids via evolution of a quadruplet-decoding ribosome".
756:
Degeneracy is the redundancy of the genetic code. This term was given by
Bernfield and Nirenberg. The genetic code has redundancy but no ambiguity (see the
7625:
177:
of the
University of Cambridge, hypothesied that information flows from DNA and that there is a link between DNA and proteins. Soviet-American physicist
2031:
Despite these differences, all known naturally occurring codes are very similar. The coding mechanism is the same for all organisms: three-base codons,
7798:
3517:
546:(in bacteria, mitochondria, and plastids). Alternative start codons depending on the organism include "GUG" or "UUG"; these codons normally represent
7086:
Tlusty T (September 2010). "A colorful origin for the genetic code: information theory, statistical mechanics and the emergence of molecular codes".
8178:
7747:
2114:
the small set of only 20 amino acids (instead of a number approaching 64), and the relation of stop codon patterns to amino acid coding patterns.
830:. The codon varies by organism; for example, most common proline codon in E. coli is CCG, whereas in humans this is the least used proline codon.
9297:
8478:
702:
as their genetic material have rapid mutation rates, which can be an advantage, since these viruses thereby evolve rapidly, and thus evade the
7966:
7282:
7261:
5716:
4354:
4327:
4067:
3752:
3642:
3289:
2677:
493:
are defined, three in the forward orientation on one strand and three reverse on the opposite strand. Protein-coding frames are defined by a
311:
5854:
Di Giulio M (October 1989). "The extension reached by the minimization of the polarity distances during the evolution of the genetic code".
9146:
7605:
2217:
737:
805:; NAN encodes average size hydrophilic residues. The genetic code is so well-structured for hydropathicity that a mathematical analysis (
7449:
4369:
Füllen G, Youvan DC (1994). "Genetic
Algorithms and Recursive Ensemble Mutagenesis in Protein Engineering". Complexity International 1.
9286:
8171:
7850:
7830:
7620:
7414:
6410:"Evolution of amino acid frequencies in proteins over deep time: inferred order of introduction of amino acids into the genetic code"
8535:
7938:
7305:
3976:
3959:
3143:
2427:
2563:
3726:
7272:
6226:
Brown, Sean M.; Voráček, Václav; Freeland, Stephen (5 April 2023). "What Would an Alien Amino Acid Alphabet Look Like and Why?".
3159:
Kubyshkin, V.; Budisa, N. (2017). "Synthetic alienation of microbial organisms by using genetic code engineering: Why and how?".
223:
119:. With some exceptions, a three-nucleotide codon in a nucleic acid sequence specifies a single amino acid. The vast majority of
9101:
8092:
8036:
4143:
2199:
prevent "deceptive" use of the genetic code, i.e. preventing the ancient equivalent of viruses from overwhelming the RNA world.
2094:. Under this hypothesis, any model for the emergence of the genetic code is intimately related to a model of the transfer from
4486:
9220:
8031:
5733:"Mathematica function for # possible arrangements of items in bins? – Online Technical Discussion Groups—Wolfram Community"
3960:"Two novel frameshift mutations causing premature stop codons in a patient with the severe form of Maroteaux-Lamy syndrome"
9503:
8119:
8050:
7791:
6898:
Tlusty T (November 2007). "A model for the emergence of the genetic code as a transition in a noisy information channel".
748:
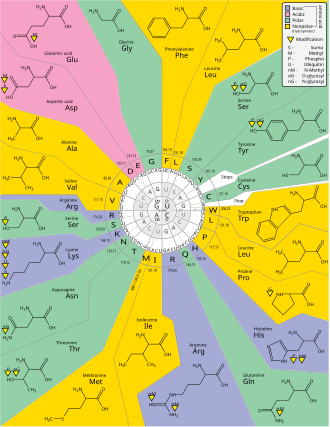
Axes 1, 2, 3 are the first, second, and third positions in the codon. The 20 amino acids and stop codons (X) are shown in
449:
432:
112:
at a time. The genetic code is highly similar among all organisms and can be expressed in a simple table with 64 entries.
9231:
8528:
8468:
7855:
7706:
4464:
1993:
806:
690:
organisms, or reproduce more quickly. In these cases a mutation will tend to become more common in a population through
1956:
can expand its genetic code from 20 to 21 amino acids (by including pyrrolysine) under different conditions of growth.
9493:
9253:
9201:
8680:
7375:
7035:
Sella G, Ardell DH (September 2006). "The coevolution of genes and genetic codes: Crick's frozen accident revisited".
2136:
Optimality: the genetic code continued to evolve after its initial creation, so that the current code maximizes some
2791:"The dependence of cell-free protein synthesis in E. coli upon naturally occurring or synthetic polyribonucleotides"
2242:
Turanov AA, Lobanov AV, Fomenko DE, Morrison HG, Sogin ML, Klobutcher LA, Hatfield DL, Gladyshev VN (January 2009).
9498:
9242:
9131:
8070:
7454:
7251:
6730:"Low complexity regions in the proteins of prokaryotes perform important functional roles and are highly conserved"
4224:
Holland J, Spindler K, Horodyski F, Grabau E, Nichol S, VandePol S (March 1982). "Rapid evolution of RNA genomes".
3700:
9136:
9081:
8252:
7610:
7595:
2099:
162:
2117:
Three main hypotheses address the origin of the genetic code. Many models belong to one of them or to a hybrid:
671:
to be read, which truncates the protein. These mutations may impair the protein's function and are thus rare in
9483:
9111:
8194:
8114:
7902:
7815:
7784:
7615:
7434:
6974:
Tlusty T (February 2008). "Rate-distortion scenario for the emergence and evolution of noisy molecular codes".
6117:"Origin of the genetic code: a testable hypothesis based on tRNA structure, sequence, and kinetic proofreading"
1952:
1932:
757:
217:
97:
9447:
2207:
error. In contrast, some stereochemical molecular models explain the origin of stop codons as "unassignable".
2129:
were coded for, any major random change in the genetic code would have been lethal; hence it became "frozen".
667:. These mutations usually result in a completely different translation from the original, and likely cause a
9411:
9275:
9176:
8646:
8075:
7892:
7877:
5180:"Novel Ciliate Genetic Code Variants Including the Reassignment of All Three Stop Codons to Sense Codons in
3506:
2593:"Crick's Adaptor Hypothesis and the Discovery of Transfer RNA: Experiment Surpassing Theoretical Prediction"
2090:, according to one version of which self-replicating RNA molecules preceded life as we know it. This is the
325:
revealed the code's triplet nature and deciphered its codons. In these experiments, various combinations of
5231:
Lobanov AV, Heaphy SM, Turanov AA, Gerashchenko MV, Pucciarelli S, Devaraj RR, et al. (January 2017).
9310:
9191:
9141:
8685:
8488:
8080:
8005:
7897:
7407:
6278:
2179:
357:
210:
4980:"The functional readthrough extension of malate dehydrogenase reveals a modification of the genetic code"
679:
9435:
9186:
9038:
9016:
8959:
8690:
8611:
8566:
8483:
8473:
7995:
7980:
7860:
7742:
7576:
7541:
7198:"The genetic code is nearly optimal for allowing additional information within protein-coding sequences"
6788:
Freeland SJ, Wu T, Keulmann N (October 2003). "The case for an error minimizing standard genetic code".
5333:
Sengupta S, Higgs PG (June 2015). "Pathways of Genetic Code Evolution in Ancient and Modern Organisms".
3359:
3051:
2966:
1965:
827:
777:
351:
334:
245:
194:
73:
2641:
2006:
species, and translation of CUG as a serine rather than leucine in yeasts of the "CTG clade" (such as
436:
Reading frames in the DNA sequence of a region of the human mitochondrial genome coding for the genes
186:
amino acids. He named this DNA–protein interaction (the original genetic code) as the "diamond code".
9264:
9106:
9086:
8675:
8586:
8493:
8102:
8000:
7918:
7641:
7459:
7328:
7105:
7044:
6993:
6917:
6852:
6797:
6621:
6513:
6458:
6366:
6235:
6190:
6128:
5984:
5918:
5907:"Role of minimization of chemical distances between amino acids in the evolution of the genetic code"
5863:
5820:
5769:
5653:
5342:
5142:
4883:
4655:
4539:
4233:
4178:
4096:
4016:
3871:
3554:
3457:
3398:
3005:
2920:
2861:
2802:
2304:
2188:
2091:
2043:
237:
174:
116:
3660:"Generation of protein isoform diversity by alternative initiation of translation at non-AUG codons"
776:(UUA, UUG, CUU, CUC, CUA, or CUG) codons (difference in the first or third position indicated using
9121:
8653:
8596:
8591:
8297:
7928:
7721:
7711:
7671:
7571:
7551:
7444:
7439:
6600:"A Thermodynamic Basis for Prebiotic Amino Acid Synthesis and the Nature of the First Genetic Code"
6357:
Sengupta S, Higgs PG (2015). "Pathways of genetic code evolution in ancient and modern organisms".
4772:"A fungal phylogeny based on 42 complete genomes derived from supertree and combined gene analysis"
711:
664:
636:
9454:
4705:
3740:
9071:
8774:
8758:
8733:
8728:
8665:
8606:
8244:
8063:
7946:
7681:
7566:
7561:
7536:
7129:
7095:
7068:
7017:
6983:
6941:
6907:
6821:
6653:
6611:
6560:
6482:
6449:
Amirnovin R (May 1997). "An analysis of the metabolic theory of the origin of the genetic code".
6390:
6259:
6097:
5887:
5793:
5452:
5409:
5366:
4752:
4679:
4421:
4040:
3989:
3588:
3512:
2771:
2622:
2488:
2480:
2336:
2174:
543:
478:
299:
233:
156:
4269:"Clonal interference and the periodic selection of new beneficial mutations in Escherichia coli"
2592:
2162:
Natural selection has led to codon assignments of the genetic code that minimize the effects of
728:
397:
residues (UGG codons) with unnatural thienopyrrole-alanine in the genetic code of the bacterium
7149:"What can information-asymmetric games tell us about the context of Crick's 'frozen accident'?"
9488:
9386:
9166:
9161:
9156:
9091:
8856:
8743:
8631:
8601:
8508:
8330:
8147:
7765:
7696:
7686:
7676:
7510:
7400:
7356:
7301:
7278:
7257:
7227:
7178:
7121:
7060:
7009:
6933:
6880:
6813:
6767:
6749:
6710:
6692:
6645:
6637:
6580:
6541:
6474:
6431:
6382:
6339:
6298:
6251:
6208:
6156:
6089:
6054:
6010:
5946:
5879:
5836:
5785:
5712:
5681:
5622:
5573:
5522:
5505:
Chin JW (February 2014). "Expanding and reprogramming the genetic code of cells and animals".
5487:
5444:
5401:
5384:
Xie J, Schultz PG (August 2006). "A chemical toolkit for proteins--an expanded genetic code".
5358:
5315:
5266:
5213:
5160:
5111:
5060:
5009:
4960:
4909:
4852:
4803:
4744:
4671:
4619:
4567:
4508:
4413:
4350:
4323:
4298:
4249:
4206:
4124:
4063:
4032:
3981:
3940:
3899:
3840:
3799:
3748:
3681:
3638:
3580:
3483:
3426:
3340:
3285:
3279:
3260:
3211:
3176:
3139:
3108:
3033:
2948:
2889:
2830:
2763:
2722:
2683:
2673:
2614:
2545:
2527:
2472:
2433:
2423:
2393:
2375:
2328:
2320:
2273:
2137:
691:
628:
624:
535:
524:. The start codon alone is not sufficient to begin the process. Nearby sequences such as the
420:
368:
48:. The nucleotides are abbreviated with the letters A, U, G and C. This is mRNA, which uses U (
4085:"Prevalence of positive selection among nearly neutral amino acid replacements in Drosophila"
2415:
538:
are also required to start translation. The most common start codon is AUG, which is read as
193:, as suggested by Watson, for scientists of different persuasions who were interested in how
9459:
9332:
9151:
9126:
8641:
7726:
7701:
7346:
7336:
7217:
7209:
7168:
7160:
7113:
7052:
7001:
6925:
6870:
6860:
6805:
6757:
6741:
6700:
6684:
6629:
6572:
6531:
6521:
6466:
6421:
6374:
6329:
6290:
6243:
6198:
6146:
6136:
6081:
6044:
6000:
5992:
5936:
5926:
5871:
5828:
5777:
5671:
5661:
5612:
5604:
5563:
5553:
5514:
5479:
5436:
5393:
5350:
5305:
5297:
5256:
5248:
5203:
5195:
5150:
5101:
5091:
5050:
5040:
4999:
4991:
4950:
4940:
4899:
4891:
4842:
4834:
4793:
4783:
4736:
4663:
4609:
4601:
4557:
4547:
4498:
4403:
4395:
4288:
4280:
4241:
4196:
4186:
4114:
4104:
4024:
3971:
3930:
3889:
3879:
3830:
3789:
3781:
3671:
3570:
3562:
3545:
3505:
3473:
3465:
3416:
3406:
3330:
3322:
3250:
3242:
3203:
3168:
3131:
3098:
3090:
3023:
3013:
2938:
2928:
2879:
2869:
2820:
2810:
2753:
2714:
2604:
2535:
2519:
2464:
2383:
2367:
2312:
2263:
2255:
2008:
823:
723:
660:
530:
415:
375:
303:
241:
4646:
Barrell BG, Bankier AT, Drouin J (1979). "A different genetic code in human mitochondria".
2992:
Nirenberg M, Leder P, Bernfield M, Brimacombe R, Trupin J, Rottman F, O'Neal C (May 1965).
706:
defensive responses. In large populations of asexually reproducing organisms, for example,
554:, respectively, but as start codons they are translated as methionine or formylmethionine.
9321:
9196:
8124:
7961:
7807:
7586:
6426:
6409:
6049:
6032:
5518:
5483:
3306:
602:
386:
127:). That scheme is often referred to as the canonical or standard genetic code, or simply
7332:
7109:
7048:
6997:
6921:
6856:
6801:
6625:
6517:
6462:
6370:
6239:
6194:
6132:
5988:
5922:
5867:
5824:
5773:
5657:
5346:
5146:
5029:"Peroxisomal lactate dehydrogenase is generated by translational readthrough in mammals"
4887:
4727:
Jukes TH, Osawa S (December 1990). "The genetic code in mitochondria and chloroplasts".
4659:
4543:
4439:
4237:
4182:
4100:
4020:
3875:
3558:
3461:
3402:
3009:
2924:
2865:
2806:
2308:
1970:
147:
9416:
9391:
9371:
9366:
9171:
9116:
8799:
8670:
8430:
7951:
7923:
7716:
7656:
7556:
7351:
7316:
7222:
7197:
7173:
7148:
6875:
6840:
6762:
6729:
6705:
6672:
6005:
5972:
5617:
5592:
5568:
5541:
5310:
5285:
5261:
5232:
5208:
5179:
5106:
5079:
5055:
5028:
5004:
4979:
4955:
4928:
4904:
4871:
4798:
4771:
4614:
4589:
4562:
4527:
4408:
4383:
4293:
4268:
4119:
4084:
3575:
3540:
3478:
3445:
3421:
3386:
3385:
Zhang Y, Lamb BM, Feldman AW, Zhou AX, Lavergne T, Li L, Romesberg FE (February 2017).
3335:
3310:
3255:
3230:
3103:
3078:
2540:
2388:
2355:
2268:
2243:
2195:
2148:
Chemical principles govern specific RNA interaction with amino acids. Experiments with
2087:
1943:
802:
797:
793:
760:
below for the full correlation). For example, although codons GAA and GAG both specify
733:
656:
632:
618:
606:
578:
525:
490:
486:
482:
390:
69:
8163:
6599:
6334:
6317:
6294:
6151:
6116:
5941:
5906:
4847:
4822:
3894:
3859:
3794:
3769:
3676:
3659:
3387:"A semisynthetic organism engineered for the stable expansion of the genetic alphabet"
3028:
2993:
2943:
2908:
2884:
2849:
2825:
2790:
27:
Rules by which information encoded within genetic material is translated into proteins
17:
9477:
9181:
9052:
8993:
8976:
8971:
8929:
8916:
8503:
8424:
8418:
8320:
8152:
7839:
7294:
6536:
6501:
6263:
6085:
5832:
5702:
5676:
5641:
5155:
5130:
4201:
4166:
3592:
2626:
2492:
2411:
1975:
761:
703:
257:
166:
136:
101:
37:
7021:
6945:
6825:
6486:
6394:
5891:
5797:
5456:
5413:
5370:
4756:
4605:
4425:
4399:
4151:
4044:
3993:
2775:
9406:
9096:
8821:
8700:
8695:
8658:
8636:
8621:
8551:
8026:
7956:
7546:
7133:
7072:
7005:
6657:
6101:
5732:
4683:
3609:
3246:
3194:
Xie J, Schultz PG (December 2005). "Adding amino acids to the genetic repertoire".
2340:
2222:
2037:
2032:
918:
898:
878:
858:
826:, can vary from species to species with functional implications for the control of
322:
307:
264:
190:
178:
170:
124:
105:
4526:
Prat L, Heinemann IU, Aerni HR, Rinehart J, O'Donoghue P, Söll D (December 2012).
2848:
Gardner RS, Wahba AJ, Basilio C, Miller RS, Lengyel P, Speyer JF (December 1962).
468:
gene starts with the ATG codon (blue circle for the M amino acid) in the +3 frame.
260:. They thereby deduced that the codon UUU specified the amino acid phenylalanine.
6865:
6576:
5706:
5096:
4344:
4317:
3935:
3918:
3835:
3818:
3632:
3094:
2994:"RNA codewords and protein synthesis, VII. On the general nature of the RNA code"
2907:
Wahba AJ, Gardner RS, Basilio C, Miller RS, Speyer JF, Lengyel P (January 1963).
2667:
2642:"On Degenerate Templates and the Adaptor Hypothesis: A Note for the RNA Tie Club"
9396:
9033:
9011:
9006:
9001:
8981:
8954:
8949:
8900:
8871:
8831:
8811:
8789:
8462:
8395:
8129:
8058:
7646:
7117:
5760:
Freeland SJ, Hurst LD (September 1998). "The genetic code is one in a million".
4823:"The CUG codon is decoded in vivo as serine and not leucine in Candida albicans"
4284:
3501:
2523:
2204:
1947:
640:
589:
521:
494:
253:
8228:
7321:
Proceedings of the National Academy of Sciences of the United States of America
6929:
6506:
Proceedings of the National Academy of Sciences of the United States of America
5911:
Proceedings of the National Academy of Sciences of the United States of America
5646:
Proceedings of the National Academy of Sciences of the United States of America
4978:
Hofhuis J, Schueren F, Nötzel C, Lingner T, Gärtner J, Jahn O, Thoms S (2016).
4532:
Proceedings of the National Academy of Sciences of the United States of America
4171:
Proceedings of the National Academy of Sciences of the United States of America
4089:
Proceedings of the National Academy of Sciences of the United States of America
4007:
Crow JF (1993). "How much do we know about spontaneous human mutation rates?".
3864:
Proceedings of the National Academy of Sciences of the United States of America
3658:
Touriol C, Bornes S, Bonnal S, Audigier S, Prats H, Prats AC, Vagner S (2003).
3391:
Proceedings of the National Academy of Sciences of the United States of America
3207:
2998:
Proceedings of the National Academy of Sciences of the United States of America
2913:
Proceedings of the National Academy of Sciences of the United States of America
2854:
Proceedings of the National Academy of Sciences of the United States of America
2795:
Proceedings of the National Academy of Sciences of the United States of America
2758:
2741:
1982:
mitochondrial genome by FACIL. The program is able to correctly infer that the
9376:
8966:
8934:
8784:
8779:
8710:
8498:
8339:
8315:
8272:
8261:
8087:
7651:
7520:
7500:
7475:
7292:
Lodish HF, Berk A, Zipursky SL, Matsudaira P, Baltimore D, Darnell JE (2000).
7056:
6809:
6378:
6203:
6178:
5996:
5973:"A model of proto-anti-codon RNA enzymes requiring L-amino acid homochirality"
5542:"A computational screen for alternative genetic codes in over 250,000 genomes"
5354:
3566:
2507:
2437:
2156:
2002:
1997:
749:
744:
668:
645:
558:
539:
410:
In May 2019, researchers reported the creation of a new "Syn61" strain of the
394:
315:
292:
109:
45:
41:
7387:
6753:
6696:
6641:
5666:
5608:
4872:"Evolution of pathogenicity and sexual reproduction in eight Candida genomes"
4689:
4191:
3785:
2618:
2531:
2476:
2379:
2324:
9356:
8571:
8389:
8354:
8344:
8305:
7341:
5199:
5178:
Heaphy SM, Mariotti M, Gladyshev VN, Atkins JF, Baranov PV (November 2016).
4838:
4552:
4245:
4109:
3411:
2874:
2815:
2701:
Barciszewska, Mirosława Z.; Perrigue, Patrick M.; Barciszewski, Jan (2016).
2687:
2259:
2013:
687:
614:
501:
411:
374:
H. Murakami and M. Sisido extended some codons to have four and five bases.
7360:
7231:
7182:
7164:
7125:
7064:
7013:
6937:
6884:
6817:
6771:
6714:
6649:
6584:
6545:
6526:
6435:
6386:
6302:
6255:
6212:
6141:
6058:
6014:
5931:
5626:
5577:
5526:
5491:
5448:
5405:
5362:
5319:
5270:
5233:"Position-dependent termination and widespread obligatory frameshifting in
5217:
5164:
5115:
5064:
5013:
4964:
4913:
4807:
4788:
4623:
4571:
4512:
4503:
4417:
4384:"Global importance of RNA secondary structures in protein coding sequences"
4302:
4210:
4128:
3944:
3884:
3844:
3803:
3739:
Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, et al., eds. (2000).
3685:
3584:
3487:
3430:
3344:
3326:
3264:
3215:
3180:
3172:
3112:
3018:
2952:
2893:
2834:
2767:
2726:
2702:
2549:
2397:
2332:
2292:
2277:
6688:
6633:
6478:
6343:
6247:
6093:
5950:
5883:
5840:
5789:
5685:
5593:"Stop or Not: Genome-Wide Profiling of Reassigned Stop Codons in Ciliates"
4856:
4748:
4253:
4036:
4028:
3985:
3037:
2933:
2371:
2244:"Genetic code supports targeted insertion of two amino acids by one codon"
9025:
8986:
8924:
8864:
8816:
8748:
8738:
8705:
8442:
8436:
8383:
8310:
8107:
8097:
8021:
7691:
7661:
7505:
7495:
7423:
6745:
6160:
5470:
Liu CC, Schultz PG (2010). "Adding new chemistries to the genetic code".
4675:
2468:
2184:
2167:
2163:
2121:
Random freeze: the genetic code was randomly created. For example, early
2095:
2075:
2048:
2017:
610:
594:
330:
276:
93:
57:
5558:
5045:
4995:
4895:
3903:
3469:
3305:
Hoesl, M. G.; Oehm, S.; Durkin, P.; Darmon, E.; Peil, L.; Aerni, H.-R.;
2609:
2484:
2452:
710:, multiple beneficial mutations may co-occur. This phenomenon is called
464:
genes terminates with the TAG stop codon (red dot) in the +1 frame. The
9361:
8846:
8841:
8836:
8826:
8753:
8576:
8287:
8282:
8277:
8266:
8213:
8208:
7843:
7213:
6470:
6279:"Selection, history and chemistry: the three faces of the genetic code"
5875:
5781:
5640:
Ribas de Pouplana L, Turner RJ, Steer BA, Schimmel P (September 1998).
5027:
Schueren F, Lingner T, George R, Hofhuis J, Gartner J, Thoms S (2014).
4740:
3977:
10.1002/(SICI)1098-1004(1996)7:4<361::AID-HUMU12>3.0.CO;2-0
3135:
2718:
2149:
2126:
2021:
673:
551:
444:
438:
288:
280:
268:
89:
53:
32:
8520:
6502:"Testing a biosynthetic theory of the genetic code: fact or artifact?"
5252:
8941:
8851:
8377:
8349:
8256:
7515:
7490:
5591:
Chen, W; Geng, Y; Zhang, B; Yan, Y; Zhao, F; Miao, M (4 April 2023).
5080:"Functional Translational Readthrough: A Systems Biology Perspective"
4945:
4667:
3770:"Lesion (in)tolerance reveals insights into DNA replication fidelity"
2316:
781:
652:
547:
509:
284:
272:
249:
49:
7250:
Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gilbert WM (1999).
5440:
5397:
5301:
4636:
Francis Crick, 1968. "The Origin of the Genetic Code". J. Mol. Biol.
240:
were the first to reveal the nature of a codon in 1961. They used a
7271:
Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002).
6179:"RNA-amino acid binding: a stereochemical era for the genetic code"
4528:"Carbon source-dependent expansion of the genetic code in bacteria"
4487:"Pyrrolysine and selenocysteine use dissimilar decoding strategies"
2102:, so the latter cannot be part of the explanation of its patterns.
7865:
7776:
7100:
6988:
6912:
6616:
6561:"The origin of the genetic code and of the earliest oligopeptides"
3610:"Revised Cambridge Reference Sequence (rCRS): accession NC_012920"
2293:"Genetical implications of the structure of deoxyribonucleic acid"
1969:
743:
727:
695:
588:
431:
165:
was discovered in 1953. The key discoverers, English biophysicist
146:
85:
31:
7381:
6318:"Rhyme or reason: RNA-arginine interactions and the genetic code"
4770:
Fitzpatrick DA, Logue ME, Stajich JE, Butler G (1 January 2006).
4346:
Reaction centers of photosynthetic bacteria: Feldafing-II-Meeting
3919:"ALS: a disease of motor neurons and their nonneuronal neighbors"
3819:"ALS: A disease of motor neurons and their nonneuronal neighbors"
3725:
References for the image are found in Wikimedia Commons page at:
108:(tRNA) molecules to carry amino acids and to read the mRNA three
9057:
6841:"Codon size reduction as the origin of the triplet genetic code"
6673:"The complex evolutionary history of aminoacyl-tRNA synthetases"
5286:"Origin and Evolution of the Genetic Code: The Universal Enigma"
3444:
Han S, Yang A, Lee S, Lee HW, Park CB, Park HS (February 2017).
2122:
2070:
505:
326:
120:
8524:
8167:
7780:
7396:
4929:"Virus-host co-evolution under a modified nuclear genetic code"
497:, usually the first AUG (ATG) codon in the RNA (DNA) sequence.
189:
In 1954, Gamow created an informal scientific organisation the
8881:
8876:
7835:
7485:
7480:
699:
115:
The codons specify which amino acid will be added next during
81:
77:
6031:
Freeland SJ, Knight RD, Landweber LF, Hurst LD (April 2000).
4699:
4697:
4062:(6th ed.). Boston, Mass: McGraw Hill. pp. 227–228.
302:
identified the rest of the genetic code. Shortly thereafter,
6072:
Crick FH (December 1968). "The origin of the genetic code".
3768:
Freisinger E, Grollman AP, Miller H, Kisker C (April 2004).
2356:"'Genetic Coding' Reconsidered: An Analysis of Actual Usage"
393:
and co-workers reported the full substitution of all 20,899
7392:
5642:"Genetic code origins: tRNAs older than their synthetases?"
3541:"Total synthesis of Escherichia coli with a recoded genome"
3284:. Springer Science & Business Media. pp. 105–106.
3054:(Press release). The Royal Swedish Academy of Science. 1968
2969:(Press release). The Royal Swedish Academy of Science. 1959
256:
that they had synthesized consisted of only the amino acid
161:
Efforts to understand how proteins are encoded began after
5811:
Taylor FJ, Coates D (1989). "The code within the codons".
4590:"FACIL: Fast and Accurate Genetic Code Inference and Logo"
3360:"First stable semisynthetic organism created | KurzweilAI"
2178:
accurately and efficiently translating information? These
732:
Grouping of codons by amino acid residue molar volume and
7147:
Jee J, Sundstrom A, Massey SE, Mishra B (November 2013).
6500:
Ronneberg TA, Landweber LF, Freeland SJ (December 2000).
5129:
Kubyshkin V, Acevedo-Rocha CG, Budisa N (February 2018).
4870:
Butler G, Rasmussen MD, Lin MF, et al. (June 2009).
4583:
4581:
4485:
Zhang Y, Baranov PV, Atkins JF, Gladyshev VN (May 2005).
2909:"Synthetic polynucleotides and the amino acid code. VIII"
2420:
What Mad Pursuit: A Personal View of Scientific Discovery
2016:
have adapted to the host's genetic code modification. In
6839:
Baranov PV, Venin M, Provan G (2009). Gemmell NJ (ed.).
3309:; Rinehart, J.; Leach, D.; Söll, D.; Budisa, N. (2015).
2850:"Synthetic polynucleotides and the amino acid code. VII"
6408:
Brooks DJ, Fresco JR, Lesk AM, Singh M (October 2002).
3958:
Isbrandt D, Hopwood JJ, von Figura K, Peters C (1996).
3917:
Boillée S, Vande Velde C, Cleveland DW (October 2006).
663:) of a non-multiple of 3 nucleotide bases are known as
279:
RNA sequence (CCCCC...) coded for the polypeptide poly-
271:
RNA sequence (AAAAA...) coded for the polypeptide poly-
4927:
Taylor DJ, Ballinger MJ, Bowman SM, Bruenn JA (2013).
4708:. National Center for Biotechnology Information (NCBI)
3626:
3624:
3079:"The genome of bacteriophage T4: an archeological dig"
2453:"Computing Science: The Invention of the Genetic Code"
371:
and function or to create novel or enhanced proteins.
252:
RNA sequence (i.e., UUUUU...) and discovered that the
4083:
Sawyer SA, Parsch J, Zhang Z, Hartl DL (April 2007).
3817:
Boillée, S; Vande Velde, C; Cleveland, D. W. (2006).
3631:
King RC, Mulligan P, Stansfield W (10 January 2013).
2742:"Establishing the Triplet Nature of the Genetic Code"
2187:
defined by the probable errors and is related to the
651:
Mutations that disrupt the reading frame sequence by
5697:
5695:
2564:"Francis Crick - Profiles in Science Search Results"
2508:"Martynas Yčas: The "Archivist" of the RNA Tie Club"
520:
Translation starts with a chain-initiation codon or
283:. Therefore, the codon AAA specified the amino acid
9344:
9210:
9070:
8899:
8798:
8767:
8719:
8620:
8559:
8404:
8363:
8329:
8296:
8243:
8236:
8201:
8140:
8049:
8014:
7988:
7979:
7937:
7911:
7885:
7876:
7814:
7735:
7634:
7585:
7529:
7468:
3231:"Expanding the genetic code for biological studies"
2669:
Avoid Boring People: Lessons from a Life in Science
2183:nonrandom. The code's emergence is governed by the
295:most of the remaining codons were then determined.
218:
DNA and RNA codon tables § Translation table 1
7293:
6277:Knight RD, Freeland SJ, Landweber LF (June 1999).
5131:"On universal coding events in protein biogenesis"
2144:Hypotheses have addressed a variety of scenarios:
2140:function, usually some kind of error minimization.
1933:DNA and RNA codon tables § Alternative codons
224:Crick, Brenner, Barnett and Watts-Tobin experiment
2360:The British Journal for the Philosophy of Science
6783:
6781:
3860:"beta 0 thalassemia, a nonsense mutation in man"
3281:Emergent Computation: Emphasizing Bioinformatics
3052:"The Nobel Prize in Physiology or Medicine 1968"
2967:"The Nobel Prize in Physiology or Medicine 1959"
2703:"tRNA--the golden standard in molecular biology"
6959:Sonneborn TM (1965). Bryson V, Vogel H (eds.).
6598:Higgs, Paul G.; Pudritz, Ralph E. (June 2009).
6177:Yarus M, Widmann JJ, Knight R (November 2009).
6172:
6170:
6026:
6024:
5966:
5964:
5962:
5960:
60:to synthesize a protein according to this code.
6790:Origins of Life and Evolution of the Biosphere
5755:
5753:
4440:"Codon Usage Frequency Table(chart)-Genscript"
2170:, such as before cells began making ribosomes.
678:may result in severe genetic diseases such as
267:'s laboratory that demonstrated that the poly-
56:) instead. This mRNA molecule will instruct a
40:(mRNA) molecule. Each codon consists of three
8536:
8179:
7792:
7408:
7300:(4th ed.). San Francisco: W.H. Freeman.
7256:(7th ed.). San Francisco: W.H. Freeman.
6963:. New York: Academic Press. pp. 377–397.
3727:Commons:File:Notable mutations.svg#References
3615:National Center for Biotechnology Information
287:, and the codon CCC specified the amino acid
76:information encoded within genetic material (
8:
4480:
4478:
3539:Fredens, Julius; et al. (15 May 2019).
3446:"Expanding the genetic code of Mus musculus"
3311:"Chemical evolution of a bacterial proteome"
714:and causes competition among the mutations.
329:were passed through a filter that contained
209:"Codon" redirects here. For other uses, see
7384:— Codon frequency tables for many organisms
7277:(4th ed.). New York: Garland Science.
6033:"Early fixation of an optimal genetic code"
5708:Life from an RNA World: The Ancestor Within
2597:Philosophy, Theory, and Practice in Biology
2291:Watson, J. D.; Crick, F. H. (30 May 1953).
8907:
8804:
8543:
8529:
8521:
8240:
8186:
8172:
8164:
7985:
7947:Precursor mRNA (pre-mRNA / hnRNA)
7882:
7799:
7785:
7777:
7415:
7401:
7393:
6316:Knight RD, Landweber LF (September 1998).
5284:Koonin EV, Novozhilov AS (February 2009).
2789:Nirenberg MW, Matthaei JH (October 1961).
833:
346:Expanded genetic codes (synthetic biology)
123:are encoded with a single scheme (see the
7350:
7340:
7221:
7172:
7099:
6987:
6911:
6874:
6864:
6761:
6704:
6615:
6535:
6525:
6425:
6333:
6202:
6150:
6140:
6048:
6004:
5940:
5930:
5675:
5665:
5616:
5567:
5557:
5540:Shulgina, Y; Eddy, SR (9 November 2021).
5309:
5260:
5241:Nature Structural & Molecular Biology
5207:
5154:
5105:
5095:
5054:
5044:
5003:
4954:
4944:
4903:
4846:
4797:
4787:
4613:
4561:
4551:
4502:
4407:
4377:
4375:
4292:
4200:
4190:
4118:
4108:
4060:Human Genetics: Concepts and Applications
3975:
3934:
3893:
3883:
3834:
3793:
3747:(7th ed.). New York: W. H. Freeman.
3675:
3574:
3477:
3420:
3410:
3334:
3254:
3229:Wang Q, Parrish AR, Wang L (March 2009).
3102:
3027:
3017:
2942:
2932:
2883:
2873:
2824:
2814:
2757:
2608:
2539:
2387:
2267:
635:that can cause genetic diseases such as
4343:Michel-Beyerle, Maria Elisabeth (1990).
3701:"How nonsense mutations got their names"
2672:. Oxford University Press. p. 112.
2354:Stegmann, Ulrich E. (1 September 2016).
844:
7748:List of genetics research organizations
7388:History of deciphering the genetic code
7153:Journal of the Royal Society, Interface
4704:Elzanowski A, Ostell J (7 April 2008).
4009:Environmental and Molecular Mutagenesis
3315:Angewandte Chemie International Edition
2234:
2133:matching by aminoacyl-tRNA synthetases.
822:The frequency of codons, also known as
7376:The Genetic Codes: Genetic Code Tables
7317:"The RNA code: nature's Rosetta Stone"
6559:Trifonov, Edward N. (September 2009).
4165:Drake JW, Holland JJ (November 1999).
2586:
2584:
2086:The genetic code is a key part of the
7967:Histone acetylation and deacetylation
6427:10.1093/oxfordjournals.molbev.a003988
6050:10.1093/oxfordjournals.molbev.a026331
5519:10.1146/annurev-biochem-060713-035737
5484:10.1146/annurev.biochem.052308.105824
5386:Nature Reviews Molecular Cell Biology
4267:de Visser JA, Rozen DE (April 2006).
3126:Budisa, Nediljko (23 December 2005).
312:Nobel Prize in Physiology or Medicine
92:. Translation is accomplished by the
84:sequences of nucleotide triplets, or
7:
9442:
8032:Ribosome-nascent chain complex (RNC)
3128:The book at the Wiley Online Library
2218:List of genetic engineering software
2026:functional translational readthrough
609:of the second strand. These errors,
263:This was followed by experiments in
44:, usually corresponding to a single
8195:Encoded (proteinogenic) amino acids
7253:An Introduction to genetic analysis
4491:The Journal of Biological Chemistry
3745:An Introduction to Genetic Analysis
3520:from the original on 2 January 2022
3196:Current Opinion in Chemical Biology
2506:Strauss, Bernard S (1 March 2019).
605:, errors occasionally occur in the
321:Extending this work, Nirenberg and
68:is the set of rules used by living
4167:"Mutation rates among RNA viruses"
2740:Yanofsky, Charles (9 March 2007).
838:Human genome codon frequency table
25:
7315:Caskey CT, Leder P (April 2014).
5078:F. Schueren und S. Thoms (2016).
581:to bind to the ribosome instead.
9453:
9441:
9430:
9429:
8227:
7761:
7760:
5156:10.1016/j.biosystems.2017.10.004
4821:Santos MA, Tuite MF (May 1995).
3618:. Retrieved on 27 December 2017.
3608:mitochondrion, complete genome.
2422:. Basic Books. pp. 89–101.
456:(in red) or in the +3 frame for
36:A series of codons in part of a
8037:Post-translational modification
6414:Molecular Biology and Evolution
6037:Molecular Biology and Evolution
5597:Molecular Biology and Evolution
5188:Molecular Biology and Evolution
378:constructed a functional 65th (
333:, the components of cells that
132:
7037:Journal of Molecular Evolution
7006:10.1103/PhysRevLett.100.048101
6900:Journal of Theoretical Biology
6451:Journal of Molecular Evolution
6359:Journal of Molecular Evolution
6283:Trends in Biochemical Sciences
6183:Journal of Molecular Evolution
6074:Journal of Molecular Evolution
5977:Journal of Molecular Evolution
5856:Journal of Molecular Evolution
5762:Journal of Molecular Evolution
5335:Journal of Molecular Evolution
3858:Chang JC, Kan YW (June 1979).
3247:10.1016/j.chembiol.2009.03.001
1980:Globobulimina pseudospinescens
1:
9298:Amino acid metabolism enzymes
7274:Molecular biology of the cell
6335:10.1016/S1074-5521(98)90001-1
6295:10.1016/S0968-0004(99)01392-4
5507:Annual Review of Biochemistry
5472:Annual Review of Biochemistry
4606:10.1093/bioinformatics/btr316
4400:10.1093/bioinformatics/bty678
4322:. Pearson/Benjamin Cummings.
4319:Molecular Biology of the Gene
3677:10.1016/S0248-4900(03)00033-9
2416:"Chapter 8: The Genetic Code"
2194:Game theory: Models based on
452:, either in the +1 frame for
450:vertebrate mitochondrial code
230:consist of three DNA bases.
7707:Missing heritability problem
7196:Itzkovitz S, Alon U (2007).
6866:10.1371/journal.pone.0005708
6577:10.1016/j.resmic.2009.05.004
6086:10.1016/0022-2836(68)90392-6
5833:10.1016/0303-2647(89)90059-2
5711:. Harvard University Press.
5097:10.1371/journal.pgen.1006196
3936:10.1016/j.neuron.2006.09.018
3836:10.1016/j.neuron.2006.09.018
3707:. San Diego State University
3699:Maloy S (29 November 2003).
2646:National Library of Medicine
1984:Protozoan Mitochondrial Code
807:Singular Value Decomposition
306:determined the structure of
9202:Volume combustion synthesis
8681:Bioorganometallic chemistry
7118:10.1016/j.plrev.2010.06.002
6961:Evolving genes and proteins
4285:10.1534/genetics.105.052373
2524:10.1534/genetics.118.301754
1987:support for the prediction.
613:, can affect an organism's
9520:
9132:Enantioselective synthesis
8253:Branched-chain amino acids
6930:10.1016/j.jtbi.2007.07.029
4144:"Malaria and the Red Cell"
3278:Simon M (7 January 2005).
3208:10.1016/j.cbpa.2005.10.011
3095:10.1093/genetics/168.2.575
2759:10.1016/j.cell.2007.02.029
2100:aminoacyl-tRNA synthetases
1963:
1930:
905:
885:
865:
721:
419:. This strain has a fully
355:
349:
215:
208:
173:, working together at the
154:
9425:
9287:Protein primary structure
9137:Fully automated synthesis
9082:Artificial gene synthesis
8910:
8807:
8456:
8225:
7756:
7430:
7057:10.1007/s00239-004-0176-7
6379:10.1007/s00239-015-9686-8
6204:10.1007/s00239-009-9270-1
5997:10.1007/s00239-011-9453-4
5905:Wong JT (February 1980).
5355:10.1007/s00239-015-9686-8
4316:Watson, James D. (2008).
3705:Microbial Genetics Course
3567:10.1038/s41586-019-1192-5
2666:Watson, James D. (2007).
1994:human mitochondrial genes
1927:Alternative genetic codes
508:are often interrupted by
195:proteins were synthesised
100:in an order specified by
98:proteinogenic amino acids
9112:Custom peptide synthesis
8098:sequestration (P-bodies)
6565:Research in Microbiology
5971:Erives A (August 2011).
5667:10.1073/pnas.95.19.11295
4776:BMC Evolutionary Biology
4192:10.1073/pnas.96.24.13910
3786:10.1038/sj.emboj.7600158
3637:. OUP USA. p. 608.
3634:A Dictionary of Genetics
3077:Edgar B (October 2004).
1953:Acetohalobium arabaticum
1938:Non-standard amino acids
780:), while the amino acid
597:that can occur in humans
314:in 1959 for work on the
226:first demonstrated that
9412:Antonie van Leeuwenhoek
8647:Bioorthogonal chemistry
8076:Gene regulatory network
7342:10.1073/pnas.1404819111
7088:Physics of Life Reviews
6976:Physical Review Letters
6810:10.1023/A:1025771327614
6322:Chemistry & Biology
4553:10.1073/pnas.1218613110
4382:Fricke, Markus (2019).
4246:10.1126/science.7041255
4110:10.1073/pnas.0701572104
3741:"Spontaneous mutations"
3412:10.1073/pnas.1616443114
3235:Chemistry & Biology
2875:10.1073/pnas.48.12.2087
2816:10.1073/pnas.47.10.1588
2640:Crick, Francis (1955).
2260:10.1126/science.1164748
2044:ribosomal frameshifting
569:(sometimes also called
169:and American biologist
9192:Solvothermal synthesis
9142:Hydrothermal synthesis
8686:Bioinorganic chemistry
8081:cis-regulatory element
7296:Molecular cell biology
7165:10.1098/rsif.2013.0614
6734:Nucleic Acids Research
6677:Nucleic Acids Research
6527:10.1073/pnas.250403097
6142:10.1073/pnas.75.9.4334
5932:10.1073/pnas.77.2.1083
5609:10.1093/molbev/msad064
4827:Nucleic Acids Research
4789:10.1186/1471-2148-6-99
4504:10.1074/jbc.M501458200
3885:10.1073/pnas.76.6.2886
3327:10.1002/anie.201502868
3173:10.1002/biot.201600097
3019:10.1073/pnas.53.5.1161
2173:Information channels:
1988:
753:
741:
601:During the process of
598:
469:
358:Nucleic acid analogues
211:Codon (disambiguation)
152:
61:
18:Universal genetic code
9221:Branches of chemistry
9187:Solid-phase synthesis
9039:Amino acid metabolism
9017:Nucleotide metabolism
8960:Fatty-acid metabolism
8691:Biophysical chemistry
8612:Theoretical chemistry
8567:List of life sciences
7743:List of genetic codes
6634:10.1089/ast.2008.0280
6248:10.1089/ast.2022.0107
5737:community.wolfram.com
5200:10.1093/molbev/msw166
4839:10.1093/nar/23.9.1481
4029:10.1002/em.2850210205
3450:Nature Communications
3161:Biotechnology Journal
2934:10.1073/pnas.49.1.116
2591:Fry, Michael (2022).
2451:Hayes, Brian (1998).
2175:Information-theoretic
1973:
1966:List of genetic codes
1946:and UAG can code for
747:
738:more detailed version
731:
592:
516:Start and stop codons
435:
352:Expanded genetic code
155:Further information:
150:
131:genetic code, though
35:
9504:Protein biosynthesis
9107:Convergent synthesis
9087:Biomimetic synthesis
8676:Bioorganic chemistry
8587:Analytical chemistry
8103:alternative splicing
8093:Post-transcriptional
7919:Transcription factor
7642:Behavioural genetics
7382:Codon Usage Database
6115:Hopfield JJ (1978).
4154:on 27 November 2011.
3349:NIHMSID: NIHMS711205
2707:Molecular BioSystems
2568:profiles.nlm.nih.gov
2469:10.1511/1998.17.3338
2189:map coloring problem
2092:RNA world hypothesis
811:without translation.
665:frameshift mutations
593:Examples of notable
238:J. Heinrich Matthaei
175:Cavendish Laboratory
117:protein biosynthesis
9402:Notable scientists:
9232:Cellular structures
9122:Divergent synthesis
8654:Medicinal chemistry
8597:Medicinal chemistry
8592:Inorganic chemistry
8027:Transfer RNA (tRNA)
7722:Population genomics
7712:Molecular evolution
7672:Genetic engineering
7333:2014PNAS..111.5758C
7110:2010PhLRv...7..362T
7049:2006JMolE..63..297S
6998:2008PhRvL.100d8101T
6922:2007JThBi.249..331T
6857:2009PLoSO...4.5708B
6802:2003OLEB...33..457F
6689:10.1093/nar/gkw1182
6626:2009AsBio...9..483H
6518:2000PNAS...9713690R
6463:1997JMolE..44..473A
6371:2015JMolE..80..229S
6240:2023AsBio..23..536B
6195:2009JMolE..69..406Y
6133:1978PNAS...75.4334H
5989:2011JMolE..73...10E
5923:1980PNAS...77.1083W
5868:1989JMolE..29..288D
5825:1989BiSys..22..177T
5774:1998JMolE..47..238F
5658:1998PNAS...9511295D
5559:10.7554/eLife.71402
5347:2015JMolE..80..229S
5182:Condylostoma magnum
5147:2018BiSys.164...16K
5046:10.7554/eLife.03640
4996:10.1098/rsob.160246
4896:10.1038/nature08064
4888:2009Natur.459..657B
4706:"The Genetic Codes"
4660:1979Natur.282..189B
4544:2012PNAS..10921070P
4465:"Codon usage table"
4349:. Springer-Verlag.
4238:1982Sci...215.1577H
4183:1999PNAS...9613910D
4142:Bridges KR (2002).
4101:2007PNAS..104.6504S
4021:1993EnvMM..21..122C
3876:1979PNAS...76.2886C
3664:Biology of the Cell
3559:2019Natur.569..514F
3470:10.1038/ncomms14568
3462:2017NatCo...814568H
3403:2017PNAS..114.1317Z
3321:(34): 10030–10034.
3010:1965PNAS...53.1161N
2925:1963PNAS...49..116W
2866:1962PNAS...48.2087G
2807:1961PNAS...47.1588N
2610:10.3998/ptpbio.2628
2372:10.1093/bjps/axv007
2309:1953Natur.171..964W
712:clonal interference
637:sickle-cell disease
585:Effect of mutations
561:have names: UAG is
298:Subsequent work by
9494:Molecular genetics
9254:Chemical synthesis
8775:Structural biology
8759:Reaction mechanism
8734:Chemical synthesis
8729:Chemical reactions
8666:Clinical chemistry
8607:Physical chemistry
8509:Dehydroamino acids
8469:Encoded (proteins)
8405:Negative charge (p
8364:Positive charge (p
8141:Influential people
8120:Post-translational
7939:Post-transcription
7682:Genetic monitoring
7214:10.1101/gr.5987307
6746:10.1093/nar/gkz730
6740:(19): 9998–10009.
6471:10.1007/PL00006170
5876:10.1007/BF02103616
5782:10.1007/PL00006381
4741:10.1007/BF01936921
4735:(11–12): 1117–26.
3513:The New York Times
3364:www.kurzweilai.net
3136:10.1002/3527607188
2719:10.1039/c5mb00557d
2457:American Scientist
1989:
784:is specified by UC
754:
750:single letter code
742:
629:nonsense mutations
625:Missense mutations
599:
536:initiation factors
479:open reading frame
470:
318:of RNA synthesis.
300:Har Gobind Khorana
275:and that the poly-
234:Marshall Nirenberg
157:Adaptor hypothesis
153:
62:
9499:Molecular biology
9469:
9468:
9387:Scientific method
9243:Organic reactions
9167:Peptide synthesis
9162:Organic synthesis
9157:One-pot synthesis
9092:Bioretrosynthesis
9066:
9065:
8895:
8894:
8857:Metabolic pathway
8744:Organic reactions
8632:Molecular biology
8602:Organic chemistry
8582:Chemistry fields:
8518:
8517:
8479:Non-proteinogenic
8452:
8451:
8161:
8160:
8045:
8044:
7975:
7974:
7851:Special transfers
7774:
7773:
7697:He Jiankui affair
7687:Genetic genealogy
7677:Genetic diversity
7606:the British Isles
7511:Genetic variation
7284:978-0-8153-3218-3
7263:978-0-7167-3771-1
5718:978-0-674-05075-4
5652:(19): 11295–300.
5253:10.1038/nsmb.3330
5194:(11): 2885–2889.
4654:(5735): 189–194.
4444:www.genscript.com
4356:978-3-540-53420-4
4329:978-0-8053-9592-1
4232:(4540): 1577–85.
4069:978-0-07-111156-0
3754:978-0-7167-3520-5
3644:978-0-19-976644-4
3553:(7757): 514–518.
3366:. 3 February 2017
3291:978-0-387-22046-8
2679:978-0-19-280273-6
2303:(4361): 964–967.
2180:"rate-distortion"
1923:
1922:
1919:
1918:
692:natural selection
680:Tay–Sachs disease
483:5' → 3' direction
369:protein structure
16:(Redirected from
9511:
9457:
9445:
9444:
9433:
9432:
9337:
9331:
9326:
9320:
9315:
9309:
9302:
9296:
9291:
9285:
9280:
9274:
9269:
9263:
9258:
9252:
9247:
9241:
9236:
9230:
9225:
9219:
9152:Mechanosynthesis
9127:Electrosynthesis
8908:
8805:
8642:Chemical biology
8545:
8538:
8531:
8522:
8241:
8231:
8188:
8181:
8174:
8165:
7986:
7883:
7801:
7794:
7787:
7778:
7764:
7763:
7727:Reverse genetics
7702:Medical genetics
7417:
7410:
7403:
7394:
7364:
7354:
7344:
7311:
7299:
7288:
7267:
7236:
7235:
7225:
7193:
7187:
7186:
7176:
7159:(88): 20130614.
7144:
7138:
7137:
7103:
7083:
7077:
7076:
7032:
7026:
7025:
6991:
6971:
6965:
6964:
6956:
6950:
6949:
6915:
6895:
6889:
6888:
6878:
6868:
6836:
6830:
6829:
6785:
6776:
6775:
6765:
6725:
6719:
6718:
6708:
6683:(3): 1059–1068.
6668:
6662:
6661:
6619:
6595:
6589:
6588:
6556:
6550:
6549:
6539:
6529:
6497:
6491:
6490:
6446:
6440:
6439:
6429:
6405:
6399:
6398:
6365:(5–6): 229–243.
6354:
6348:
6347:
6337:
6313:
6307:
6306:
6274:
6268:
6267:
6223:
6217:
6216:
6206:
6174:
6165:
6164:
6154:
6144:
6127:(9): 4334–4338.
6112:
6106:
6105:
6069:
6063:
6062:
6052:
6028:
6019:
6018:
6008:
5968:
5955:
5954:
5944:
5934:
5902:
5896:
5895:
5851:
5845:
5844:
5808:
5802:
5801:
5757:
5748:
5747:
5745:
5743:
5729:
5723:
5722:
5699:
5690:
5689:
5679:
5669:
5637:
5631:
5630:
5620:
5588:
5582:
5581:
5571:
5561:
5537:
5531:
5530:
5502:
5496:
5495:
5467:
5461:
5460:
5435:(464): 441–444.
5424:
5418:
5417:
5381:
5375:
5374:
5341:(5–6): 229–243.
5330:
5324:
5323:
5313:
5281:
5275:
5274:
5264:
5228:
5222:
5221:
5211:
5175:
5169:
5168:
5158:
5126:
5120:
5119:
5109:
5099:
5075:
5069:
5068:
5058:
5048:
5024:
5018:
5017:
5007:
4975:
4969:
4968:
4958:
4948:
4946:10.7717/peerj.50
4924:
4918:
4917:
4907:
4882:(7247): 657–62.
4867:
4861:
4860:
4850:
4818:
4812:
4811:
4801:
4791:
4767:
4761:
4760:
4724:
4718:
4717:
4715:
4713:
4701:
4692:
4687:
4668:10.1038/282189a0
4643:
4637:
4634:
4628:
4627:
4617:
4585:
4576:
4575:
4565:
4555:
4523:
4517:
4516:
4506:
4497:(21): 20740–51.
4482:
4473:
4472:
4469:www.kazusa.or.jp
4461:
4455:
4454:
4452:
4450:
4436:
4430:
4429:
4411:
4379:
4370:
4367:
4361:
4360:
4340:
4334:
4333:
4313:
4307:
4306:
4296:
4264:
4258:
4257:
4221:
4215:
4214:
4204:
4194:
4162:
4156:
4155:
4150:. Archived from
4139:
4133:
4132:
4122:
4112:
4080:
4074:
4073:
4058:Lewis R (2005).
4055:
4049:
4048:
4004:
3998:
3997:
3979:
3955:
3949:
3948:
3938:
3914:
3908:
3907:
3897:
3887:
3855:
3849:
3848:
3838:
3814:
3808:
3807:
3797:
3774:The EMBO Journal
3765:
3759:
3758:
3736:
3730:
3723:
3717:
3716:
3714:
3712:
3696:
3690:
3689:
3679:
3655:
3649:
3648:
3628:
3619:
3603:
3597:
3596:
3578:
3536:
3530:
3529:
3527:
3525:
3509:
3498:
3492:
3491:
3481:
3441:
3435:
3434:
3424:
3414:
3397:(6): 1317–1322.
3382:
3376:
3375:
3373:
3371:
3356:
3350:
3348:
3338:
3302:
3296:
3295:
3275:
3269:
3268:
3258:
3226:
3220:
3219:
3191:
3185:
3184:
3156:
3150:
3149:
3123:
3117:
3116:
3106:
3074:
3068:
3067:
3061:
3059:
3048:
3042:
3041:
3031:
3021:
2989:
2983:
2982:
2976:
2974:
2963:
2957:
2956:
2946:
2936:
2904:
2898:
2897:
2887:
2877:
2845:
2839:
2838:
2828:
2818:
2801:(10): 1588–602.
2786:
2780:
2779:
2761:
2737:
2731:
2730:
2698:
2692:
2691:
2663:
2657:
2656:
2654:
2652:
2637:
2631:
2630:
2612:
2588:
2579:
2578:
2576:
2574:
2560:
2554:
2553:
2543:
2503:
2497:
2496:
2448:
2442:
2441:
2414:(10 July 1990).
2408:
2402:
2401:
2391:
2351:
2345:
2344:
2317:10.1038/171964b0
2288:
2282:
2281:
2271:
2254:(5911): 259–61.
2239:
2009:Candida albicans
845:
834:
824:codon usage bias
818:Codon usage bias
724:Codon degeneracy
631:are examples of
544:formylmethionine
416:Escherichia coli
399:Escherichia coli
376:Steven A. Benner
304:Robert W. Holley
291:. Using various
242:cell-free system
185:
151:The genetic code
21:
9519:
9518:
9514:
9513:
9512:
9510:
9509:
9508:
9484:Gene expression
9474:
9473:
9470:
9465:
9421:
9340:
9335:
9329:
9324:
9318:
9313:
9307:
9300:
9294:
9289:
9283:
9278:
9272:
9267:
9261:
9256:
9250:
9245:
9239:
9234:
9228:
9223:
9217:
9206:
9197:Total synthesis
9073:
9062:
8903:
8891:
8794:
8763:
8721:
8715:
8624:
8616:
8560:Science fields
8555:
8549:
8519:
8514:
8513:
8494:Secondary amino
8448:
8411:
8400:
8370:
8359:
8325:
8292:
8232:
8223:
8197:
8192:
8162:
8157:
8136:
8071:Transcriptional
8041:
8010:
7971:
7962:Polyadenylation
7933:
7907:
7872:
7866:Protein→Protein
7817:
7810:
7808:Gene expression
7805:
7775:
7770:
7752:
7731:
7630:
7621:the Middle East
7587:Archaeogenetics
7581:
7525:
7464:
7426:
7421:
7372:
7367:
7314:
7308:
7291:
7285:
7270:
7264:
7249:
7245:
7243:Further reading
7240:
7239:
7202:Genome Research
7195:
7194:
7190:
7146:
7145:
7141:
7085:
7084:
7080:
7034:
7033:
7029:
6973:
6972:
6968:
6958:
6957:
6953:
6897:
6896:
6892:
6838:
6837:
6833:
6796:(4–5): 457–77.
6787:
6786:
6779:
6727:
6726:
6722:
6670:
6669:
6665:
6597:
6596:
6592:
6558:
6557:
6553:
6512:(25): 13690–5.
6499:
6498:
6494:
6448:
6447:
6443:
6420:(10): 1645–55.
6407:
6406:
6402:
6356:
6355:
6351:
6315:
6314:
6310:
6276:
6275:
6271:
6225:
6224:
6220:
6176:
6175:
6168:
6114:
6113:
6109:
6071:
6070:
6066:
6030:
6029:
6022:
5970:
5969:
5958:
5904:
5903:
5899:
5853:
5852:
5848:
5810:
5809:
5805:
5759:
5758:
5751:
5741:
5739:
5731:
5730:
5726:
5719:
5701:
5700:
5693:
5639:
5638:
5634:
5590:
5589:
5585:
5539:
5538:
5534:
5504:
5503:
5499:
5469:
5468:
5464:
5441:10.1038/nrm2005
5426:
5425:
5421:
5398:10.1038/nrm2005
5392:(10): 775–782.
5383:
5382:
5378:
5332:
5331:
5327:
5302:10.1002/iub.146
5283:
5282:
5278:
5230:
5229:
5225:
5177:
5176:
5172:
5128:
5127:
5123:
5090:(8): e1006196.
5077:
5076:
5072:
5026:
5025:
5021:
4977:
4976:
4972:
4926:
4925:
4921:
4869:
4868:
4864:
4820:
4819:
4815:
4769:
4768:
4764:
4726:
4725:
4721:
4711:
4709:
4703:
4702:
4695:
4645:
4644:
4640:
4635:
4631:
4600:(14): 1929–33.
4587:
4586:
4579:
4538:(51): 21070–5.
4525:
4524:
4520:
4484:
4483:
4476:
4463:
4462:
4458:
4448:
4446:
4438:
4437:
4433:
4381:
4380:
4373:
4368:
4364:
4357:
4342:
4341:
4337:
4330:
4315:
4314:
4310:
4279:(4): 2093–100.
4266:
4265:
4261:
4223:
4222:
4218:
4177:(24): 13910–3.
4164:
4163:
4159:
4141:
4140:
4136:
4095:(16): 6504–10.
4082:
4081:
4077:
4070:
4057:
4056:
4052:
4006:
4005:
4001:
3957:
3956:
3952:
3916:
3915:
3911:
3857:
3856:
3852:
3816:
3815:
3811:
3780:(7): 1494–505.
3767:
3766:
3762:
3755:
3738:
3737:
3733:
3724:
3720:
3710:
3708:
3698:
3697:
3693:
3670:(3–4): 169–78.
3657:
3656:
3652:
3645:
3630:
3629:
3622:
3604:
3600:
3538:
3537:
3533:
3523:
3521:
3504:(15 May 2019).
3500:
3499:
3495:
3443:
3442:
3438:
3384:
3383:
3379:
3369:
3367:
3358:
3357:
3353:
3304:
3303:
3299:
3292:
3277:
3276:
3272:
3228:
3227:
3223:
3193:
3192:
3188:
3158:
3157:
3153:
3146:
3125:
3124:
3120:
3076:
3075:
3071:
3057:
3055:
3050:
3049:
3045:
2991:
2990:
2986:
2972:
2970:
2965:
2964:
2960:
2906:
2905:
2901:
2860:(12): 2087–94.
2847:
2846:
2842:
2788:
2787:
2783:
2739:
2738:
2734:
2700:
2699:
2695:
2680:
2665:
2664:
2660:
2650:
2648:
2639:
2638:
2634:
2590:
2589:
2582:
2572:
2570:
2562:
2561:
2557:
2505:
2504:
2500:
2450:
2449:
2445:
2430:
2410:
2409:
2405:
2353:
2352:
2348:
2290:
2289:
2285:
2241:
2240:
2236:
2231:
2214:
2196:signaling games
2088:history of life
2084:
2062:
1968:
1962:
1940:
1935:
1929:
1924:
839:
820:
726:
720:
633:point mutations
619:DNA polymerases
603:DNA replication
587:
518:
475:
460:(in blue). The
430:
360:
354:
348:
220:
214:
207:
183:
163:DNA's structure
159:
145:
125:RNA codon table
52:). DNA uses T (
28:
23:
22:
15:
12:
11:
5:
9517:
9515:
9507:
9506:
9501:
9496:
9491:
9486:
9476:
9475:
9467:
9466:
9464:
9463:
9451:
9439:
9426:
9423:
9422:
9420:
9419:
9417:Francesco Redi
9414:
9409:
9404:
9399:
9394:
9392:Periodic table
9389:
9384:
9379:
9374:
9372:Microbiologist
9369:
9367:Bacteriologist
9364:
9359:
9354:
9348:
9346:
9342:
9341:
9339:
9338:
9327:
9316:
9304:
9303:
9292:
9281:
9276:Chemical bonds
9270:
9259:
9248:
9237:
9226:
9214:
9212:
9208:
9207:
9205:
9204:
9199:
9194:
9189:
9184:
9179:
9177:Retrosynthesis
9174:
9172:Radiosynthesis
9169:
9164:
9159:
9154:
9149:
9144:
9139:
9134:
9129:
9124:
9119:
9117:Direct process
9114:
9109:
9104:
9102:Chemosynthesis
9099:
9094:
9089:
9084:
9078:
9076:
9068:
9067:
9064:
9063:
9061:
9060:
9055:
9050:
9043:
9042:
9041:
9031:
9021:
9020:
9019:
9009:
9004:
8999:
8989:
8984:
8979:
8974:
8969:
8964:
8963:
8962:
8952:
8947:
8937:
8932:
8927:
8922:
8911:
8905:
8897:
8896:
8893:
8892:
8890:
8889:
8884:
8879:
8874:
8869:
8860:
8859:
8854:
8849:
8844:
8839:
8834:
8829:
8824:
8819:
8814:
8808:
8802:
8796:
8795:
8793:
8792:
8787:
8782:
8777:
8771:
8769:
8765:
8764:
8762:
8761:
8756:
8751:
8746:
8741:
8736:
8731:
8725:
8723:
8717:
8716:
8714:
8713:
8708:
8703:
8698:
8693:
8688:
8683:
8678:
8673:
8671:Neurochemistry
8668:
8663:
8662:
8661:
8651:
8650:
8649:
8639:
8634:
8628:
8626:
8618:
8617:
8615:
8614:
8609:
8604:
8599:
8594:
8589:
8584:
8579:
8574:
8569:
8563:
8561:
8557:
8556:
8550:
8548:
8547:
8540:
8533:
8525:
8516:
8515:
8512:
8511:
8506:
8501:
8496:
8491:
8486:
8481:
8476:
8471:
8458:
8457:
8454:
8453:
8450:
8449:
8447:
8446:
8440:
8434:
8431:Selenocysteine
8428:
8422:
8415:
8413:
8409:
8402:
8401:
8399:
8398:
8393:
8387:
8381:
8374:
8372:
8368:
8361:
8360:
8358:
8357:
8352:
8347:
8342:
8336:
8334:
8327:
8326:
8324:
8323:
8318:
8313:
8308:
8302:
8300:
8294:
8293:
8291:
8290:
8285:
8280:
8275:
8270:
8264:
8259:
8249:
8247:
8238:
8234:
8233:
8226:
8224:
8222:
8221:
8216:
8211:
8205:
8203:
8202:General topics
8199:
8198:
8193:
8191:
8190:
8183:
8176:
8168:
8159:
8158:
8156:
8155:
8150:
8148:François Jacob
8144:
8142:
8138:
8137:
8135:
8134:
8133:
8132:
8127:
8117:
8112:
8111:
8110:
8105:
8100:
8090:
8085:
8084:
8083:
8078:
8068:
8067:
8066:
8055:
8053:
8047:
8046:
8043:
8042:
8040:
8039:
8034:
8029:
8024:
8018:
8016:
8012:
8011:
8009:
8008:
8003:
7998:
7992:
7990:
7983:
7977:
7976:
7973:
7972:
7970:
7969:
7964:
7959:
7954:
7949:
7943:
7941:
7935:
7934:
7932:
7931:
7926:
7924:RNA polymerase
7921:
7915:
7913:
7909:
7908:
7906:
7905:
7900:
7895:
7889:
7887:
7880:
7874:
7873:
7871:
7870:
7869:
7868:
7863:
7858:
7848:
7847:
7846:
7828:
7822:
7820:
7812:
7811:
7806:
7804:
7803:
7796:
7789:
7781:
7772:
7771:
7769:
7768:
7757:
7754:
7753:
7751:
7750:
7745:
7739:
7737:
7733:
7732:
7730:
7729:
7724:
7719:
7717:Plant genetics
7714:
7709:
7704:
7699:
7694:
7689:
7684:
7679:
7674:
7669:
7664:
7659:
7657:Genome editing
7654:
7649:
7644:
7638:
7636:
7635:Related topics
7632:
7631:
7629:
7628:
7623:
7618:
7613:
7608:
7603:
7598:
7592:
7590:
7583:
7582:
7580:
7579:
7574:
7569:
7564:
7559:
7557:Immunogenetics
7554:
7549:
7544:
7539:
7533:
7531:
7527:
7526:
7524:
7523:
7518:
7513:
7508:
7503:
7498:
7493:
7488:
7483:
7478:
7472:
7470:
7469:Key components
7466:
7465:
7463:
7462:
7457:
7452:
7447:
7442:
7437:
7431:
7428:
7427:
7422:
7420:
7419:
7412:
7405:
7397:
7391:
7390:
7385:
7378:
7371:
7370:External links
7368:
7366:
7365:
7327:(16): 5758–9.
7312:
7306:
7289:
7283:
7268:
7262:
7246:
7244:
7241:
7238:
7237:
7208:(4): 405–412.
7188:
7139:
7078:
7043:(3): 297–313.
7027:
6966:
6951:
6890:
6831:
6777:
6720:
6663:
6610:(5): 483–490.
6590:
6571:(7): 481–486.
6551:
6492:
6441:
6400:
6349:
6328:(9): R215–20.
6308:
6269:
6234:(5): 536–549.
6218:
6166:
6107:
6064:
6020:
5983:(1–2): 10–22.
5956:
5897:
5846:
5803:
5749:
5724:
5717:
5703:Yarus, Michael
5691:
5632:
5583:
5532:
5497:
5462:
5419:
5376:
5325:
5276:
5223:
5170:
5121:
5070:
5019:
4990:(11): 160246.
4970:
4919:
4862:
4813:
4762:
4719:
4693:
4638:
4629:
4594:Bioinformatics
4577:
4518:
4474:
4456:
4431:
4394:(4): 579–583.
4388:Bioinformatics
4371:
4362:
4355:
4335:
4328:
4308:
4259:
4216:
4157:
4134:
4075:
4068:
4050:
3999:
3964:Human Mutation
3950:
3909:
3850:
3809:
3760:
3753:
3731:
3718:
3691:
3650:
3643:
3620:
3598:
3531:
3493:
3436:
3377:
3351:
3307:Rappsilber, J.
3297:
3290:
3270:
3221:
3186:
3167:(8): 1600097.
3151:
3144:
3118:
3069:
3043:
2984:
2958:
2899:
2840:
2781:
2752:(5): 815–818.
2732:
2693:
2678:
2658:
2632:
2580:
2555:
2518:(3): 789–795.
2498:
2443:
2428:
2412:Crick, Francis
2403:
2366:(3): 707–730.
2346:
2283:
2233:
2232:
2230:
2227:
2226:
2225:
2220:
2213:
2210:
2209:
2208:
2200:
2192:
2171:
2160:
2153:
2142:
2141:
2134:
2130:
2083:
2080:
2061:
2058:
1961:
1958:
1944:selenocysteine
1939:
1936:
1928:
1925:
1921:
1920:
1917:
1916:
1913:
1910:
1907:
1904:
1901:
1898:
1895:
1892:
1889:
1886:
1883:
1880:
1877:
1874:
1871:
1868:
1865:
1862:
1859:
1855:
1854:
1851:
1848:
1845:
1842:
1839:
1836:
1833:
1830:
1827:
1824:
1821:
1818:
1815:
1812:
1809:
1806:
1803:
1800:
1797:
1793:
1792:
1789:
1786:
1783:
1780:
1777:
1774:
1771:
1768:
1765:
1762:
1759:
1756:
1753:
1750:
1747:
1744:
1741:
1738:
1735:
1731:
1730:
1727:
1724:
1721:
1718:
1715:
1712:
1709:
1706:
1703:
1700:
1697:
1694:
1691:
1688:
1685:
1682:
1679:
1676:
1673:
1669:
1668:
1665:
1662:
1659:
1656:
1653:
1650:
1647:
1644:
1641:
1638:
1635:
1632:
1629:
1626:
1623:
1620:
1617:
1614:
1611:
1607:
1606:
1603:
1600:
1597:
1594:
1591:
1588:
1585:
1582:
1579:
1576:
1573:
1570:
1567:
1564:
1561:
1558:
1555:
1552:
1549:
1545:
1544:
1541:
1538:
1535:
1532:
1529:
1526:
1523:
1520:
1517:
1514:
1511:
1508:
1505:
1502:
1499:
1496:
1493:
1490:
1487:
1483:
1482:
1479:
1476:
1473:
1470:
1467:
1464:
1461:
1458:
1455:
1452:
1449:
1446:
1443:
1440:
1437:
1434:
1431:
1428:
1425:
1421:
1420:
1417:
1414:
1411:
1408:
1405:
1402:
1399:
1396:
1393:
1390:
1387:
1384:
1381:
1378:
1375:
1372:
1369:
1366:
1363:
1359:
1358:
1355:
1352:
1349:
1346:
1343:
1340:
1337:
1334:
1331:
1328:
1325:
1322:
1319:
1316:
1313:
1310:
1307:
1304:
1301:
1297:
1296:
1293:
1290:
1287:
1284:
1281:
1278:
1275:
1272:
1269:
1266:
1263:
1260:
1257:
1254:
1251:
1248:
1245:
1242:
1239:
1235:
1234:
1231:
1228:
1225:
1222:
1219:
1216:
1213:
1210:
1207:
1204:
1201:
1198:
1195:
1192:
1189:
1186:
1183:
1180:
1177:
1173:
1172:
1169:
1166:
1163:
1160:
1157:
1154:
1151:
1148:
1145:
1142:
1139:
1136:
1133:
1130:
1127:
1124:
1121:
1118:
1115:
1111:
1110:
1107:
1104:
1101:
1098:
1095:
1092:
1089:
1086:
1083:
1080:
1077:
1074:
1071:
1068:
1065:
1062:
1059:
1056:
1053:
1049:
1048:
1045:
1042:
1039:
1036:
1033:
1030:
1027:
1024:
1021:
1018:
1015:
1012:
1009:
1006:
1003:
1000:
997:
994:
991:
987:
986:
983:
980:
977:
974:
971:
968:
965:
962:
959:
956:
953:
950:
947:
944:
941:
938:
935:
932:
929:
925:
924:
921:
915:
912:
909:
906:
904:
901:
895:
892:
889:
886:
884:
881:
875:
872:
869:
866:
864:
861:
855:
852:
849:
841:
840:
837:
832:
819:
816:
803:hydropathicity
798:hydrophobicity
794:hydrophilicity
778:IUPAC notation
734:hydropathicity
722:Main article:
719:
716:
607:polymerization
586:
583:
579:release factor
573:), and UAA is
526:Shine-Dalgarno
517:
514:
491:reading frames
487:reading frames
474:
471:
429:
426:
350:Main article:
347:
344:
206:
203:
144:
141:
104:(mRNA), using
96:, which links
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
9516:
9505:
9502:
9500:
9497:
9495:
9492:
9490:
9487:
9485:
9482:
9481:
9479:
9472:
9462:
9461:
9456:
9452:
9450:
9449:
9440:
9438:
9437:
9428:
9427:
9424:
9418:
9415:
9413:
9410:
9408:
9405:
9403:
9400:
9398:
9395:
9393:
9390:
9388:
9385:
9383:
9380:
9378:
9375:
9373:
9370:
9368:
9365:
9363:
9360:
9358:
9355:
9353:
9350:
9349:
9347:
9343:
9334:
9328:
9323:
9317:
9312:
9311:Life on Earth
9306:
9305:
9299:
9293:
9288:
9282:
9277:
9271:
9266:
9260:
9255:
9249:
9244:
9238:
9233:
9227:
9222:
9216:
9215:
9213:
9209:
9203:
9200:
9198:
9195:
9193:
9190:
9188:
9185:
9183:
9182:Semisynthesis
9180:
9178:
9175:
9173:
9170:
9168:
9165:
9163:
9160:
9158:
9155:
9153:
9150:
9148:
9145:
9143:
9140:
9138:
9135:
9133:
9130:
9128:
9125:
9123:
9120:
9118:
9115:
9113:
9110:
9108:
9105:
9103:
9100:
9098:
9095:
9093:
9090:
9088:
9085:
9083:
9080:
9079:
9077:
9075:
9069:
9059:
9056:
9054:
9053:Tetrapyrroles
9051:
9049:
9048:
9044:
9040:
9037:
9036:
9035:
9032:
9030:
9029:
9027:
9022:
9018:
9015:
9014:
9013:
9010:
9008:
9005:
9003:
9000:
8998:
8997:
8995:
8994:Nucleic acids
8990:
8988:
8985:
8983:
8980:
8978:
8977:Sphingolipids
8975:
8973:
8972:Phospholipids
8970:
8968:
8965:
8961:
8958:
8957:
8956:
8953:
8951:
8948:
8946:
8945:
8943:
8938:
8936:
8933:
8931:
8930:Glycoproteins
8928:
8926:
8923:
8921:
8920:
8918:
8917:Carbohydrates
8913:
8912:
8909:
8906:
8902:
8898:
8888:
8885:
8883:
8880:
8878:
8875:
8873:
8870:
8868:
8866:
8862:
8861:
8858:
8855:
8853:
8850:
8848:
8845:
8843:
8840:
8838:
8835:
8833:
8830:
8828:
8825:
8823:
8820:
8818:
8815:
8813:
8810:
8809:
8806:
8803:
8801:
8797:
8791:
8788:
8786:
8783:
8781:
8778:
8776:
8773:
8772:
8770:
8766:
8760:
8757:
8755:
8752:
8750:
8747:
8745:
8742:
8740:
8737:
8735:
8732:
8730:
8727:
8726:
8724:
8718:
8712:
8709:
8707:
8704:
8702:
8699:
8697:
8694:
8692:
8689:
8687:
8684:
8682:
8679:
8677:
8674:
8672:
8669:
8667:
8664:
8660:
8657:
8656:
8655:
8652:
8648:
8645:
8644:
8643:
8640:
8638:
8635:
8633:
8630:
8629:
8627:
8623:
8619:
8613:
8610:
8608:
8605:
8603:
8600:
8598:
8595:
8593:
8590:
8588:
8585:
8583:
8580:
8578:
8575:
8573:
8570:
8568:
8565:
8564:
8562:
8558:
8553:
8546:
8541:
8539:
8534:
8532:
8527:
8526:
8523:
8510:
8507:
8505:
8504:D-amino acids
8502:
8500:
8497:
8495:
8492:
8490:
8487:
8485:
8482:
8480:
8477:
8475:
8472:
8470:
8466:
8464:
8460:
8459:
8455:
8444:
8441:
8438:
8435:
8432:
8429:
8426:
8425:Glutamic acid
8423:
8420:
8419:Aspartic acid
8417:
8416:
8414:
8408:
8403:
8397:
8394:
8391:
8388:
8385:
8382:
8379:
8376:
8375:
8373:
8367:
8362:
8356:
8353:
8351:
8348:
8346:
8343:
8341:
8338:
8337:
8335:
8332:
8328:
8322:
8321:Phenylalanine
8319:
8317:
8314:
8312:
8309:
8307:
8304:
8303:
8301:
8299:
8295:
8289:
8286:
8284:
8281:
8279:
8276:
8274:
8271:
8268:
8265:
8263:
8260:
8258:
8254:
8251:
8250:
8248:
8246:
8242:
8239:
8237:By properties
8235:
8230:
8220:
8217:
8215:
8212:
8210:
8207:
8206:
8204:
8200:
8196:
8189:
8184:
8182:
8177:
8175:
8170:
8169:
8166:
8154:
8153:Jacques Monod
8151:
8149:
8146:
8145:
8143:
8139:
8131:
8128:
8126:
8123:
8122:
8121:
8118:
8116:
8115:Translational
8113:
8109:
8106:
8104:
8101:
8099:
8096:
8095:
8094:
8091:
8089:
8086:
8082:
8079:
8077:
8074:
8073:
8072:
8069:
8065:
8062:
8061:
8060:
8057:
8056:
8054:
8052:
8048:
8038:
8035:
8033:
8030:
8028:
8025:
8023:
8020:
8019:
8017:
8013:
8007:
8004:
8002:
7999:
7997:
7994:
7993:
7991:
7987:
7984:
7982:
7978:
7968:
7965:
7963:
7960:
7958:
7955:
7953:
7950:
7948:
7945:
7944:
7942:
7940:
7936:
7930:
7927:
7925:
7922:
7920:
7917:
7916:
7914:
7910:
7904:
7901:
7899:
7896:
7894:
7891:
7890:
7888:
7884:
7881:
7879:
7878:Transcription
7875:
7867:
7864:
7862:
7859:
7857:
7854:
7853:
7852:
7849:
7845:
7841:
7837:
7834:
7833:
7832:
7831:Central dogma
7829:
7827:
7824:
7823:
7821:
7819:
7813:
7809:
7802:
7797:
7795:
7790:
7788:
7783:
7782:
7779:
7767:
7759:
7758:
7755:
7749:
7746:
7744:
7741:
7740:
7738:
7734:
7728:
7725:
7723:
7720:
7718:
7715:
7713:
7710:
7708:
7705:
7703:
7700:
7698:
7695:
7693:
7690:
7688:
7685:
7683:
7680:
7678:
7675:
7673:
7670:
7668:
7665:
7663:
7660:
7658:
7655:
7653:
7650:
7648:
7645:
7643:
7640:
7639:
7637:
7633:
7627:
7624:
7622:
7619:
7617:
7614:
7612:
7609:
7607:
7604:
7602:
7599:
7597:
7594:
7593:
7591:
7588:
7584:
7578:
7575:
7573:
7570:
7568:
7565:
7563:
7560:
7558:
7555:
7553:
7550:
7548:
7545:
7543:
7540:
7538:
7535:
7534:
7532:
7528:
7522:
7519:
7517:
7514:
7512:
7509:
7507:
7504:
7502:
7499:
7497:
7494:
7492:
7489:
7487:
7484:
7482:
7479:
7477:
7474:
7473:
7471:
7467:
7461:
7458:
7456:
7453:
7451:
7448:
7446:
7443:
7441:
7438:
7436:
7433:
7432:
7429:
7425:
7418:
7413:
7411:
7406:
7404:
7399:
7398:
7395:
7389:
7386:
7383:
7379:
7377:
7374:
7373:
7369:
7362:
7358:
7353:
7348:
7343:
7338:
7334:
7330:
7326:
7322:
7318:
7313:
7309:
7307:9780716737063
7303:
7298:
7297:
7290:
7286:
7280:
7276:
7275:
7269:
7265:
7259:
7255:
7254:
7248:
7247:
7242:
7233:
7229:
7224:
7219:
7215:
7211:
7207:
7203:
7199:
7192:
7189:
7184:
7180:
7175:
7170:
7166:
7162:
7158:
7154:
7150:
7143:
7140:
7135:
7131:
7127:
7123:
7119:
7115:
7111:
7107:
7102:
7097:
7094:(3): 362–76.
7093:
7089:
7082:
7079:
7074:
7070:
7066:
7062:
7058:
7054:
7050:
7046:
7042:
7038:
7031:
7028:
7023:
7019:
7015:
7011:
7007:
7003:
6999:
6995:
6990:
6985:
6982:(4): 048101.
6981:
6977:
6970:
6967:
6962:
6955:
6952:
6947:
6943:
6939:
6935:
6931:
6927:
6923:
6919:
6914:
6909:
6906:(2): 331–42.
6905:
6901:
6894:
6891:
6886:
6882:
6877:
6872:
6867:
6862:
6858:
6854:
6850:
6846:
6842:
6835:
6832:
6827:
6823:
6819:
6815:
6811:
6807:
6803:
6799:
6795:
6791:
6784:
6782:
6778:
6773:
6769:
6764:
6759:
6755:
6751:
6747:
6743:
6739:
6735:
6731:
6724:
6721:
6716:
6712:
6707:
6702:
6698:
6694:
6690:
6686:
6682:
6678:
6674:
6667:
6664:
6659:
6655:
6651:
6647:
6643:
6639:
6635:
6631:
6627:
6623:
6618:
6613:
6609:
6605:
6601:
6594:
6591:
6586:
6582:
6578:
6574:
6570:
6566:
6562:
6555:
6552:
6547:
6543:
6538:
6533:
6528:
6523:
6519:
6515:
6511:
6507:
6503:
6496:
6493:
6488:
6484:
6480:
6476:
6472:
6468:
6464:
6460:
6456:
6452:
6445:
6442:
6437:
6433:
6428:
6423:
6419:
6415:
6411:
6404:
6401:
6396:
6392:
6388:
6384:
6380:
6376:
6372:
6368:
6364:
6360:
6353:
6350:
6345:
6341:
6336:
6331:
6327:
6323:
6319:
6312:
6309:
6304:
6300:
6296:
6292:
6288:
6284:
6280:
6273:
6270:
6265:
6261:
6257:
6253:
6249:
6245:
6241:
6237:
6233:
6229:
6222:
6219:
6214:
6210:
6205:
6200:
6196:
6192:
6189:(5): 406–29.
6188:
6184:
6180:
6173:
6171:
6167:
6162:
6158:
6153:
6148:
6143:
6138:
6134:
6130:
6126:
6122:
6118:
6111:
6108:
6103:
6099:
6095:
6091:
6087:
6083:
6080:(3): 367–79.
6079:
6075:
6068:
6065:
6060:
6056:
6051:
6046:
6043:(4): 511–18.
6042:
6038:
6034:
6027:
6025:
6021:
6016:
6012:
6007:
6002:
5998:
5994:
5990:
5986:
5982:
5978:
5974:
5967:
5965:
5963:
5961:
5957:
5952:
5948:
5943:
5938:
5933:
5928:
5924:
5920:
5917:(2): 1083–6.
5916:
5912:
5908:
5901:
5898:
5893:
5889:
5885:
5881:
5877:
5873:
5869:
5865:
5862:(4): 288–93.
5861:
5857:
5850:
5847:
5842:
5838:
5834:
5830:
5826:
5822:
5819:(3): 177–87.
5818:
5814:
5807:
5804:
5799:
5795:
5791:
5787:
5783:
5779:
5775:
5771:
5768:(3): 238–48.
5767:
5763:
5756:
5754:
5750:
5738:
5734:
5728:
5725:
5720:
5714:
5710:
5709:
5704:
5698:
5696:
5692:
5687:
5683:
5678:
5673:
5668:
5663:
5659:
5655:
5651:
5647:
5643:
5636:
5633:
5628:
5624:
5619:
5614:
5610:
5606:
5602:
5598:
5594:
5587:
5584:
5579:
5575:
5570:
5565:
5560:
5555:
5551:
5547:
5543:
5536:
5533:
5528:
5524:
5520:
5516:
5512:
5508:
5501:
5498:
5493:
5489:
5485:
5481:
5477:
5473:
5466:
5463:
5458:
5454:
5450:
5446:
5442:
5438:
5434:
5430:
5423:
5420:
5415:
5411:
5407:
5403:
5399:
5395:
5391:
5387:
5380:
5377:
5372:
5368:
5364:
5360:
5356:
5352:
5348:
5344:
5340:
5336:
5329:
5326:
5321:
5317:
5312:
5307:
5303:
5299:
5296:(2): 91–111.
5295:
5291:
5287:
5280:
5277:
5272:
5268:
5263:
5258:
5254:
5250:
5246:
5242:
5238:
5236:
5227:
5224:
5219:
5215:
5210:
5205:
5201:
5197:
5193:
5189:
5185:
5183:
5174:
5171:
5166:
5162:
5157:
5152:
5148:
5144:
5140:
5136:
5132:
5125:
5122:
5117:
5113:
5108:
5103:
5098:
5093:
5089:
5085:
5084:PLOS Genetics
5081:
5074:
5071:
5066:
5062:
5057:
5052:
5047:
5042:
5038:
5034:
5030:
5023:
5020:
5015:
5011:
5006:
5001:
4997:
4993:
4989:
4985:
4981:
4974:
4971:
4966:
4962:
4957:
4952:
4947:
4942:
4938:
4934:
4930:
4923:
4920:
4915:
4911:
4906:
4901:
4897:
4893:
4889:
4885:
4881:
4877:
4873:
4866:
4863:
4858:
4854:
4849:
4844:
4840:
4836:
4833:(9): 1481–6.
4832:
4828:
4824:
4817:
4814:
4809:
4805:
4800:
4795:
4790:
4785:
4781:
4777:
4773:
4766:
4763:
4758:
4754:
4750:
4746:
4742:
4738:
4734:
4730:
4723:
4720:
4707:
4700:
4698:
4694:
4690:
4685:
4681:
4677:
4673:
4669:
4665:
4661:
4657:
4653:
4649:
4642:
4639:
4633:
4630:
4625:
4621:
4616:
4611:
4607:
4603:
4599:
4595:
4591:
4584:
4582:
4578:
4573:
4569:
4564:
4559:
4554:
4549:
4545:
4541:
4537:
4533:
4529:
4522:
4519:
4514:
4510:
4505:
4500:
4496:
4492:
4488:
4481:
4479:
4475:
4470:
4466:
4460:
4457:
4445:
4441:
4435:
4432:
4427:
4423:
4419:
4415:
4410:
4405:
4401:
4397:
4393:
4389:
4385:
4378:
4376:
4372:
4366:
4363:
4358:
4352:
4348:
4347:
4339:
4336:
4331:
4325:
4321:
4320:
4312:
4309:
4304:
4300:
4295:
4290:
4286:
4282:
4278:
4274:
4270:
4263:
4260:
4255:
4251:
4247:
4243:
4239:
4235:
4231:
4227:
4220:
4217:
4212:
4208:
4203:
4198:
4193:
4188:
4184:
4180:
4176:
4172:
4168:
4161:
4158:
4153:
4149:
4145:
4138:
4135:
4130:
4126:
4121:
4116:
4111:
4106:
4102:
4098:
4094:
4090:
4086:
4079:
4076:
4071:
4065:
4061:
4054:
4051:
4046:
4042:
4038:
4034:
4030:
4026:
4022:
4018:
4014:
4010:
4003:
4000:
3995:
3991:
3987:
3983:
3978:
3973:
3969:
3965:
3961:
3954:
3951:
3946:
3942:
3937:
3932:
3928:
3924:
3920:
3913:
3910:
3905:
3901:
3896:
3891:
3886:
3881:
3877:
3873:
3870:(6): 2886–9.
3869:
3865:
3861:
3854:
3851:
3846:
3842:
3837:
3832:
3828:
3824:
3820:
3813:
3810:
3805:
3801:
3796:
3791:
3787:
3783:
3779:
3775:
3771:
3764:
3761:
3756:
3750:
3746:
3742:
3735:
3732:
3728:
3722:
3719:
3706:
3702:
3695:
3692:
3687:
3683:
3678:
3673:
3669:
3665:
3661:
3654:
3651:
3646:
3640:
3636:
3635:
3627:
3625:
3621:
3617:
3616:
3611:
3607:
3602:
3599:
3594:
3590:
3586:
3582:
3577:
3572:
3568:
3564:
3560:
3556:
3552:
3548:
3547:
3542:
3535:
3532:
3519:
3515:
3514:
3508:
3503:
3497:
3494:
3489:
3485:
3480:
3475:
3471:
3467:
3463:
3459:
3455:
3451:
3447:
3440:
3437:
3432:
3428:
3423:
3418:
3413:
3408:
3404:
3400:
3396:
3392:
3388:
3381:
3378:
3365:
3361:
3355:
3352:
3346:
3342:
3337:
3332:
3328:
3324:
3320:
3316:
3312:
3308:
3301:
3298:
3293:
3287:
3283:
3282:
3274:
3271:
3266:
3262:
3257:
3252:
3248:
3244:
3241:(3): 323–36.
3240:
3236:
3232:
3225:
3222:
3217:
3213:
3209:
3205:
3202:(6): 548–54.
3201:
3197:
3190:
3187:
3182:
3178:
3174:
3170:
3166:
3162:
3155:
3152:
3147:
3145:9783527312436
3141:
3137:
3133:
3129:
3122:
3119:
3114:
3110:
3105:
3100:
3096:
3092:
3089:(2): 575–82.
3088:
3084:
3080:
3073:
3070:
3066:
3053:
3047:
3044:
3039:
3035:
3030:
3025:
3020:
3015:
3011:
3007:
3004:(5): 1161–8.
3003:
2999:
2995:
2988:
2985:
2981:
2968:
2962:
2959:
2954:
2950:
2945:
2940:
2935:
2930:
2926:
2922:
2919:(1): 116–22.
2918:
2914:
2910:
2903:
2900:
2895:
2891:
2886:
2881:
2876:
2871:
2867:
2863:
2859:
2855:
2851:
2844:
2841:
2836:
2832:
2827:
2822:
2817:
2812:
2808:
2804:
2800:
2796:
2792:
2785:
2782:
2777:
2773:
2769:
2765:
2760:
2755:
2751:
2747:
2743:
2736:
2733:
2728:
2724:
2720:
2716:
2712:
2708:
2704:
2697:
2694:
2689:
2685:
2681:
2675:
2671:
2670:
2662:
2659:
2647:
2643:
2636:
2633:
2628:
2624:
2620:
2616:
2611:
2606:
2602:
2598:
2594:
2587:
2585:
2581:
2569:
2565:
2559:
2556:
2551:
2547:
2542:
2537:
2533:
2529:
2525:
2521:
2517:
2513:
2509:
2502:
2499:
2494:
2490:
2486:
2482:
2478:
2474:
2470:
2466:
2462:
2458:
2454:
2447:
2444:
2439:
2435:
2431:
2429:9780465091386
2425:
2421:
2417:
2413:
2407:
2404:
2399:
2395:
2390:
2385:
2381:
2377:
2373:
2369:
2365:
2361:
2357:
2350:
2347:
2342:
2338:
2334:
2330:
2326:
2322:
2318:
2314:
2310:
2306:
2302:
2298:
2294:
2287:
2284:
2279:
2275:
2270:
2265:
2261:
2257:
2253:
2249:
2245:
2238:
2235:
2228:
2224:
2221:
2219:
2216:
2215:
2211:
2206:
2201:
2197:
2193:
2190:
2186:
2181:
2176:
2172:
2169:
2165:
2161:
2158:
2154:
2151:
2147:
2146:
2145:
2139:
2135:
2131:
2128:
2124:
2120:
2119:
2118:
2115:
2111:
2107:
2103:
2101:
2097:
2093:
2089:
2081:
2079:
2077:
2072:
2066:
2059:
2057:
2053:
2051:
2050:
2045:
2041:
2039:
2034:
2029:
2027:
2023:
2019:
2015:
2011:
2010:
2005:
2004:
1999:
1995:
1985:
1981:
1977:
1974:Genetic code
1972:
1967:
1959:
1957:
1955:
1954:
1949:
1945:
1937:
1934:
1926:
1914:
1911:
1908:
1905:
1902:
1899:
1896:
1893:
1890:
1887:
1884:
1881:
1878:
1875:
1872:
1869:
1866:
1863:
1860:
1857:
1856:
1852:
1849:
1846:
1843:
1840:
1837:
1834:
1831:
1828:
1825:
1822:
1819:
1816:
1813:
1810:
1807:
1804:
1801:
1798:
1795:
1794:
1790:
1787:
1784:
1781:
1778:
1775:
1772:
1769:
1766:
1763:
1760:
1757:
1754:
1751:
1748:
1745:
1742:
1739:
1736:
1733:
1732:
1728:
1725:
1722:
1719:
1716:
1713:
1710:
1707:
1704:
1701:
1698:
1695:
1692:
1689:
1686:
1683:
1680:
1677:
1674:
1671:
1670:
1666:
1663:
1660:
1657:
1654:
1651:
1648:
1645:
1642:
1639:
1636:
1633:
1630:
1627:
1624:
1621:
1618:
1615:
1612:
1609:
1608:
1604:
1601:
1598:
1595:
1592:
1589:
1586:
1583:
1580:
1577:
1574:
1571:
1568:
1565:
1562:
1559:
1556:
1553:
1550:
1547:
1546:
1542:
1539:
1536:
1533:
1530:
1527:
1524:
1521:
1518:
1515:
1512:
1509:
1506:
1503:
1500:
1497:
1494:
1491:
1488:
1485:
1484:
1480:
1477:
1474:
1471:
1468:
1465:
1462:
1459:
1456:
1453:
1450:
1447:
1444:
1441:
1438:
1435:
1432:
1429:
1426:
1423:
1422:
1418:
1415:
1412:
1409:
1406:
1403:
1400:
1397:
1394:
1391:
1388:
1385:
1382:
1379:
1376:
1373:
1370:
1367:
1364:
1361:
1360:
1356:
1353:
1350:
1347:
1344:
1341:
1338:
1335:
1332:
1329:
1326:
1323:
1320:
1317:
1314:
1311:
1308:
1305:
1302:
1299:
1298:
1294:
1291:
1288:
1285:
1282:
1279:
1276:
1273:
1270:
1267:
1264:
1261:
1258:
1255:
1252:
1249:
1246:
1243:
1240:
1237:
1236:
1232:
1229:
1226:
1223:
1220:
1217:
1214:
1211:
1208:
1205:
1202:
1199:
1196:
1193:
1190:
1187:
1184:
1181:
1178:
1175:
1174:
1170:
1167:
1164:
1161:
1158:
1155:
1152:
1149:
1146:
1143:
1140:
1137:
1134:
1131:
1128:
1125:
1122:
1119:
1116:
1113:
1112:
1108:
1105:
1102:
1099:
1096:
1093:
1090:
1087:
1084:
1081:
1078:
1075:
1072:
1069:
1066:
1063:
1060:
1057:
1054:
1051:
1050:
1046:
1043:
1040:
1037:
1034:
1031:
1028:
1025:
1022:
1019:
1016:
1013:
1010:
1007:
1004:
1001:
998:
995:
992:
989:
988:
984:
981:
978:
975:
972:
969:
966:
963:
960:
957:
954:
951:
948:
945:
942:
939:
936:
933:
930:
927:
926:
922:
920:
916:
913:
910:
907:
902:
900:
896:
893:
890:
887:
882:
880:
876:
873:
870:
867:
862:
860:
856:
853:
850:
847:
846:
843:
842:
836:
835:
831:
829:
825:
817:
815:
812:
808:
804:
799:
795:
791:
787:
783:
779:
775:
771:
767:
763:
762:glutamic acid
759:
751:
746:
740:is available.
739:
735:
730:
725:
717:
715:
713:
709:
705:
704:immune system
701:
697:
693:
689:
683:
681:
676:
675:
670:
666:
662:
658:
654:
649:
647:
642:
638:
634:
630:
626:
622:
620:
616:
612:
608:
604:
596:
591:
584:
582:
580:
576:
572:
568:
564:
560:
555:
553:
549:
545:
541:
537:
533:
532:
527:
523:
515:
513:
511:
507:
503:
498:
496:
492:
488:
484:
480:
473:Reading frame
472:
467:
463:
459:
455:
451:
447:
446:
441:
440:
434:
427:
425:
422:
418:
417:
413:
408:
405:
402:
400:
396:
392:
388:
383:
381:
377:
372:
370:
364:
359:
353:
345:
343:
339:
336:
332:
328:
324:
319:
317:
313:
309:
305:
301:
296:
294:
290:
286:
282:
278:
274:
270:
266:
261:
259:
258:phenylalanine
255:
251:
247:
243:
239:
235:
231:
229:
225:
219:
212:
204:
202:
198:
196:
192:
187:
180:
176:
172:
168:
167:Francis Crick
164:
158:
149:
142:
140:
138:
134:
133:variant codes
130:
126:
122:
118:
113:
111:
107:
103:
102:messenger RNA
99:
95:
91:
87:
83:
79:
75:
71:
67:
59:
55:
51:
47:
43:
39:
38:messenger RNA
34:
30:
19:
9471:
9458:
9446:
9434:
9407:Robert Hooke
9401:
9381:
9351:
9336:}}
9330:{{
9325:}}
9319:{{
9314:}}
9308:{{
9301:}}
9295:{{
9290:}}
9284:{{
9279:}}
9273:{{
9268:}}
9262:{{
9257:}}
9251:{{
9246:}}
9240:{{
9235:}}
9229:{{
9224:}}
9218:{{
9097:Biosynthesis
9046:
9045:
9024:
9023:
8992:
8991:
8940:
8939:
8915:
8914:
8887:Genetic code
8886:
8863:
8701:parasitology
8696:Bacteriology
8659:Pharmacology
8637:Cell biology
8622:Biochemistry
8581:
8552:Biochemistry
8461:
8406:
8365:
8219:Genetic code
8218:
8130:irreversible
8015:Key elements
7912:Key elements
7826:Genetic code
7825:
7816:Introduction
7667:Genetic code
7666:
7601:the Americas
7577:Quantitative
7547:Cytogenetics
7542:Conservation
7435:Introduction
7324:
7320:
7295:
7273:
7252:
7205:
7201:
7191:
7156:
7152:
7142:
7091:
7087:
7081:
7040:
7036:
7030:
6979:
6975:
6969:
6960:
6954:
6903:
6899:
6893:
6851:(5): e5708.
6848:
6844:
6834:
6793:
6789:
6737:
6733:
6723:
6680:
6676:
6666:
6607:
6604:Astrobiology
6603:
6593:
6568:
6564:
6554:
6509:
6505:
6495:
6457:(5): 473–6.
6454:
6450:
6444:
6417:
6413:
6403:
6362:
6358:
6352:
6325:
6321:
6311:
6289:(6): 241–7.
6286:
6282:
6272:
6231:
6228:Astrobiology
6227:
6221:
6186:
6182:
6124:
6120:
6110:
6077:
6073:
6067:
6040:
6036:
5980:
5976:
5914:
5910:
5900:
5859:
5855:
5849:
5816:
5812:
5806:
5765:
5761:
5740:. Retrieved
5736:
5727:
5707:
5649:
5645:
5635:
5600:
5596:
5586:
5549:
5545:
5535:
5510:
5506:
5500:
5475:
5471:
5465:
5432:
5428:
5422:
5389:
5385:
5379:
5338:
5334:
5328:
5293:
5289:
5279:
5247:(1): 61–68.
5244:
5240:
5237:translation"
5234:
5226:
5191:
5187:
5181:
5173:
5138:
5134:
5124:
5087:
5083:
5073:
5036:
5032:
5022:
4987:
4983:
4973:
4936:
4932:
4922:
4879:
4875:
4865:
4830:
4826:
4816:
4779:
4775:
4765:
4732:
4728:
4722:
4710:. Retrieved
4651:
4647:
4641:
4632:
4597:
4593:
4535:
4531:
4521:
4494:
4490:
4468:
4459:
4447:. Retrieved
4443:
4434:
4391:
4387:
4365:
4345:
4338:
4318:
4311:
4276:
4272:
4262:
4229:
4225:
4219:
4174:
4170:
4160:
4152:the original
4147:
4137:
4092:
4088:
4078:
4059:
4053:
4015:(2): 122–9.
4012:
4008:
4002:
3970:(4): 361–3.
3967:
3963:
3953:
3929:(1): 39–59.
3926:
3922:
3912:
3867:
3863:
3853:
3829:(1): 39–59.
3826:
3822:
3812:
3777:
3773:
3763:
3744:
3734:
3721:
3709:. Retrieved
3704:
3694:
3667:
3663:
3653:
3633:
3613:
3606:Homo sapiens
3605:
3601:
3550:
3544:
3534:
3522:. Retrieved
3511:
3502:Zimmer, Carl
3496:
3453:
3449:
3439:
3394:
3390:
3380:
3368:. Retrieved
3363:
3354:
3318:
3314:
3300:
3280:
3273:
3238:
3234:
3224:
3199:
3195:
3189:
3164:
3160:
3154:
3127:
3121:
3086:
3082:
3072:
3063:
3056:. Retrieved
3046:
3001:
2997:
2987:
2978:
2971:. Retrieved
2961:
2916:
2912:
2902:
2857:
2853:
2843:
2798:
2794:
2784:
2749:
2745:
2735:
2713:(1): 12–17.
2710:
2706:
2696:
2668:
2661:
2649:. Retrieved
2645:
2635:
2600:
2596:
2571:. Retrieved
2567:
2558:
2515:
2511:
2501:
2460:
2456:
2446:
2419:
2406:
2363:
2359:
2349:
2300:
2296:
2286:
2251:
2247:
2237:
2223:Codon tables
2143:
2116:
2112:
2108:
2104:
2085:
2067:
2063:
2054:
2047:
2038:Condylostoma
2036:
2030:
2025:
2007:
2001:
1990:
1979:
1951:
1941:
821:
810:
789:
785:
773:
769:
765:
758:codon tables
755:
707:
684:
672:
650:
623:
600:
574:
570:
566:
562:
556:
529:
528:sequence in
519:
499:
476:
465:
461:
457:
453:
443:
437:
414:
409:
406:
403:
398:
384:
379:
373:
365:
361:
340:
323:Philip Leder
320:
308:transfer RNA
297:
265:Severo Ochoa
262:
232:
227:
221:
199:
191:RNA Tie Club
188:
179:George Gamow
171:James Watson
160:
137:mitochondria
135:(such as in
128:
114:
106:transfer RNA
66:genetic code
65:
63:
29:
9397:Experiments
9352:Occupations
9265:Amino acids
9034:Amino acids
9012:Nucleotides
9007:Nucleosides
9002:Nucleobases
8982:Cholesterol
8955:Fatty acids
8950:Eicosanoids
8901:Biomolecule
8872:Chromosomes
8832:Amino acids
8812:Cell theory
8790:Biomolecule
8499:Imino acids
8463:Amino acids
8396:Pyrrolysine
8333:, uncharged
7981:Translation
7818:to genetics
7647:Epigenetics
5813:Bio Systems
5513:: 379–408.
5478:: 413–444.
5135:Bio Systems
4729:Experientia
3065:synthesis'.
3058:27 February
2973:27 February
2463:(1): 8–14.
2205:frame shift
2042:or trigger
2014:totiviruses
1948:pyrrolysine
828:translation
641:thalassemia
559:stop codons
522:start codon
495:start codon
254:polypeptide
110:nucleotides
42:nucleotides
9478:Categories
9377:Microscopy
8967:Glycerides
8935:Glycosides
8785:Metabolism
8780:Enzymology
8711:immunology
8489:Glucogenic
8340:Asparagine
8316:Tryptophan
8273:Methionine
8262:Isoleucine
8125:reversible
8088:lac operon
8064:imprinting
8059:Epigenetic
8051:Regulation
8006:Eukaryotic
7952:5' capping
7903:Eukaryotic
7652:Geneticist
7626:South Asia
7572:Population
7552:Ecological
7521:Amino acid
7501:Nucleotide
7476:Chromosome
5742:3 February
5290:IUBMB Life
5039:: e03640.
4449:4 February
3370:9 February
2438:1020240407
2229:References
2157:metabolism
2003:Mycoplasma
1998:tryptophan
1964:See also:
1960:Variations
1931:See also:
718:Degeneracy
669:stop codon
657:insertions
646:stop codon
557:The three
540:methionine
504:, ORFs in
502:eukaryotes
395:tryptophan
356:See also:
316:enzymology
293:copolymers
216:See also:
46:amino acid
9382:Concepts:
9357:Biologist
9211:Templates
9074:synthesis
9072:Chemical
8722:chemistry
8572:Chemistry
8484:Ketogenic
8474:Essential
8390:Histidine
8355:Threonine
8345:Glutamine
8306:Histidine
8245:Aliphatic
7996:Bacterial
7893:Bacterial
7567:Molecular
7562:Microbial
7537:Classical
7101:1007.3906
6989:1007.4149
6913:1007.4122
6754:0305-1048
6697:0305-1048
6642:1531-1074
6617:0904.0402
6264:257983174
5141:: 16–25.
4984:Open Biol
3593:205571025
3456:: 14568.
2627:249112573
2619:2475-3025
2532:1943-2631
2493:121907709
2477:0003-0996
2380:0007-0882
2325:0028-0836
2164:mutations
2096:ribozymes
2060:Inference
1900:1,609,975
1870:1,143,534
1838:1,177,632
1776:1,020,595
1761:1,127,679
1652:1,295,568
1404:1,391,973
1374:1,611,801
698:that use
688:wild type
661:deletions
615:phenotype
611:mutations
595:mutations
565:, UGA is
485:in three
421:synthetic
412:bacterium
387:N. Budisa
382:) codon.
335:translate
331:ribosomes
246:translate
139:) exist.
74:translate
9489:Genetics
9436:Category
9026:Proteins
8987:Steroids
8925:Alcohols
8904:families
8865:Genetics
8817:Ribosome
8768:Concepts
8749:Catalyst
8739:Molecule
8720:General
8706:virology
8443:Tyrosine
8437:Cysteine
8384:Arginine
8311:Tyrosine
8298:Aromatic
8108:microRNA
8022:Ribosome
8001:Archaeal
7957:Splicing
7929:Promoter
7898:Archaeal
7842: →
7838: →
7766:Category
7692:Heredity
7662:Genomics
7506:Mutation
7496:Heredity
7460:Glossary
7450:Timeline
7424:Genetics
7361:24756939
7232:17293451
7183:23985735
7126:20558115
7065:16838217
7022:12246664
7014:18352335
6946:12206140
6938:17826800
6885:19479032
6845:PLOS ONE
6826:18823745
6818:14604186
6772:31504783
6715:28180287
6650:19566427
6585:19524038
6546:11087835
6487:23334860
6436:12270892
6395:15542587
6387:26054480
6303:10366854
6256:37022727
6213:19795157
6059:10742043
6015:21779963
5892:20803686
5798:20130470
5705:(2010).
5627:36952281
5578:34751130
5527:24555827
5492:20307192
5457:19385756
5449:16926858
5414:19385756
5406:16926858
5371:15542587
5363:26054480
5320:19117371
5271:27870834
5235:Euplotes
5218:27501944
5165:29030023
5116:27490485
5065:25247702
5014:27881739
4965:23638388
4914:19465905
4808:17121679
4757:19264964
4712:10 March
4624:21653513
4572:23185002
4513:15788401
4426:51968530
4418:30101307
4303:16489229
4273:Genetics
4211:10570172
4129:17409186
4045:32918971
3994:22693748
3945:17015226
3845:17015226
3804:15057282
3711:10 March
3686:12867081
3585:31092918
3518:Archived
3488:28220771
3431:28115716
3345:26136259
3265:19318213
3216:16260173
3181:28671771
3113:15514035
3083:Genetics
2953:13998282
2894:13946552
2835:14479932
2776:14249277
2768:17350564
2727:26549858
2688:47716375
2550:30846543
2512:Genetics
2485:27856930
2398:27924115
2333:13063483
2278:19131629
2212:See also
2185:topology
2168:ribosome
2150:aptamers
2127:peptides
2076:ciliates
2049:Euplotes
2018:bacteria
1915:669,768
1853:669,873
1791:903,565
1729:437,126
1667:486,463
1605:494,682
1543:791,383
1481:493,429
1419:464,485
1357:250,760
1295:423,516
1233:184,609
1171:535,595
1047:513,028
985:430,311
914:Fraction
894:Fraction
874:Fraction
854:Fraction
428:Features
385:In 2015
277:cytosine
94:ribosome
90:proteins
58:ribosome
9448:Commons
9362:Chemist
9345:Related
9333:Zoology
8847:Hormone
8842:Peptide
8837:Protein
8827:Peptide
8822:Nucleus
8754:Reagent
8577:Biology
8445:(≈10.1)
8386:(≈12.5)
8380:(≈10.8)
8288:Glycine
8283:Proline
8278:Alanine
8267:Leucine
8214:Peptide
8209:Protein
7861:RNA→DNA
7856:RNA→RNA
7844:Protein
7445:History
7440:Outline
7352:4000803
7329:Bibcode
7223:1832087
7174:3785830
7134:1845965
7106:Bibcode
7073:1260806
7045:Bibcode
6994:Bibcode
6918:Bibcode
6876:2682656
6853:Bibcode
6798:Bibcode
6763:6821194
6706:5388404
6658:9039622
6622:Bibcode
6514:Bibcode
6479:9115171
6459:Bibcode
6367:Bibcode
6344:9751648
6236:Bibcode
6191:Bibcode
6129:Bibcode
6102:4144681
6094:4887876
6006:3223571
5985:Bibcode
5951:6928661
5919:Bibcode
5884:2514270
5864:Bibcode
5841:2650752
5821:Bibcode
5790:9732450
5770:Bibcode
5686:9736730
5654:Bibcode
5618:1008964
5569:8629427
5343:Bibcode
5311:3293468
5262:5295771
5209:5062323
5143:Bibcode
5107:4973966
5056:4359377
5005:5133446
4956:3628385
4939:: e50.
4905:2834264
4884:Bibcode
4857:7784200
4799:1679813
4749:2253709
4684:4335828
4656:Bibcode
4615:3129529
4563:3529041
4540:Bibcode
4409:7109657
4294:1456385
4254:7041255
4234:Bibcode
4226:Science
4179:Bibcode
4148:Harvard
4120:1871816
4097:Bibcode
4037:8444142
4017:Bibcode
3986:8723688
3872:Bibcode
3576:7039709
3555:Bibcode
3479:5321798
3458:Bibcode
3422:5307467
3399:Bibcode
3336:4782924
3256:2696486
3104:1448817
3038:5330357
3006:Bibcode
2921:Bibcode
2862:Bibcode
2803:Bibcode
2651:21 July
2573:21 July
2541:6404253
2389:4990703
2341:4256010
2305:Bibcode
2269:3088105
2248:Science
2138:fitness
2022:archaea
1978:of the
1885:299,495
1823:643,471
1808:287,712
1746:588,138
1714:885,429
1699:750,096
1684:448,607
1637:246,105
1622:896,005
1590:993,621
1575:614,523
1560:304,565
1528:776,603
1513:768,147
1498:846,466
1466:689,701
1451:533,609
1436:650,473
1389:281,570
1342:501,911
1327:688,038
1312:290,751
1280:613,713
1265:804,620
1250:796,638
1218:441,711
1203:713,233
1188:536,515
1141:179,419
1126:525,688
1109:63,237
1079:496,448
1064:311,881
1032:622,407
1017:718,892
1002:824,692
970:495,699
955:618,711
940:714,298
923:Number
903:Number
883:Number
863:Number
708:E. coli
696:Viruses
674:in vivo
552:leucine
531:E. coli
510:introns
466:MT-ATP6
462:MT-ATP8
458:MT-ATP6
454:MT-ATP8
445:MT-ATP6
439:MT-ATP8
391:D. Söll
380:in vivo
289:proline
281:proline
269:adenine
248:a poly-
143:History
88:) into
54:thymine
9460:Portal
9322:Botany
9047:Other:
8942:Lipids
8852:Enzyme
8625:fields
8554:topics
8439:(≈8.3)
8433:(≈5.4)
8427:(≈4.1)
8421:(≈3.9)
8392:(≈6.1)
8378:Lysine
8350:Serine
8257:Valine
7611:Europe
7596:Africa
7530:Fields
7516:Allele
7491:Genome
7359:
7349:
7304:
7281:
7260:
7230:
7220:
7181:
7171:
7132:
7124:
7071:
7063:
7020:
7012:
6944:
6936:
6883:
6873:
6824:
6816:
6770:
6760:
6752:
6713:
6703:
6695:
6656:
6648:
6640:
6583:
6544:
6534:
6485:
6477:
6434:
6393:
6385:
6342:
6301:
6262:
6254:
6211:
6161:279919
6159:
6152:336109
6149:
6100:
6092:
6057:
6013:
6003:
5949:
5942:348428
5939:
5890:
5882:
5839:
5796:
5788:
5715:
5684:
5674:
5625:
5615:
5576:
5566:
5525:
5490:
5455:
5447:
5429:Nature
5412:
5404:
5369:
5361:
5318:
5308:
5269:
5259:
5216:
5206:
5163:
5114:
5104:
5063:
5053:
5012:
5002:
4963:
4953:
4912:
4902:
4876:Nature
4855:
4848:306886
4845:
4806:
4796:
4782:: 99.
4755:
4747:
4682:
4676:226894
4674:
4648:Nature
4622:
4612:
4570:
4560:
4511:
4424:
4416:
4406:
4353:
4326:
4301:
4291:
4252:
4209:
4199:
4127:
4117:
4066:
4043:
4035:
3992:
3984:
3943:
3923:Neuron
3902:
3895:383714
3892:
3843:
3823:Neuron
3802:
3795:391067
3792:
3751:
3684:
3641:
3591:
3583:
3573:
3546:Nature
3524:16 May
3486:
3476:
3429:
3419:
3343:
3333:
3288:
3263:
3253:
3214:
3179:
3142:
3111:
3101:
3036:
3029:301388
3026:
2980:acid'.
2951:
2944:300638
2941:
2892:
2885:221128
2882:
2833:
2826:223178
2823:
2774:
2766:
2725:
2686:
2676:
2625:
2617:
2548:
2538:
2530:
2491:
2483:
2475:
2436:
2426:
2396:
2386:
2378:
2339:
2331:
2323:
2297:Nature
2276:
2266:
2082:Origin
2046:as in
2040:magnum
1156:32,109
1094:40,285
782:serine
653:indels
548:valine
542:or as
285:lysine
273:lysine
250:uracil
228:codons
205:Codons
184:4 = 64
86:codons
50:uracil
9147:LASiS
8800:Cells
8465:types
8331:Polar
7989:Types
7886:Types
7736:Lists
7616:Italy
7455:Index
7130:S2CID
7096:arXiv
7069:S2CID
7018:S2CID
6984:arXiv
6942:S2CID
6908:arXiv
6822:S2CID
6654:S2CID
6612:arXiv
6537:17637
6483:S2CID
6391:S2CID
6260:S2CID
6098:S2CID
5888:S2CID
5794:S2CID
5677:21636
5603:(4).
5546:eLife
5453:S2CID
5410:S2CID
5367:S2CID
5033:eLife
4933:PeerJ
4753:S2CID
4680:S2CID
4422:S2CID
4202:24164
4041:S2CID
3990:S2CID
3904:88735
3589:S2CID
2772:S2CID
2623:S2CID
2489:S2CID
2481:JSTOR
2337:S2CID
917:Freq
908:Codon
897:Freq
888:Codon
877:Freq
868:Codon
857:Freq
848:Codon
788:or AG
772:or CU
736:. A
575:ochre
571:umber
563:amber
506:exons
121:genes
70:cells
9058:Heme
7380:The
7357:PMID
7302:ISBN
7279:ISBN
7258:ISBN
7228:PMID
7179:PMID
7122:PMID
7061:PMID
7010:PMID
6934:PMID
6881:PMID
6814:PMID
6768:PMID
6750:ISSN
6711:PMID
6693:ISSN
6646:PMID
6638:ISSN
6581:PMID
6542:PMID
6475:PMID
6432:PMID
6383:PMID
6340:PMID
6299:PMID
6252:PMID
6209:PMID
6157:PMID
6121:PNAS
6090:PMID
6055:PMID
6011:PMID
5947:PMID
5880:PMID
5837:PMID
5786:PMID
5744:2017
5713:ISBN
5682:PMID
5623:PMID
5574:PMID
5523:PMID
5488:PMID
5445:PMID
5402:PMID
5359:PMID
5316:PMID
5267:PMID
5214:PMID
5161:PMID
5112:PMID
5061:PMID
5010:PMID
4961:PMID
4910:PMID
4853:PMID
4804:PMID
4745:PMID
4714:2010
4672:PMID
4620:PMID
4568:PMID
4509:PMID
4451:2022
4414:PMID
4351:ISBN
4324:ISBN
4299:PMID
4250:PMID
4207:PMID
4125:PMID
4064:ISBN
4033:PMID
3982:PMID
3941:PMID
3900:PMID
3841:PMID
3800:PMID
3749:ISBN
3713:2010
3682:PMID
3639:ISBN
3581:PMID
3526:2019
3484:PMID
3427:PMID
3372:2017
3341:PMID
3286:ISBN
3261:PMID
3212:PMID
3177:PMID
3140:ISBN
3109:PMID
3060:2010
3034:PMID
2975:2010
2949:PMID
2890:PMID
2831:PMID
2764:PMID
2746:Cell
2723:PMID
2684:OCLC
2674:ISBN
2653:2022
2615:ISSN
2575:2022
2546:PMID
2528:ISSN
2473:ISSN
2434:OCLC
2424:ISBN
2394:PMID
2376:ISSN
2329:PMID
2321:ISSN
2274:PMID
2123:tRNA
2071:Pfam
2033:tRNA
2020:and
1976:logo
1912:16.5
1909:0.25
1897:39.6
1894:0.58
1879:0.11
1867:28.1
1864:0.46
1850:16.5
1847:0.25
1835:29.0
1832:0.42
1820:15.8
1817:0.23
1802:0.12
1788:22.2
1785:0.34
1773:25.1
1770:0.54
1758:27.7
1755:0.40
1743:14.5
1740:0.24
1726:10.8
1723:0.16
1711:21.8
1708:0.46
1696:18.4
1693:0.27
1681:11.0
1678:0.18
1664:12.0
1661:0.21
1649:31.9
1646:0.57
1631:0.11
1619:22.0
1616:1.00
1602:12.2
1599:0.21
1587:24.4
1584:0.43
1572:15.1
1569:0.28
1554:0.17
1540:19.5
1537:0.24
1525:19.1
1522:0.53
1510:18.9
1507:0.36
1495:20.8
1492:0.47
1478:12.1
1475:0.15
1463:17.0
1460:0.47
1448:13.1
1445:0.25
1433:16.0
1430:0.36
1416:11.4
1413:0.20
1401:34.2
1398:0.73
1383:0.11
1371:39.6
1368:0.40
1351:0.11
1339:12.3
1336:0.27
1324:16.9
1321:0.28
1306:0.07
1292:10.4
1289:0.18
1277:15.1
1274:0.58
1262:19.8
1259:0.32
1247:19.6
1244:0.20
1227:0.08
1215:10.9
1212:0.42
1200:17.5
1197:0.29
1185:13.2
1182:0.13
1168:13.2
1165:1.00
1150:0.24
1135:0.05
1123:12.9
1120:0.13
1103:0.47
1088:0.30
1076:12.2
1073:0.15
1058:0.08
1044:12.6
1041:0.54
1029:15.3
1026:0.56
1014:17.7
1011:0.22
999:20.3
996:0.54
982:10.6
979:0.46
967:12.2
964:0.44
952:15.2
949:0.19
937:17.6
934:0.46
639:and
627:and
567:opal
550:and
534:and
442:and
327:mRNA
236:and
222:The
64:The
8882:DNA
8877:RNA
7840:RNA
7836:DNA
7486:RNA
7481:DNA
7347:PMC
7337:doi
7325:111
7218:PMC
7210:doi
7169:PMC
7161:doi
7114:doi
7053:doi
7002:doi
6980:100
6926:doi
6904:249
6871:PMC
6861:doi
6806:doi
6758:PMC
6742:doi
6701:PMC
6685:doi
6630:doi
6573:doi
6569:160
6532:PMC
6522:doi
6467:doi
6422:doi
6375:doi
6330:doi
6291:doi
6244:doi
6199:doi
6147:PMC
6137:doi
6082:doi
6045:doi
6001:PMC
5993:doi
5937:PMC
5927:doi
5872:doi
5829:doi
5778:doi
5672:PMC
5662:doi
5613:PMC
5605:doi
5564:PMC
5554:doi
5515:doi
5480:doi
5437:doi
5394:doi
5351:doi
5306:PMC
5298:doi
5257:PMC
5249:doi
5204:PMC
5196:doi
5151:doi
5139:164
5102:PMC
5092:doi
5051:PMC
5041:doi
5000:PMC
4992:doi
4951:PMC
4941:doi
4900:PMC
4892:doi
4880:459
4843:PMC
4835:doi
4794:PMC
4784:doi
4737:doi
4664:doi
4652:282
4610:PMC
4602:doi
4558:PMC
4548:doi
4536:109
4499:doi
4495:280
4404:PMC
4396:doi
4289:PMC
4281:doi
4277:172
4242:doi
4230:215
4197:PMC
4187:doi
4115:PMC
4105:doi
4093:104
4025:doi
3972:doi
3931:doi
3890:PMC
3880:doi
3831:doi
3790:PMC
3782:doi
3672:doi
3571:PMC
3563:doi
3551:569
3474:PMC
3466:doi
3417:PMC
3407:doi
3395:114
3331:PMC
3323:doi
3251:PMC
3243:doi
3204:doi
3169:doi
3132:doi
3099:PMC
3091:doi
3087:168
3024:PMC
3014:doi
2939:PMC
2929:doi
2880:PMC
2870:doi
2821:PMC
2811:doi
2754:doi
2750:128
2715:doi
2605:doi
2536:PMC
2520:doi
2516:211
2465:doi
2384:PMC
2368:doi
2313:doi
2301:171
2264:PMC
2256:doi
2252:323
2000:in
1903:GGG
1888:GAG
1882:7.4
1873:GCG
1858:GUG
1841:GGA
1826:GAA
1811:GCA
1805:7.1
1796:GUA
1779:GGC
1764:GAC
1749:GCC
1734:GUC
1717:GGU
1702:GAU
1687:GCU
1672:GUU
1655:AGG
1640:AAG
1634:6.1
1625:ACG
1610:AUG
1593:AGA
1578:AAA
1563:ACA
1557:7.5
1548:AUA
1531:AGC
1516:AAC
1501:ACC
1486:AUC
1469:AGU
1454:AAU
1439:ACU
1424:AUU
1407:CGG
1392:CAG
1386:6.9
1377:CCG
1362:CUG
1354:6.2
1345:CGA
1330:CAA
1315:CCA
1309:7.2
1300:CUA
1283:CGC
1268:CAC
1253:CCC
1238:CUC
1230:4.5
1221:CGU
1206:CAU
1191:CCU
1176:CUU
1159:UGG
1153:0.8
1144:UAG
1138:4.4
1129:UCG
1114:UUG
1106:1.6
1097:UGA
1091:1.0
1082:UAA
1067:UCA
1061:7.7
1052:UUA
1035:UGC
1020:UAC
1005:UCC
990:UUC
973:UGU
958:UAU
943:UCU
928:UUU
796:or
700:RNA
659:or
500:In
244:to
129:the
82:RNA
80:or
78:DNA
72:to
9480::
8467::
7589:of
7355:.
7345:.
7335:.
7323:.
7319:.
7226:.
7216:.
7206:17
7204:.
7200:.
7177:.
7167:.
7157:10
7155:.
7151:.
7128:.
7120:.
7112:.
7104:.
7090:.
7067:.
7059:.
7051:.
7041:63
7039:.
7016:.
7008:.
7000:.
6992:.
6978:.
6940:.
6932:.
6924:.
6916:.
6902:.
6879:.
6869:.
6859:.
6847:.
6843:.
6820:.
6812:.
6804:.
6794:33
6792:.
6780:^
6766:.
6756:.
6748:.
6738:47
6736:.
6732:.
6709:.
6699:.
6691:.
6681:45
6679:.
6675:.
6652:.
6644:.
6636:.
6628:.
6620:.
6606:.
6602:.
6579:.
6567:.
6563:.
6540:.
6530:.
6520:.
6510:97
6508:.
6504:.
6481:.
6473:.
6465:.
6455:44
6453:.
6430:.
6418:19
6416:.
6412:.
6389:.
6381:.
6373:.
6363:80
6361:.
6338:.
6324:.
6320:.
6297:.
6287:24
6285:.
6281:.
6258:.
6250:.
6242:.
6232:23
6230:.
6207:.
6197:.
6187:69
6185:.
6181:.
6169:^
6155:.
6145:.
6135:.
6125:75
6123:.
6119:.
6096:.
6088:.
6078:38
6076:.
6053:.
6041:17
6039:.
6035:.
6023:^
6009:.
5999:.
5991:.
5981:73
5979:.
5975:.
5959:^
5945:.
5935:.
5925:.
5915:77
5913:.
5909:.
5886:.
5878:.
5870:.
5860:29
5858:.
5835:.
5827:.
5817:22
5815:.
5792:.
5784:.
5776:.
5766:47
5764:.
5752:^
5735:.
5694:^
5680:.
5670:.
5660:.
5650:95
5648:.
5644:.
5621:.
5611:.
5601:40
5599:.
5595:.
5572:.
5562:.
5552:.
5550:10
5548:.
5544:.
5521:.
5511:83
5509:.
5486:.
5476:79
5474:.
5451:.
5443:.
5433:18
5431:.
5408:.
5400:.
5388:.
5365:.
5357:.
5349:.
5339:80
5337:.
5314:.
5304:.
5294:61
5292:.
5288:.
5265:.
5255:.
5245:24
5243:.
5239:.
5212:.
5202:.
5192:33
5190:.
5186:.
5159:.
5149:.
5137:.
5133:.
5110:.
5100:.
5088:12
5086:.
5082:.
5059:.
5049:.
5035:.
5031:.
5008:.
4998:.
4986:.
4982:.
4959:.
4949:.
4935:.
4931:.
4908:.
4898:.
4890:.
4878:.
4874:.
4851:.
4841:.
4831:23
4829:.
4825:.
4802:.
4792:.
4778:.
4774:.
4751:.
4743:.
4733:46
4731:.
4696:^
4678:.
4670:.
4662:.
4650:.
4618:.
4608:.
4598:27
4596:.
4592:.
4580:^
4566:.
4556:.
4546:.
4534:.
4530:.
4507:.
4493:.
4489:.
4477:^
4467:.
4442:.
4420:.
4412:.
4402:.
4392:35
4390:.
4386:.
4374:^
4297:.
4287:.
4275:.
4271:.
4248:.
4240:.
4228:.
4205:.
4195:.
4185:.
4175:96
4173:.
4169:.
4146:.
4123:.
4113:.
4103:.
4091:.
4087:.
4039:.
4031:.
4023:.
4013:21
4011:.
3988:.
3980:.
3966:.
3962:.
3939:.
3927:52
3925:.
3921:.
3898:.
3888:.
3878:.
3868:76
3866:.
3862:.
3839:.
3827:52
3825:.
3821:.
3798:.
3788:.
3778:23
3776:.
3772:.
3743:.
3703:.
3680:.
3668:95
3666:.
3662:.
3623:^
3612:,
3587:.
3579:.
3569:.
3561:.
3549:.
3543:.
3516:.
3510:.
3482:.
3472:.
3464:.
3452:.
3448:.
3425:.
3415:.
3405:.
3393:.
3389:.
3362:.
3339:.
3329:.
3319:54
3317:.
3313:.
3259:.
3249:.
3239:16
3237:.
3233:.
3210:.
3198:.
3175:.
3165:12
3163:.
3138:.
3130:.
3107:.
3097:.
3085:.
3081:.
3062:.
3032:.
3022:.
3012:.
3002:53
3000:.
2996:.
2977:.
2947:.
2937:.
2927:.
2917:49
2915:.
2911:.
2888:.
2878:.
2868:.
2858:48
2856:.
2852:.
2829:.
2819:.
2809:.
2799:47
2797:.
2793:.
2770:.
2762:.
2748:.
2744:.
2721:.
2711:12
2709:.
2705:.
2682:.
2644:.
2621:.
2613:.
2603:.
2601:14
2599:.
2595:.
2583:^
2566:.
2544:.
2534:.
2526:.
2514:.
2510:.
2487:.
2479:.
2471:.
2461:86
2459:.
2455:.
2432:.
2418:.
2392:.
2382:.
2374:.
2364:67
2362:.
2358:.
2335:.
2327:.
2319:.
2311:.
2299:.
2295:.
2272:.
2262:.
2250:.
2246:.
2078:.
2052:.
2028:.
911:AA
891:AA
871:AA
851:AA
694:.
682:.
648:.
621:.
512:.
401:.
389:,
9028::
8996::
8944::
8919::
8867::
8544:e
8537:t
8530:v
8412:)
8410:a
8407:K
8371:)
8369:a
8366:K
8269:)
8255:(
8187:e
8180:t
8173:v
7800:e
7793:t
7786:v
7416:e
7409:t
7402:v
7363:.
7339::
7331::
7310:.
7287:.
7266:.
7234:.
7212::
7185:.
7163::
7136:.
7116::
7108::
7098::
7092:7
7075:.
7055::
7047::
7024:.
7004::
6996::
6986::
6948:.
6928::
6920::
6910::
6887:.
6863::
6855::
6849:4
6828:.
6808::
6800::
6774:.
6744::
6717:.
6687::
6660:.
6632::
6624::
6614::
6608:9
6587:.
6575::
6548:.
6524::
6516::
6489:.
6469::
6461::
6438:.
6424::
6397:.
6377::
6369::
6346:.
6332::
6326:5
6305:.
6293::
6266:.
6246::
6238::
6215:.
6201::
6193::
6163:.
6139::
6131::
6104:.
6084::
6061:.
6047::
6017:.
5995::
5987::
5953:.
5929::
5921::
5894:.
5874::
5866::
5843:.
5831::
5823::
5800:.
5780::
5772::
5746:.
5721:.
5688:.
5664::
5656::
5629:.
5607::
5580:.
5556::
5529:.
5517::
5494:.
5482::
5459:.
5439::
5416:.
5396::
5390:7
5373:.
5353::
5345::
5322:.
5300::
5273:.
5251::
5220:.
5198::
5184:"
5167:.
5153::
5145::
5118:.
5094::
5067:.
5043::
5037:3
5016:.
4994::
4988:6
4967:.
4943::
4937:1
4916:.
4894::
4886::
4859:.
4837::
4810:.
4786::
4780:6
4759:.
4739::
4716:.
4691:)
4688:(
4686:.
4666::
4658::
4626:.
4604::
4574:.
4550::
4542::
4515:.
4501::
4471:.
4453:.
4428:.
4398::
4359:.
4332:.
4305:.
4283::
4256:.
4244::
4236::
4213:.
4189::
4181::
4131:.
4107::
4099::
4072:.
4047:.
4027::
4019::
3996:.
3974::
3968:7
3947:.
3933::
3906:.
3882::
3874::
3847:.
3833::
3806:.
3784::
3757:.
3729:.
3715:.
3688:.
3674::
3647:.
3595:.
3565::
3557::
3528:.
3490:.
3468::
3460::
3454:8
3433:.
3409::
3401::
3374:.
3347:.
3325::
3294:.
3267:.
3245::
3218:.
3206::
3200:9
3183:.
3171::
3148:.
3134::
3115:.
3093::
3040:.
3016::
3008::
2955:.
2931::
2923::
2896:.
2872::
2864::
2837:.
2813::
2805::
2778:.
2756::
2729:.
2717::
2690:.
2655:.
2629:.
2607::
2577:.
2552:.
2522::
2495:.
2467::
2440:.
2400:.
2370::
2343:.
2315::
2307::
2280:.
2258::
2191:.
1906:G
1891:E
1876:A
1861:V
1844:G
1829:E
1814:A
1799:V
1782:G
1767:D
1752:A
1737:V
1720:G
1705:D
1690:A
1675:V
1658:R
1643:K
1628:T
1613:M
1596:R
1581:K
1566:T
1551:I
1534:S
1519:N
1504:T
1489:I
1472:S
1457:N
1442:T
1427:I
1410:R
1395:Q
1380:P
1365:L
1348:R
1333:Q
1318:P
1303:L
1286:R
1271:H
1256:P
1241:L
1224:R
1209:H
1194:P
1179:L
1162:W
1147:*
1132:S
1117:L
1100:*
1085:*
1070:S
1055:L
1038:C
1023:Y
1008:S
993:F
976:C
961:Y
946:S
931:F
919:‰
899:‰
879:‰
859:‰
790:Y
786:N
774:N
770:R
768:U
766:Y
752:.
655:(
213:.
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.