383:, which adds methyl residues to the same glutamate residues. If the level of an attractant remains high, the level of phosphorylation of CheA (and, therefore, CheY and CheB) will remain low, the cell will swim smoothly, and the level of methylation of the MCPs will increase (because CheB-P is not present to demethylate). The MCPs no longer respond to the attractant when they are fully methylated; therefore, even though the level of attractant might remain high, the level of CheA-P (and CheB-P) increases and the cell begins to tumble. The MCPs can be demethylated by CheB-P, and, when this happens, the receptors can once again respond to attractants. The situation is the opposite with regard to repellents: fully methylated MCPs respond best to repellents, while least-methylated MCPs respond worst to repellents. This regulation allows the bacterium to 'remember' chemical concentrations from the recent past, a few seconds, and compare them to those it is currently experiencing, thus 'know' whether it is traveling up or down a gradient. that bacteria have to chemical gradients, other mechanisms are involved in increasing the absolute value of the sensitivity on a given background. Well-established examples are the ultra-sensitive response of the motor to the CheY-P signal, and the clustering of chemoreceptors.
191:
879:
889:
584:. Eukaryotic chemotaxis involves detecting a concentration gradient spatially by comparing the asymmetric activation of these receptors at the different ends of the cell. Activation of these receptors results in migration towards chemoattractants, or away from chemorepellants. In mating yeast, which are non-motile, patches of polarity proteins on the cell cortex can relocate in a chemotactic fashion up pheromone gradients.
710:
294:
407:
5873:
358:
1148:
950:). Investigations of the three-dimensional structures of chemokines provided evidence that a characteristic composition of beta-sheets and an alpha helix provides expression of sequences required for interaction with the chemokine receptors. Formation of dimers and their increased biological activity was demonstrated by crystallography of several chemokines, e.g. IL-8.
1197:) pathogens itself represents a significant clinical target. Modification of endogenous chemotactic ability of these microorganisms by pharmaceutical agents can decrease or inhibit the ratio of infections or spreading of infectious diseases. Apart from infections, there are some other diseases wherein impaired chemotaxis is the primary etiological factor, as in
849:
33:
595:(MCPs). This results in their desensitization and allows prokaryotes to "remember" and adapt to a chemical gradient. In contrast, chemotactic memory in eukaryotes can be explained by the Local Excitation Global Inhibition (LEGI) model. LEGI involves the balance between a fast excitation and delayed inhibition which controls downstream signaling such as
1970:
349:, and it is a common form of signal transduction in bacteria. CheY induces tumbling by interacting with the flagellar switch protein FliM, inducing a change from counter-clockwise to clockwise rotation of the flagellum. Change in the rotation state of a single flagellum can disrupt the entire flagella bundle and cause a tumble.
668:
chemoattractant. However, these molecules apparently are activated independently of the motility of the cell. That is, even an immnobilized cell is still able to detect the direction of a chemoattractant. There appear to be mechanisms by which an external chemotactic gradient is sensed and turned into an intracellular Ras and
555:
177:
in the 1950s. In the 1960s and 1970s, the revolution of modern cell biology and biochemistry provided a series of novel techniques that became available to investigate the migratory responder cells and subcellular fractions responsible for chemotactic activity. The availability of this technology led
906:
are di-, tri-, tetrapeptides of bacterial origin, formylated on the N-terminus of the peptide. They are released from bacteria in vivo or after decomposition of the cell, a typical member of this group is the N-formylmethionyl-leucyl-phenylalanine (abbreviated fMLF or fMLP). Bacterial fMLF is a key
258:
bacteria will chemotax, or direct their overall motion based on the gradient. If the bacterium senses that it is moving in the correct direction (toward attractant/away from repellent), it will keep swimming in a straight line for a longer time before tumbling; however, if it is moving in the wrong
2099:
of soil microorganisms is a function of their chemotactic sensitivities towards substrate and fellow organisms. The chemotactic behavior of the bacteria was proven to lead to non-trivial population patterns even in the absence of environmental heterogeneities. The presence of structural pore scale
667:
The specific molecule/s that allow a eukaryotic cells detect a gradient of chemoattractant ligands (that is, a sort of the molecular compass that detects the direction of a chemoattractant) seems to change depending on the cell and chemoattractant receptor involved or even the concentration of the
441:
Chemoattractants or chemorepellents bind MCPs at its extracellular domain; an intracellular signaling domain relays the changes in concentration of these chemotactic ligands to downstream proteins like that of CheA which then relays this signal to flagellar motors via phosphorylated CheY (CheY-P).
925:
are intermediate products of the complement cascade. Their synthesis is joined to the three alternative pathways (classical, lectin-dependent, and alternative) of complement activation by a convertase enzyme. The main target cells of these derivatives are neutrophil granulocytes and monocytes as
36:
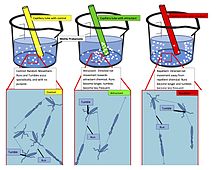
Capillary tube assay for chemotaxis. Motile prokaryotes sense chemicals in their environment and change their motility accordingly. Absent chemicals, movement is completely random. When an attractant or repellent is present, runs become longer and tumbles become less frequent. The result is net
2210:
that use artificial chemotaxis to navigate autonomously have been designed. Applications include targeted delivery of drugs in the body. More recently, enzyme molecules have also shown positive chemotactic behavior in the gradient of their substrates. The thermodynamically favorable binding of
307:, called methyl-accepting chemotaxis proteins (MCPs), which vary in the molecules that they detect. Thousands of MCP receptors are known to be encoded across the bacterial kingdom. These receptors may bind attractants or repellents directly or indirectly through interaction with proteins of
269:
This biased random walk is a result of simply choosing between two methods of random movement; namely tumbling and straight swimming. The helical nature of the individual flagellar filament is critical for this movement to occur. The protein structure that makes up the flagellar filament,
341:, CheA, at a single highly conserved histidine residue. CheA, in turn, transfers phosphoryl groups to conserved aspartate residues in the response regulators CheB and CheY; CheA is a histidine kinase and it does not actively transfer the phosphoryl group, rather, the response regulator
1772:
1371:
Although interactions of the factors listed above make the behavior of the solutions of mathematical models of chemotaxis rather complex, it is possible to describe the basic phenomenon of chemotaxis-driven motion in a straightforward way. Indeed, let us denote with
285:, are monoflagellated and have a single flagellum at one pole of the cell. Their method of chemotaxis is different. Others possess a single flagellum that is kept inside the cell wall. These bacteria move by spinning the whole cell, which is shaped like a corkscrew.
472:
cause MCPs to have lower affinity to chemoattractants which causes increased activity of CheA and CheY-P resulting in tumbles. In this way cells are able to adapt to the immediate chemoattractant concentration and detect further changes to modulate cell motility.
115:. The aberrant chemotaxis of leukocytes and lymphocytes also contribute to inflammatory diseases such as atherosclerosis, asthma, and arthritis. Sub-cellular components, such as the polarity patch generated by mating yeast, may also display chemotactic behavior.
2546:
Planagumà A, Domènech T, Pont M, Calama E, García-González V, López R, et al. (October 2015). "Combined anti CXC receptors 1 and 2 therapy is a promising anti-inflammatory treatment for respiratory diseases by reducing neutrophil migration and activation".
606:
Levels of receptors, intracellular signalling pathways and the effector mechanisms all represent diverse, eukaryotic-type components. In eukaryotic unicellular cells, amoeboid movement and cilium or the eukaryotic flagellum are the main effectors (e.g.,
700:
and the beat of the 9 + 2 microtubules within cilia. The orchestrated beating of hundreds of cilia is synchronized by a submembranous system built between basal bodies. The details of the signaling pathways are still not totally clear.
251:; in other words, bacteria "forget" the direction in which they are going. By repeatedly evaluating their course, and adjusting if they are moving in the wrong direction, bacteria can direct their random walk motion toward favorable locations.
169:. The significance of chemotaxis in biology and clinical pathology was widely accepted in the 1930s, and the most fundamental definitions underlying the phenomenon were drafted by this time. The most important aspects in quality control of
1070:-eicosatrirenoid acid); they stimulate leukocyte chemotaxis through the oxoeicosanoid receptor 1 with 5-oxoeicosatrienoic acid being as potent as its arachidonic acid-derived analog, 5-oxo-eicosatetraenoic acid, in stimulating human blood
37:
movement towards or away from the chemical (i.e., up or down the chemical gradient). The net movement can be seen in the beaker, where the bacteria accumulate around the origin of the attractant, and away from the origin of the repellent.
576:
cannot directly detect a concentration gradient. Instead, they employ temporal gradient sensing, where they move over larger distances several times their own width and measure the rate at which perceived chemical concentration changes.
4648:
Rodríguez-Fernández JL, Criado-García O (October 2022). "A meta-analysis indicates that the regulation of cell motility is a non-intrinsic function of chemoattractant receptors that is governed independently of directional sensing".
2214:
Apart from active enzymes, non-reacting molecules also show chemotactic behavior. This has been demonstrated by using dye molecules that move directionally in gradients of polymer solution through favorable hydrophobic interactions.
728:
refers to an increase in cellular motility in response to chemicals in the surrounding environment. Unlike chemotaxis, the migration stimulated by chemokinesis lacks directionality, and instead increases environmental scanning
1719:. This general equation applies to both the cell density and the chemo-attractant. Therefore, incorporating a diffusion flux into the total flux term, the interactions between these quantities are governed by a set of coupled
742:
of the chemoattractant is expressed or bound on a surface, in contrast to the classical model of chemotaxis, in which the gradient develops in a soluble fluid. The most common biologically active haptotactic surface is the
907:
component of inflammation has characteristic chemoattractant effects in neutrophil granulocytes and monocytes. The chemotactic factor ligands and receptors related to formyl peptides are summarized in the related article,
1681:
898:
The number of molecules capable of eliciting chemotactic responses is relatively high, and we can distinguish primary and secondary chemotactic molecules. The main groups of the primary ligands are as follows:
1965:{\displaystyle {\begin{aligned}{\partial C \over {\partial t}}&=f(C)+\nabla \cdot \left\\{\partial \varphi \over {\partial t}}&=g(\varphi ,C)+\nabla \cdot (D_{\varphi }\nabla \varphi )\end{aligned}}}
265:
use temporal sensing to decide whether their situation is improving or not, and in this way, find the location with the highest concentration of attractant, detecting even small differences in concentration.
2114:
A wide range of techniques is available to evaluate chemotactic activity of cells or the chemoattractant and chemorepellent character of ligands. The basic requirements of the measurement are as follows:
2582:
Rana AK, Li Y, Dang Q, Yang F (December 2018). "Monocytes in rheumatoid arthritis: Circulating precursors of macrophages and osteoclasts and, their heterogeneity and plasticity role in RA pathogenesis".
2136:
is still not available, there are several protocols and pieces of equipment that offer good correspondence with the conditions described above. The most commonly used are summarised in the table below:
1003:
eicosanoids are arachidonic acid metabolites also formed by ALOX5. Three members of the family form naturally and have prominent chemotactic activity. These, listed in order of decreasing potency, are:
1488:
2993:
680:
filaments. The growing distal end of actin filaments develops connections with the internal surface of the plasma membrane via different sets of peptides and results in the formation of anterior
1777:
862:
in the presence of the ligand. The diverse features of the chemotaxis receptors and ligands allows for the possibility of selecting chemotactic responder cells with a simple chemotaxis assay By
858:
While some chemotaxis receptors are expressed in the surface membrane with long-term characteristics, as they are determined genetically, others have short-term dynamics, as they are assembled
942:
C chemokines) represent structurally related molecules with a special arrangement of disulfide bridges but also their target cell specificity is diverse. CC chemokines act on monocytes (e.g.,
499:, causing movement toward infection sites. Non-acylated methioninyl peptides do not act as chemoattractants to neutrophils and macrophages. Leukocytes also move toward chemoattractants C5a, a
5452:
Kohidai L, Lang O and Csaba G (2003). "Chemotactic-range-fitting of amino acids and its correlations to physicochemical parameters in
Tetrahymena pyriformis - Evolutionary consequences".
1189:
A changed migratory potential of cells has relatively high importance in the development of several clinical symptoms and syndromes. Altered chemotactic activity of extracellular (e.g.,
2744:
2211:
enzymes to their specific substrates is recognized as the origin of enzymatic chemotaxis. Additionally, enzymes in cascades have also shown substrate-driven chemotactic aggregation.
771:
cells. Depending on the chemical character of released substances, necrotaxis can accumulate or repel cells, which underlines the pathophysiological significance of this phenomenon.
4725:
Lin Y, Pal DS, Banerjee P, Banerjee T, Qin G, Deng Y, et al. (July 2024). "Ras suppression potentiates rear actomyosin contractility-driven cell polarization and migration".
3701:"A multi-scale model of Escherichia coli chemotaxis from intracellular signaling pathway to motility and nutrient uptake in nutrient gradient and isotropic fluid environments"
1713:
1413:
1122:
is a non-eicosanoid metabolite of arachidonic acid made by cyclooxygenase 1 or cyclooxygenase 2 that stimulates leukocyte chemotaxis though the leukotriene B4 receptor, BLT2.
2037:
2091:
1595:
1437:
1764:
1390:
5120:
Köhidai L, Csaba G (July 1998). "Chemotaxis and chemotactic selection induced with cytokines (IL-8, RANTES and TNF-alpha) in the unicellular
Tetrahymena pyriformis".
182:
modernized
Pfeffer's capillary assay and represented a significant turning point in understanding the whole process of intracellular signal transduction of bacteria.
2064:
1571:
1551:
1531:
719:
Chemotaxis refers to the directional migration of cells in response to chemical gradients; several variations of chemical-induced migration exist as listed below.
2002:
5221:"Biosynthesis, biological effects, and receptors of hydroxyeicosatetraenoic acids (HETEs) and oxoeicosatetraenoic acids (oxo-ETEs) derived from arachidonic acid"
1744:
1615:
1511:
2689:
2286:
2986:
600:
1044:
infections. After leaving nearby blood vessels, these cells recognize chemicals produced by bacteria in a cut or scratch and migrate "toward the smell".
870:
is also used to designate a technique that separates eukaryotic or prokaryotic cells according to their chemotactic responsiveness to selector ligands.
510:
Mechanisms concerning chemorepellents are less known than chemoattractants. Although chemorepellents work to confer an avoidance response in organisms,
5790:
Zhao X, Palacci H, Yadav V, Spiering MM, Gilson MK, Butler PJ, et al. (March 2018). "Substrate-driven chemotactic assembly in an enzyme cascade".
4527:
Kedrin D, van
Rheenen J, Hernandez L, Condeelis J, Segall JE (September 2007). "Cell motility and cytoskeletal regulation in invasion and metastasis".
221:
Clockwise rotation breaks the flagella bundle apart such that each flagellum points in a different direction, causing the bacterium to tumble in place.
6346:
3504:"The two-component signaling pathway of bacterial chemotaxis: a molecular view of signal transduction by receptors, kinases, and adaptation enzymes"
4323:
Kutscher B, Devreotes P, Iglesias PA (February 2004). "Local excitation, global inhibition mechanism for gradient sensing: an interactive applet".
190:
1627:
484:
587:
It has also been shown that both prokaryotic and eukaryotic cells are capable of chemotactic memory. In prokaryotes, this mechanism involves the
2039:
is the kinetics/source term for the chemo-attractant, and the diffusion coefficients for cell density and the chemo-attractant are respectively
554:
5527:
Gharasoo M, Centler F, Fetzer I, Thullner M (2014). "How the chemotactic characteristics of bacteria can determine their population patterns".
247:
are unable to choose the direction in which they swim, and are unable to swim in a straight line for more than a few seconds due to rotational
2782:"Interactions of the complement system with endotoxic lipopolysaccharide. Generation of a factor chemotactic for polymorphonuclear leukocytes"
6159:
6070:
6051:
6032:
5993:
5914:
5833:
Guha R, Mohajerani F, Collins M, Ghosh S, Sen A, Velegol D (November 2017). "Chemotaxis of
Molecular Dyes in Polymer Gradients in Solution".
5504:
5104:
5067:
4025:
3483:
3355:
3119:
3603:
Cluzel P, Surette M, Leibler S (March 2000). "An ultrasensitive bacterial motor revealed by monitoring signaling proteins in single cells".
2755:
572:; however, sensing of chemical gradients is still a crucial step in the process. Due to their small size and other biophysical constraints,
104:
during injury or infection). In addition, it has been recognized that mechanisms that allow chemotaxis in animals can be subverted during
5510:
780:
In general, eukaryotic cells sense the presence of chemotactic stimuli through the use of 7-transmembrane (or serpentine) heterotrimeric
72:
organisms direct their movements according to certain chemicals in their environment. This is important for bacteria to find food (e.g.,
592:
415:
274:, is conserved among all flagellated bacteria. Vertebrates seem to have taken advantage of this fact by possessing an immune receptor (
6097:
980:, which elicits adhesion, chemotaxis, and aggregation of leukocytes. The chemoattractant action of LTB4 is induced via either of two
6186:
5737:
Mohajerani F, Zhao X, Somasundar A, Velegol D, Sen A (October 2018). "A Theory of Enzyme
Chemotaxis: From Experiments to Modeling".
4935:
3587:
2903:
1367:
Other environmental effects possessing direct or indirect influence on the migration (lighting, temperature, magnetic fields, etc.)
218:
Counter-clockwise rotation aligns the flagella into a single rotating bundle, causing the bacterium to swim in a straight line; and
6549:
6314:
5702:
Sengupta S, Dey KK, Muddana HS, Tabouillot T, Ibele ME, Butler PJ, et al. (January 2013). "Enzyme molecules as nanomotors".
1118:
342:
3425:"A census of membrane-bound and intracellular signal transduction proteins in bacteria: bacterial IQ, extroverts and introverts"
1248:
1198:
888:
2829:
Adler J, Tso WW (June 1974). ""Decision"-making in bacteria: chemotactic response of
Escherichia coli to conflicting stimuli".
1445:
1133:
1082:
346:
293:
981:
1016:
1720:
866:, we can determine whether a still-uncharacterized molecule acts via the long- or the short-term receptor pathway. The term
403:
create chemical concentration gradients that organisms, prokaryotic and eukaryotic, move toward or away from, respectively.
6339:
1723:
1364:
Assay systems applied to evaluate chemotaxis (see incubation times, development, and stability of concentration gradients)
1000:
878:
709:
3552:
165:
also contributed to the study of the field during 1882 to 1886, with investigations of the process as an initial step of
6554:
635:) a large group of cells—considered previously to be fixed into tissues—are also motile in special physiological (e.g.,
546:. These organisms avoid these molecules by producing avoiding reactions to re-orient themselves away from the gradient.
5563:
5405:"Stimulating properties of 5-oxo-eicosanoids for human monocytes: synergism with monocyte chemotactic protein-1 and -3"
6309:
5903:"Bacterial Chemotaxis Depends on a Two-Component Signaling Pathway Activated by Histidine-Kinase-associated Receptors"
1006:
462:
the binding of chemoattractants to MCPs inhibit CheA and therefore CheY-P activity, resulting in smooth runs, but for
2700:
2255:
1107:
954:
145:, a Caltech lecture regarding chemotaxis propounds that 'erudite description of chemotaxis was only first made by
4008:
Köhidai L (2016). "Chemotaxis as an
Expression of Communication of Tetrahymena". In Witzany G, Nowacki M (eds.).
2743:
Roberts B, Chung E, Yu SH, Li SZ, et al. (Mathematical Method of
Bioengineering Group Presentation) (2012).
2401:"An excitable Ras/PI3K/ERK signaling network controls migration and oncogenic transformation in epithelial cells"
1151:
357:
225:
The directions of rotation are given for an observer outside the cell looking down the flagella toward the cell.
179:
146:
6169:
Miller LD, Russell MH, Alexandre G (2009). "Diversity in bacterial chemotactic responses and niche adaptation".
6544:
6332:
1033:
908:
174:
154:
6206:"Design and diversity in bacterial chemotaxis: a comparative study in Escherichia coli and Bacillus subtilis"
5607:
4484:
Köhidai L (1999). "Chemotaxis: the proper physiological response to evaluate phylogeny of signal molecules".
5491:. Interdisciplinary Applied Mathematics. Vol. 17 (3rd ed.). New York: Springer. pp. 395–417.
5010:"Antagonists of chemoattractants reveal separate receptors for cAMP, folic acid and pterin in Dictyostelium"
1162:
920:
801:
692:
of eukaryotic cells can also produce chemotaxis; in this case, it is mainly a Ca-dependent induction of the
457:
178:
to the discovery of C5a, a major chemotactic factor involved in acute inflammation. The pioneering works of
3010:
Berg HC, Brown DA (October 1972). "Chemotaxis in
Escherichia coli analysed by Three-dimensional Tracking".
241:
with relatively straight swims interrupted by random tumbles that reorient the bacterium. Bacteria such as
111:, and the aberrant change of the overall property of these networks, which control chemotaxis, can lead to
5944:
5888:
4438:
3663:
1194:
1136:). The receptor or other mechanism by which this metabolite stimulates chemotaxis has not been elucidated.
533:
521:
304:
281:
As in many instances in biology, there are bacteria that do not follow this rule. Many bacteria, such as
6534:
1158:
863:
234:
580:
Eukaryotic cells are much larger than prokaryotes and have receptors embedded uniformly throughout the
517:
6512:
6012:
5799:
5619:
5536:
4792:
4379:
4277:
4118:
4054:
3862:
3612:
3197:
3019:
2940:
2838:
2642:
1689:
1395:
1036:, which like the receptors for leukotriene B4, is a G protein-coupled receptor. Aside from the skin,
821:
744:
672:
gradients, which results in a gradient and the activation of a signaling pathway, culminating in the
65:
6324:
5009:
3668:
6539:
2007:
1618:
1252:
685:
308:
162:
5949:
4443:
2069:
1339:
Migration (e.g., basic differences of bacterial swimming, movement of unicellular eukaryotes with
321:
5772:
5746:
5684:
5643:
5581:
5434:
5145:
5040:
4900:
4816:
4595:
4552:
4509:
4348:
4194:
3636:
3405:
3043:
2909:
2862:
2608:
1287:
1278:
848:
811:
275:
141:
Although migration of cells was detected from the early days of the development of microscopy by
46:
2119:
Concentration gradients can develop relatively quickly and persist for a long time in the system
1165:
interactions vary with the concentration of the ligand. Investigations of ligand families (e.g.
233:
The overall movement of a bacterium is the result of alternating tumble and swim phases, called
121:
chemotaxis occurs if the movement is toward a higher concentration of the chemical in question;
2887:
1576:
1418:
333:
The proteins CheW and CheA bind to the receptor. The absence of receptor activation results in
6280:
6237:
6192:
6182:
6155:
6138:
6093:
6066:
6047:
6028:
5989:
5972:
5910:
5850:
5815:
5764:
5719:
5635:
5500:
5483:
5461:
5426:
5403:
Sozzani S, Zhou D, Locati M, Bernasconi S, Luini W, Mantovani A, et al. (November 1996).
5385:
5350:
5299:
5250:
5194:
5137:
5100:
5063:
5032:
4990:
4941:
4931:
4892:
4857:
4808:
4765:
4738:
4707:
4658:
4630:
4587:
4544:
4501:
4466:
4407:
4340:
4305:
4243:
4186:
4144:
4080:
4021:
3990:
3941:
3890:
3826:
3774:
3681:
3654:
Sourjik V (December 2004). "Receptor clustering and signal processing in E. coli chemotaxis".
3628:
3583:
3533:
3479:
3456:
3397:
3351:
3325:
3296:"Flagellin: a unique microbe-associated molecular pattern and a multi-faceted immunomodulator"
3276:
3225:
3166:
3115:
3092:
3035:
2968:
2899:
2854:
2811:
2670:
2600:
2564:
2528:
2479:
2430:
2381:
2332:
1749:
1375:
1314:
976:(also termed 5-lipoxygenase). Their most prominent member with chemotactic factor activity is
829:
644:
616:
525:
500:
464:
126:
84:). In multicellular organisms, chemotaxis is critical to early development (e.g., movement of
5631:
5084:
6432:
6270:
6262:
6227:
6217:
6174:
6128:
6120:
6085:
6020:
5962:
5954:
5842:
5807:
5756:
5711:
5674:
5627:
5571:
5544:
5492:
5416:
5377:
5340:
5330:
5289:
5281:
5240:
5232:
5184:
5176:
5129:
5092:
5024:
4980:
4972:
4923:
4884:
4847:
4800:
4730:
4697:
4689:
4678:"Actuation of single downstream nodes in growth factor network steers immune cell migration"
4622:
4579:
4536:
4493:
4456:
4448:
4397:
4387:
4332:
4295:
4285:
4233:
4225:
4178:
4134:
4126:
4070:
4062:
4013:
3980:
3972:
3931:
3921:
3880:
3870:
3816:
3808:
3764:
3756:
3712:
3673:
3620:
3523:
3515:
3446:
3436:
3389:
3315:
3307:
3266:
3256:
3215:
3205:
3156:
3148:
3082:
3074:
3027:
2958:
2948:
2891:
2846:
2801:
2793:
2660:
2650:
2592:
2556:
2518:
2510:
2469:
2461:
2420:
2412:
2371:
2363:
2322:
2314:
2225:
2133:
2109:
1553:
is often not constant, but a decreasing function of the chemo-attractant. For some quantity
1295:
1282:
1190:
1103:
1099:
1094:
969:
488:
406:
396:
338:
261:
243:
206:
170:
101:
4756:
Becker EL (October 1977). "Stimulated neutrophil locomotion: chemokinesis and chemotaxis".
2929:"Asymmetry in the clockwise and counterclockwise rotation of the bacterial flagellar motor"
2042:
1556:
1536:
1516:
763:
embodies a special type of chemotaxis when the chemoattractant molecules are released from
237:. As a result, the trajectory of a bacterium swimming in a uniform environment will form a
4836:"Social senses: G-protein-coupled receptor signaling pathways in Dictyostelium discoideum"
4676:
Pal DS, Banerjee T, Lin Y, de Trogoff F, Borleis J, Iglesias PA, et al. (July 2023).
4583:
2280:
2096:
1978:
1291:
1262:
1174:
916:
334:
312:
150:
3380:
Wadhams GH, Armitage JP (December 2004). "Making sense of it all: bacterial chemotaxis".
6124:
6089:
6016:
5878:
5803:
5623:
5540:
4796:
4702:
4677:
4383:
4281:
4122:
4058:
3866:
3616:
3201:
3023:
2944:
2842:
2725:
2646:
1361:
Biological characteristics of the ligands (attractant, neutral, and repellent molecules)
345:
takes the phosphoryl group from CheA. This mechanism of signal transduction is called a
6466:
6275:
6250:
6133:
6108:
5967:
5932:
5345:
5318:
5294:
5269:
5245:
5220:
5189:
5165:"Ability of polymorphonuclear leukocytes to orient in gradients of chemotactic factors"
5164:
4985:
4960:
4461:
4426:
4402:
4367:
4300:
4265:
4238:
4213:
4139:
4106:
4075:
4042:
3985:
3960:
3936:
3909:
3528:
3503:
3451:
3424:
3320:
3295:
3271:
3244:
3161:
3136:
3087:
3062:
2963:
2928:
2806:
2781:
2665:
2630:
2523:
2498:
2474:
2449:
2425:
2400:
2376:
2352:"Cell motility in cancer invasion and metastasis: insights from simple model organisms"
2351:
2327:
2302:
1729:
1600:
1496:
1178:
977:
673:
608:
442:
CheY-P can then control flagellar rotation influencing the direction of cell motility.
325:
are activated. The Che proteins alter the tumbling frequency, and alter the receptors.
112:
6232:
6205:
6178:
4852:
4835:
4264:
Skoge M, Yue H, Erickstad M, Bae A, Levine H, Groisman A, et al. (October 2014).
4066:
3885:
3850:
3821:
3796:
3769:
3744:
3220:
3185:
2881:
418:(MCP). MCPs in E.coli include Tar, Tsr, Trg and Tap. Chemoattracttants to Trg include
6528:
5647:
5585:
5028:
4888:
4513:
4198:
3519:
3409:
2399:
Zhan H, Bhattacharya S, Cai H, Iglesias PA, Huang CH, Devreotes PN (September 2020).
2275:
1270:
1111:
947:
652:
620:
596:
581:
89:
69:
5902:
5776:
5688:
5438:
5368:
Yokomizo T (February 2015). "Two distinct leukotriene B4 receptors, BLT1 and BLT2".
5149:
5044:
4904:
4599:
4570:
Solnica-Krezel L, Sepich DS (2012). "Gastrulation: making and shaping germ layers".
4556:
4368:"Cells navigate with a local-excitation, global-inhibition-biased excitable network"
3797:"Repellent response functions of the Trg and Tap chemoreceptors of Escherichia coli"
2913:
2612:
2125:
Migration of cells is free toward and away on the axis of the concentration gradient
6461:
4820:
4352:
3908:
Kuruvilla H, Schmidt B, Song S, Bhajjan M, Merical M, Alley C, et al. (2016).
3760:
3640:
3047:
2866:
2245:
1226:
1170:
989:
960:
817:
752:
724:
399:
substances possessing chemotaxis-inducer effect in motile cells. These chemotactic
166:
130:
57:
5548:
5285:
3812:
3624:
2850:
1335:
Several mathematical models of chemotaxis were developed depending on the type of
76:) by swimming toward the highest concentration of food molecules, or to flee from
6222:
5421:
5404:
5236:
4693:
4626:
3580:
An Introductory Text to Bioengineering (Advanced Series in Biomechanics - Vol. 4)
3114:(Expanded, rev. ed.). Princeton, NJ: Princeton Univ. Press. pp. 83–94.
2631:"Chemotactic movement of a polarity site enables yeast cells to find their mates"
2596:
2416:
6502:
6497:
5760:
5096:
4169:
Vladimirov N, Sourjik V (November 2009). "Chemotaxis: how bacteria use memory".
4017:
3717:
3700:
2250:
1235:
1166:
1090:
which stimulates leukocyte chemotaxis through the leukotriene B4 receptor, BLT2.
1037:
1025:
1021:
833:
693:
656:
624:
612:
588:
511:
451:
380:
368:
238:
142:
4927:
4734:
4372:
Proceedings of the National Academy of Sciences of the United States of America
4270:
Proceedings of the National Academy of Sciences of the United States of America
3855:
Proceedings of the National Academy of Sciences of the United States of America
3190:
Proceedings of the National Academy of Sciences of the United States of America
2933:
Proceedings of the National Academy of Sciences of the United States of America
2635:
Proceedings of the National Academy of Sciences of the United States of America
2560:
2514:
2465:
1439:
that is generated by the chemotaxis is linked to the above gradient by the law:
56:) is the movement of an organism or entity in response to a chemical stimulus.
6490:
6473:
6439:
5679:
5662:
5576:
4540:
3976:
3961:"Responses of the ciliates Tetrahymena and Paramecium to external ATP and GTP"
3677:
3556:
3245:"Bacterial Flagellar Filament: A Supramolecular Multifunctional Nanostructure"
3152:
3078:
2303:"Neutrophil migration in infection and wound repair: going forward in reverse"
2240:
1716:
1309:
1075:
1071:
965:
930:
807:
789:
759:
734:
697:
681:
648:
640:
632:
565:
542:
529:
520:, within 10 minutes of exposure; however, exposure to chemorepellents such as
496:
492:
108:
97:
5639:
4613:
Shellard A, Mayor R (July 2016). "Chemotaxis during neural crest migration".
125:
chemotaxis if the movement is in the opposite direction. Chemically prompted
6485:
6418:
6413:
6369:
4976:
4392:
4336:
4290:
4229:
2953:
2886:. Biological and Medical Physics, Biomedical Engineering. Springer. p.
2655:
2235:
1676:{\displaystyle {\partial \rho \over {\partial t}}+\nabla \cdot {\bf {J}}=S}
1355:
1344:
1274:
1266:
1055:
1041:
1029:
935:
793:
781:
768:
636:
431:
423:
392:
372:
271:
248:
211:
6284:
6266:
6241:
6196:
6142:
5976:
5854:
5819:
5768:
5723:
5465:
5389:
5354:
5303:
5254:
5133:
4994:
4945:
4896:
4783:
Carter SB (January 1967). "Haptotaxis and the mechanism of cell motility".
4742:
4711:
4662:
4634:
4591:
4548:
4505:
4470:
4411:
4344:
4309:
4247:
4190:
4148:
3994:
3945:
3926:
3875:
3778:
3685:
3632:
3460:
3441:
3401:
3329:
3280:
3210:
3170:
3096:
2972:
2674:
2604:
2568:
2532:
2483:
2434:
2385:
2336:
17:
5430:
5225:
Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids
5141:
5036:
4861:
4812:
3894:
3830:
3537:
3229:
3039:
2858:
2815:
2797:
1415:
as its gradient. Then the chemotactic cellular flow (also called current)
6454:
6406:
6394:
5846:
5335:
5198:
5180:
4769:
4084:
3261:
1318:
1305:
764:
739:
628:
504:
255:
201:
61:
5381:
4214:"Mechanistic insights into actin-driven polarity site movement in yeast"
4182:
3311:
2499:"Importance of the leukotriene B4-BLT1 and LTB4-BLT2 pathways in asthma"
2367:
2318:
2100:
heterogeneities has an extra impact on the emerging bacterial patterns.
1201:, where giant intracellular vesicles inhibit normal migration of cells.
476:
Chemoattractants in eukaryotes are well characterized for immune cells.
32:
6444:
5958:
5811:
4497:
4452:
2230:
1348:
993:
837:
796:(smelling). The main classes of chemotaxis receptors are triggered by:
748:
480:
376:
316:
158:
73:
6024:
5715:
4918:
Antunes G, Simoes de Souza FM (2016). "Olfactory receptor signaling".
4130:
2290:. Vol. 6 (11th ed.). Cambridge University Press. p. 77.
784:-coupled receptors, a class representing a significant portion of the
6374:
4804:
3031:
1129:
1087:
943:
785:
689:
477:
435:
427:
419:
400:
105:
93:
81:
77:
6304:
3582:. Singapore: World Scientific Publishing Co. Pte. Ltd. p. 418.
3393:
1392:
the spatially non-uniform concentration of the chemo-attractant and
1147:
1020:. This family of agonists stimulates chemotactic responses in human
623:
also move to where they need to be. Besides immune competent cells (
414:
Effects of chemoattractants are elicited via chemoreceptors such as
27:
Movement of an organism or entity in response to a chemical stimulus
6319:
5751:
5496:
4920:
G Protein-Coupled Receptors - Signaling, Trafficking and Regulation
3745:"Diversity in chemotaxis mechanisms among the bacteria and archaea"
2895:
379:
side of the receptor; it works antagonistically with CheR, a methyl
6478:
6356:
6299:
6044:
Living at micro scale : the unexpected physics of being small
3476:
Cognitive Biology: Dealing with Information from Bacteria to Minds
1340:
1322:
1146:
1110:. It elicits chemotactic responses in eosinophils, basophils, and
973:
677:
85:
52:
31:
5988:(Expanded, rev. ed.). Princeton, NJ: Princeton Univ. Press.
3910:"Netrin-1 Peptide Is a Chemorepellent in Tetrahymena thermophila"
2279:
946:), and CXC chemokines are neutrophil granulocyte-specific (e.g.,
6384:
4875:
Montell C (November 1999). "Visual transduction in Drosophila".
3502:
Falke JJ, Bass RB, Butler SL, Chervitz SA, Danielson MA (1997).
2690:"How Cells Decide Where to Go: The Case of Bacterial Chemotaxis"
1231:
985:
669:
6328:
5270:"The eosinophil chemoattractant 5-oxo-ETE and the OXE receptor"
3851:"N-formylmethionyl peptides as chemoattractants for leucocytes"
2128:
Detected responses are the results of active migration of cells
751:
is responsible for induction of transendothelial migration and
100:) as well as in normal function and health (e.g., migration of
828:
However, induction of a wide set of membrane receptors (e.g.,
788:. Some members of this gene superfamily are used in eyesight (
311:. The signals from these receptors are transmitted across the
5085:"Chemotaxis as an Expression of Communication of Tetrahymena"
4366:
Xiong Y, Huang CH, Iglesias PA, Devreotes PN (October 2010).
1132:; it has weak chemotactic activity for human monocytes (sees
6109:"Bacterial chemotaxis: the early years of molecular studies"
5608:"Olfactory Sensing and Navigation in Turbulent Environments"
887:
877:
847:
708:
559:
Difference of gradient sensing in prokaryotes and eukaryotes
553:
405:
356:
292:
189:
4922:. Methods in Cell Biology. Vol. 132. pp. 127–45.
2629:
Ghose D, Jacobs K, Ramirez S, Elston T, Lew D (June 2021).
651:). Chemotaxis has high significance in the early phases of
1098:
is an eicosanoid metabolite of arachidononic acid made by
840:, vasoactive peptides) also elicit migration of the cell.
568:
cells employ is quite different from that in the bacteria
536:
are chemorepellents in micro-molar concentrations to both
5901:
Alberts B, Johnson A, Lewis J, Walter P, Raff MC (2002).
2122:
Chemotactic and chemokinetic activities are distinguished
1483:{\displaystyle {\bf {J}}=C\chi (\varphi )\nabla \varphi }
1354:
Physico-chemical characteristics of the chemicals (e.g.,
214:
per cell (4–10 typically). These can rotate in two ways:
6251:"Chemotaxis on the move - active learning teaching tool"
3186:"The gradient-sensing mechanism in bacterial chemotaxis"
3137:"Responding to Chemical Gradients: Bacterial Chemotaxis"
3063:"Responding to chemical gradients: bacterial chemotaxis"
1128:
is an eicosanoid metabolite of arachidonic acid made my
1086:
is an eicosanoid metabolite of arachidonic acid made by
195:
Correlation of swimming behaviour and flagellar rotation
3478:. United States: Oxford University Press. p. 266.
2301:
de Oliveira S, Rosowski EE, Huttenlocher A (May 2016).
2754:. University of California - San Diego. Archived from
6080:
Eisenbach M (December 2011). "Bacterial Chemotaxis".
5008:
van Haastert PJ, De Wit RJ, Konijn TM (August 1982).
2780:
Snyderman R, Gewurz H, Mergenhagen SE (August 1968).
2072:
2045:
2010:
1981:
1775:
1752:
1732:
1692:
1630:
1603:
1579:
1559:
1539:
1519:
1499:
1448:
1421:
1398:
1378:
6320:
Bacterial Chemotaxis Interactive Simulator (web-app)
129:(randomly directed or nondirectional) can be called
3243:Nedeljković M, Sastre DE, Sundberg EJ (July 2021).
2927:Yuan J, Fahrner KA, Turner L, Berg HC (July 2010).
5606:Reddy G, Murthy VN, Vergassola M (10 March 2022).
2454:Arteriosclerosis, Thrombosis, and Vascular Biology
2450:"Lymphocyte migration into atherosclerotic plaque"
2085:
2058:
2031:
1996:
1964:
1758:
1738:
1707:
1675:
1609:
1589:
1565:
1545:
1525:
1505:
1482:
1431:
1407:
1384:
988:, which are highly expressed in cells involved in
3849:Schiffmann E, Corcoran BA, Wahl SM (March 1975).
468:, CheA activity increases. Methylation events in
4959:Thomas MA, Kleist AB, Volkman BF (August 2018).
4834:Kim JY, Haastert PV, Devreotes PN (April 1996).
278:) designed to recognize this conserved protein.
259:direction, it will tumble sooner. Bacteria like
6255:Journal of Microbiology & Biology Education
5214:
5212:
5210:
5208:
4877:Annual Review of Cell and Developmental Biology
4758:Archives of Pathology & Laboratory Medicine
4572:Annual Review of Cell and Developmental Biology
3508:Annual Review of Cell and Developmental Biology
2350:Stuelten CH, Parent CA, Montell DJ (May 2018).
303:Chemical gradients are sensed through multiple
298:Domain structure of chemotaxis receptor for Asp
6046:. Cambridge, Mass.: Harvard University Press.
4529:Journal of Mammary Gland Biology and Neoplasia
3795:Yamamoto K, Macnab RM, Imae Y (January 1990).
6340:
5663:"Chemical Robotics—Chemotactic Drug Carriers"
5319:"Prostaglandin receptor signaling in disease"
4100:
4098:
4096:
4094:
3705:Computers & Mathematics with Applications
3497:
3495:
1533:is the so-called 'Chemotactic coefficient' -
1040:are the body's first line of defense against
8:
6204:Rao CV, Kirby JR, Arkin AP (February 2004).
4615:Seminars in Cell & Developmental Biology
4259:
4257:
3844:
3842:
3840:
2732:. Encyclopædia Britannica, Inc. 12 May 2024.
659:is guided by gradients of signal molecules.
186:Bacterial chemotaxis—general characteristics
4164:
4162:
4160:
4158:
3699:Xu F, Bierman R, Healy F, Nguyen H (2016).
3249:International Journal of Molecular Sciences
430:as a chemorepellent. Tap and Tsr recognize
6347:
6333:
6325:
4266:"Cellular memory in eukaryotic chemotaxis"
3749:Microbiology and Molecular Biology Reviews
1134:15-Hydroxyeicosatetraenoic acid#15-oxo-ETE
968:lipid mediators made by the metabolism of
938:; not only do their groups (C, CC, CXC, CX
663:Detection of a gradient of chemoattractant
6274:
6231:
6221:
6132:
5966:
5948:
5750:
5678:
5612:Annual Review of Condensed Matter Physics
5575:
5420:
5344:
5334:
5317:Matsuoka T, Narumiya S (September 2007).
5293:
5244:
5188:
4984:
4851:
4701:
4460:
4442:
4401:
4391:
4299:
4289:
4237:
4138:
4074:
3984:
3935:
3925:
3884:
3874:
3820:
3768:
3738:
3736:
3734:
3732:
3730:
3728:
3716:
3667:
3527:
3450:
3440:
3319:
3270:
3260:
3219:
3209:
3184:Macnab RM, Koshland DE (September 1972).
3160:
3086:
2962:
2952:
2805:
2664:
2654:
2549:Pulmonary Pharmacology & Therapeutics
2522:
2473:
2424:
2375:
2326:
2077:
2071:
2050:
2044:
2009:
1980:
1943:
1892:
1882:
1837:
1790:
1780:
1776:
1774:
1751:
1731:
1691:
1661:
1660:
1641:
1631:
1629:
1602:
1581:
1580:
1578:
1558:
1538:
1518:
1498:
1450:
1449:
1447:
1423:
1422:
1420:
1397:
1377:
1177:activity occurs over a wide range, while
893:Three dimensional structure of chemokines
391:Chemoattractants and chemorepellents are
6315:Downloadable Matlab chemotaxis simulator
5835:Journal of the American Chemical Society
5704:Journal of the American Chemical Society
5632:10.1146/annurev-conmatphys-031720-032754
2139:
1513:is the spatial density of the cells and
1203:
367:CheB, when activated by CheA, acts as a
6061:Eisenbach M (2004). Lengeler JW (ed.).
5485:Mathematical Biology I: An Introduction
3790:
3788:
2624:
2622:
2267:
1106:that stimulates chemotaxis through the
4107:"The physics of eukaryotic chemotaxis"
3382:Nature Reviews. Molecular Cell Biology
2004:describes the growth in cell density,
1012:5-oxo-15-hydroxy-eicosatetraenoic acid
714:Chemotaxis related migratory responses
705:Chemotaxis-related migratory responses
92:) and development (e.g., migration of
6003:Berg HC (2003). "E. coli in motion".
5477:
5475:
4584:10.1146/annurev-cellbio-092910-154043
4105:Levine H, Rappel WJ (February 2013).
4041:Berg HC, Purcell EM (November 1977).
3375:
3373:
3371:
3369:
3367:
3294:Zhong M, Yan H, Li Y (October 2017).
3061:Sourjik V, Wingreen NS (April 2012).
7:
5931:Bagorda A, Parent CA (August 2008).
5667:Central European Journal of Medicine
5268:Powell WS, Rokach J (October 2013).
4425:Bagorda A, Parent CA (August 2008).
3135:Sourjik V, Wingreen N (April 2012).
2786:The Journal of Experimental Medicine
2745:"Keller-Segel Models for Chemotaxis"
647:) or pathological conditions (e.g.,
593:methyl-accepting chemotaxis proteins
416:methyl-accepting chemotaxis proteins
387:Chemoattractants and chemorepellents
6125:10.1146/annurev-micro-092611-150120
6090:10.1002/9780470015902.a0001251.pub3
5933:"Eukaryotic chemotaxis at a glance"
4427:"Eukaryotic chemotaxis at a glance"
3300:Cellular & Molecular Immunology
2752:Integrated Systems Neuroengineering
438:as chemoattractants, respectively.
6359:(directional movements in biology)
6065:. London: Imperial College Press.
5219:Powell WS, Rokach J (April 2015).
5087:. In Witzany G, Nowacki M (eds.).
3743:Szurmant H, Ordal GW (June 2004).
1949:
1930:
1893:
1885:
1867:
1843:
1822:
1791:
1783:
1693:
1654:
1642:
1634:
1474:
1399:
1157:Chemotactic responses elicited by
532:show no such adaptations. GTP and
25:
6152:Chemotaxis: Methods and Protocols
6107:Hazelbauer GL (13 October 2012).
4961:"Decoding the chemotactic signal"
3914:International Journal of Peptides
1242:Chemotaxis results in the disease
564:The mechanism of chemotaxis that
6249:Williams AH (20 December 2010).
6173:. Vol. 66. pp. 53–75.
6171:Advances in Applied Microbiology
6011:(2). New York: Springer: 64–65.
5871:
5516:from the original on 6 May 2022.
4889:10.1146/annurev.cellbio.15.1.231
3578:Shu C, Chen PC, Fung YC (2008).
3520:10.1146/annurev.cellbio.13.1.457
2999:from the original on 6 May 2017.
2585:International Immunopharmacology
1662:
1617:, it is possible to formulate a
1597:and generation/destruction term
1582:
1451:
1424:
1119:12-Hydroxyheptadecatrienoic acid
5564:"How animals follow their nose"
3141:Current Opinion in Cell Biology
3067:Current Opinion in Cell Biology
2132:Despite the fact that an ideal
1708:{\displaystyle \nabla \cdot ()}
1408:{\displaystyle \nabla \varphi }
1181:activities have narrow ranges.
1083:12-Hydroxyeicosatetraenoic acid
507:-specific ligands on bacteria.
5909:. Taylor & Francis Group.
5454:Cellular and Molecular Biology
4657:(eCollection 2022): 10011086.
3761:10.1128/MMBR.68.2.301-319.2004
2880:Berg H (2004). Berg HC (ed.).
2203:Artificial chemotactic systems
2026:
2014:
1991:
1985:
1955:
1936:
1924:
1912:
1864:
1858:
1816:
1810:
1724:partial differential equations
1702:
1699:
1573:that is subject to total flux
1471:
1465:
1017:5-Hydroxyeicosatetraenoic acid
883:Structure of chemokine classes
371:, removing methyl groups from
254:In the presence of a chemical
137:History of chemotaxis research
1:
6179:10.1016/S0065-2164(08)00803-4
6113:Annual Review of Microbiology
6082:Encyclopedia of Life Sciences
5907:Molecular Biology of the Cell
5549:10.1016/j.soilbio.2013.11.019
5529:Soil Biology and Biochemistry
5286:10.1016/j.plipres.2013.09.001
5058:Witzany G, Nowacki M (2016).
4853:10.1016/s1074-5521(96)90103-9
4218:Molecular Biology of the Cell
4067:10.1016/s0006-3495(77)85544-6
3813:10.1128/jb.172.1.383-388.1990
3625:10.1126/science.287.5458.1652
2851:10.1126/science.184.4143.1292
2032:{\displaystyle g(\varphi ,C)}
1001:5-Hydroxyicosatetraenoic acid
934:belong to a special class of
747:(ECM); the presence of bound
362:Signalling pathways of E.coli
6223:10.1371/journal.pbio.0020049
5562:Mackenzie D (6 March 2023).
5422:10.4049/jimmunol.157.10.4664
5237:10.1016/j.bbalip.2014.10.008
5163:Zigmond SH (November 1977).
5091:. Springer. pp. 65–82.
5089:Biocommunication of Ciliates
5060:Biocommunication of Ciliates
5029:10.1016/0014-4827(82)90139-2
4965:Journal of Leukocyte Biology
4694:10.1016/j.devcel.2023.04.019
4627:10.1016/j.semcdb.2016.01.031
4010:Biocommunication of Ciliates
2597:10.1016/j.intimp.2018.10.016
2448:Li J, Ley K (January 2015).
2417:10.1016/j.devcel.2020.08.001
2086:{\displaystyle D_{\varphi }}
1126:15-oxo-eicosatetraenoic acid
1048:5-hydroxyeicosatrienoic acid
615:). Some eukaryotic cells of
161:'. The Nobel Prize laureate
5761:10.1021/acs.biochem.8b00801
5169:The Journal of Cell Biology
5097:10.1007/978-3-319-32211-7_5
4212:Ghose D, Lew D (May 2020).
4043:"Physics of chemoreception"
4018:10.1007/978-3-319-32211-7_5
3718:10.1016/j.camwa.2015.12.019
2497:Gelfand EW (October 2017).
1007:5-oxo-eicosatetraenoic acid
982:G protein–coupled receptors
955:polyunsaturated fatty acids
516:adapt to a chemorepellent,
6571:
5274:Progress in Lipid Research
5017:Experimental Cell Research
4928:10.1016/bs.mcb.2015.11.003
4735:10.1038/s41556-024-01453-4
3959:Hennessey TM (June 2005).
3350:. New York, NY: Springer.
2561:10.1016/j.pupt.2015.08.002
2515:10.1016/j.smim.2017.08.005
2466:10.1161/ATVBAHA.114.303227
2307:Nature Reviews. Immunology
2256:Thin layers (oceanography)
2107:
1193:) or intracellular (e.g.,
1108:Prostaglandin DP2 receptor
6364:
5680:10.2478/s11536-012-0130-9
5577:10.1146/knowable-030623-4
5323:TheScientificWorldJournal
4541:10.1007/s10911-007-9046-4
3977:10.1007/s11302-005-6213-1
3678:10.1016/j.tim.2004.10.003
3423:Galperin MY (June 2005).
3153:10.1016/j.ceb.2011.11.008
3079:10.1016/j.ceb.2011.11.008
2104:Measurement of chemotaxis
1726:describing the change in
1590:{\displaystyle {\bf {J}}}
1432:{\displaystyle {\bf {J}}}
1152:Chemotactic range fitting
1143:Chemotactic range fitting
6150:Jin T, Hereld D (2016).
4486:Acta Biologica Hungarica
1759:{\displaystyle \varphi }
1385:{\displaystyle \varphi }
1249:Chédiak–Higashi syndrome
1199:Chédiak–Higashi syndrome
1052:5-oxoeicosatrienoic acid
1034:Oxoeicosanoid receptor 1
909:Formyl peptide receptors
802:formyl peptide receptors
153:(1884) in bacteria, and
6550:Transmembrane receptors
5986:Random walks in biology
5937:Journal of Cell Science
5370:Journal of Biochemistry
4977:10.1002/JLB.1MR0218-044
4840:Chemistry & Biology
4431:Journal of Cell Science
4393:10.1073/pnas.1011271107
4337:10.1126/stke.2192004pl3
4291:10.1073/pnas.1412197111
4230:10.1091/mbc.e20-01-0040
3801:Journal of Bacteriology
3551:ToxCafe (2 June 2011).
3112:Random walks in biology
2954:10.1073/pnas.1007333107
2730:Encyclopædia Britannica
2656:10.1073/pnas.2025445118
2287:Encyclopædia Britannica
513:Tetrahymena thermophila
305:transmembrane receptors
88:towards the egg during
6310:Cell Migration Gateway
6267:10.1128/jmbe.v11i2.216
5134:10.1006/cyto.1997.0328
3876:10.1073/pnas.72.3.1059
3656:Trends in Microbiology
3442:10.1186/1471-2180-5-35
3211:10.1073/pnas.69.9.2509
2987:"Bacterial Chemotaxis"
2503:Seminars in Immunology
2356:Nature Reviews. Cancer
2087:
2060:
2033:
1998:
1966:
1760:
1740:
1709:
1677:
1611:
1591:
1567:
1547:
1527:
1507:
1484:
1433:
1409:
1386:
1259:Chemotaxis is affected
1206:Chemotaxis in diseases
1195:Listeria monocytogenes
1154:
895:
885:
855:
716:
561:
411:
364:
300:
197:
38:
6305:Neutrophil Chemotaxis
6042:Dusenbery DB (2009).
5409:Journal of Immunology
3965:Purinergic Signalling
2798:10.1084/jem.128.2.259
2191:Opalescence technique
2088:
2061:
2059:{\displaystyle D_{C}}
2034:
1999:
1967:
1761:
1741:
1710:
1678:
1612:
1592:
1568:
1566:{\displaystyle \rho }
1548:
1546:{\displaystyle \chi }
1528:
1526:{\displaystyle \chi }
1508:
1485:
1434:
1410:
1387:
1218:Chemotaxis decreased
1185:Clinical significance
1150:
919:) and complement 5a (
891:
881:
868:chemotactic selection
864:chemotactic selection
853:Chemotactic selection
851:
844:Chemotactic selection
822:leukotriene receptors
712:
557:
550:Eukaryotic chemotaxis
409:
360:
296:
235:run-and-tumble motion
193:
35:
5847:10.1021/jacs.7b08783
5336:10.1100/tsw.2007.182
5181:10.1083/jcb.75.2.606
4688:(13): 1170–1188.e7.
4171:Biological Chemistry
3927:10.1155/2016/7142868
3567:– via YouTube.
3262:10.3390/ijms22147521
2181:Capillary techniques
2070:
2043:
2008:
1997:{\displaystyle f(C)}
1979:
1773:
1750:
1730:
1690:
1628:
1601:
1577:
1557:
1537:
1517:
1497:
1446:
1419:
1396:
1376:
1358:) working as ligands
1215:Chemotaxis increased
1173:) demonstrates that
745:extracellular matrix
591:of receptors called
347:two-component system
329:Flagellum regulation
6555:Transport phenomena
6017:2005PhT....58b..64B
5841:(44): 15588–15591.
5804:2018NatCh..10..311Z
5624:2022ARCMP..13..191R
5541:2014SBiBi..69..346G
4797:1967Natur.213..256C
4727:Nature Cell Biology
4384:2010PNAS..10717079X
4282:2014PNAS..11114448S
4183:10.1515/BC.2009.130
4123:2013PhT....66b..24L
4059:1977BpJ....20..193B
4047:Biophysical Journal
3867:1975PNAS...72.1059S
3617:2000Sci...287.1652C
3312:10.1038/cmi.2017.78
3202:1972PNAS...69.2509M
3024:1972Natur.239..500B
2945:2010PNAS..10712846Y
2843:1974Sci...184.1292A
2688:Phillips R (2007).
2647:2021PNAS..11825445G
2641:(22): e2025445118.
2368:10.1038/nrc.2018.15
2319:10.1038/nri.2016.49
2178:Multi-well chambers
2149:Two-chamber assays
1619:continuity equation
1331:Mathematical models
1253:Kartagener syndrome
1208:
1114:of the Th2 subtype.
1054:are metabolites of
874:Chemotactic ligands
812:chemokine receptors
353:Receptor regulation
335:autophosphorylation
309:periplasmatic space
289:Signal transduction
5959:10.1242/jcs.018077
5812:10.1038/nchem.2905
5570:. Annual Reviews.
5482:Murray JD (2002).
5175:(2 Pt 1): 606–16.
5083:Köhidai L (2016).
4682:Developmental Cell
4498:10.1007/BF03543060
4453:10.1242/jcs.018077
4012:. pp. 65–82.
3474:Auletta G (2011).
2726:"Élie Metchnikoff"
2697:Chemotaxis Lecture
2405:Developmental Cell
2281:"Chemotaxis"
2194:Orientation assays
2146:Agar-plate assays
2083:
2056:
2029:
1994:
1962:
1960:
1756:
1736:
1721:reaction-diffusion
1705:
1673:
1607:
1587:
1563:
1543:
1523:
1503:
1480:
1429:
1405:
1382:
1288:Multiple sclerosis
1279:reperfusion injury
1204:
1155:
1032:by binding to the
896:
886:
856:
830:cyclic nucleotides
814:(CCR or CXCR), and
800:Formyl peptides -
717:
655:as development of
562:
412:
365:
301:
198:
173:were described by
39:
6522:
6521:
6161:978-1-4939-3480-5
6072:978-1-86094-413-0
6053:978-0-674-03116-6
6034:978-0-387-00888-2
6025:10.1063/1.1897527
5995:978-0-691-00064-0
5943:(Pt 16): 2621–4.
5916:978-0-8153-4069-0
5745:(43): 6256–6263.
5716:10.1021/ja3091615
5568:Knowable Magazine
5506:978-0-387-95223-9
5382:10.1093/jb/mvu078
5106:978-3-319-32211-7
5069:978-3-319-32211-7
4437:(Pt 16): 2621–4.
4224:(10): 1085–1102.
4177:(11): 1097–1104.
4131:10.1063/PT.3.1884
4027:978-3-319-32211-7
3711:(11): 2466–2478.
3485:978-0-19-960848-5
3388:(12): 1024–1037.
3357:978-0-387-00888-2
3121:978-0-691-00064-0
3018:(5374): 500–504.
2883:E. coli in Motion
2761:on 29 August 2017
2200:
2199:
1900:
1798:
1739:{\displaystyle C}
1649:
1610:{\displaystyle S}
1506:{\displaystyle C}
1328:
1327:
1283:metastatic tumors
645:endothelial cells
617:higher vertebrate
171:chemotaxis assays
16:(Redirected from
6562:
6513:physical contact
6433:electric current
6373:(stimulation by
6349:
6342:
6335:
6326:
6288:
6278:
6245:
6235:
6225:
6200:
6165:
6154:. Humana Press.
6146:
6136:
6103:
6076:
6057:
6038:
5999:
5984:Berg HC (1993).
5980:
5970:
5952:
5927:
5925:
5923:
5875:
5874:
5859:
5858:
5830:
5824:
5823:
5792:Nature Chemistry
5787:
5781:
5780:
5754:
5734:
5728:
5727:
5710:(4): 1406–1414.
5699:
5693:
5692:
5682:
5661:Lagzi I (2013).
5658:
5652:
5651:
5603:
5597:
5596:
5594:
5592:
5579:
5559:
5553:
5552:
5524:
5518:
5517:
5515:
5490:
5479:
5470:
5469:
5449:
5443:
5442:
5424:
5400:
5394:
5393:
5365:
5359:
5358:
5348:
5338:
5314:
5308:
5307:
5297:
5265:
5259:
5258:
5248:
5216:
5203:
5202:
5192:
5160:
5154:
5153:
5117:
5111:
5110:
5080:
5074:
5073:
5055:
5049:
5048:
5014:
5005:
4999:
4998:
4988:
4956:
4950:
4949:
4915:
4909:
4908:
4872:
4866:
4865:
4855:
4831:
4825:
4824:
4805:10.1038/213256a0
4791:(5073): 256–60.
4780:
4774:
4773:
4753:
4747:
4746:
4722:
4716:
4715:
4705:
4673:
4667:
4666:
4645:
4639:
4638:
4610:
4604:
4603:
4567:
4561:
4560:
4524:
4518:
4517:
4481:
4475:
4474:
4464:
4446:
4422:
4416:
4415:
4405:
4395:
4378:(40): 17079–86.
4363:
4357:
4356:
4320:
4314:
4313:
4303:
4293:
4276:(40): 14448–53.
4261:
4252:
4251:
4241:
4209:
4203:
4202:
4166:
4153:
4152:
4142:
4102:
4089:
4088:
4078:
4038:
4032:
4031:
4005:
3999:
3998:
3988:
3956:
3950:
3949:
3939:
3929:
3905:
3899:
3898:
3888:
3878:
3846:
3835:
3834:
3824:
3792:
3783:
3782:
3772:
3740:
3723:
3722:
3720:
3696:
3690:
3689:
3671:
3651:
3645:
3644:
3611:(5458): 1652–5.
3600:
3594:
3593:
3575:
3569:
3568:
3566:
3564:
3555:. Archived from
3548:
3542:
3541:
3531:
3499:
3490:
3489:
3471:
3465:
3464:
3454:
3444:
3429:BMC Microbiology
3420:
3414:
3413:
3377:
3362:
3361:
3343:Berg HC (2003).
3340:
3334:
3333:
3323:
3291:
3285:
3284:
3274:
3264:
3240:
3234:
3233:
3223:
3213:
3196:(9): 2509–2512.
3181:
3175:
3174:
3164:
3132:
3126:
3125:
3110:Berg HC (1993).
3107:
3101:
3100:
3090:
3058:
3052:
3051:
3032:10.1038/239500a0
3007:
3001:
3000:
2998:
2991:
2983:
2977:
2976:
2966:
2956:
2924:
2918:
2917:
2877:
2871:
2870:
2837:(4143): 1292–4.
2826:
2820:
2819:
2809:
2777:
2771:
2770:
2768:
2766:
2760:
2749:
2740:
2734:
2733:
2722:
2716:
2715:
2713:
2711:
2705:
2699:. Archived from
2694:
2685:
2679:
2678:
2668:
2658:
2626:
2617:
2616:
2579:
2573:
2572:
2543:
2537:
2536:
2526:
2494:
2488:
2487:
2477:
2445:
2439:
2438:
2428:
2396:
2390:
2389:
2379:
2347:
2341:
2340:
2330:
2298:
2292:
2291:
2283:
2272:
2226:McCutcheon index
2188:T-maze technique
2140:
2134:chemotaxis assay
2110:Chemotaxis assay
2092:
2090:
2089:
2084:
2082:
2081:
2065:
2063:
2062:
2057:
2055:
2054:
2038:
2036:
2035:
2030:
2003:
2001:
2000:
1995:
1971:
1969:
1968:
1963:
1961:
1948:
1947:
1901:
1899:
1891:
1883:
1877:
1873:
1842:
1841:
1799:
1797:
1789:
1781:
1765:
1763:
1762:
1757:
1745:
1743:
1742:
1737:
1714:
1712:
1711:
1706:
1682:
1680:
1679:
1674:
1666:
1665:
1650:
1648:
1640:
1632:
1616:
1614:
1613:
1608:
1596:
1594:
1593:
1588:
1586:
1585:
1572:
1570:
1569:
1564:
1552:
1550:
1549:
1544:
1532:
1530:
1529:
1524:
1512:
1510:
1509:
1504:
1489:
1487:
1486:
1481:
1455:
1454:
1438:
1436:
1435:
1430:
1428:
1427:
1414:
1412:
1411:
1406:
1391:
1389:
1388:
1383:
1296:male infertility
1209:
1191:Escherichia coli
1104:cyclooxygenase 2
1100:cyclooxygenase 1
1095:Prostaglandin D2
970:arachidonic acid
792:) as well as in
619:origin, such as
518:Netrin-1 peptide
375:residues on the
339:histidine kinase
21:
6570:
6569:
6565:
6564:
6563:
6561:
6560:
6559:
6545:Taxes (biology)
6525:
6524:
6523:
6518:
6501:(by changes in
6360:
6353:
6296:
6291:
6248:
6203:
6189:
6168:
6162:
6149:
6106:
6100:
6079:
6073:
6060:
6054:
6041:
6035:
6002:
5996:
5983:
5930:
5921:
5919:
5917:
5900:
5896:
5895:
5894:
5876:
5872:
5867:
5865:Further reading
5862:
5832:
5831:
5827:
5789:
5788:
5784:
5736:
5735:
5731:
5701:
5700:
5696:
5660:
5659:
5655:
5605:
5604:
5600:
5590:
5588:
5561:
5560:
5556:
5526:
5525:
5521:
5513:
5507:
5488:
5481:
5480:
5473:
5451:
5450:
5446:
5415:(10): 4664–71.
5402:
5401:
5397:
5367:
5366:
5362:
5316:
5315:
5311:
5267:
5266:
5262:
5218:
5217:
5206:
5162:
5161:
5157:
5119:
5118:
5114:
5107:
5082:
5081:
5077:
5070:
5057:
5056:
5052:
5012:
5007:
5006:
5002:
4958:
4957:
4953:
4938:
4917:
4916:
4912:
4874:
4873:
4869:
4833:
4832:
4828:
4782:
4781:
4777:
4755:
4754:
4750:
4724:
4723:
4719:
4675:
4674:
4670:
4647:
4646:
4642:
4612:
4611:
4607:
4569:
4568:
4564:
4535:(2–3): 143–52.
4526:
4525:
4521:
4483:
4482:
4478:
4424:
4423:
4419:
4365:
4364:
4360:
4322:
4321:
4317:
4263:
4262:
4255:
4211:
4210:
4206:
4168:
4167:
4156:
4104:
4103:
4092:
4040:
4039:
4035:
4028:
4007:
4006:
4002:
3958:
3957:
3953:
3907:
3906:
3902:
3848:
3847:
3838:
3794:
3793:
3786:
3742:
3741:
3726:
3698:
3697:
3693:
3669:10.1.1.318.4824
3653:
3652:
3648:
3602:
3601:
3597:
3590:
3577:
3576:
3572:
3562:
3560:
3559:on 11 July 2015
3550:
3549:
3545:
3501:
3500:
3493:
3486:
3473:
3472:
3468:
3422:
3421:
3417:
3394:10.1038/nrm1524
3379:
3378:
3365:
3358:
3342:
3341:
3337:
3306:(10): 862–864.
3293:
3292:
3288:
3242:
3241:
3237:
3183:
3182:
3178:
3134:
3133:
3129:
3122:
3109:
3108:
3104:
3060:
3059:
3055:
3009:
3008:
3004:
2996:
2989:
2985:
2984:
2980:
2939:(29): 12846–9.
2926:
2925:
2921:
2906:
2879:
2878:
2874:
2828:
2827:
2823:
2779:
2778:
2774:
2764:
2762:
2758:
2747:
2742:
2741:
2737:
2724:
2723:
2719:
2709:
2707:
2706:on 19 June 2010
2703:
2692:
2687:
2686:
2682:
2628:
2627:
2620:
2581:
2580:
2576:
2545:
2544:
2540:
2496:
2495:
2491:
2447:
2446:
2442:
2398:
2397:
2393:
2349:
2348:
2344:
2300:
2299:
2295:
2274:
2273:
2269:
2265:
2260:
2221:
2208:Chemical robots
2205:
2172:Zigmond chamber
2112:
2106:
2097:Spatial ecology
2073:
2068:
2067:
2046:
2041:
2040:
2006:
2005:
1977:
1976:
1973:
1959:
1958:
1939:
1902:
1884:
1879:
1878:
1833:
1832:
1828:
1800:
1782:
1771:
1770:
1748:
1747:
1728:
1727:
1688:
1687:
1633:
1626:
1625:
1599:
1598:
1575:
1574:
1555:
1554:
1535:
1534:
1515:
1514:
1495:
1494:
1491:
1444:
1443:
1417:
1416:
1394:
1393:
1374:
1373:
1333:
1292:Hodgkin disease
1263:Atherosclerosis
1212:Type of disease
1187:
1175:chemoattractant
1145:
953:Metabolites of
941:
915:Complement 3a (
904:Formyl peptides
894:
884:
876:
854:
846:
778:
715:
707:
665:
599:activation and
560:
552:
503:component, and
389:
363:
355:
331:
313:plasma membrane
299:
291:
231:
210:, have several
196:
188:
147:T. W. Engelmann
139:
28:
23:
22:
15:
12:
11:
5:
6568:
6566:
6558:
6557:
6552:
6547:
6542:
6537:
6527:
6526:
6520:
6519:
6517:
6516:
6506:
6494:
6482:
6470:
6467:magnetic field
6458:
6448:
6436:
6422:
6410:
6398:
6388:
6378:
6365:
6362:
6361:
6354:
6352:
6351:
6344:
6337:
6329:
6323:
6322:
6317:
6312:
6307:
6302:
6295:
6294:External links
6292:
6290:
6289:
6246:
6201:
6187:
6166:
6160:
6147:
6119:(1): 285–303.
6104:
6099:978-0470016176
6098:
6077:
6071:
6058:
6052:
6039:
6033:
6000:
5994:
5981:
5928:
5915:
5897:
5877:
5870:
5869:
5868:
5866:
5863:
5861:
5860:
5825:
5798:(3): 311–317.
5782:
5729:
5694:
5673:(4): 377–382.
5653:
5618:(1): 191–213.
5598:
5554:
5519:
5505:
5497:10.1007/b98868
5471:
5444:
5395:
5360:
5309:
5260:
5204:
5155:
5112:
5105:
5075:
5068:
5050:
5000:
4971:(2): 359–374.
4951:
4936:
4910:
4883:(1): 231–268.
4867:
4846:(4): 239–243.
4826:
4775:
4764:(10): 509–13.
4748:
4717:
4668:
4640:
4605:
4562:
4519:
4476:
4417:
4358:
4325:Science's STKE
4315:
4253:
4204:
4154:
4090:
4053:(2): 193–219.
4033:
4026:
4000:
3951:
3900:
3861:(3): 1059–62.
3836:
3784:
3724:
3691:
3662:(12): 569–76.
3646:
3595:
3588:
3570:
3543:
3491:
3484:
3466:
3415:
3363:
3356:
3335:
3286:
3235:
3176:
3127:
3120:
3102:
3073:(2): 262–268.
3053:
3002:
2978:
2919:
2904:
2896:10.1007/b97370
2872:
2821:
2772:
2735:
2717:
2680:
2618:
2574:
2538:
2489:
2440:
2411:(5): 608–623.
2391:
2362:(5): 296–312.
2342:
2293:
2278:, ed. (1911).
2266:
2264:
2261:
2259:
2258:
2253:
2248:
2243:
2238:
2233:
2228:
2222:
2220:
2217:
2204:
2201:
2198:
2197:
2196:
2195:
2192:
2189:
2184:
2183:
2182:
2179:
2176:
2173:
2170:
2169:Boyden chamber
2165:
2164:
2163:
2158:
2154:
2153:
2150:
2147:
2144:
2143:Type of assay
2130:
2129:
2126:
2123:
2120:
2108:Main article:
2105:
2102:
2080:
2076:
2053:
2049:
2028:
2025:
2022:
2019:
2016:
2013:
1993:
1990:
1987:
1984:
1957:
1954:
1951:
1946:
1942:
1938:
1935:
1932:
1929:
1926:
1923:
1920:
1917:
1914:
1911:
1908:
1905:
1903:
1898:
1895:
1890:
1887:
1881:
1880:
1876:
1872:
1869:
1866:
1863:
1860:
1857:
1854:
1851:
1848:
1845:
1840:
1836:
1831:
1827:
1824:
1821:
1818:
1815:
1812:
1809:
1806:
1803:
1801:
1796:
1793:
1788:
1785:
1779:
1778:
1768:
1755:
1735:
1704:
1701:
1698:
1695:
1684:
1683:
1672:
1669:
1664:
1659:
1656:
1653:
1647:
1644:
1639:
1636:
1606:
1584:
1562:
1542:
1522:
1502:
1479:
1476:
1473:
1470:
1467:
1464:
1461:
1458:
1453:
1441:
1426:
1404:
1401:
1381:
1369:
1368:
1365:
1362:
1359:
1352:
1332:
1329:
1326:
1325:
1312:
1303:
1299:
1298:
1285:
1260:
1256:
1255:
1246:
1243:
1239:
1238:
1229:
1224:
1220:
1219:
1216:
1213:
1186:
1183:
1179:chemorepellent
1144:
1141:
1140:
1139:
1138:
1137:
1123:
1115:
1112:T helper cells
1091:
1079:
1045:
999:The family of
997:
978:leukotriene B4
951:
939:
927:
912:
892:
882:
875:
872:
852:
845:
842:
826:
825:
815:
805:
777:
774:
773:
772:
756:
730:
713:
706:
703:
696:system of the
684:and posterior
674:polymerisation
664:
661:
558:
551:
548:
388:
385:
369:methylesterase
361:
354:
351:
330:
327:
297:
290:
287:
230:
227:
223:
222:
219:
194:
187:
184:
163:I. Metchnikoff
155:H. S. Jennings
138:
135:
113:carcinogenesis
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
6567:
6556:
6553:
6551:
6548:
6546:
6543:
6541:
6538:
6536:
6533:
6532:
6530:
6514:
6510:
6507:
6504:
6500:
6499:
6495:
6492:
6488:
6487:
6483:
6480:
6476:
6475:
6471:
6468:
6464:
6463:
6459:
6456:
6452:
6449:
6446:
6442:
6441:
6437:
6434:
6430:
6426:
6423:
6420:
6416:
6415:
6411:
6408:
6404:
6403:
6399:
6396:
6392:
6389:
6386:
6382:
6379:
6376:
6372:
6371:
6367:
6366:
6363:
6358:
6350:
6345:
6343:
6338:
6336:
6331:
6330:
6327:
6321:
6318:
6316:
6313:
6311:
6308:
6306:
6303:
6301:
6298:
6297:
6293:
6286:
6282:
6277:
6272:
6268:
6264:
6260:
6256:
6252:
6247:
6243:
6239:
6234:
6229:
6224:
6219:
6215:
6211:
6207:
6202:
6198:
6194:
6190:
6188:9780123747884
6184:
6180:
6176:
6172:
6167:
6163:
6157:
6153:
6148:
6144:
6140:
6135:
6130:
6126:
6122:
6118:
6114:
6110:
6105:
6101:
6095:
6091:
6087:
6083:
6078:
6074:
6068:
6064:
6059:
6055:
6049:
6045:
6040:
6036:
6030:
6026:
6022:
6018:
6014:
6010:
6006:
6005:Physics Today
6001:
5997:
5991:
5987:
5982:
5978:
5974:
5969:
5964:
5960:
5956:
5951:
5950:10.1.1.515.32
5946:
5942:
5938:
5934:
5929:
5918:
5912:
5908:
5904:
5899:
5898:
5892:
5891:
5890:
5884:
5880:
5864:
5856:
5852:
5848:
5844:
5840:
5836:
5829:
5826:
5821:
5817:
5813:
5809:
5805:
5801:
5797:
5793:
5786:
5783:
5778:
5774:
5770:
5766:
5762:
5758:
5753:
5748:
5744:
5740:
5733:
5730:
5725:
5721:
5717:
5713:
5709:
5705:
5698:
5695:
5690:
5686:
5681:
5676:
5672:
5668:
5664:
5657:
5654:
5649:
5645:
5641:
5637:
5633:
5629:
5625:
5621:
5617:
5613:
5609:
5602:
5599:
5587:
5583:
5578:
5573:
5569:
5565:
5558:
5555:
5550:
5546:
5542:
5538:
5534:
5530:
5523:
5520:
5512:
5508:
5502:
5498:
5494:
5487:
5486:
5478:
5476:
5472:
5467:
5463:
5459:
5455:
5448:
5445:
5440:
5436:
5432:
5428:
5423:
5418:
5414:
5410:
5406:
5399:
5396:
5391:
5387:
5383:
5379:
5375:
5371:
5364:
5361:
5356:
5352:
5347:
5342:
5337:
5332:
5328:
5324:
5320:
5313:
5310:
5305:
5301:
5296:
5291:
5287:
5283:
5280:(4): 651–65.
5279:
5275:
5271:
5264:
5261:
5256:
5252:
5247:
5242:
5238:
5234:
5231:(4): 340–55.
5230:
5226:
5222:
5215:
5213:
5211:
5209:
5205:
5200:
5196:
5191:
5186:
5182:
5178:
5174:
5170:
5166:
5159:
5156:
5151:
5147:
5143:
5139:
5135:
5131:
5127:
5123:
5116:
5113:
5108:
5102:
5098:
5094:
5090:
5086:
5079:
5076:
5071:
5065:
5061:
5054:
5051:
5046:
5042:
5038:
5034:
5030:
5026:
5022:
5018:
5011:
5004:
5001:
4996:
4992:
4987:
4982:
4978:
4974:
4970:
4966:
4962:
4955:
4952:
4947:
4943:
4939:
4937:9780128035955
4933:
4929:
4925:
4921:
4914:
4911:
4906:
4902:
4898:
4894:
4890:
4886:
4882:
4878:
4871:
4868:
4863:
4859:
4854:
4849:
4845:
4841:
4837:
4830:
4827:
4822:
4818:
4814:
4810:
4806:
4802:
4798:
4794:
4790:
4786:
4779:
4776:
4771:
4767:
4763:
4759:
4752:
4749:
4744:
4740:
4736:
4732:
4728:
4721:
4718:
4713:
4709:
4704:
4699:
4695:
4691:
4687:
4683:
4679:
4672:
4669:
4664:
4660:
4656:
4652:
4651:Front Immunol
4644:
4641:
4636:
4632:
4628:
4624:
4620:
4616:
4609:
4606:
4601:
4597:
4593:
4589:
4585:
4581:
4577:
4573:
4566:
4563:
4558:
4554:
4550:
4546:
4542:
4538:
4534:
4530:
4523:
4520:
4515:
4511:
4507:
4503:
4499:
4495:
4492:(4): 375–94.
4491:
4487:
4480:
4477:
4472:
4468:
4463:
4458:
4454:
4450:
4445:
4444:10.1.1.515.32
4440:
4436:
4432:
4428:
4421:
4418:
4413:
4409:
4404:
4399:
4394:
4389:
4385:
4381:
4377:
4373:
4369:
4362:
4359:
4354:
4350:
4346:
4342:
4338:
4334:
4330:
4326:
4319:
4316:
4311:
4307:
4302:
4297:
4292:
4287:
4283:
4279:
4275:
4271:
4267:
4260:
4258:
4254:
4249:
4245:
4240:
4235:
4231:
4227:
4223:
4219:
4215:
4208:
4205:
4200:
4196:
4192:
4188:
4184:
4180:
4176:
4172:
4165:
4163:
4161:
4159:
4155:
4150:
4146:
4141:
4136:
4132:
4128:
4124:
4120:
4116:
4112:
4111:Physics Today
4108:
4101:
4099:
4097:
4095:
4091:
4086:
4082:
4077:
4072:
4068:
4064:
4060:
4056:
4052:
4048:
4044:
4037:
4034:
4029:
4023:
4019:
4015:
4011:
4004:
4001:
3996:
3992:
3987:
3982:
3978:
3974:
3971:(2): 101–10.
3970:
3966:
3962:
3955:
3952:
3947:
3943:
3938:
3933:
3928:
3923:
3919:
3915:
3911:
3904:
3901:
3896:
3892:
3887:
3882:
3877:
3872:
3868:
3864:
3860:
3856:
3852:
3845:
3843:
3841:
3837:
3832:
3828:
3823:
3818:
3814:
3810:
3806:
3802:
3798:
3791:
3789:
3785:
3780:
3776:
3771:
3766:
3762:
3758:
3755:(2): 301–19.
3754:
3750:
3746:
3739:
3737:
3735:
3733:
3731:
3729:
3725:
3719:
3714:
3710:
3706:
3702:
3695:
3692:
3687:
3683:
3679:
3675:
3670:
3665:
3661:
3657:
3650:
3647:
3642:
3638:
3634:
3630:
3626:
3622:
3618:
3614:
3610:
3606:
3599:
3596:
3591:
3589:9789812707932
3585:
3581:
3574:
3571:
3558:
3554:
3547:
3544:
3539:
3535:
3530:
3525:
3521:
3517:
3513:
3509:
3505:
3498:
3496:
3492:
3487:
3481:
3477:
3470:
3467:
3462:
3458:
3453:
3448:
3443:
3438:
3434:
3430:
3426:
3419:
3416:
3411:
3407:
3403:
3399:
3395:
3391:
3387:
3383:
3376:
3374:
3372:
3370:
3368:
3364:
3359:
3353:
3349:
3345:
3339:
3336:
3331:
3327:
3322:
3317:
3313:
3309:
3305:
3301:
3297:
3290:
3287:
3282:
3278:
3273:
3268:
3263:
3258:
3254:
3250:
3246:
3239:
3236:
3231:
3227:
3222:
3217:
3212:
3207:
3203:
3199:
3195:
3191:
3187:
3180:
3177:
3172:
3168:
3163:
3158:
3154:
3150:
3146:
3142:
3138:
3131:
3128:
3123:
3117:
3113:
3106:
3103:
3098:
3094:
3089:
3084:
3080:
3076:
3072:
3068:
3064:
3057:
3054:
3049:
3045:
3041:
3037:
3033:
3029:
3025:
3021:
3017:
3013:
3006:
3003:
2995:
2988:
2982:
2979:
2974:
2970:
2965:
2960:
2955:
2950:
2946:
2942:
2938:
2934:
2930:
2923:
2920:
2915:
2911:
2907:
2905:0-387-00888-8
2901:
2897:
2893:
2889:
2885:
2884:
2876:
2873:
2868:
2864:
2860:
2856:
2852:
2848:
2844:
2840:
2836:
2832:
2825:
2822:
2817:
2813:
2808:
2803:
2799:
2795:
2792:(2): 259–75.
2791:
2787:
2783:
2776:
2773:
2757:
2753:
2746:
2739:
2736:
2731:
2727:
2721:
2718:
2702:
2698:
2691:
2684:
2681:
2676:
2672:
2667:
2662:
2657:
2652:
2648:
2644:
2640:
2636:
2632:
2625:
2623:
2619:
2614:
2610:
2606:
2602:
2598:
2594:
2590:
2586:
2578:
2575:
2570:
2566:
2562:
2558:
2554:
2550:
2542:
2539:
2534:
2530:
2525:
2520:
2516:
2512:
2508:
2504:
2500:
2493:
2490:
2485:
2481:
2476:
2471:
2467:
2463:
2459:
2455:
2451:
2444:
2441:
2436:
2432:
2427:
2422:
2418:
2414:
2410:
2406:
2402:
2395:
2392:
2387:
2383:
2378:
2373:
2369:
2365:
2361:
2357:
2353:
2346:
2343:
2338:
2334:
2329:
2324:
2320:
2316:
2313:(6): 378–91.
2312:
2308:
2304:
2297:
2294:
2289:
2288:
2282:
2277:
2271:
2268:
2262:
2257:
2254:
2252:
2249:
2247:
2244:
2242:
2239:
2237:
2234:
2232:
2229:
2227:
2224:
2223:
2218:
2216:
2212:
2209:
2202:
2193:
2190:
2187:
2186:
2185:
2180:
2177:
2175:Dunn chambers
2174:
2171:
2168:
2167:
2166:
2161:
2160:
2159:
2156:
2155:
2151:
2148:
2145:
2142:
2141:
2138:
2135:
2127:
2124:
2121:
2118:
2117:
2116:
2111:
2103:
2101:
2098:
2094:
2078:
2074:
2051:
2047:
2023:
2020:
2017:
2011:
1988:
1982:
1972:
1952:
1944:
1940:
1933:
1927:
1921:
1918:
1915:
1909:
1906:
1904:
1896:
1888:
1874:
1870:
1861:
1855:
1852:
1849:
1846:
1838:
1834:
1829:
1825:
1819:
1813:
1807:
1804:
1802:
1794:
1786:
1767:
1753:
1733:
1725:
1722:
1718:
1696:
1670:
1667:
1657:
1651:
1645:
1637:
1624:
1623:
1622:
1620:
1604:
1560:
1540:
1520:
1500:
1490:
1477:
1468:
1462:
1459:
1456:
1440:
1402:
1379:
1366:
1363:
1360:
1357:
1353:
1350:
1346:
1342:
1338:
1337:
1336:
1330:
1324:
1320:
1316:
1313:
1311:
1307:
1304:
1302:Intoxications
1301:
1300:
1297:
1293:
1289:
1286:
1284:
1280:
1276:
1272:
1271:periodontitis
1268:
1264:
1261:
1258:
1257:
1254:
1250:
1247:
1244:
1241:
1240:
1237:
1233:
1230:
1228:
1227:Inflammations
1225:
1222:
1221:
1217:
1214:
1211:
1210:
1207:
1202:
1200:
1196:
1192:
1184:
1182:
1180:
1176:
1172:
1171:oligopeptides
1168:
1164:
1160:
1153:
1149:
1142:
1135:
1131:
1127:
1124:
1121:
1120:
1116:
1113:
1109:
1105:
1101:
1097:
1096:
1092:
1089:
1085:
1084:
1080:
1077:
1073:
1069:
1065:
1061:
1057:
1053:
1049:
1046:
1043:
1039:
1035:
1031:
1027:
1023:
1019:
1018:
1013:
1009:
1008:
1002:
998:
995:
991:
987:
983:
979:
975:
971:
967:
963:
962:
958:
957:
956:
952:
949:
945:
937:
933:
932:
928:
924:
922:
918:
913:
910:
905:
902:
901:
900:
890:
880:
873:
871:
869:
865:
861:
850:
843:
841:
839:
835:
831:
823:
819:
816:
813:
809:
806:
803:
799:
798:
797:
795:
791:
787:
783:
775:
770:
766:
762:
761:
757:
754:
750:
746:
741:
737:
736:
731:
727:
726:
722:
721:
720:
711:
704:
702:
699:
695:
691:
687:
683:
679:
675:
671:
662:
660:
658:
654:
653:embryogenesis
650:
646:
642:
638:
634:
630:
626:
622:
618:
614:
610:
604:
602:
598:
594:
590:
585:
583:
582:cell membrane
578:
575:
571:
567:
556:
549:
547:
545:
544:
539:
535:
531:
527:
523:
519:
515:
514:
508:
506:
502:
498:
494:
490:
486:
482:
479:
474:
471:
467:
466:
461:
459:
458:R. spheroides
454:
453:
448:
443:
439:
437:
433:
429:
425:
421:
417:
408:
404:
402:
398:
394:
386:
384:
382:
378:
374:
370:
359:
352:
350:
348:
344:
340:
336:
328:
326:
324:
323:
318:
314:
310:
306:
295:
288:
286:
284:
279:
277:
273:
267:
264:
263:
257:
252:
250:
246:
245:
240:
236:
228:
226:
220:
217:
216:
215:
213:
209:
208:
203:
192:
185:
183:
181:
176:
172:
168:
164:
160:
156:
152:
151:W. F. Pfeffer
148:
144:
136:
134:
132:
128:
124:
120:
116:
114:
110:
107:
103:
99:
95:
91:
90:fertilization
87:
83:
79:
75:
71:
70:multicellular
67:
63:
59:
58:Somatic cells
55:
54:
49:
48:
43:
34:
30:
19:
6535:Motile cells
6508:
6496:
6484:
6472:
6462:Magnetotaxis
6460:
6450:
6438:
6429:Galvanotaxis
6428:
6425:Electrotaxis
6424:
6412:
6401:
6400:
6390:
6380:
6368:
6261:(2): 177–8.
6258:
6254:
6213:
6210:PLOS Biology
6209:
6170:
6151:
6116:
6112:
6081:
6062:
6043:
6008:
6004:
5985:
5940:
5936:
5922:18 September
5920:. Retrieved
5906:
5887:
5886:
5885:profile for
5882:
5838:
5834:
5828:
5795:
5791:
5785:
5742:
5739:Biochemistry
5738:
5732:
5707:
5703:
5697:
5670:
5666:
5656:
5615:
5611:
5601:
5589:. Retrieved
5567:
5557:
5532:
5528:
5522:
5484:
5460:: OL487–95.
5457:
5453:
5447:
5412:
5408:
5398:
5376:(2): 65–71.
5373:
5369:
5363:
5326:
5322:
5312:
5277:
5273:
5263:
5228:
5224:
5172:
5168:
5158:
5128:(7): 481–6.
5125:
5121:
5115:
5088:
5078:
5062:. Springer.
5059:
5053:
5023:(2): 453–6.
5020:
5016:
5003:
4968:
4964:
4954:
4919:
4913:
4880:
4876:
4870:
4843:
4839:
4829:
4788:
4784:
4778:
4761:
4757:
4751:
4726:
4720:
4685:
4681:
4671:
4654:
4650:
4643:
4618:
4614:
4608:
4575:
4571:
4565:
4532:
4528:
4522:
4489:
4485:
4479:
4434:
4430:
4420:
4375:
4371:
4361:
4331:(219): pl3.
4328:
4324:
4318:
4273:
4269:
4221:
4217:
4207:
4174:
4170:
4117:(2): 24–30.
4114:
4110:
4050:
4046:
4036:
4009:
4003:
3968:
3964:
3954:
3917:
3913:
3903:
3858:
3854:
3807:(1): 383–8.
3804:
3800:
3752:
3748:
3708:
3704:
3694:
3659:
3655:
3649:
3608:
3604:
3598:
3579:
3573:
3561:. Retrieved
3557:the original
3553:"Chemotaxis"
3546:
3511:
3507:
3475:
3469:
3432:
3428:
3418:
3385:
3381:
3347:
3344:
3338:
3303:
3299:
3289:
3255:(14): 7521.
3252:
3248:
3238:
3193:
3189:
3179:
3147:(2): 262–8.
3144:
3140:
3130:
3111:
3105:
3070:
3066:
3056:
3015:
3011:
3005:
2981:
2936:
2932:
2922:
2882:
2875:
2834:
2830:
2824:
2789:
2785:
2775:
2763:. Retrieved
2756:the original
2751:
2738:
2729:
2720:
2708:. Retrieved
2701:the original
2696:
2683:
2638:
2634:
2588:
2584:
2577:
2552:
2548:
2541:
2506:
2502:
2492:
2457:
2453:
2443:
2408:
2404:
2394:
2359:
2355:
2345:
2310:
2306:
2296:
2285:
2270:
2246:Mechanotaxis
2213:
2207:
2206:
2131:
2113:
2095:
1974:
1769:
1685:
1492:
1442:
1370:
1334:
1205:
1188:
1156:
1125:
1117:
1093:
1081:
1067:
1063:
1059:
1051:
1047:
1015:
1011:
1005:
990:inflammation
961:Leukotrienes
959:
929:
914:
903:
897:
867:
859:
857:
827:
818:Leukotrienes
779:
758:
753:angiogenesis
733:
725:Chemokinesis
723:
718:
694:microtubular
666:
621:immune cells
605:
603:production.
586:
579:
573:
569:
563:
541:
537:
512:
509:
475:
469:
465:B. substilis
463:
456:
450:
446:
444:
440:
413:
390:
366:
332:
322:Che proteins
320:
302:
282:
280:
268:
260:
253:
242:
232:
224:
205:
199:
167:phagocytosis
140:
131:chemokinesis
122:
118:
117:
64:, and other
51:
45:
41:
40:
29:
6509:Thigmotaxis
6503:temperature
6498:Thermotaxis
5535:: 346–358.
5329:: 1329–47.
4578:: 687–717.
3920:: 7142868.
3514:: 457–512.
2591:: 348–359.
2460:(1): 40–9.
2251:Plithotaxis
1236:Brucellosis
1167:amino acids
1078:chemotaxis.
1038:neutrophils
1026:neutrophils
1022:eosinophils
984:, BLT1 and
834:amino acids
657:germ layers
625:granulocyte
613:Tetrahymena
589:methylation
538:Tetrahymena
497:macrophages
493:neutrophils
452:S. meliloti
381:transferase
239:random walk
149:(1881) and
143:Leeuwenhoek
98:lymphocytes
66:single-cell
18:Chemotactic
6540:Perception
6529:Categories
6491:fluid flow
6474:Phototaxis
6451:Hydrotaxis
6440:Gravitaxis
6402:Chemotaxis
6381:Anemotaxis
6300:Chemotaxis
6216:(2): E49.
6063:Chemotaxis
5889:Chemotaxis
5752:1809.02530
2276:Chisholm H
2263:References
2241:Haptotaxis
2162:PP-chamber
1717:divergence
1351:migration)
1310:benzpyrene
1223:Infections
1076:neutrophil
1072:eosinophil
966:eicosanoid
931:Chemokines
808:Chemokines
790:rhodopsins
760:Necrotaxis
735:haptotaxis
729:behaviors.
698:basal body
682:pseudopods
649:metastases
641:fibroblast
633:lymphocyte
566:eukaryotic
543:Paramecium
530:nociceptin
501:complement
489:leukocytes
487:, attract
483:, such as
432:dipeptides
204:, such as
157:(1906) in
109:metastasis
102:leukocytes
42:Chemotaxis
6486:Rheotaxis
6419:stiffness
6414:Durotaxis
6407:chemicals
6391:Barotaxis
6370:Aerotaxis
6355:Types of
5945:CiteSeerX
5648:243966350
5640:1947-5454
5586:257388244
4621:: 111–8.
4514:248703226
4439:CiteSeerX
4199:207440927
3664:CiteSeerX
3410:205493118
3348:in motion
2890:, 19–29.
2555:: 37–45.
2509:: 44–51.
2236:Durotaxis
2157:Examples
2079:φ
2018:φ
1953:φ
1950:∇
1945:φ
1934:⋅
1931:∇
1916:φ
1894:∂
1889:φ
1886:∂
1871:φ
1868:∇
1862:φ
1856:χ
1850:−
1844:∇
1826:⋅
1823:∇
1792:∂
1784:∂
1754:φ
1697:⋅
1694:∇
1658:⋅
1655:∇
1643:∂
1638:ρ
1635:∂
1561:ρ
1541:χ
1521:χ
1478:φ
1475:∇
1469:φ
1463:χ
1403:φ
1400:∇
1380:φ
1356:diffusion
1345:flagellum
1275:psoriasis
1267:arthritis
1056:Mead acid
1042:bacterial
1030:monocytes
936:cytokines
794:olfaction
782:G-protein
776:Receptors
769:apoptotic
637:mast cell
424:galactose
393:inorganic
377:cytosolic
373:glutamate
315:into the
272:flagellin
249:diffusion
175:H. Harris
6455:moisture
6395:pressure
6285:23653726
6242:14966542
6197:19203648
6143:22994495
5977:18685153
5855:29064685
5820:29461522
5777:52816076
5769:30251529
5724:23308365
5689:84150518
5591:13 March
5511:Archived
5466:14995080
5439:23499393
5390:25480980
5355:17767353
5304:24056189
5255:25449650
5150:33755476
5122:Cytokine
5045:27784085
4995:29873835
4946:26928542
4905:14193715
4897:10611962
4743:38951708
4729:: 1–15.
4712:37220748
4703:10524337
4663:36341452
4635:26820523
4600:11331182
4592:22804578
4557:31704677
4549:17557195
4506:10735174
4471:18685153
4412:20864631
4345:14872096
4310:25249632
4248:32186970
4191:19747082
4149:24363460
3995:18404496
3946:27123011
3779:15187186
3686:15539117
3633:10698740
3563:23 March
3461:15955239
3402:15573139
3330:28845044
3281:34299141
3171:22169400
3097:22169400
2994:Archived
2973:20615986
2914:35733036
2710:15 April
2675:34050026
2613:53116963
2605:30366278
2569:26271598
2533:29042028
2484:25301842
2435:32877650
2386:29546880
2337:27231052
2219:See also
1349:amoeboid
1306:Asbestos
1163:receptor
765:necrotic
740:gradient
629:monocyte
526:PACAP-38
505:pathogen
491:such as
481:peptides
319:, where
256:gradient
229:Behavior
212:flagella
202:bacteria
180:J. Adler
159:ciliates
123:negative
119:Positive
62:bacteria
6445:gravity
6276:3577161
6134:3989901
6013:Bibcode
5968:7213762
5879:Scholia
5800:Bibcode
5620:Bibcode
5537:Bibcode
5431:8906847
5346:5901339
5295:5710732
5246:5710736
5190:2109936
5142:9702410
5037:7117406
4986:6099250
4862:8807851
4821:4212997
4813:6030602
4793:Bibcode
4462:7213762
4403:2951443
4380:Bibcode
4353:4660870
4301:4210025
4278:Bibcode
4239:7346724
4140:3867297
4119:Bibcode
4076:1473391
4055:Bibcode
3986:2096533
3937:4830718
3895:1093163
3863:Bibcode
3831:2403544
3641:5334523
3613:Bibcode
3605:Science
3538:9442881
3529:2899694
3452:1183210
3346:E. coli
3321:5649114
3272:8306008
3230:4560688
3198:Bibcode
3162:3320702
3088:3320702
3048:1909173
3040:4563019
3020:Bibcode
2964:2919929
2941:Bibcode
2867:7221477
2859:4598187
2839:Bibcode
2831:Science
2816:4873021
2807:2138524
2765:1 April
2666:8179161
2643:Bibcode
2524:5679233
2475:4429868
2426:7505206
2377:6790333
2328:5367630
2231:Tropism
2152:Others
1715:is the
1321:salts,
994:allergy
838:insulin
749:ligands
686:uropods
574:E. coli
570:E. coli
401:ligands
397:organic
337:in the
317:cytosol
262:E. coli
244:E. coli
207:E. coli
127:kinesis
94:neurons
80:(e.g.,
78:poisons
74:glucose
6375:oxygen
6283:
6273:
6240:
6233:340952
6230:
6195:
6185:
6158:
6141:
6131:
6096:
6069:
6050:
6031:
5992:
5975:
5965:
5947:
5913:
5881:has a
5853:
5818:
5775:
5767:
5722:
5687:
5646:
5638:
5584:
5503:
5464:
5437:
5429:
5388:
5353:
5343:
5302:
5292:
5253:
5243:
5199:264125
5197:
5187:
5148:
5140:
5103:
5066:
5043:
5035:
4993:
4983:
4944:
4934:
4903:
4895:
4860:
4819:
4811:
4785:Nature
4770:199132
4768:
4741:
4710:
4700:
4661:
4633:
4598:
4590:
4555:
4547:
4512:
4504:
4469:
4459:
4441:
4410:
4400:
4351:
4343:
4308:
4298:
4246:
4236:
4197:
4189:
4147:
4137:
4085:911982
4083:
4073:
4024:
3993:
3983:
3944:
3934:
3893:
3886:432465
3883:
3829:
3822:208443
3819:
3777:
3770:419924
3767:
3684:
3666:
3639:
3631:
3586:
3536:
3526:
3482:
3459:
3449:
3435:: 35.
3408:
3400:
3354:
3328:
3318:
3279:
3269:
3228:
3221:426976
3218:
3169:
3159:
3118:
3095:
3085:
3046:
3038:
3012:Nature
2971:
2961:
2912:
2902:
2865:
2857:
2814:
2804:
2673:
2663:
2611:
2603:
2567:
2531:
2521:
2482:
2472:
2433:
2423:
2384:
2374:
2335:
2325:
1975:where
1686:where
1493:where
1159:ligand
1130:ALOX15
1088:ALOX12
1028:, and
1014:, and
944:RANTES
860:ad hoc
824:(BLT).
804:(FPR),
786:genome
609:Amoeba
528:, and
478:Formyl
470:E.coli
455:, and
447:E.coli
436:serine
428:phenol
420:ribose
283:Vibrio
106:cancer
82:phenol
47:chemo-
44:(from
6479:light
6357:taxes
5883:topic
5773:S2CID
5747:arXiv
5685:S2CID
5644:S2CID
5582:S2CID
5514:(PDF)
5489:(PDF)
5435:S2CID
5146:S2CID
5041:S2CID
5013:(PDF)
4901:S2CID
4817:S2CID
4596:S2CID
4553:S2CID
4510:S2CID
4349:S2CID
4195:S2CID
3637:S2CID
3406:S2CID
3044:S2CID
2997:(PDF)
2990:(PDF)
2910:S2CID
2863:S2CID
2759:(PDF)
2748:(PDF)
2704:(PDF)
2693:(PDF)
2609:S2CID
1341:cilia
1323:ozone
974:ALOX5
926:well.
690:Cilia
678:actin
426:with
410:float
200:Some
86:sperm
53:taxis
6511:(by
6489:(by
6477:(by
6465:(by
6453:(by
6443:(by
6431:(by
6417:(by
6405:(by
6393:(by
6385:wind
6383:(by
6281:PMID
6238:PMID
6193:PMID
6183:ISBN
6156:ISBN
6139:PMID
6094:ISBN
6067:ISBN
6048:ISBN
6029:ISBN
5990:ISBN
5973:PMID
5924:2017
5911:ISBN
5851:PMID
5816:PMID
5765:PMID
5720:PMID
5636:ISSN
5593:2023
5501:ISBN
5462:PMID
5427:PMID
5386:PMID
5351:PMID
5300:PMID
5251:PMID
5229:1851
5195:PMID
5138:PMID
5101:ISBN
5064:ISBN
5033:PMID
4991:PMID
4942:PMID
4932:ISBN
4893:PMID
4858:PMID
4809:PMID
4766:PMID
4739:PMID
4708:PMID
4659:PMID
4631:PMID
4588:PMID
4545:PMID
4502:PMID
4467:PMID
4408:PMID
4341:PMID
4329:2004
4306:PMID
4244:PMID
4187:PMID
4145:PMID
4081:PMID
4022:ISBN
3991:PMID
3942:PMID
3918:2016
3891:PMID
3827:PMID
3775:PMID
3682:PMID
3629:PMID
3584:ISBN
3565:2017
3534:PMID
3480:ISBN
3457:PMID
3398:PMID
3352:ISBN
3326:PMID
3277:PMID
3226:PMID
3167:PMID
3116:ISBN
3093:PMID
3036:PMID
2969:PMID
2900:ISBN
2855:PMID
2812:PMID
2767:2017
2712:2017
2671:PMID
2601:PMID
2565:PMID
2529:PMID
2480:PMID
2431:PMID
2382:PMID
2333:PMID
2066:and
1746:and
1347:and
1317:and
1232:AIDS
1074:and
1050:and
992:and
986:BLT2
964:are
948:IL-8
738:the
670:PIP3
601:PIP3
540:and
495:and
485:fMLF
445:For
434:and
422:and
343:CheB
276:TLR5
6427:or
6271:PMC
6263:doi
6228:PMC
6218:doi
6175:doi
6129:PMC
6121:doi
6086:doi
6021:doi
5963:PMC
5955:doi
5941:121
5843:doi
5839:139
5808:doi
5757:doi
5712:doi
5708:135
5675:doi
5628:doi
5572:doi
5545:doi
5493:doi
5417:doi
5413:157
5378:doi
5374:157
5341:PMC
5331:doi
5290:PMC
5282:doi
5241:PMC
5233:doi
5185:PMC
5177:doi
5130:doi
5093:doi
5025:doi
5021:140
4981:PMC
4973:doi
4969:104
4924:doi
4885:doi
4848:doi
4801:doi
4789:213
4762:101
4731:doi
4698:PMC
4690:doi
4623:doi
4580:doi
4537:doi
4494:doi
4457:PMC
4449:doi
4435:121
4398:PMC
4388:doi
4376:107
4333:doi
4296:PMC
4286:doi
4274:111
4234:PMC
4226:doi
4179:doi
4175:390
4135:PMC
4127:doi
4071:PMC
4063:doi
4014:doi
3981:PMC
3973:doi
3932:PMC
3922:doi
3881:PMC
3871:doi
3817:PMC
3809:doi
3805:172
3765:PMC
3757:doi
3713:doi
3674:doi
3621:doi
3609:287
3524:PMC
3516:doi
3447:PMC
3437:doi
3390:doi
3316:PMC
3308:doi
3267:PMC
3257:doi
3216:PMC
3206:doi
3157:PMC
3149:doi
3083:PMC
3075:doi
3028:doi
3016:239
2959:PMC
2949:doi
2937:107
2892:doi
2847:doi
2835:184
2802:PMC
2794:doi
2790:128
2661:PMC
2651:doi
2639:118
2593:doi
2557:doi
2519:PMC
2511:doi
2470:PMC
2462:doi
2421:PMC
2413:doi
2372:PMC
2364:doi
2323:PMC
2315:doi
1169:or
1102:or
1066:,11
972:by
921:C5a
917:C3a
767:or
732:In
676:of
611:or
597:Ras
534:ATP
522:GTP
395:or
96:or
68:or
6531::
6279:.
6269:.
6259:11
6257:.
6253:.
6236:.
6226:.
6212:.
6208:.
6191:.
6181:.
6137:.
6127:.
6117:66
6115:.
6111:.
6092:.
6084:.
6027:.
6019:.
6009:58
6007:.
5971:.
5961:.
5953:.
5939:.
5935:.
5905:.
5849:.
5837:.
5814:.
5806:.
5796:10
5794:.
5771:.
5763:.
5755:.
5743:57
5741:.
5718:.
5706:.
5683:.
5669:.
5665:.
5642:.
5634:.
5626:.
5616:13
5614:.
5610:.
5580:.
5566:.
5543:.
5533:69
5531:.
5509:.
5499:.
5474:^
5458:49
5456:.
5433:.
5425:.
5411:.
5407:.
5384:.
5372:.
5349:.
5339:.
5325:.
5321:.
5298:.
5288:.
5278:52
5276:.
5272:.
5249:.
5239:.
5227:.
5223:.
5207:^
5193:.
5183:.
5173:75
5171:.
5167:.
5144:.
5136:.
5126:10
5124:.
5099:.
5039:.
5031:.
5019:.
5015:.
4989:.
4979:.
4967:.
4963:.
4940:.
4930:.
4899:.
4891:.
4881:15
4879:.
4856:.
4842:.
4838:.
4815:.
4807:.
4799:.
4787:.
4760:.
4737:.
4706:.
4696:.
4686:58
4684:.
4680:.
4655:13
4653:.
4629:.
4619:55
4617:.
4594:.
4586:.
4576:28
4574:.
4551:.
4543:.
4533:12
4531:.
4508:.
4500:.
4490:50
4488:.
4465:.
4455:.
4447:.
4433:.
4429:.
4406:.
4396:.
4386:.
4374:.
4370:.
4347:.
4339:.
4327:.
4304:.
4294:.
4284:.
4272:.
4268:.
4256:^
4242:.
4232:.
4222:31
4220:.
4216:.
4193:.
4185:.
4173:.
4157:^
4143:.
4133:.
4125:.
4115:66
4113:.
4109:.
4093:^
4079:.
4069:.
4061:.
4051:20
4049:.
4045:.
4020:.
3989:.
3979:.
3967:.
3963:.
3940:.
3930:.
3916:.
3912:.
3889:.
3879:.
3869:.
3859:72
3857:.
3853:.
3839:^
3825:.
3815:.
3803:.
3799:.
3787:^
3773:.
3763:.
3753:68
3751:.
3747:.
3727:^
3709:71
3707:.
3703:.
3680:.
3672:.
3660:12
3658:.
3635:.
3627:.
3619:.
3607:.
3532:.
3522:.
3512:13
3510:.
3506:.
3494:^
3455:.
3445:.
3431:.
3427:.
3404:.
3396:.
3384:.
3366:^
3324:.
3314:.
3304:14
3302:.
3298:.
3275:.
3265:.
3253:22
3251:.
3247:.
3224:.
3214:.
3204:.
3194:69
3192:.
3188:.
3165:.
3155:.
3145:24
3143:.
3139:.
3091:.
3081:.
3071:24
3069:.
3065:.
3042:.
3034:.
3026:.
3014:.
2992:.
2967:.
2957:.
2947:.
2935:.
2931:.
2908:.
2898:.
2888:15
2861:.
2853:.
2845:.
2833:.
2810:.
2800:.
2788:.
2784:.
2750:.
2728:.
2695:.
2669:.
2659:.
2649:.
2637:.
2633:.
2621:^
2607:.
2599:.
2589:65
2587:.
2563:.
2553:34
2551:.
2527:.
2517:.
2507:33
2505:.
2501:.
2478:.
2468:.
2458:35
2456:.
2452:.
2429:.
2419:.
2409:54
2407:.
2403:.
2380:.
2370:.
2360:18
2358:.
2354:.
2331:.
2321:.
2311:16
2309:.
2305:.
2284:.
2093:.
1621::
1319:Cr
1315:Hg
1308:,
1294:,
1290:,
1281:,
1277:,
1273:,
1269:,
1265:,
1251:,
1234:,
1062:,8
1058:(5
1024:,
1010:,
836:,
832:,
820:-
810:-
688:.
643:,
639:,
631:,
627:,
524:,
449:,
133:.
60:,
50:+
6515:)
6505:)
6493:)
6481:)
6469:)
6457:)
6447:)
6435:)
6421:)
6409:)
6397:)
6387:)
6377:)
6348:e
6341:t
6334:v
6287:.
6265::
6244:.
6220::
6214:2
6199:.
6177::
6164:.
6145:.
6123::
6102:.
6088::
6075:.
6056:.
6037:.
6023::
6015::
5998:.
5979:.
5957::
5926:.
5893:.
5857:.
5845::
5822:.
5810::
5802::
5779:.
5759::
5749::
5726:.
5714::
5691:.
5677::
5671:8
5650:.
5630::
5622::
5595:.
5574::
5551:.
5547::
5539::
5495::
5468:.
5441:.
5419::
5392:.
5380::
5357:.
5333::
5327:7
5306:.
5284::
5257:.
5235::
5201:.
5179::
5152:.
5132::
5109:.
5095::
5072:.
5047:.
5027::
4997:.
4975::
4948:.
4926::
4907:.
4887::
4864:.
4850::
4844:3
4823:.
4803::
4795::
4772:.
4745:.
4733::
4714:.
4692::
4665:.
4637:.
4625::
4602:.
4582::
4559:.
4539::
4516:.
4496::
4473:.
4451::
4414:.
4390::
4382::
4355:.
4335::
4312:.
4288::
4280::
4250:.
4228::
4201:.
4181::
4151:.
4129::
4121::
4087:.
4065::
4057::
4030:.
4016::
3997:.
3975::
3969:1
3948:.
3924::
3897:.
3873::
3865::
3833:.
3811::
3781:.
3759::
3721:.
3715::
3688:.
3676::
3643:.
3623::
3615::
3592:.
3540:.
3518::
3488:.
3463:.
3439::
3433:5
3412:.
3392::
3386:5
3360:.
3332:.
3310::
3283:.
3259::
3232:.
3208::
3200::
3173:.
3151::
3124:.
3099:.
3077::
3050:.
3030::
3022::
2975:.
2951::
2943::
2916:.
2894::
2869:.
2849::
2841::
2818:.
2796::
2769:.
2714:.
2677:.
2653::
2645::
2615:.
2595::
2571:.
2559::
2535:.
2513::
2486:.
2464::
2437:.
2415::
2388:.
2366::
2339:.
2317::
2075:D
2052:C
2048:D
2027:)
2024:C
2021:,
2015:(
2012:g
1992:)
1989:C
1986:(
1983:f
1956:)
1941:D
1937:(
1928:+
1925:)
1922:C
1919:,
1913:(
1910:g
1907:=
1897:t
1875:]
1865:)
1859:(
1853:C
1847:C
1839:C
1835:D
1830:[
1820:+
1817:)
1814:C
1811:(
1808:f
1805:=
1795:t
1787:C
1766::
1734:C
1703:)
1700:(
1671:S
1668:=
1663:J
1652:+
1646:t
1605:S
1583:J
1501:C
1472:)
1466:(
1460:C
1457:=
1452:J
1425:J
1343:/
1245:—
1161:-
1068:Z
1064:Z
1060:Z
996:.
940:3
923:)
911:.
755:.
460:,
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.