259:, respectively. Many transamination reactions occur in tissues, catalysed by transaminases specific for a particular amino/keto acid pair. The reactions are readily reversible, the direction being determined by which of the reactants are in excess. This reversibility can be exploited for synthetic chemistry applications to achieve the synthesis of valuable chiral amines. The specific enzymes are named from one of the reactant pairs, for example; the reaction between glutamic acid and pyruvic acid to make alpha ketoglutaric acid and alanine is called
29:
1210:
279:
270:
with various amino/keto acid pairs. Transamination is demonstrated if the corresponding new amino acid and keto acid are formed, as revealed by paper chromatography. Reversibility is demonstrated by using the complementary keto/amino acid pair as starting reactants. After chromatogram has been taken
389:(ALT), also called alanine aminotransferase (ALAT) or serum glutamate-pyruvate transaminase (SGPT). These transaminases were discovered in 1954 and their clinical importance was described in 1955.
766:
129:
846:
709:
Ghany M, Hoofnagle JH (2005). "Approach to the
Patient With Liver Disease". In Kasper DL, Fauci AS, Longo DL, Braunwald E, Hauser SL, Jameson JL (eds.).
220:
group on one molecule is exchanged with the =O group on the other molecule. The amino acid becomes a keto acid, and the keto acid becomes an amino acid.
563:
Ladue JS, Wroblewski F, Karmen A (September 1954). "Serum glutamic oxaloacetic transaminase activity in human acute transmural myocardial infarction".
836:
759:
679:
851:
896:
929:
752:
630:
826:
402:
1085:
149:
1200:
841:
1070:
1186:
1173:
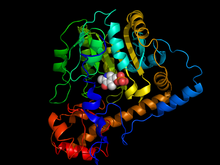
1160:
1147:
1134:
1121:
1108:
887:
868:
831:
792:
1080:
1034:
977:
783:
738:
137:
982:
654:
804:
382:
378:
33:
1230:
1003:
922:
1075:
133:
821:
572:
386:
358:
260:
1240:
1039:
369:
The transaminase enzymes are important in the production of various amino acids, and measuring the
320:
244:
224:
85:
972:
877:
1235:
588:
545:
496:
457:
124:
1018:
1013:
987:
915:
580:
535:
527:
516:"A note on the spectrometric assay of glutamic-oxalacetic transaminase in human blood serum"
488:
447:
439:
744:
607:"Biblioteca Nazionale di Napoli. News: Serata in onore di Mario Coltorti e Giuseppe Giusti"
335:
for conversion to urea for excretion of nitrogen. In similar manner, in muscles the use of
231:
in the first half-reaction, when an amino acid is converted into a keto acid. Enzyme-bound
116:
1065:
1049:
962:
350:
287:
734:
349:). Here other transaminases regenerate pyruvate, which provides a valuable precursor for
576:
381:
can be an indicator of liver and cardiac damage. Two important transaminase enzymes are
1214:
1103:
1044:
373:
of various transaminases in the blood is important in the diagnosing and tracking many
213:
177:
28:
540:
515:
452:
427:
343:, which is carried by the bloodstream to the liver (the overall reaction being termed
1224:
1008:
967:
398:
370:
345:
256:
252:
66:
957:
680:"Il Resto Del Carlino - Macerata - E' morto Mario Coltorti: scoprì la transaminasi"
316:
240:
232:
228:
112:
606:
90:
584:
78:
1181:
1116:
952:
775:
308:
267:
1209:
354:
324:
283:
278:
189:
181:
500:
477:"Application of Paper Chromatography to the Study of the Transaminase System"
1155:
1129:
332:
328:
272:
185:
592:
549:
476:
461:
263:
and was originally called glutamic-pyruvic transaminase or GPT for short.
779:
722:(3rd ed.). New York: Worth Publishers. pp. 628–31, 634, 828–30.
385:(AST), also known as serum glutamic oxaloacetic transaminase (SGOT); and
336:
304:
236:
173:
73:
374:
340:
248:
38:
531:
443:
1168:
938:
492:
209:
169:
144:
266:
Tissue transaminase activities can be investigated by incubating a
1142:
312:
277:
201:
856:
814:
809:
106:
61:
911:
748:
907:
271:
out of the solvent the chromatogram is then treated with
713:(16th ed.). New York: McGraw-Hill. pp. 1814–5.
307:
to amino acids, at the expense of muscle tissue, when
1198:
631:"E' morto il prof. Coltorti: scoprì le transaminasi"
1094:
1058:
1027:
996:
945:
886:
867:
791:
143:
123:
105:
100:
84:
72:
60:
52:
47:
21:
426:Karmen A, Wroblewski F, Ladue JS (January 1955).
475:Tulpule, P. G.; Patwardhan, V. N. (April 1952).
421:
419:
294:) group and the keto (=O) group are exchanged.
923:
760:
8:
323:plays a key role in funneling nitrogen from
930:
916:
908:
847:Branched-chain amino acid aminotransferase
767:
753:
745:
711:Harrison's Principles of Internal Medicine
97:
27:
737:at the U.S. National Library of Medicine
635:notizie-segreteria-liver-pool.blogspot.it
539:
451:
353:. This alanine cycle is analogous to the
188:. They are important in the synthesis of
1205:
415:
428:"Transaminase activity in human blood"
18:
520:The Journal of Clinical Investigation
432:The Journal of Clinical Investigation
7:
837:Kynurenine—oxoglutarate transaminase
720:Lehninger Principles of Biochemistry
42:with Pyridoxal 5' Phosphate cofactor
223:Transaminases require the coenzyme
282:Aminotransfer reaction between an
14:
1208:
897:Pyridoxine 5'-phosphate synthase
299:Amino acid metabolism in animals
208:) group. A keto acid contains a
852:Alanine—glyoxylate transaminase
377:. For example, the presence of
1:
101:Available protein structures:
678:MonrifNet (3 January 2009).
585:10.1126/science.120.3117.497
1257:
842:Ornithine aminotransferase
718:Nelson DL, Cox MM (2000).
311:is low. The preference of
227:, which is converted into
200:An amino acid contains an
1086:Michaelis–Menten kinetics
832:Tyrosine aminotransferase
514:Karmen A (January 1955).
339:for transamination gives
96:
26:
978:Diffusion-limited enzyme
739:Medical Subject Headings
684:www.ilrestodelcarlino.it
659:www.istitutobioetica.org
611:vecchiosito.bnnonline.it
303:Animals must metabolize
192:, which form proteins.
805:Aspartate transaminase
655:"Campania su Coltorti"
383:aspartate transaminase
379:elevated transaminases
295:
196:Function and mechanism
34:Aspartate transaminase
1071:Eadie–Hofstee diagram
1004:Allosteric regulation
871:: Oximinotransferases
346:glucose-alanine cycle
281:
275:to locate the spots..
1081:Lineweaver–Burk plot
822:Alanine transaminase
387:alanine transaminase
359:anaerobic metabolism
261:alanine transaminase
235:in turn reacts with
180:reaction between an
577:1954Sci...120..497L
321:alpha-ketoglutarate
245:alpha-ketoglutarate
225:pyridoxal phosphate
1040:Enzyme superfamily
973:Enzyme promiscuity
878:Oximinotransferase
315:transaminases for
296:
1196:
1195:
905:
904:
827:GABA transaminase
532:10.1172/JCI103055
487:(4303): 671–671.
444:10.1172/jci103055
403:GABA transaminase
166:aminotransferases
159:
158:
155:
154:
150:structure summary
1248:
1213:
1212:
1204:
1076:Hanes–Woolf plot
1019:Enzyme activator
1014:Enzyme inhibitor
988:Enzyme catalysis
932:
925:
918:
909:
769:
762:
755:
746:
723:
714:
695:
694:
692:
691:
675:
669:
668:
666:
665:
651:
645:
644:
642:
641:
627:
621:
620:
618:
617:
603:
597:
596:
560:
554:
553:
543:
511:
505:
504:
493:10.1038/169671a0
472:
466:
465:
455:
423:
98:
56:Aminotransferase
31:
22:Aminotransferase
19:
16:Class of enzymes
1256:
1255:
1251:
1250:
1249:
1247:
1246:
1245:
1221:
1220:
1219:
1207:
1199:
1197:
1192:
1104:Oxidoreductases
1090:
1066:Enzyme kinetics
1054:
1050:List of enzymes
1023:
992:
963:Catalytic triad
941:
936:
906:
901:
882:
863:
787:
773:
731:
726:
717:
708:
704:
702:Further reading
699:
698:
689:
687:
677:
676:
672:
663:
661:
653:
652:
648:
639:
637:
629:
628:
624:
615:
613:
605:
604:
600:
571:(3117): 497–9.
562:
561:
557:
513:
512:
508:
474:
473:
469:
425:
424:
417:
412:
395:
367:
365:Diagnostic uses
357:, which allows
351:gluconeogenesis
301:
293:
290:. The amino (NH
288:alpha-keto acid
219:
212:(=O) group. In
207:
198:
43:
17:
12:
11:
5:
1254:
1252:
1244:
1243:
1238:
1233:
1223:
1222:
1218:
1217:
1194:
1193:
1191:
1190:
1177:
1164:
1151:
1138:
1125:
1112:
1098:
1096:
1092:
1091:
1089:
1088:
1083:
1078:
1073:
1068:
1062:
1060:
1056:
1055:
1053:
1052:
1047:
1042:
1037:
1031:
1029:
1028:Classification
1025:
1024:
1022:
1021:
1016:
1011:
1006:
1000:
998:
994:
993:
991:
990:
985:
980:
975:
970:
965:
960:
955:
949:
947:
943:
942:
937:
935:
934:
927:
920:
912:
903:
902:
900:
899:
893:
891:
884:
883:
881:
880:
874:
872:
865:
864:
862:
861:
860:
859:
849:
844:
839:
834:
829:
824:
819:
818:
817:
812:
801:
799:
789:
788:
774:
772:
771:
764:
757:
749:
743:
742:
730:
729:External links
727:
725:
724:
715:
705:
703:
700:
697:
696:
670:
646:
622:
598:
555:
506:
467:
414:
413:
411:
408:
407:
406:
394:
391:
371:concentrations
366:
363:
327:metabolism to
300:
297:
291:
217:
214:transamination
205:
197:
194:
178:transamination
157:
156:
153:
152:
147:
141:
140:
127:
121:
120:
110:
103:
102:
94:
93:
88:
82:
81:
76:
70:
69:
64:
58:
57:
54:
50:
49:
45:
44:
32:
24:
23:
15:
13:
10:
9:
6:
4:
3:
2:
1253:
1242:
1239:
1237:
1234:
1232:
1229:
1228:
1226:
1216:
1211:
1206:
1202:
1188:
1184:
1183:
1178:
1175:
1171:
1170:
1165:
1162:
1158:
1157:
1152:
1149:
1145:
1144:
1139:
1136:
1132:
1131:
1126:
1123:
1119:
1118:
1113:
1110:
1106:
1105:
1100:
1099:
1097:
1093:
1087:
1084:
1082:
1079:
1077:
1074:
1072:
1069:
1067:
1064:
1063:
1061:
1057:
1051:
1048:
1046:
1045:Enzyme family
1043:
1041:
1038:
1036:
1033:
1032:
1030:
1026:
1020:
1017:
1015:
1012:
1010:
1009:Cooperativity
1007:
1005:
1002:
1001:
999:
995:
989:
986:
984:
981:
979:
976:
974:
971:
969:
968:Oxyanion hole
966:
964:
961:
959:
956:
954:
951:
950:
948:
944:
940:
933:
928:
926:
921:
919:
914:
913:
910:
898:
895:
894:
892:
889:
885:
879:
876:
875:
873:
870:
866:
858:
855:
854:
853:
850:
848:
845:
843:
840:
838:
835:
833:
830:
828:
825:
823:
820:
816:
813:
811:
808:
807:
806:
803:
802:
800:
798:
797:Transaminases
794:
790:
785:
781:
777:
770:
765:
763:
758:
756:
751:
750:
747:
740:
736:
735:Transaminases
733:
732:
728:
721:
716:
712:
707:
706:
701:
685:
681:
674:
671:
660:
656:
650:
647:
636:
632:
626:
623:
612:
608:
602:
599:
594:
590:
586:
582:
578:
574:
570:
566:
559:
556:
551:
547:
542:
537:
533:
529:
525:
521:
517:
510:
507:
502:
498:
494:
490:
486:
482:
478:
471:
468:
463:
459:
454:
449:
445:
441:
438:(1): 126–31.
437:
433:
429:
422:
420:
416:
409:
404:
400:
399:Valproic acid
397:
396:
392:
390:
388:
384:
380:
376:
372:
364:
362:
360:
356:
352:
348:
347:
342:
338:
334:
330:
326:
322:
318:
314:
310:
306:
298:
289:
285:
280:
276:
274:
269:
264:
262:
258:
257:glutamic acid
254:
253:aspartic acid
250:
246:
242:
238:
234:
230:
226:
221:
215:
211:
203:
195:
193:
191:
187:
183:
179:
175:
171:
167:
163:
162:Transaminases
151:
148:
146:
142:
139:
135:
131:
128:
126:
122:
118:
114:
111:
108:
104:
99:
95:
92:
89:
87:
83:
80:
77:
75:
71:
68:
65:
63:
59:
55:
51:
46:
41:
40:
35:
30:
25:
20:
1231:Transferases
1182:Translocases
1179:
1166:
1153:
1140:
1127:
1117:Transferases
1114:
1101:
958:Binding site
796:
719:
710:
688:. Retrieved
686:(in Italian)
683:
673:
662:. Retrieved
658:
649:
638:. Retrieved
634:
625:
614:. Retrieved
610:
601:
568:
564:
558:
526:(1): 131–3.
523:
519:
509:
484:
480:
470:
435:
431:
368:
361:by muscles.
344:
317:oxaloacetate
302:
265:
241:oxaloacetate
233:pyridoxamine
229:pyridoxamine
222:
199:
165:
161:
160:
37:
953:Active site
780:nitrogenous
776:Transferase
309:blood sugar
190:amino acids
48:Identifiers
1241:Hepatology
1225:Categories
1156:Isomerases
1130:Hydrolases
997:Regulation
690:2017-09-10
664:2017-09-10
640:2017-09-10
616:2017-09-10
410:References
355:Cori cycle
325:amino acid
284:amino acid
268:homogenate
182:amino acid
113:structures
86:Membranome
1035:EC number
501:1476-4687
405:inhibitor
333:glutamate
329:aspartate
273:ninhydrin
247:, giving
186:keto acid
184:and an α-
79:IPR004839
1236:EC 2.6.1
1059:Kinetics
983:Cofactor
946:Activity
782:groups (
593:13195683
550:13221664
462:13221663
393:See also
375:diseases
337:pyruvate
305:proteins
237:pyruvate
216:, the NH
174:catalyze
130:RCSB PDB
74:InterPro
1215:Biology
1169:Ligases
939:Enzymes
890:: Other
573:Bibcode
565:Science
341:alanine
286:and an
249:alanine
170:enzymes
67:PF00155
39:E. coli
1201:Portal
1143:Lyases
888:2.6.99
741:(MeSH)
591:
548:
541:438594
538:
499:
481:Nature
460:
453:438594
450:
145:PDBsum
119:
109:
53:Symbol
1095:Types
869:2.6.3
793:2.6.1
313:liver
255:, or
243:, or
202:amino
172:that
36:from
1187:list
1180:EC7
1174:list
1167:EC6
1161:list
1154:EC5
1148:list
1141:EC4
1135:list
1128:EC3
1122:list
1115:EC2
1109:list
1102:EC1
857:AGXT
815:GOT2
810:GOT1
786:2.6)
589:PMID
546:PMID
497:ISSN
458:PMID
401:- a
331:and
210:keto
168:are
138:PDBj
134:PDBe
117:ECOD
107:Pfam
62:Pfam
581:doi
569:120
536:PMC
528:doi
489:doi
485:169
448:PMC
440:doi
319:or
204:(NH
164:or
125:PDB
91:273
1227::
795::
784:EC
778::
682:.
657:.
633:.
609:.
587:.
579:.
567:.
544:.
534:.
524:34
522:.
518:.
495:.
483:.
479:.
456:.
446:.
436:34
434:.
430:.
418:^
251:,
239:,
176:a
136:;
132:;
115:/
1203::
1189:)
1185:(
1176:)
1172:(
1163:)
1159:(
1150:)
1146:(
1137:)
1133:(
1124:)
1120:(
1111:)
1107:(
931:e
924:t
917:v
768:e
761:t
754:v
693:.
667:.
643:.
619:.
595:.
583::
575::
552:.
530::
503:.
491::
464:.
442::
292:2
218:2
206:2
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.