563:). Suitable enzymes can be isolated from natural sources or produced recombinantly. As an alternative, whole cell-based systems using either endogenous glycosyl donors or cell-based systems containing cloned and expressed systems for synthesis of glycosyl donors have been developed. In cell-free approaches, the large-scale application of glycosyltransferases for glycoconjugate synthesis has required access to large quantities of the glycosyl donors. On the flip-side, nucleotide recycling systems that allow the resynthesis of glycosyl donors from the released nucleotide have been developed. The nucleotide recycling approach has a further benefit of reducing the amount of nucleotide formed as a by-product, thereby reducing the amount of inhibition caused to the glycosyltransferase of interest – a commonly observed feature of the nucleotide byproduct.
266:
31:
2256:
272:
Glycosyltransferases can be segregated into "retaining" or "inverting" enzymes according to whether the stereochemistry of the donor's anomeric bond is retained (α→α) or inverted (α→β) during the transfer. The inverting mechanism is straightforward, requiring a single nucleophilic attack from the
294:
Sequence-based classification methods have proven to be a powerful way of generating hypotheses for protein function based on sequence alignment to related proteins. The carbohydrate-active enzyme database presents a sequence-based classification of glycosyltransferases into over 90 families. The
530:
to D-galactose end of H antigen, producing the A antigen. The B allele encodes 1-3-galactosyltransferase that joins α-D-galactose bonded to D-galactose end of H antigen, creating the B antigen. In case of O allele the exon 6 contains a deletion that results in a loss of enzymatic activity. The O
276:
The retaining mechanism has been a matter of debate, but there exists strong evidence against a double displacement mechanism (which would cause two inversions about the anomeric carbon for a net retention of stereochemistry) or a dissociative mechanism (a prevalent variant of which was known as
1190:
Kocev, A; Melamed, J; Wang, S; Kong, X; Vlahakis, JZ; Xu, Y; Szarek, WA; Brockhausen, I (June 2020). "Inhibition of bacterial growth and galactosyltransferase activity of WbwC by α, ω-bis(3-alkyl-1H-imidazolium)alkane salts: Effect of varying carbon content".
277:
SNi). An "orthogonal associative" mechanism has been proposed which, akin to the inverting enzymes, requires only a single nucleophilic attack from an acceptor from a non-linear angle (as observed in many crystal structures) to achieve anomer retention.
311:
database, only three different folds have been observed for glycosyltransferases Very recently, a new glycosyltransferase fold was identified for the glycosyltransferases involved in the biosynthesis of the NAG-NAM polymer backbone of
285:
The recent discovery of the reversibility of many reactions catalyzed by inverting glycosyltransferases served as a paradigm shift in the field and raises questions regarding the designation of sugar nucleotides as 'activated' donors.
550:
Glycosyltransferases have been widely used in both the targeted synthesis of specific glycoconjugates as well as the synthesis of differentially glycosylated libraries of drugs, biological probes or natural products in the context of
818:
Zhang, C; Albermann, C; Fu, X; Thorson, JS (27 December 2006). "The in vitro characterization of the iterative avermectin glycosyltransferase AveBI reveals reaction reversibility and sugar nucleotide flexibility".
510:
1401:
1396:
854:
Zhang, C; Fu, Q; Albermann, C; Li, L; Thorson, JS (5 March 2007). "The in vitro characterization of the erythronolide mycarosyltransferase EryBV and its utility in macrolide diversification".
767:
Zhang, C; Griffith, BR; Fu, Q; Albermann, C; Fu, X; Lee, IK; Li, L; Thorson, JS (1 September 2006). "Exploiting the reversibility of natural product glycosyltransferase-catalyzed reactions".
1774:
1650:
1441:
477:
539:
and results in translation of an almost entirely different protein that lacks enzymatic activity. This results in H antigen remaining unchanged in case of O groups.
1755:
1823:
1272:
421:
409:
1932:
1902:
1140:
Lovering AL, de Castro LH, Lim D, Strynadka NC (March 2007). "Structural insight into the transglycosylation step of bacterial cell-wall biosynthesis".
526:
expressing the glycosyltransferases has three main allelic forms: A, B, and O. The A allele encodes 1-3-N-acetylgalactosaminyltransferase that bonds α-
1577:
1572:
247:. The phosphate(s) of these donor molecules are usually coordinated by divalent cations such as manganese, however metal independent enzymes exist.
1220:
Gantt, RW; Peltier-Pain, P; Thorson, JS (October 2011). "Enzymatic methods for glyco(diversification/randomization) of drugs and small molecules".
1937:
2306:
676:
308:
542:
The combination of glycosyltransferases by both alleles present in each person determines whether there is an AB, A, B or O blood type.
1265:
1975:
1818:
643:
251:
340:. Some glycosyltransferase inhibitors are of use as drugs or antibiotics. Moenomycin is used in animal feed as a growth promoter.
1828:
2281:
1849:
1833:
2131:
497:
1258:
2246:
1368:
356:-based synthetic inhibitors of glycosyltransferases have been designed for use as antimicrobial and antiseptic agents.
1353:
1793:
2116:
2232:
2219:
2206:
2193:
2180:
2167:
2154:
1918:
1810:
1723:
1358:
1298:
1006:
607:
2126:
2080:
2023:
1363:
1336:
1289:
190:
54:
485:
337:
2028:
1853:
1416:
582:
159:
2276:
1331:
577:
572:
516:
175:
2311:
2286:
2049:
1968:
1897:
1383:
1373:
527:
2121:
626:
Williams, GJ; Thorson, JS (2009). "Natural
Product Glycosyltransferases: Properties and Applications".
481:
1765:
1592:
1326:
1149:
776:
721:
244:
193:
for his work on carbohydrate metabolism. Glycosyltransferases that use non-nucleotide donors such as
348:
is an inhibitor of mycobacterial arabinotransferases and is used for the treatment of tuberculosis.
2085:
1727:
1642:
1540:
1313:
695:
433:
304:
232:
123:
1082:"Glycosyltransferase structural biology and its role in the design of catalysts for glycosylation"
630:. Advances in Enzymology - and Related Areas of Molecular Biology. Vol. 76. pp. 55–119.
265:
2018:
1880:
1564:
1302:
1173:
879:
800:
560:
69:
240:
228:
224:
324:
Many inhibitors of glycosyltransferases are known. Some of these are natural products, such as
2301:
2296:
1922:
1348:
1250:
1237:
1165:
1111:
1062:
977:
928:
871:
836:
792:
749:
672:
649:
639:
592:
587:
472:
84:
1200:
710:"Geometric Attributes of Retaining Glycosyltransferase Enzymes Favor an Orthogonal Mechanism"
2064:
2059:
2033:
1961:
1760:
1747:
1391:
1341:
1229:
1157:
1101:
1093:
1052:
1044:
967:
959:
918:
910:
863:
828:
784:
739:
729:
631:
556:
523:
73:
1632:
1627:
464:
30:
2291:
2111:
2095:
2008:
1779:
994:
255:
115:
58:
307:, glycosyltransferase have a much smaller range of structures. In fact, according to the
1153:
948:"Using simple donors to drive the equilibria of glycosyltransferase-catalyzed reactions"
780:
725:
2260:
2149:
2090:
1106:
1081:
1057:
1032:
972:
947:
923:
898:
744:
709:
597:
552:
414:
186:
119:
77:
2270:
2054:
2013:
1784:
1321:
602:
333:
313:
216:
201:
1177:
883:
804:
390:
352:
is an inhibitor of insect chitin syntheses and is used to control fleas in animals.
189:, the scientist who discovered the first sugar nucleotide and who received the 1970
2003:
460:
147:
107:
81:
426:
1097:
734:
402:
2227:
2162:
1998:
1281:
899:"The in vitro characterization of polyene glycosyltransferases AmphDI and NysDI"
344:
has been developed from the echinocandins and is in use as an antifungal agent.
341:
236:
212:
167:
17:
2255:
531:
allele differs slightly from the A allele by deletion of a single nucleotide -
635:
536:
519:
is determined by what type of glycosyltransferases are expressed in the body.
345:
329:
325:
295:
same three-dimensional fold is expected to occur within each of the families.
220:
198:
155:
151:
66:
1033:"The structural biology of enzymes involved in natural product glycosylation"
438:
2201:
2175:
1207:
1161:
788:
691:
353:
349:
143:
111:
1241:
1169:
1115:
1066:
981:
932:
914:
875:
867:
840:
796:
753:
667:
Etzler ME, Varki A, Cummings RL, Esko JD, Freeze HH, Hart GW, eds. (2008).
653:
963:
946:
Gantt, RW; Peltier-Pain, P; Cournoyer, WJ; Thorson, JS (21 August 2011).
397:
194:
171:
135:
96:
62:
162:, which is relatively abundant in eukaryotes. Transferases may also use
1942:
1800:
1533:
1528:
1523:
1518:
1513:
1481:
1405:
1233:
1048:
532:
131:
1010:
832:
154:
to give N-linked glycoproteins. Mannosyl groups may be transferred to
2214:
1984:
1738:
1508:
1503:
1498:
1493:
1488:
1476:
1471:
1466:
1461:
1456:
1451:
1446:
1434:
1429:
1424:
671:(2nd ed.). Plainview, N.Y: Cold Spring Harbor Laboratory Press.
492:
211:
Mammals use only 9 sugar nucleotide donors for glycosyltransferases:
163:
139:
100:
92:
88:
50:
34:
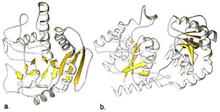
Most glycosyltransferase enzymes form one of two folds: GT-A or GT-B
511:
Glycoprotein-fucosylgalactoside a-N-acetylgalactosaminyltransferase
2188:
1890:
1885:
1858:
1709:
1660:
1655:
1080:
Chang, A; Singh, S; Phillips GN, Jr; Thorson, JS (December 2011).
264:
127:
29:
1704:
1699:
1694:
1689:
1684:
1679:
1672:
1667:
1622:
1617:
1612:
1607:
1602:
1597:
1587:
1582:
1554:
1549:
1544:
1204:
454:
385:
1957:
1402:
Glycoprotein-N-acetylgalactosamine 3-beta-galactosyltransferase
1397:
B-N-acetylglucosaminyl-glycopeptide b-1,4-galactosyltransferase
1254:
897:
Zhang, C; Moretti, R; Jiang, J; Thorson, JS (13 October 2008).
1128:
170:, and even use lipid-linked sugar phosphate donors, such as
303:
In contrast to the diversity of 3D structures observed for
1953:
328:, an inhibitor of peptidoglycan glycosyltransferases, the
181:
Glycosyltransferases that use sugar nucleotide donors are
1031:
Singh, S; Phillips GN, Jr; Thorson, JS (October 2012).
1651:
Dolichyl-phosphate-mannose-protein mannosyltransferase
2244:
2140:
2104:
2073:
2042:
1991:
1917:
1873:
1842:
1809:
1746:
1737:
1722:
1641:
1563:
1415:
1382:
1312:
1297:
491:
471:
453:
448:
432:
420:
408:
396:
384:
376:
371:
366:
122:. Some glycosyltransferases catalyse transfer to
254:, and they are usually anchored to membranes of
1824:Hypoxanthine-guanine phosphoribosyltransferase
1969:
1266:
708:Schuman B, Evans SV, Fyles TM (August 2013).
8:
1933:Beta-galactoside alpha-2,6-sialyltransferase
1903:Indolylacetylinositol arabinosyltransferase
1129:SCOP: Structural Classification of Proteins
1976:
1962:
1954:
1743:
1734:
1309:
1273:
1259:
1251:
445:
273:accepting atom to invert stereochemistry.
87:molecule, the nucleophile of which can be
1105:
1056:
971:
922:
743:
733:
332:, inhibitors of chitin synthase, and the
106:The result of glycosyl transfer can be a
821:Journal of the American Chemical Society
2251:
618:
535:at position 261. The deletion causes a
1938:Monosialoganglioside sialyltransferase
1775:NAD(P):arginine ADP-ribosyltransferase
1756:NAD:diphthamide ADP-ribosyltransferase
363:
174:phosphates in eukaryotic organism, or
130:. Glycosyl transfer can also occur to
309:Structural Classification of Proteins
7:
1193:Bioorganic and Medicinal Chemistry
252:single-pass transmembrane proteins
25:
1819:Adenine phosphoribosyltransferase
2254:
1829:Uracil phosphoribosyltransferase
1086:Current Opinion in Biotechnology
1850:Purine nucleoside phosphorylase
206:non-Leloir glycosyltransferases
1834:Amidophosphoribosyltransferase
995:CAZypedia Glycosyltransferases
250:Many glycosyltransferases are
1:
1201:doi:10.1016/j.bmc.2020.115494
449:Available protein structures:
2307:Peripheral membrane proteins
1369:Ceramide glucosyltransferase
1098:10.1016/j.copbio.2011.04.013
735:10.1371/journal.pone.0071077
367:Glycosyltransferase family 6
1007:"CAZy Glycosyl Transferase"
2328:
1794:Poly ADP ribose polymerase
669:Essentials of Glycobiology
508:
290:Classification by sequence
2132:Michaelis–Menten kinetics
1811:Phosphoribosyltransferase
636:10.1002/9780470392881.ch2
608:Oligosaccharyltransferase
444:
360:Determinant of blood type
57:) that establish natural
2024:Diffusion-limited enzyme
1364:1,3-Beta-glucan synthase
191:Nobel Prize in Chemistry
166:as an acceptor, forming
1854:Thymidine phosphorylase
1222:Natural Product Reports
1162:10.1126/science.1136611
1037:Natural Product Reports
952:Nature Chemical Biology
789:10.1126/science.1130028
583:Glucuronosyltransferase
336:, inhibitors of fungal
2282:Carbohydrate chemistry
1748:ADP-ribosyltransferase
915:10.1002/cbic.200800349
868:10.1002/cbic.200600509
628:Advances in Enzymology
578:Chemical glycosylation
573:Carbohydrate chemistry
517:ABO blood group system
338:β-1,3-glucan synthases
281:Reaction reversibility
269:
176:undecaprenyl phosphate
35:
2117:Eadie–Hofstee diagram
2050:Allosteric regulation
1898:Arabinosyltransferase
1374:N-glycosyltransferase
528:N-acetylgalactosamine
268:
160:C-mannosyl tryptophan
134:residues, usually to
33:
2127:Lineweaver–Burk plot
1766:Pseudomonas exotoxin
1286:glycosyltransferases
964:10.1038/nchembio.638
559:(a process known as
305:glycoside hydrolases
76:(also known as the "
39:Glycosyltransferases
1541:Hyaluronan synthase
1154:2007Sci...315.1402L
781:2006Sci...313.1291Z
726:2013PLoSO...871077S
696:Membranome database
233:UDP-glucuronic acid
124:inorganic phosphate
59:glycosidic linkages
2086:Enzyme superfamily
2019:Enzyme promiscuity
1881:Xylosyltransferase
1354:Debranching enzyme
1234:10.1039/c1np00045d
1049:10.1039/c2np20039b
561:glycorandomization
270:
72:from an activated
36:
2242:
2241:
1951:
1950:
1913:
1912:
1869:
1868:
1718:
1717:
1349:Glycogen synthase
833:10.1021/ja065950k
678:978-0-87969-770-9
593:Glycosyl acceptor
588:Glycogen synthase
507:
506:
503:
502:
498:structure summary
146:to give O-linked
85:glycosyl acceptor
16:(Redirected from
2319:
2259:
2258:
2250:
2122:Hanes–Woolf plot
2065:Enzyme activator
2060:Enzyme inhibitor
2034:Enzyme catalysis
1978:
1971:
1964:
1955:
1761:Diphtheria toxin
1744:
1735:
1392:Lactose synthase
1359:Branching enzyme
1310:
1275:
1268:
1261:
1252:
1246:
1245:
1217:
1211:
1188:
1182:
1181:
1148:(5817): 1402–5.
1137:
1131:
1126:
1120:
1119:
1109:
1077:
1071:
1070:
1060:
1028:
1022:
1021:
1019:
1018:
1009:. Archived from
1003:
997:
992:
986:
985:
975:
943:
937:
936:
926:
894:
888:
887:
851:
845:
844:
815:
809:
808:
775:(5791): 1291–4.
764:
758:
757:
747:
737:
705:
699:
689:
683:
682:
664:
658:
657:
623:
557:drug development
446:
364:
74:nucleotide sugar
65:the transfer of
27:Class of enzymes
21:
18:Transglycosylase
2327:
2326:
2322:
2321:
2320:
2318:
2317:
2316:
2267:
2266:
2265:
2253:
2245:
2243:
2238:
2150:Oxidoreductases
2136:
2112:Enzyme kinetics
2100:
2096:List of enzymes
2069:
2038:
2009:Catalytic triad
1987:
1982:
1952:
1947:
1924:
1909:
1865:
1838:
1805:
1780:Pertussis toxin
1729:
1714:
1637:
1559:
1411:
1378:
1304:
1293:
1279:
1249:
1228:(11): 1811–53.
1219:
1218:
1214:
1189:
1185:
1139:
1138:
1134:
1127:
1123:
1079:
1078:
1074:
1043:(10): 1201–37.
1030:
1029:
1025:
1016:
1014:
1005:
1004:
1000:
993:
989:
945:
944:
940:
909:(15): 2506–14.
896:
895:
891:
853:
852:
848:
827:(51): 16420–1.
817:
816:
812:
766:
765:
761:
707:
706:
702:
690:
686:
679:
666:
665:
661:
646:
625:
624:
620:
616:
569:
548:
513:
410:OPM superfamily
362:
322:
301:
292:
283:
263:
256:Golgi apparatus
245:CMP-sialic acid
116:oligosaccharide
28:
23:
22:
15:
12:
11:
5:
2325:
2323:
2315:
2314:
2309:
2304:
2299:
2294:
2289:
2284:
2279:
2269:
2268:
2264:
2263:
2240:
2239:
2237:
2236:
2223:
2210:
2197:
2184:
2171:
2158:
2144:
2142:
2138:
2137:
2135:
2134:
2129:
2124:
2119:
2114:
2108:
2106:
2102:
2101:
2099:
2098:
2093:
2088:
2083:
2077:
2075:
2074:Classification
2071:
2070:
2068:
2067:
2062:
2057:
2052:
2046:
2044:
2040:
2039:
2037:
2036:
2031:
2026:
2021:
2016:
2011:
2006:
2001:
1995:
1993:
1989:
1988:
1983:
1981:
1980:
1973:
1966:
1958:
1949:
1948:
1946:
1945:
1940:
1935:
1929:
1927:
1915:
1914:
1911:
1910:
1908:
1907:
1906:
1905:
1895:
1894:
1893:
1888:
1877:
1875:
1871:
1870:
1867:
1866:
1864:
1863:
1862:
1861:
1846:
1844:
1840:
1839:
1837:
1836:
1831:
1826:
1821:
1815:
1813:
1807:
1806:
1804:
1803:
1797:
1796:
1790:
1789:
1788:
1787:
1782:
1771:
1770:
1769:
1768:
1763:
1752:
1750:
1741:
1732:
1720:
1719:
1716:
1715:
1713:
1712:
1707:
1702:
1697:
1692:
1687:
1682:
1676:
1675:
1670:
1665:
1664:
1663:
1658:
1647:
1645:
1639:
1638:
1636:
1635:
1630:
1625:
1620:
1615:
1610:
1605:
1600:
1595:
1590:
1585:
1580:
1575:
1569:
1567:
1561:
1560:
1558:
1557:
1552:
1547:
1537:
1536:
1531:
1526:
1521:
1516:
1511:
1506:
1501:
1496:
1491:
1485:
1484:
1479:
1474:
1469:
1464:
1459:
1454:
1449:
1444:
1438:
1437:
1432:
1427:
1421:
1419:
1413:
1412:
1410:
1409:
1399:
1394:
1388:
1386:
1380:
1379:
1377:
1376:
1371:
1366:
1361:
1356:
1351:
1346:
1345:
1344:
1339:
1334:
1329:
1318:
1316:
1307:
1295:
1294:
1280:
1278:
1277:
1270:
1263:
1255:
1248:
1247:
1212:
1199:(11): 115494.
1183:
1132:
1121:
1072:
1023:
998:
987:
958:(10): 685–91.
938:
889:
846:
810:
759:
700:
684:
677:
659:
644:
617:
615:
612:
611:
610:
605:
600:
598:Glycosyl donor
595:
590:
585:
580:
575:
568:
565:
553:drug discovery
547:
544:
509:Main article:
505:
504:
501:
500:
495:
489:
488:
475:
469:
468:
458:
451:
450:
442:
441:
436:
430:
429:
424:
418:
417:
412:
406:
405:
400:
394:
393:
388:
382:
381:
378:
374:
373:
369:
368:
361:
358:
321:
318:
300:
297:
291:
288:
282:
279:
262:
259:
187:Luis F. Leloir
183:Leloir enzymes
120:polysaccharide
78:glycosyl donor
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
2324:
2313:
2310:
2308:
2305:
2303:
2300:
2298:
2295:
2293:
2290:
2288:
2285:
2283:
2280:
2278:
2277:Carbohydrates
2275:
2274:
2272:
2262:
2257:
2252:
2248:
2234:
2230:
2229:
2224:
2221:
2217:
2216:
2211:
2208:
2204:
2203:
2198:
2195:
2191:
2190:
2185:
2182:
2178:
2177:
2172:
2169:
2165:
2164:
2159:
2156:
2152:
2151:
2146:
2145:
2143:
2139:
2133:
2130:
2128:
2125:
2123:
2120:
2118:
2115:
2113:
2110:
2109:
2107:
2103:
2097:
2094:
2092:
2091:Enzyme family
2089:
2087:
2084:
2082:
2079:
2078:
2076:
2072:
2066:
2063:
2061:
2058:
2056:
2055:Cooperativity
2053:
2051:
2048:
2047:
2045:
2041:
2035:
2032:
2030:
2027:
2025:
2022:
2020:
2017:
2015:
2014:Oxyanion hole
2012:
2010:
2007:
2005:
2002:
2000:
1997:
1996:
1994:
1990:
1986:
1979:
1974:
1972:
1967:
1965:
1960:
1959:
1956:
1944:
1941:
1939:
1936:
1934:
1931:
1930:
1928:
1926:
1920:
1916:
1904:
1901:
1900:
1899:
1896:
1892:
1889:
1887:
1884:
1883:
1882:
1879:
1878:
1876:
1872:
1860:
1857:
1856:
1855:
1851:
1848:
1847:
1845:
1841:
1835:
1832:
1830:
1827:
1825:
1822:
1820:
1817:
1816:
1814:
1812:
1808:
1802:
1799:
1798:
1795:
1792:
1791:
1786:
1785:Cholera toxin
1783:
1781:
1778:
1777:
1776:
1773:
1772:
1767:
1764:
1762:
1759:
1758:
1757:
1754:
1753:
1751:
1749:
1745:
1742:
1740:
1736:
1733:
1731:
1725:
1721:
1711:
1708:
1706:
1703:
1701:
1698:
1696:
1693:
1691:
1688:
1686:
1683:
1681:
1678:
1677:
1674:
1671:
1669:
1666:
1662:
1659:
1657:
1654:
1653:
1652:
1649:
1648:
1646:
1644:
1640:
1634:
1631:
1629:
1626:
1624:
1621:
1619:
1616:
1614:
1611:
1609:
1606:
1604:
1601:
1599:
1596:
1594:
1591:
1589:
1586:
1584:
1581:
1579:
1576:
1574:
1571:
1570:
1568:
1566:
1562:
1556:
1553:
1551:
1548:
1546:
1542:
1539:
1538:
1535:
1532:
1530:
1527:
1525:
1522:
1520:
1517:
1515:
1512:
1510:
1507:
1505:
1502:
1500:
1497:
1495:
1492:
1490:
1487:
1486:
1483:
1480:
1478:
1475:
1473:
1470:
1468:
1465:
1463:
1460:
1458:
1455:
1453:
1450:
1448:
1445:
1443:
1440:
1439:
1436:
1433:
1431:
1428:
1426:
1423:
1422:
1420:
1418:
1417:Glucuronosyl-
1414:
1407:
1403:
1400:
1398:
1395:
1393:
1390:
1389:
1387:
1385:
1381:
1375:
1372:
1370:
1367:
1365:
1362:
1360:
1357:
1355:
1352:
1350:
1347:
1343:
1340:
1338:
1335:
1333:
1330:
1328:
1325:
1324:
1323:
1322:Phosphorylase
1320:
1319:
1317:
1315:
1311:
1308:
1306:
1300:
1296:
1291:
1287:
1283:
1276:
1271:
1269:
1264:
1262:
1257:
1256:
1253:
1243:
1239:
1235:
1231:
1227:
1223:
1216:
1213:
1209:
1206:
1202:
1198:
1194:
1187:
1184:
1179:
1175:
1171:
1167:
1163:
1159:
1155:
1151:
1147:
1143:
1136:
1133:
1130:
1125:
1122:
1117:
1113:
1108:
1103:
1099:
1095:
1091:
1087:
1083:
1076:
1073:
1068:
1064:
1059:
1054:
1050:
1046:
1042:
1038:
1034:
1027:
1024:
1013:on 2009-03-23
1012:
1008:
1002:
999:
996:
991:
988:
983:
979:
974:
969:
965:
961:
957:
953:
949:
942:
939:
934:
930:
925:
920:
916:
912:
908:
904:
900:
893:
890:
885:
881:
877:
873:
869:
865:
862:(4): 385–90.
861:
857:
850:
847:
842:
838:
834:
830:
826:
822:
814:
811:
806:
802:
798:
794:
790:
786:
782:
778:
774:
770:
763:
760:
755:
751:
746:
741:
736:
731:
727:
723:
720:(8): e71077.
719:
715:
711:
704:
701:
697:
693:
688:
685:
680:
674:
670:
663:
660:
655:
651:
647:
645:9780470392881
641:
637:
633:
629:
622:
619:
613:
609:
606:
604:
603:Glycosylation
601:
599:
596:
594:
591:
589:
586:
584:
581:
579:
576:
574:
571:
570:
566:
564:
562:
558:
554:
545:
543:
540:
538:
534:
529:
525:
520:
518:
512:
499:
496:
494:
490:
487:
483:
479:
476:
474:
470:
466:
462:
459:
456:
452:
447:
443:
440:
437:
435:
431:
428:
425:
423:
419:
416:
413:
411:
407:
404:
401:
399:
395:
392:
389:
387:
383:
379:
375:
370:
365:
359:
357:
355:
351:
347:
343:
339:
335:
334:echinocandins
331:
327:
319:
317:
315:
314:peptidoglycan
310:
306:
298:
296:
289:
287:
280:
278:
274:
267:
260:
258:
257:
253:
248:
246:
242:
238:
234:
230:
226:
222:
218:
217:UDP-galactose
214:
209:
207:
203:
202:pyrophosphate
200:
196:
192:
188:
184:
179:
178:in bacteria.
177:
173:
169:
165:
161:
157:
153:
149:
148:glycoproteins
145:
141:
137:
133:
129:
125:
121:
117:
113:
109:
104:
102:
98:
94:
90:
86:
83:
79:
75:
71:
68:
64:
60:
56:
52:
48:
44:
40:
32:
19:
2312:Glycobiology
2287:Transferases
2228:Translocases
2225:
2212:
2199:
2186:
2173:
2163:Transferases
2160:
2147:
2004:Binding site
1925:transferases
1730:transferases
1305:transferases
1285:
1282:Transferases
1225:
1221:
1215:
1196:
1192:
1186:
1145:
1141:
1135:
1124:
1092:(6): 800–8.
1089:
1085:
1075:
1040:
1036:
1026:
1015:. Retrieved
1011:the original
1001:
990:
955:
951:
941:
906:
902:
892:
859:
855:
849:
824:
820:
813:
772:
768:
762:
717:
713:
703:
692:Transferases
687:
668:
662:
627:
621:
549:
541:
521:
514:
323:
302:
293:
284:
275:
271:
249:
210:
205:
182:
180:
158:to generate
108:carbohydrate
105:
82:nucleophilic
46:
42:
38:
37:
1999:Active site
1384:Galactosyl-
903:ChemBioChem
856:ChemBioChem
422:OPM protein
372:Identifiers
354:Imidazolium
342:Caspofungin
330:nikkomycins
237:GDP-mannose
213:UDP-glucose
168:glycolipids
2271:Categories
2202:Isomerases
2176:Hydrolases
2043:Regulation
1337:Cellobiose
1017:2009-08-25
614:References
537:frameshift
524:gene locus
461:structures
434:Membranome
346:Ethambutol
326:moenomycin
320:Inhibitors
241:GDP-fucose
229:UDP-xylose
225:UDP-GalNAc
221:UDP-GlcNAc
199:polyprenol
156:tryptophan
152:asparagine
67:saccharide
2081:EC number
1728:Pentosyl-
1643:Mannosyl-
1314:Glucosyl-
403:IPR005076
350:Lufenuron
299:Structure
261:Mechanism
144:threonine
112:glycoside
2302:EC 2.4.2
2297:EC 2.4.1
2105:Kinetics
2029:Cofactor
1992:Activity
1565:Fucosyl-
1332:Glycogen
1303:Hexosyl-
1242:21901218
1208:32312486
1178:41658295
1170:17347437
1116:21592771
1067:22688446
982:21857660
933:18798210
884:45058028
876:17262863
841:17177349
805:38072017
797:16946071
754:23936487
714:PLOS ONE
654:18990828
567:See also
522:The ABO
478:RCSB PDB
398:InterPro
195:dolichol
185:, after
172:dolichol
150:, or to
136:tyrosine
103:-based.
97:nitrogen
80:") to a
70:moieties
63:catalyze
2261:Biology
2215:Ligases
1985:Enzymes
1943:ST8SIA4
1801:Sirtuin
1534:UGT2B28
1529:UGT2B17
1524:UGT2B15
1519:UGT2B11
1514:UGT2B10
1482:UGT1A10
1406:C1GALT1
1150:Bibcode
1142:Science
1107:3163058
1058:3627186
973:3177962
924:2947747
777:Bibcode
769:Science
745:3731257
722:Bibcode
533:Guanine
391:PF03414
132:protein
118:, or a
61:. They
51:enzymes
2292:EC 2.4
2247:Portal
2189:Lyases
1923:Sialyl
1919:2.4.99
1739:Ribose
1578:POFUT2
1573:POFUT1
1509:UGT2B7
1504:UGT2B4
1499:UGT2A3
1494:UGT2A2
1489:UGT2A1
1477:UGT1A9
1472:UGT1A8
1467:UGT1A7
1462:UGT1A6
1457:UGT1A5
1452:UGT1A4
1447:UGT1A3
1442:UGT1A1
1435:B3GAT3
1430:B3GAT2
1425:B3GAT1
1327:Starch
1240:
1176:
1168:
1114:
1104:
1065:
1055:
980:
970:
931:
921:
882:
874:
839:
803:
795:
752:
742:
675:
652:
642:
493:PDBsum
467:
457:
377:Symbol
243:, and
164:lipids
140:serine
101:sulfur
99:-, or
93:carbon
89:oxygen
55:EC 2.4
49:) are
2141:Types
1891:XYLT2
1886:XYLT1
1874:Other
1859:ECGF1
1843:Other
1724:2.4.2
1710:ALG12
1661:POMT2
1656:POMT1
1633:FUT11
1628:FUT10
1299:2.4.1
1174:S2CID
880:S2CID
801:S2CID
142:, or
128:water
2233:list
2226:EC7
2220:list
2213:EC6
2207:list
2200:EC5
2194:list
2187:EC4
2181:list
2174:EC3
2168:list
2161:EC2
2155:list
2148:EC1
1705:ALG9
1700:ALG8
1695:ALG6
1690:ALG3
1685:ALG2
1680:ALG1
1673:DPM3
1668:DPM1
1623:FUT9
1618:FUT8
1613:FUT7
1608:FUT6
1603:FUT5
1598:FUT4
1593:FUT3
1588:FUT2
1583:FUT1
1555:HAS3
1550:HAS2
1545:HAS1
1342:Myo-
1292:2.4)
1238:PMID
1205:PMID
1166:PMID
1112:PMID
1063:PMID
978:PMID
929:PMID
872:PMID
837:PMID
793:PMID
750:PMID
673:ISBN
650:PMID
640:ISBN
555:and
546:Uses
515:The
486:PDBj
482:PDBe
465:ECOD
455:Pfam
427:2rj6
386:Pfam
204:are
47:Gtfs
43:GTFs
1230:doi
1158:doi
1146:315
1102:PMC
1094:doi
1053:PMC
1045:doi
968:PMC
960:doi
919:PMC
911:doi
864:doi
829:doi
825:128
785:doi
773:313
740:PMC
730:doi
694:in
632:doi
473:PDB
439:468
415:199
380:GT6
197:or
126:or
95:-,
2273::
1921::
1852::
1726::
1543::
1301::
1290:EC
1284::
1236:.
1226:28
1224:.
1203:.
1197:28
1195:.
1172:.
1164:.
1156:.
1144:.
1110:.
1100:.
1090:22
1088:.
1084:.
1061:.
1051:.
1041:29
1039:.
1035:.
976:.
966:.
954:.
950:.
927:.
917:.
905:.
901:.
878:.
870:.
858:.
835:.
823:.
799:.
791:.
783:.
771:.
748:.
738:.
728:.
716:.
712:.
648:.
638:.
484:;
480:;
463:/
316:.
239:,
235:,
231:,
227:,
223:,
219:,
215:,
208:.
138:,
114:,
110:,
91:-
45:,
2249::
2235:)
2231:(
2222:)
2218:(
2209:)
2205:(
2196:)
2192:(
2183:)
2179:(
2170:)
2166:(
2157:)
2153:(
1977:e
1970:t
1963:v
1408:)
1404:(
1288:(
1274:e
1267:t
1260:v
1244:.
1232::
1210:.
1180:.
1160::
1152::
1118:.
1096::
1069:.
1047::
1020:.
984:.
962::
956:7
935:.
913::
907:9
886:.
866::
860:8
843:.
831::
807:.
787::
779::
756:.
732::
724::
718:8
698:.
681:.
656:.
634::
53:(
41:(
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.