1655:
87:
58:
3169:
2071:
3157:
2011:
infection is higher because many patients have weakened immune systems or have undergone procedures. In healthcare, the risk of more serious staph infection is higher for patients in intensive care units (ICUs), patients who have undergone certain types of surgeries and patients with medical devices inserted in their bodies.
2010:
It is said that anyone can develop a staph infection, although certain groups of people are at greater risk, including people with chronic conditions such as diabetes, cancer, vascular disease, eczema, lung disease, and people who inject drugs. In healthcare facilities, the risk of more serious staph
3142:
Stevens D, Cornmell R, Taylor D, Grimshaw SG, Riazanskaia S, Arnold DS, Fernstad SJ, Smith AM, Heaney LM, Reynolds JC, Thomas CL, Harker M. Spatial variations in the microbial community structure and diversity of the human foot is associated with the production of odorous volatiles. FEMS Microbiol
1608:. This may lead to better control of outbreak strains. A greater understanding of how the staphylococci evolve, especially due to the acquisition of mobile genetic elements encoding resistance and virulence genes is helping to identify new outbreak strains and may even prevent their emergence.
2046:
can cause a wide variety of diseases in humans and animals through either toxin production or penetration. Staphylococcal toxins are a common cause of food poisoning, for they can be produced by bacteria growing in improperly stored food items. The most common
2021:. It facilitates factors such as tissue adhesion, immune evasion, and host cell injury. In the bloodstream, these factors cause inflammation, impair immune cell function, alter coagulation, and compromise vascular integrity. When left untreated,
1500:
inhabits and sometimes infects the skin of domestic dogs and cats. This organism, too, can carry the genetic material that imparts multiple bacterial resistance. It is rarely implicated in infections in humans, as a
1587:
genomes have been submitted to the public databases, making it one of the most extensively sequenced bacteria. The use of genomic data is now widespread and provides a valuable resource for researchers working with
1604:. Relating this information to pathogenic behaviour is one of the major areas of staphylococcal research. The development of molecular typing methods has enabled the tracking of different strains of
3338:
2694:"Identification of marine sponge-associated bacteria of the Saint Martin's island of the Bay of Bengal emphasizing on the prevention of motile Aeromonas septicemia in Labeo rohita"
2025:
triggers pathophysiologic disturbances that are further amplified by the host inflammatory response, culminating in the severe clinical manifestations of sepsis and
3312:
1631:
to be much greater than previously expected, and encompasses genes with functions beyond antibiotic resistance and virulence, and beyond genes residing within the
729:. Despite strong attempts to get rid of them, staphylococcus bacteria stay present in hospitals, where they can infect people who are most at risk of infection.
3351:
2514:
Pantůček R, Sedláček I, Indráková A, Vrbovská V, Mašlaňová I, Kovařovic V, Švec P, Králová S, Krištofová L, Kekláková J, Petráš P, Doškař J (October 2017).
2084:
2093:
3286:
3325:
1378:(100 μg disc: resistance = < 15 mm zone of inhibition). Further biochemical testing is needed to identify to the species level.
2457:
Tong SY, Schaumburg F, Ellington MJ, Corander J, Pichon B, Leendertz F, Bentley SD, Parkhill J, Holt DC, Peters G, Giffard PM (January 2015).
2912:
2740:
2339:
1690:
species observed on some animals appear more rarely on more distantly related host species. Some of the observed host specificity includes:
1643:
1627:
of genes encoding antibiotic/metal resistance and virulence. A recent study demonstrated the extent of horizontal gene transfer among
1565:
3374:
3173:
3412:
2810:
Jin M, Rosario W, Watler E, Calhoun DH (March 2004). "Development of a large-scale HPLC-based purification for the urease from
2199:
1496:
473:
644:
501:
466:
2692:
Paul, Sulav Indra; Rahman, Md. Mahbubur; Salam, Mohammad Abdus; Khan, Md. Arifur Rahman; Islam, Md. Tofazzal (2021-12-15).
670:
3330:
1332:
species can be differentiated from other aerobic and facultative anaerobic, Gram-positive cocci by several simple tests.
86:
3417:
2139:
1527:
1386:
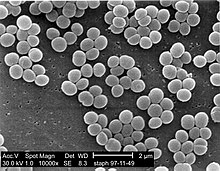
One of the most important phenotypical features used in the classification of staphylococci is their ability to produce
550:
508:
65:
2175:
736:, one has three subspecies, and one has four subspecies. Many species cannot cause disease and reside normally on the
557:
480:
459:
417:
340:
263:
2270:"Among-population variation in microbial community structure in the floral nectar of the bee-pollinated forest herb
2229:, and techniques for identifying and quantifying S. aureus adhesins in relation to adhesion to biomaterials: review"
3422:
2400:
Kloos WE, Ballard DN, George CG, Webster JA, Hubner RJ, Ludwig W, Schleifer KH, Fiedler F, Schubert K (July 1998).
2187:
2151:
1545:
1509:
1299:
has yet to be clarified. The published descriptions of these species do not appear to have been validly published.
396:
361:
305:
249:
200:
155:
3356:
3161:
1551:
515:
438:
333:
326:
270:
242:
1065:
522:
3407:
2038:
1624:
1612:
1522:
1059:
571:
487:
291:
172:
1078:
368:
298:
725:
Staphylococcus was one of the leading infections in hospitals and many strains of this bacterium have become
452:
445:
410:
277:
3402:
1632:
564:
543:
284:
235:
207:
179:
2459:"Novel staphylococcal species that form part of a Staphylococcus aureus-related complex: the non-pigmented
3200:
2693:
1686:
species are highly salt tolerant. Some species specificity has been observed in host range, such that the
661:
592:
585:
578:
424:
375:
347:
312:
221:
214:
186:
38:
382:
2056:
1557:
1456:
536:
529:
494:
431:
403:
389:
354:
256:
228:
193:
2581:"Reclassification of Staphylococcus pulvereri Zakrzewska-Czerwinska et al. 1995 as a later synonym of
2562:
Coates-Brown R, Moran JC, Pongchaikul P, Darby AC and MJ Horsburgh MJ (2018) "Comparative genomics of
3379:
3366:
3260:
2284:
1219:
726:
319:
3238:
1536:
697:
682:
1654:
2159:
688:
81:
1098:
Based on an analysis of orthologous gene content three groups (A, B and C) have been proposed.
3317:
3247:
3104:
3086:
3002:
2963:
2908:
2880:
2831:
2792:
2736:
2713:
2659:
2606:
2579:
Svec P, Vancanneyt M, Sedlácek I, Engelbeen K, Stetina V, Swings J, Petrás P (November 2004).
2545:
2496:
2439:
2382:
2335:
2312:
2250:
1518:
1484:) to water and oxygen, which makes the catalase test useful to distinguish staphylococci from
1473:
1355:
620:
138:
3179:
1340:(capable of growth both aerobically and anaerobically). All species grow in the presence of
3094:
3076:
2994:
2953:
2945:
2870:
2862:
2823:
2782:
2705:
2649:
2641:
2630:
species based on different partial gap, 16S rRNA, hsp60, rpoB, sodA, and tuf gene sequences"
2596:
2535:
2527:
2486:
2478:
2429:
2372:
2302:
2292:
2240:
1985:
1968:
1911:
1845:
1464:
strains are coagulase-positive, some may be atypical in that they do not produce coagulase.
651:
1678:
frequently colonize the skin and upper respiratory tracts of mammals and birds and also in
3065:"Igniting the Fire: Staphylococcus aureus Virulence Factors in the Pathogenesis of Sepsis"
3020:
2163:
741:
57:
1326:
type and teichoic acid presence) and G + C content of DNA in a range of 30–40 mol%.
2998:
2288:
2224:
3143:
Ecol. 2015 Jan;91(1):1-11. doi: 10.1093/femsec/fiu018. Epub 2014 Dec 8. PMID: 25764539.
3099:
3064:
2958:
2929:
2875:
2850:
2654:
2625:
2540:
2516:"mecCgene and genomic islands with suspected role in adaptation to extreme environment"
2515:
2491:
2458:
2307:
2269:
2076:
1374:(0.04 U disc: resistance = < 10 mm zone of inhibition) and susceptible to
2709:
2117:"staphylococcus | Origin and meaning of staphylococcus by Online Etymology Dictionary"
1564:
Common abbreviations for coagulase-negative staphylococci are CoNS, CNS, or CNST. The
3396:
2183:
2147:
1663:
1532:
1489:
1460:
is coagulase-positive, meaning it produces coagulase. However, while the majority of
1323:
1312:
656:
654:(1844–1929), following the pattern established five years earlier with the naming of
613:
31:
2048:
2026:
1833:
1735:
1485:
1375:
757:
3252:
2566:
reveals determinants of speciation and diversification of antimicrobial defense".
3081:
2297:
2116:
3299:
3232:
1857:
1600:
strains. Each contains different combinations of surface proteins and different
1472:-positive (meaning that it can produce the enzyme catalase) and able to convert
1231:
1046:
3168:
3038:
2787:
2766:
2070:
1351:
were once thought to be coagulase-positive, but this has since been disproven.
2827:
2434:
2401:
2377:
2356:
2066:
2051:
is caused by staphylococci, as bacterial infections. Staphylococci break down
1851:
1827:
1593:
1391:
1371:
1341:
1222:. This group is the only clade within the staphylococci to possess this gene.
1196:
1083:
733:
628:
624:
128:
17:
3223:
3090:
2717:
2162:, with the assistance of McKenzie, Roderick. Oxford: Clarendon Press. In the
3343:
3273:
1514:
1449:
1387:
1337:
1319:
108:
3122:
3108:
2967:
2884:
2835:
2796:
2663:
2610:
2549:
2500:
2482:
2386:
2316:
2254:
3156:
3006:
2601:
2580:
2443:
1539:
infections in sexually active young women. In recent years, several other
3217:
2866:
2645:
2531:
2245:
1989:
1817:
1583:
to be sequenced were those of N315 and Mu50, in 2001. Many more complete
1502:
1469:
772:
sequences, and most of the staphylococcal species fall into 11 clusters:
616:
98:
2949:
2268:
Jacquemyn H, Lenaerts M, Brys R, Willems K, Honnay O, Lievens B (2013).
3291:
2052:
1766:
1698:
1215:
745:
118:
3304:
3185:
2018:
1882:
1784:
1775:
1739:
1706:
1679:
1580:
1315:
753:
732:
Staphylococcus includes at least 44 species. Of these, nine have two
3194:
2899:
2765:
Matthews KR, Roberson J, Gillespie BE, Luther DA, Oliver SP (1997).
1397:
Seven species are currently recognised as being coagulase-positive:
1050:, the species of which are currently the closest known relatives of
3278:
3063:
Powers, Michael E.; Wardenburg, Juliane Bubeck (13 February 2014).
650:
The name was coined in 1880 by
Scottish surgeon and bacteriologist
3189:
1901:
1886:
1839:
1793:
1667:
1653:
1601:
636:
632:
609:
2589:
International
Journal of Systematic and Evolutionary Microbiology
2471:
International
Journal of Systematic and Evolutionary Microbiology
1721:
1702:
1531:, another coagulase-negative species that is part of the normal
769:
737:
3198:
3265:
1946:
1867:
1802:
1725:
2176:
2140:
1592:. Whole genome technologies, such as sequencing projects and
1642:
are available from biological research centres, such as the
710:
675:
2767:"Identification and Differentiation of Coagulase-Negative
1543:
species have been implicated in human infections, notably
3021:"Symptoms and Treatments of Staph Infection (Cellulitis)"
2624:
Ghebremedhin B, Layer F, König W, König B (March 2008).
1568:
abbreviates coagulase-negative staphylococci as "CoNS".
647:(capable of growth both aerobically and anaerobically).
1229:
group appears to be the closest relations to the genus
1069:– both of which were previously considered variants of
3123:"Sialoadenitis: inflammation of the salivary glands"
3039:"Staphylococcus aureus in Healthcare Settings | HAI"
2357:"Phylogenetic relationships of 38 taxa of the genus
1429:. These species belong to two separate groups – the
3207:
2981:Kloos WE (1980). "Natural populations of the genus
2223:Harris LG, Foster SJ, Richards RG (December 2002).
702:'spherical bacterium' (from Ancient Greek:
72:colonies: Note the grape-like clustering common to
1354:Growth can also occur in a 6.5% NaCl solution. On
756:microbes. They are also a small component of the
2930:"Lateral transfer of genes and gene fragments in
2422:International Journal of Systematic Bacteriology
2365:International Journal of Systematic Bacteriology
1517:of the skin, but can cause severe infections in
1218:-positive due to their possession of the enzyme
687:'bunch of grapes'), and suffixed by the
37:"Staph" redirects here. Not to be confused with
2849:Becker K, Heilmann C, Peters G (October 2014).
2626:"Genetic classification and distinguishing of
2463:sp. nov. and the non-human primate-associated
2355:Takahashi T, Satoh I, Kikuchi N (April 1999).
3188:, a Bioinformatics Resource Center funded by
3125:. The Medical Consumer's Advocate. 2001-01-04
2418:sp. no. and Macrococcus carouselicus sp. nov"
8:
1444:An eighth species has also been described –
1086:. This species is probably a member of the
2928:Chan CX, Beiko RG, Ragan MA (August 2011).
717:
660:. It combines the prefix "staphylo-" (from
3195:
1666:– numbered ticks on the scale are 11
56:
45:
3098:
3080:
2957:
2874:
2814:and determination of subunit structure".
2786:
2653:
2600:
2539:
2490:
2433:
2376:
2361:based on 16S rRNA gene sequence analysis"
2306:
2296:
2244:
1241:has been shown to be a junior synonym of
1362:species grow fermentatively, except for
1318:that forms clusters, has an appropriate
2108:
1256:groups appear to be related, as do the
1044:– has now been removed to a new genus,
2520:Applied and Environmental Microbiology
2158:, revised and augmented throughout by
2687:
2685:
2683:
2681:
2679:
2677:
2675:
2673:
1513:, a coagulase-negative species, is a
1057:Two species were described in 2015 –
7:
3367:6b5535fe-b96b-48b5-9d70-a268b72efa29
2330:Madigan M, Martinko J, eds. (2005).
1644:National Collection of Type Cultures
1596:, have shown an enormous variety of
1307:Assignment of a strain to the genus
3184:genomes and related information at
2999:10.1146/annurev.mi.34.100180.003015
2816:Protein Expression and Purification
1966:– South American squirrel monkeys (
1076:A new coagulase negative species –
2851:"Coagulase-negative staphylococci"
2017:has emerged as a leading agent of
1284:, but this needs to be confirmed.
25:
2755:PreTest, Surgery, 12th ed., p. 88
2710:10.1016/j.aquaculture.2021.737156
1619:, or across different species of
1566:American Society for Microbiology
1535:, is predominantly implicated in
3167:
3155:
2634:Journal of Clinical Microbiology
2334:(11th ed.). Prentice Hall.
2332:Brock Biomlogy of Microorganisms
2069:
85:
2934:extends beyond mobile elements"
722:'grain, seed, berry').
645:facultative anaerobic organisms
27:Genus of Gram-positive bacteria
2731:Ryan KJ, Ray CG, eds. (2004).
2233:European Cells & Materials
2059:, the main odor of foot odor.
1572:Genomics and molecular biology
752:species have been found to be
1:
2987:Annual Review of Microbiology
2855:Clinical Microbiology Reviews
2771:by Polymerase Chain Reaction"
2735:(4th ed.). McGraw Hill.
1270:appears to be related to the
768:The taxonomy is based on 16s
3082:10.1371/journal.ppat.1003871
2733:Sherris Medical Microbiology
2298:10.1371/journal.pone.0056917
1611:The widespread incidence of
1441:group (the remaining five).
711:
676:
66:Scanning electron micrograph
1682:. Marine sponge associated
1366:, which grows oxidatively.
1287:The taxonomic position of
3439:
2907:. Caister Academic Press.
2788:10.4315/0362-028X-60.6.686
2775:Journal of Food Protection
2465:Staphylococcus schweitzeri
2410:gen. nov., comb. nov. and
2036:
1615:across various strains of
1303:Biochemical identification
1066:Staphylococcus schweitzeri
1040:A twelfth group – that of
703:
665:
36:
29:
2828:10.1016/j.pep.2003.10.012
2435:10.1099/00207713-48-3-859
2378:10.1099/00207713-49-2-725
1370:species are resistant to
1248:Within these clades, the
1082:– has been isolated from
631:, they appear spherical (
168:
163:
82:Scientific classification
80:
64:
55:
48:
2583:Staphylococcus vitulinus
2461:Staphylococcus argenteus
2412:Macrococcus equipercicus
2408:Macrococcus caseolyticus
2039:Staphylococcal infection
2002:Populations at risk for
1625:horizontal gene transfer
1523:central venous catheters
1521:patients and those with
1390:, an enzyme that causes
1336:species are facultative
1079:Staphylococcus edaphicus
1060:Staphylococcus argenteus
30:Not to be confused with
2938:Journal of Bacteriology
2898:Lindsay J, ed. (2008).
2406:through description of
2177:
2164:Perseus Digital Library
2160:Jones, Sir Henry Stuart
2156:A Greek–English Lexicon
2141:
1633:mobile genetic elements
1623:has been attributed to
3413:Gram-positive bacteria
2483:10.1099/ijs.0.062752-0
2402:"Delimiting the genus
2272:Pulmonaria officinalis
2085:Methicillin-resistant
1671:
692:
2769:Staphylococcus aureus
2602:10.1099/ijs.0.63080-0
2227:Staphylococcus aureus
2148:Liddell, Henry George
2094:Vancomycin-resistant
2015:Staphylococcus aureus
2004:Staphylococcus aureus
1960:– humans, dogs, goats
1674:Members of the genus
1657:
1613:antibiotic resistance
1448:– from patients with
1439:S. hyicus-intermedius
1437:alone) group and the
1349:Staphylococcus aureus
1322:structure (including
1195:groups are generally
878:S. hyicus-intermedius
3164:at Wikimedia Commons
3027:. December 14, 2013.
2904:: Molecular Genetics
2867:10.1128/CMR.00109-13
2646:10.1128/JCM.02058-07
2585:Webster et al. 1994"
2532:10.1128/AEM.01746-17
2246:10.22203/ecm.v004a04
2225:"An introduction to
1382:Coagulase production
1311:requires it to be a
1297:S. pseudolugdunensis
1220:cytochrome c oxidase
744:of humans and other
727:antibiotic resistant
481:S. pseudolugdunensis
3418:Pathogenic bacteria
2950:10.1128/JB.01524-10
2812:Staphylococcus leei
2416:Macrococcus bovicus
2289:2013PLoSO...856917J
2166:, Tufts University.
1937:S. pseudintermedius
1816:– chickens, goats,
1658:Unknown variety of
1638:Various strains of
1537:genitourinary tract
1497:S. pseudintermedius
1446:Staphylococcus leei
1419:S. pseudintermedius
1356:Baird-Parker medium
1177:S. pseudintermedius
922:S. pseudintermedius
474:S. pseudintermedius
2200:"Staph infections"
2121:www.etymonline.com
1759:– chickens, humans
1672:
1280:may be related to
1199:-resistant, as is
852:S. saccharolyticus
827:S. piscifermentans
502:S. saccharolyticus
467:S. piscifermentans
3423:Staphylococcaceae
3390:
3389:
3201:Taxon identifiers
3160:Media related to
2914:978-1-904455-29-5
2742:978-0-8385-8529-0
2341:978-0-13-144329-7
1474:hydrogen peroxide
1167:Group C includes
1144:Group B includes
1101:Group A includes
754:nectar-inhabiting
721:
709:
701:
686:
674:
621:Staphylococcaceae
601:
600:
159:
139:Staphylococcaceae
16:(Redirected from
3430:
3383:
3382:
3370:
3369:
3360:
3359:
3347:
3346:
3334:
3333:
3321:
3320:
3308:
3307:
3295:
3294:
3282:
3281:
3269:
3268:
3256:
3255:
3243:
3242:
3241:
3228:
3227:
3226:
3196:
3172:Data related to
3171:
3159:
3144:
3140:
3134:
3133:
3131:
3130:
3119:
3113:
3112:
3102:
3084:
3060:
3054:
3053:
3051:
3050:
3035:
3029:
3028:
3017:
3011:
3010:
2978:
2972:
2971:
2961:
2925:
2919:
2918:
2895:
2889:
2888:
2878:
2846:
2840:
2839:
2807:
2801:
2800:
2790:
2762:
2756:
2753:
2747:
2746:
2728:
2722:
2721:
2689:
2668:
2667:
2657:
2621:
2615:
2614:
2604:
2595:(Pt 6): 2213–5.
2576:
2570:
2560:
2554:
2553:
2543:
2526:(2): e01746–17.
2511:
2505:
2504:
2494:
2454:
2448:
2447:
2437:
2397:
2391:
2390:
2380:
2352:
2346:
2345:
2327:
2321:
2320:
2310:
2300:
2265:
2259:
2258:
2248:
2220:
2214:
2213:
2211:
2210:
2196:
2190:
2180:
2173:
2167:
2144:
2137:
2131:
2130:
2128:
2127:
2113:
2079:
2074:
2073:
1986:Cercopithecoidea
1969:Saimiri sciureus
1912:Microtus arvalis
1714:– humans, cattle
1528:S. saprophyticus
1519:immunosuppressed
1364:S. saprophyticus
1189:S. saprophyticus
1158:S. saprophyticus
1088:S. saprophyticus
980:S. saprophyticus
944:S. saprophyticus
742:mucous membranes
719:
716:
714:
708:romanized:
707:
705:
696:
681:
679:
669:
667:
652:Alexander Ogston
639:-like clusters.
551:S. singaporensis
509:S. saprophyticus
154:
90:
89:
60:
46:
21:
3438:
3437:
3433:
3432:
3431:
3429:
3428:
3427:
3408:Bacteria genera
3393:
3392:
3391:
3386:
3378:
3373:
3365:
3363:
3355:
3350:
3342:
3337:
3329:
3324:
3316:
3311:
3303:
3298:
3290:
3285:
3277:
3272:
3264:
3259:
3251:
3246:
3237:
3236:
3231:
3222:
3221:
3216:
3203:
3152:
3147:
3141:
3137:
3128:
3126:
3121:
3120:
3116:
3075:(2): e1003871.
3062:
3061:
3057:
3048:
3046:
3037:
3036:
3032:
3025:Just-Health.net
3019:
3018:
3014:
2980:
2979:
2975:
2944:(15): 3964–77.
2927:
2926:
2922:
2915:
2897:
2896:
2892:
2848:
2847:
2843:
2809:
2808:
2804:
2764:
2763:
2759:
2754:
2750:
2743:
2730:
2729:
2725:
2691:
2690:
2671:
2623:
2622:
2618:
2578:
2577:
2573:
2568:Front Microbiol
2561:
2557:
2513:
2512:
2508:
2477:(Pt 1): 15–22.
2456:
2455:
2451:
2399:
2398:
2394:
2354:
2353:
2349:
2342:
2329:
2328:
2324:
2267:
2266:
2262:
2222:
2221:
2217:
2208:
2206:
2198:
2197:
2193:
2174:
2170:
2138:
2134:
2125:
2123:
2115:
2114:
2110:
2106:
2075:
2068:
2065:
2057:isovaleric acid
2041:
2035:
2008:
1931:S. pettenkoferi
1927:– humans, goats
1894:– humans, goats
1823:S. haemolyticus
1753:– goats, humans
1652:
1574:
1483:
1479:
1384:
1347:All species of
1305:
1282:S. haemolyticus
1272:S. haemolyticus
1250:S. haemolyticus
1210:Members of the
1205:novobiosepticus
1185:
1131:S. pettenkoferi
1119:S. haemolyticus
1096:
1042:S. caseolyticus
1009:S. stepanovicii
890:S. cornubiensis
869:S. haemolyticus
857:S. haemolyticus
823:S. massiliensis
766:
758:soil microbiome
635:), and form in
623:from the order
558:S. stepanovicii
460:S. pettenkoferi
418:S. massiliensis
341:S. haemolyticus
264:S. cornubiensis
243:S. caseolyticus
153:
84:
42:
35:
28:
23:
22:
15:
12:
11:
5:
3436:
3434:
3426:
3425:
3420:
3415:
3410:
3405:
3403:Staphylococcus
3395:
3394:
3388:
3387:
3385:
3384:
3371:
3361:
3348:
3344:staphylococcus
3335:
3322:
3309:
3296:
3283:
3270:
3257:
3244:
3239:Staphylococcus
3229:
3213:
3211:
3209:Staphylococcus
3205:
3204:
3199:
3193:
3192:
3181:Staphylococcus
3177:
3176:at Wikispecies
3174:Staphylococcus
3165:
3162:Staphylococcus
3151:
3150:External links
3148:
3146:
3145:
3135:
3114:
3069:PLOS Pathogens
3055:
3030:
3012:
2983:Staphylococcus
2973:
2932:Staphylococcus
2920:
2913:
2902:Staphylococcus
2890:
2861:(4): 870–926.
2841:
2802:
2757:
2748:
2741:
2723:
2669:
2640:(3): 1019–25.
2628:Staphylococcus
2616:
2571:
2564:Staphylococcus
2555:
2506:
2449:
2414:sp. nov., and
2404:Staphylococcus
2392:
2359:Staphylococcus
2347:
2340:
2322:
2260:
2215:
2204:mayoclinic.org
2191:
2168:
2132:
2107:
2105:
2102:
2101:
2100:
2091:
2081:
2080:
2077:Biology portal
2064:
2061:
2044:Staphylococcus
2037:Main article:
2034:
2031:
2007:
2000:
1999:
1998:
1992:
1979:
1973:
1961:
1955:
1949:
1940:
1934:
1928:
1922:
1916:
1904:
1895:
1892:S. lugdunensis
1889:
1876:
1870:
1861:
1820:
1811:
1805:
1796:
1787:
1781:S. epidermidis
1778:
1769:
1760:
1754:
1748:
1742:
1729:
1718:S. auricularis
1715:
1709:
1688:Staphylococcus
1684:Staphylococcus
1676:Staphylococcus
1660:Staphylococcus
1651:
1648:
1640:Staphylococcus
1629:Staphylococcus
1621:Staphylococcus
1573:
1570:
1546:S. lugdunensis
1541:Staphylococcus
1510:S. epidermidis
1481:
1477:
1411:S. intermedius
1383:
1380:
1368:Staphylococcus
1360:Staphylococcus
1334:Staphylococcus
1330:Staphylococcus
1309:Staphylococcus
1304:
1301:
1268:S. lugdunensis
1262:S. epidermidis
1184:
1181:
1173:S. intermedius
1127:S. lugdunensis
1115:S. epidermidis
1095:
1092:
1052:Staphylococcus
1038:
1037:
1024:
1015:
990:
941:
939:S. lugdunensis
935:S. lugdunensis
932:
906:S. intermedius
886:S. chromogenes
875:
854:
848:S. epidermidis
836:S. epidermidis
833:
804:
802:S. auricularis
798:S. auricularis
795:
789:S. schweitzeri
765:
762:
750:Staphylococcus
641:Staphylococcus
619:in the family
605:Staphylococcus
599:
598:
597:
596:
589:
582:
575:
568:
561:
554:
547:
540:
533:
526:
523:S. schweitzeri
519:
512:
505:
498:
491:
484:
477:
470:
463:
456:
449:
442:
435:
428:
421:
414:
407:
400:
397:S. lugdunensis
393:
386:
379:
372:
365:
362:S. intermedius
358:
351:
344:
337:
330:
323:
316:
309:
306:S. epidermidis
302:
295:
288:
281:
274:
267:
260:
253:
250:S. chromogenes
246:
239:
232:
225:
218:
211:
204:
201:S. auricularis
197:
190:
183:
176:
166:
165:
161:
160:
150:Staphylococcus
146:
142:
141:
136:
132:
131:
126:
122:
121:
116:
112:
111:
106:
102:
101:
96:
92:
91:
78:
77:
74:Staphylococcus
62:
61:
53:
52:
50:Staphylococcus
26:
24:
18:Staphylococcal
14:
13:
10:
9:
6:
4:
3:
2:
3435:
3424:
3421:
3419:
3416:
3414:
3411:
3409:
3406:
3404:
3401:
3400:
3398:
3381:
3376:
3372:
3368:
3362:
3358:
3353:
3349:
3345:
3340:
3336:
3332:
3327:
3323:
3319:
3314:
3310:
3306:
3301:
3297:
3293:
3288:
3284:
3280:
3275:
3271:
3267:
3262:
3258:
3254:
3249:
3245:
3240:
3234:
3230:
3225:
3219:
3215:
3214:
3212:
3210:
3206:
3202:
3197:
3191:
3187:
3183:
3182:
3178:
3175:
3170:
3166:
3163:
3158:
3154:
3153:
3149:
3139:
3136:
3124:
3118:
3115:
3110:
3106:
3101:
3096:
3092:
3088:
3083:
3078:
3074:
3070:
3066:
3059:
3056:
3044:
3040:
3034:
3031:
3026:
3022:
3016:
3013:
3008:
3004:
3000:
2996:
2992:
2988:
2984:
2977:
2974:
2969:
2965:
2960:
2955:
2951:
2947:
2943:
2939:
2935:
2933:
2924:
2921:
2916:
2910:
2906:
2905:
2901:
2894:
2891:
2886:
2882:
2877:
2872:
2868:
2864:
2860:
2856:
2852:
2845:
2842:
2837:
2833:
2829:
2825:
2821:
2817:
2813:
2806:
2803:
2798:
2794:
2789:
2784:
2780:
2776:
2772:
2770:
2761:
2758:
2752:
2749:
2744:
2738:
2734:
2727:
2724:
2719:
2715:
2711:
2707:
2703:
2699:
2695:
2688:
2686:
2684:
2682:
2680:
2678:
2676:
2674:
2670:
2665:
2661:
2656:
2651:
2647:
2643:
2639:
2635:
2631:
2629:
2620:
2617:
2612:
2608:
2603:
2598:
2594:
2590:
2586:
2584:
2575:
2572:
2569:
2565:
2559:
2556:
2551:
2547:
2542:
2537:
2533:
2529:
2525:
2521:
2517:
2510:
2507:
2502:
2498:
2493:
2488:
2484:
2480:
2476:
2472:
2468:
2466:
2462:
2453:
2450:
2445:
2441:
2436:
2431:
2428:(3): 859–77.
2427:
2423:
2419:
2417:
2413:
2409:
2405:
2396:
2393:
2388:
2384:
2379:
2374:
2370:
2366:
2362:
2360:
2351:
2348:
2343:
2337:
2333:
2326:
2323:
2318:
2314:
2309:
2304:
2299:
2294:
2290:
2286:
2283:(3): e56917.
2282:
2278:
2275:
2273:
2264:
2261:
2256:
2252:
2247:
2242:
2238:
2234:
2230:
2228:
2219:
2216:
2205:
2201:
2195:
2192:
2189:
2185:
2181:
2179:
2172:
2169:
2165:
2161:
2157:
2153:
2152:Scott, Robert
2149:
2145:
2143:
2136:
2133:
2122:
2118:
2112:
2109:
2103:
2098:
2097:
2092:
2089:
2088:
2083:
2082:
2078:
2072:
2067:
2062:
2060:
2058:
2054:
2050:
2045:
2040:
2032:
2030:
2028:
2024:
2020:
2016:
2012:
2005:
2001:
1996:
1993:
1991:
1987:
1983:
1980:
1977:
1974:
1971:
1970:
1965:
1962:
1959:
1956:
1953:
1952:S. schleiferi
1950:
1948:
1944:
1941:
1938:
1935:
1932:
1929:
1926:
1923:
1920:
1919:S. nepalensis
1917:
1914:
1913:
1908:
1905:
1903:
1899:
1896:
1893:
1890:
1888:
1884:
1880:
1877:
1874:
1871:
1869:
1865:
1862:
1860:
1859:
1854:
1853:
1848:
1847:
1842:
1841:
1836:
1835:
1830:
1829:
1824:
1821:
1819:
1815:
1814:S. gallinarum
1812:
1809:
1808:S. fleurettii
1806:
1804:
1800:
1797:
1795:
1791:
1788:
1786:
1785:marine sponge
1782:
1779:
1777:
1773:
1770:
1768:
1764:
1761:
1758:
1755:
1752:
1749:
1746:
1743:
1741:
1737:
1733:
1730:
1727:
1723:
1719:
1716:
1713:
1710:
1708:
1707:marine sponge
1704:
1700:
1696:
1693:
1692:
1691:
1689:
1685:
1681:
1680:marine sponge
1677:
1669:
1665:
1661:
1656:
1649:
1647:
1645:
1641:
1636:
1634:
1630:
1626:
1622:
1618:
1614:
1609:
1607:
1603:
1599:
1595:
1591:
1586:
1582:
1579:
1571:
1569:
1567:
1562:
1560:
1559:
1554:
1553:
1552:S. schleiferi
1548:
1547:
1542:
1538:
1534:
1533:vaginal flora
1530:
1529:
1524:
1520:
1516:
1512:
1511:
1506:
1504:
1499:
1498:
1493:
1491:
1487:
1475:
1471:
1467:
1463:
1459:
1458:
1453:
1451:
1447:
1442:
1440:
1436:
1432:
1428:
1424:
1423:S. schleiferi
1420:
1416:
1412:
1408:
1404:
1400:
1395:
1393:
1389:
1381:
1379:
1377:
1373:
1369:
1365:
1361:
1357:
1352:
1350:
1345:
1343:
1339:
1335:
1331:
1327:
1325:
1324:peptidoglycan
1321:
1317:
1314:
1313:Gram-positive
1310:
1302:
1300:
1298:
1294:
1290:
1285:
1283:
1279:
1275:
1273:
1269:
1265:
1263:
1259:
1255:
1251:
1246:
1244:
1240:
1236:
1234:
1233:
1228:
1223:
1221:
1217:
1213:
1208:
1206:
1202:
1198:
1194:
1190:
1182:
1180:
1178:
1174:
1170:
1165:
1163:
1159:
1155:
1151:
1147:
1142:
1140:
1136:
1132:
1128:
1124:
1120:
1116:
1112:
1108:
1104:
1099:
1093:
1091:
1089:
1085:
1081:
1080:
1074:
1072:
1068:
1067:
1062:
1061:
1055:
1053:
1049:
1048:
1043:
1036:
1032:
1028:
1025:
1023:
1019:
1016:
1014:
1010:
1006:
1002:
998:
997:S. fleurettii
994:
991:
989:
985:
981:
977:
976:S. nepalensis
973:
969:
965:
964:S. gallinarum
961:
957:
953:
949:
945:
942:
940:
936:
933:
931:
930:S. schleiferi
927:
923:
919:
915:
911:
907:
903:
899:
895:
891:
887:
883:
879:
876:
874:
870:
866:
862:
858:
855:
853:
849:
845:
841:
837:
834:
832:
828:
824:
820:
816:
815:S. condimenti
812:
808:
805:
803:
799:
796:
794:
790:
786:
782:
778:
775:
774:
773:
771:
763:
761:
759:
755:
751:
747:
743:
739:
735:
730:
728:
723:
713:
699:
694:
690:
684:
678:
672:
663:
662:Ancient Greek
659:
658:
657:Streptococcus
653:
648:
646:
642:
638:
634:
630:
626:
622:
618:
615:
614:Gram-positive
611:
607:
606:
595:
594:
590:
588:
587:
583:
581:
580:
576:
574:
573:
569:
567:
566:
562:
560:
559:
555:
553:
552:
548:
546:
545:
541:
539:
538:
534:
532:
531:
527:
525:
524:
520:
518:
517:
516:S. schleiferi
513:
511:
510:
506:
504:
503:
499:
497:
496:
492:
490:
489:
485:
483:
482:
478:
476:
475:
471:
469:
468:
464:
462:
461:
457:
455:
454:
450:
448:
447:
443:
441:
440:
439:S. nepalensis
436:
434:
433:
429:
427:
426:
422:
420:
419:
415:
413:
412:
408:
406:
405:
401:
399:
398:
394:
392:
391:
387:
385:
384:
380:
378:
377:
373:
371:
370:
366:
364:
363:
359:
357:
356:
352:
350:
349:
345:
343:
342:
338:
336:
335:
334:S. gallinarum
331:
329:
328:
327:S. fleurettii
324:
322:
321:
317:
315:
314:
310:
308:
307:
303:
301:
300:
296:
294:
293:
289:
287:
286:
282:
280:
279:
275:
273:
272:
271:S. condimenti
268:
266:
265:
261:
259:
258:
254:
252:
251:
247:
245:
244:
240:
238:
237:
233:
231:
230:
226:
224:
223:
219:
217:
216:
212:
210:
209:
205:
203:
202:
198:
196:
195:
191:
189:
188:
184:
182:
181:
177:
175:
174:
170:
169:
167:
162:
157:
152:
151:
147:
144:
143:
140:
137:
134:
133:
130:
127:
124:
123:
120:
117:
114:
113:
110:
107:
104:
103:
100:
97:
94:
93:
88:
83:
79:
75:
71:
67:
63:
59:
54:
51:
47:
44:
40:
33:
32:Streptococcus
19:
3208:
3180:
3138:
3127:. Retrieved
3117:
3072:
3068:
3058:
3047:. Retrieved
3045:. 2020-12-10
3042:
3033:
3024:
3015:
2990:
2986:
2982:
2976:
2941:
2937:
2931:
2923:
2903:
2900:
2893:
2858:
2854:
2844:
2822:(1): 111–7.
2819:
2815:
2811:
2805:
2781:(6): 686–8.
2778:
2774:
2768:
2760:
2751:
2732:
2726:
2701:
2697:
2637:
2633:
2627:
2619:
2592:
2588:
2582:
2574:
2567:
2563:
2558:
2523:
2519:
2509:
2474:
2470:
2464:
2460:
2452:
2425:
2421:
2415:
2411:
2407:
2403:
2395:
2371:(2): 725–8.
2368:
2364:
2358:
2350:
2331:
2325:
2280:
2276:
2271:
2263:
2236:
2232:
2226:
2218:
2207:. Retrieved
2203:
2194:
2171:
2155:
2135:
2124:. Retrieved
2120:
2111:
2095:
2086:
2049:sialadenitis
2043:
2042:
2027:septic shock
2022:
2014:
2013:
2009:
2003:
1994:
1981:
1975:
1967:
1963:
1957:
1951:
1942:
1936:
1930:
1924:
1918:
1910:
1906:
1897:
1891:
1878:
1872:
1863:
1856:
1850:
1844:
1838:
1834:Erythrocebus
1832:
1826:
1822:
1813:
1807:
1798:
1789:
1780:
1772:S. devriesei
1771:
1762:
1756:
1750:
1744:
1731:
1717:
1711:
1694:
1687:
1683:
1675:
1673:
1664:Gram-stained
1659:
1639:
1637:
1628:
1620:
1616:
1610:
1605:
1597:
1589:
1584:
1577:
1575:
1563:
1556:
1550:
1544:
1540:
1526:
1508:
1507:
1495:
1494:
1490:streptococci
1465:
1461:
1455:
1454:
1445:
1443:
1438:
1434:
1430:
1426:
1422:
1418:
1414:
1410:
1406:
1402:
1398:
1396:
1385:
1376:furazolidone
1367:
1363:
1359:
1353:
1348:
1346:
1333:
1329:
1328:
1308:
1306:
1296:
1292:
1288:
1286:
1281:
1277:
1276:
1271:
1267:
1266:
1261:
1257:
1253:
1249:
1247:
1243:S. vitulinus
1242:
1239:S. pulvereri
1238:
1237:
1230:
1226:
1224:
1211:
1209:
1204:
1200:
1192:
1188:
1186:
1176:
1172:
1168:
1166:
1161:
1157:
1153:
1149:
1145:
1143:
1138:
1134:
1130:
1126:
1122:
1118:
1114:
1110:
1106:
1102:
1100:
1097:
1087:
1077:
1075:
1070:
1064:
1058:
1056:
1051:
1045:
1041:
1039:
1034:
1030:
1026:
1021:
1017:
1013:S. vitulinus
1012:
1008:
1004:
1000:
996:
992:
987:
983:
979:
975:
971:
967:
963:
959:
955:
951:
947:
943:
938:
934:
929:
925:
921:
917:
913:
909:
905:
901:
897:
893:
889:
885:
881:
877:
872:
868:
865:S. devriesei
864:
860:
856:
851:
847:
843:
839:
835:
830:
826:
822:
818:
814:
810:
806:
801:
797:
792:
788:
784:
781:S. argenteus
780:
776:
767:
749:
731:
724:
655:
649:
643:species are
640:
627:. Under the
604:
603:
602:
591:
584:
577:
572:S. vitulinus
570:
563:
556:
549:
542:
535:
528:
521:
514:
507:
500:
493:
488:S. pulvereri
486:
479:
472:
465:
458:
451:
444:
437:
430:
423:
416:
409:
402:
395:
388:
381:
374:
369:S. jettensis
367:
360:
353:
346:
339:
332:
325:
318:
311:
304:
299:S. edaphicus
297:
292:S. devriesei
290:
283:
276:
269:
262:
255:
248:
241:
234:
227:
220:
213:
206:
199:
192:
185:
178:
173:S. argenteus
171:
149:
148:
73:
69:
49:
43:
3300:iNaturalist
3233:Wikispecies
2698:Aquaculture
1976:S. simulans
1925:S. pasteuri
1763:S. delphini
1732:S. borealis
1695:S. arlattae
1594:microarrays
1486:enterococci
1403:S. delphini
1394:formation.
1293:S. petrasii
1289:S. lyticans
1278:S. petrasii
1254:S. simulans
1232:Macrococcus
1169:S. delphini
1146:S. arlettae
1107:S. borealis
1047:Macrococcus
1031:S. pasteuri
1022:S. simulans
1018:S. simulans
984:S. succinus
948:S. arlettae
898:S. delphini
861:S. borealis
831:S. simulans
819:S. debuckii
811:S. carnosus
807:S. carnosus
565:S. succinus
544:S. simulans
453:S. petrasii
446:S. pasteuri
411:S. lyticans
285:S. delphini
278:S. debuckii
236:S. carnosus
208:S. borealis
180:S. arlettae
3397:Categories
3129:2011-01-04
3049:2022-04-23
2993:: 559–92.
2704:: 737156.
2209:2022-11-27
2126:2018-07-25
2104:References
1995:S. xylosus
1984:– humans,
1982:S. warneri
1907:S. microti
1852:Microcebus
1828:Cercocebus
1825:– humans,
1790:S. equorum
1783:– humans,
1745:S. capitis
1650:Host range
1576:The first
1392:blood clot
1372:bacitracin
1342:bile salts
1214:group are
1201:S. hominis
1197:novobiocin
1162:S. xylosus
1154:S. equorum
1139:S. warneri
1123:S. hominis
1111:S. capitis
1084:Antarctica
1035:S. warneri
1027:S. warneri
988:S. xylosus
968:S. kloosii
960:S. equorum
914:S. microti
882:S. agnetis
873:S. hominis
840:S. capitis
734:subspecies
629:microscope
625:Bacillales
586:S. xylosus
579:S. warneri
425:S. microti
376:S. kloosii
348:S. hominis
313:S. equorum
222:S. capitis
187:S. agnetis
129:Bacillales
3091:1553-7374
2718:0044-8486
2239:: 39–60.
2182: in
2146: in
2096:S. aureus
2087:S. aureus
2023:S. aureus
2006:infection
1964:S. simiae
1958:S. sciuri
1943:S. rostri
1909:– voles (
1898:S. lutrae
1881:– goats,
1879:S. lentus
1864:S. hyicus
1818:pheasants
1757:S. cohnii
1751:S. caprae
1712:S. aureus
1617:S. aureus
1606:S. aureus
1598:S. aureus
1590:S. aureus
1585:S. aureus
1578:S. aureus
1558:S. caprae
1515:commensal
1466:S. aureus
1462:S. aureus
1457:S. aureus
1450:gastritis
1435:S. aureus
1431:S. aureus
1427:coagulans
1415:S. lutrae
1407:S. hyicus
1399:S. aureus
1388:coagulase
1338:anaerobes
1320:cell wall
1258:S. aureus
1227:S. sciuri
1212:S. sciuri
1193:S. sciuri
1150:S. cohnii
1135:S. simiae
1103:S. aureus
1071:S. aureus
1005:S. sciuri
1001:S. lentus
993:S. sciuri
956:S. cohnii
926:S. rostri
918:S. muscae
910:S. lutrae
902:S. hyicus
844:S. caprae
793:S. simiae
785:S. aureus
777:S. aureus
689:New Latin
671:romanized
593:S. Westin
537:S. simiae
530:S. sciuri
495:S. rostri
432:S. muscae
404:S. lutrae
390:S. lentus
355:S. hyicus
257:S. cohnii
229:S. caprae
194:S. aureus
156:Rosenbach
109:Bacillota
76:species.
70:S. aureus
3218:Wikidata
3109:24550724
2968:21622749
2885:25278577
2836:14766306
2797:31195568
2664:18174295
2611:15545460
2550:29079617
2501:25269845
2467:sp. nov"
2387:10319495
2317:23536759
2277:PLOS ONE
2255:14562246
2142:stafulh/
2063:See also
2033:Clinical
1997:– humans
1990:Pongidae
1978:– humans
1954:– humans
1933:– humans
1875:– humans
1799:S. felis
1767:dolphins
1747:– humans
1728:, humans
1699:chickens
1503:zoonosis
1470:catalase
1264:groups.
1029:group –
1020:group –
995:group –
952:S. caeli
946:group –
937:group –
894:S. felis
880:group –
859:group –
838:group –
809:group –
800:group –
779:group –
764:Taxonomy
677:staphylē
617:bacteria
320:S. felis
215:S. caeli
164:Species
135:Family:
105:Phylum:
99:Bacteria
95:Domain:
3318:1032605
3292:3227654
3224:Q199695
3100:3923759
3007:7002032
2959:3147504
2876:4187637
2655:2268370
2541:5752872
2492:4298100
2444:9734040
2308:3594240
2285:Bibcode
2184:Liddell
2178:ko)kkos
2154:(1940)
2053:leucine
1921:– goats
1883:rabbits
1873:S. leei
1810:– goats
1581:genomes
1425:subsp.
1274:group.
1216:oxidase
1203:subsp.
1090:group.
972:S. leei
746:animals
720:
700:
685:
673::
666:σταφυλή
383:S. leei
145:Genus:
125:Order:
119:Bacilli
115:Class:
3380:416009
3364:NZOR:
3305:337774
3279:1STAPG
3186:PATRIC
3107:
3097:
3089:
3005:
2966:
2956:
2911:
2883:
2873:
2834:
2795:
2739:
2716:
2662:
2652:
2609:
2548:
2538:
2499:
2489:
2442:
2385:
2338:
2315:
2305:
2253:
2099:(VRSA)
2090:(MRSA)
2019:sepsis
1939:– dogs
1902:otters
1794:horses
1776:cattle
1740:cattle
1736:humans
1602:toxins
1555:, and
1421:, and
1316:coccus
1295:, and
1094:Groups
712:kókkos
704:κόκκος
693:coccus
3375:WoRMS
3313:IRMNG
3266:83232
3190:NIAID
2188:Scott
2055:into
1887:sheep
1846:Macca
1840:Lemur
1703:goats
1670:apart
1183:Notes
1137:and
637:grape
633:cocci
610:genus
608:is a
39:Staff
3357:1279
3352:NCBI
3339:LPSN
3326:ITIS
3287:GBIF
3274:EPPO
3253:7LY9
3105:PMID
3087:ISSN
3003:PMID
2964:PMID
2909:ISBN
2881:PMID
2832:PMID
2793:PMID
2737:ISBN
2714:ISSN
2660:PMID
2607:PMID
2546:PMID
2497:PMID
2440:PMID
2383:PMID
2336:ISBN
2313:PMID
2251:PMID
2186:and
1947:pigs
1868:pigs
1803:cats
1726:dogs
1722:deer
1488:and
1260:and
1252:and
1225:The
1191:and
1187:The
1175:and
1160:and
1063:and
770:rRNA
740:and
738:skin
718:lit.
698:lit.
683:lit.
158:1884
3331:359
3261:EoL
3248:CoL
3095:PMC
3077:doi
3043:CDC
2995:doi
2985:".
2954:PMC
2946:doi
2942:193
2871:PMC
2863:doi
2824:doi
2783:doi
2706:doi
2702:545
2650:PMC
2642:doi
2597:doi
2536:PMC
2528:doi
2487:PMC
2479:doi
2430:doi
2373:doi
2303:PMC
2293:doi
2274:L."
2241:doi
1858:Pan
1468:is
612:of
68:of
3399::
3377::
3354::
3341::
3328::
3315::
3302::
3289::
3276::
3263::
3250::
3235::
3220::
3103:.
3093:.
3085:.
3073:10
3071:.
3067:.
3041:.
3023:.
3001:.
2991:34
2989:.
2962:.
2952:.
2940:.
2936:.
2879:.
2869:.
2859:27
2857:.
2853:.
2830:.
2820:34
2818:.
2791:.
2779:60
2777:.
2773:.
2712:.
2700:.
2696:.
2672:^
2658:.
2648:.
2638:46
2636:.
2632:.
2605:.
2593:54
2591:.
2587:.
2544:.
2534:.
2524:84
2522:.
2518:.
2495:.
2485:.
2475:65
2473:.
2469:.
2438:.
2426:48
2424:.
2420:.
2381:.
2369:49
2367:.
2363:.
2311:.
2301:.
2291:.
2279:.
2249:.
2235:.
2231:.
2202:.
2150:;
2119:.
2029:.
1988:,
1945:–
1900:–
1885:,
1866:–
1855:,
1849:,
1843:,
1837:,
1831:,
1801:–
1792:–
1774:–
1765:–
1738:,
1734:–
1724:,
1720:–
1705:,
1701:,
1697:–
1668:μm
1662:,
1646:.
1635:.
1561:.
1549:,
1525:.
1505:.
1492:.
1476:(H
1452:.
1417:,
1413:,
1409:,
1405:,
1401:,
1358:,
1344:.
1245:.
1235:.
1207:.
1179:.
1171:,
1164:.
1156:,
1152:,
1148:,
1141:.
1133:,
1129:,
1125:,
1121:,
1117:,
1113:,
1109:,
1105:,
1073:.
1054:.
1033:,
1011:,
1007:,
1003:,
999:,
986:,
982:,
978:,
974:,
970:,
966:,
962:,
958:,
954:,
950:,
928:,
924:,
920:,
916:,
912:,
908:,
904:,
900:,
896:,
892:,
888:,
884:,
871:,
867:,
863:,
850:,
846:,
842:,
829:,
825:,
821:,
817:,
813:,
791:,
787:,
783:,
760:.
748:.
715:,
706:,
695:,
691::
680:,
668:,
664::
3132:.
3111:.
3079::
3052:.
3009:.
2997::
2970:.
2948::
2917:.
2887:.
2865::
2838:.
2826::
2799:.
2785::
2745:.
2720:.
2708::
2666:.
2644::
2613:.
2599::
2552:.
2530::
2503:.
2481::
2446:.
2432::
2389:.
2375::
2344:.
2319:.
2295::
2287::
2281:8
2257:.
2243::
2237:4
2212:.
2129:.
1972:)
1915:)
1482:2
1480:O
1478:2
1433:(
1291:,
41:.
34:.
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.