112:
1073:) can explain variation in genomic architecture among species, e.g. the size of the genome, or the mutation rate. Specifically, larger populations will have lower mutation rates, more streamlined genomic architectures, and generally more finely tuned adaptations. However, if robustness to the consequences of each possible error in processes such as transcription and translation substantially reduces the cost of making such errors, larger populations might evolve lower rates of global
121:
95:
that included both beneficial and deleterious mutations, so that no artificial "shift" of overall population fitness was necessary. According to Ohta, however, the nearly neutral theory largely fell out of favor in the late 1980s, because the mathematically simpler neutral theory for the widespread
929:
is constant (in this sense, the argument in the previous paragraphs can be regarded as based on the “shift model”). This assumption can lead to indefinite improvement or deterioration of protein function. Alternatively, the later “fixed model” fixes the distribution of mutations’ effect on protein
94:
Between then and the early 1990s, many studies of molecular evolution used a "shift model" in which the negative effect on the fitness of a population due to deleterious mutations shifts back to an original value when a mutation reaches fixation. In the early 1990s, Ohta developed a "fixed model"
86:
In this case, the faster rate of neutral evolution in proteins expected in small populations (due to a more lenient threshold for purging deleterious mutations) is offset by longer generation times (and vice versa), but in large populations with short generation times, noncoding DNA evolves faster
1035:: a large product corresponds to adaptive evolution, an intermediate product corresponds to nearly neutral evolution, and a small product corresponds to almost neutral evolution. According to this classification, slightly advantageous mutations can contribute to nearly neutral evolution.
66:
According to the neutral theory of molecular evolution, the rate at which molecular changes accumulate between species should be equal to the rate of neutral mutations and hence relatively constant across species. However, this is a per-generation rate. Since larger organisms have longer
115:
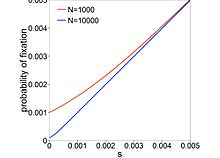
The probability of fixation depends strongly on N for deleterious mutations (note the log scale on the y-axis) relative to the neutral case of s=0. Dashed lines show the probability of fixation of a mutation with s=-1/N. Note that larger populations have more deleterious mutations (not
984:
populations, advantageous mutations are quickly picked up by selection, increasing the mean fitness of the population. In response, the mutation rate of nearly neutral mutations is reduced because these mutations are restricted to the tail of the distribution of selection coefficients.
103:. As more detailed systematics studies started to compare the evolution of genome regions subject to strong selection versus weaker selection in the 1990s, the nearly neutral theory and the interaction between selection and drift have once again become an important focus of research.
82:
substitutions tend to be more neutral, independent of population size, their rate of evolution is correctly predicted to depend on population size / generation time, unlike the rate of non-synonymous changes.
71:, the neutral theory predicts that their rate of molecular evolution should be slower. However, molecular evolutionists found that rates of protein evolution were fairly independent of generation time.
78:
substitutions are slightly deleterious, this would increase the rate of effectively neutral mutation rate in small populations, which could offset the effect of long generation times. However, because
450:
838:
213:
727:
581:
528:
624:
91:
suggesting that a wide variety of molecular evidence supported the theory that most mutation events at the molecular level are slightly deleterious rather than strictly neutral.
657:
1071:
1013:
982:
865:
785:
754:
684:
355:
284:
146:
1033:
952:
927:
907:
887:
474:
328:
308:
257:
237:
124:
The probability of fixation of beneficial mutations is fairly insensitive to N. Note that larger populations have more beneficial mutations (not illustrated).
87:
while protein evolution is retarded by selection (which is more significant than drift for large populations) In 1973, Ohta published a short letter in
909:
can vary between generations but the mean fitness of the population is reset to zero after fixation. This basically assumes the distribution of
1093:
depend on protein abundance (which is responsible for modulating the locus-specific strength of selection), but do so only for high-error-rate
31:
are either so deleterious such that they can be ignored, or else neutral. Slightly deleterious mutations are reliably purged only when their
24:
1500:"Correction for Traverse and Ochman, Conserved rates and patterns of transcription errors across bacterial growth states and lifestyles"
988:
The “fixed model” expands the nearly neutral theory. Tachida classified evolution under the “fixed model” based on the product of
1662:
363:
1117:
287:
44:
787:
populations, these mutations are purged by selection. If nearly neutral mutations are common, then the proportion for which
111:
1683:
1678:
74:
Noting that population size is generally inversely proportional to generation time, Tomoko Ohta proposed that if most
1043:
790:
154:
55:
689:
36:
533:
1085:
483:
586:
54:
in 1973. The population-size-dependent threshold for purging mutations has been called the "drift barrier" by
1688:
1256:
957:
The “fixed model” provides a slightly different explanation for the rate of protein evolution. In large
477:
96:
32:
1046:
has proposed that variation in the ability to purge slightly deleterious mutations (i.e. variation in
1511:
1453:
1200:
1146:
1074:
1261:
286:
is the effective population size. The last term is the probability that a new mutation will become
1224:
1170:
1292:
Ohta T (August 1996). "The current significance and standing of neutral and neutral theories".
120:
1641:
1590:
1539:
1481:
1407:
1358:
1309:
1274:
1216:
1162:
629:
1631:
1621:
1580:
1570:
1529:
1519:
1471:
1461:
1397:
1389:
1348:
1340:
1301:
1266:
1208:
1154:
1079:
1049:
991:
960:
843:
763:
732:
662:
333:
262:
131:
68:
39:. In larger populations, a higher proportion of mutations exceed this threshold for which
1329:"Theoretical study of near neutrality. I. Heterozygosity and rate of mutant substitution"
1247:
Ohta T, Gillespie JH (April 1996). "Development of
Neutral and Nearly Neutral Theories".
1515:
1457:
1204:
1150:
1636:
1609:
1585:
1558:
1534:
1499:
1476:
1441:
1402:
1377:
1353:
1328:
1018:
937:
912:
892:
872:
459:
313:
293:
242:
222:
100:
1672:
757:
79:
40:
1228:
1174:
931:
357:. Kimura’s equation for the probability of fixation in a haploid population gives:
1191:
Ohta T (November 1973). "Slightly deleterious mutant substitutions in evolution".
1393:
1344:
1575:
1559:"Drift Barriers to Quality Control When Genes Are Expressed at Different Levels"
1101:
51:
1504:
Proceedings of the
National Academy of Sciences of the United States of America
1446:
Proceedings of the
National Academy of Sciences of the United States of America
1610:"High Transcriptional Error Rates Vary as a Function of Gene Expression Level"
75:
1524:
1466:
1645:
1594:
1543:
1485:
1305:
1270:
1442:"Evolution of molecular error rates and the consequences for evolvability"
1411:
1362:
1313:
1278:
1220:
1166:
1626:
1094:
58:, and used to explain differences in genomic architecture among species.
28:
1137:
Kimura M (February 1968). "Evolutionary rate at the molecular level".
729:
are called nearly neutral mutations. These mutations can fix in small-
1212:
1158:
1098:
869:
The effect of nearly neutral mutations can depend on fluctuations in
1089:. This is supported by the fact that transcriptional error rates in
1378:"A study on a nearly neutral mutation model in finite populations"
119:
110:
1077:, and hence have higher rates of error. This may explain why
1557:
Xiong K, McEntee JP, Porfirio DJ, Masel J (January 2017).
934:
of population to evolve. This allows the distribution of
445:{\displaystyle P_{fix}={\frac {1-e^{-s}}{1-e^{-sN_{e}}}}}
1608:
Meer KM, Nelson PG, Xiong K, Masel J (January 2020).
1052:
1021:
994:
963:
940:
915:
895:
875:
846:
793:
766:
735:
692:
665:
632:
589:
536:
486:
462:
366:
336:
316:
296:
265:
245:
225:
157:
134:
99:research that flourished after the advent of rapid
1065:
1027:
1007:
976:
946:
921:
901:
881:
859:
832:
779:
748:
721:
678:
651:
618:
575:
522:
468:
444:
349:
322:
302:
278:
251:
231:
207:
140:
1663:The Nearly Neutral Theory of Molecular Evolution
954:to change with the mean fitness of population.
1083:has higher rates of transcription error than
833:{\displaystyle P_{fix}\ll {\frac {1}{N_{e}}}}
208:{\displaystyle \rho =ugN_{e}{\bar {P}}_{fix}}
43:cannot overpower selection, leading to fewer
8:
21:nearly neutral theory of molecular evolution
1242:
1240:
1238:
1186:
1184:
889:. Early work used a “shift model” in which
722:{\displaystyle -s\simeq {\frac {1}{N_{e}}}}
576:{\displaystyle P_{fix}={\frac {1}{N_{e}}}}
50:The nearly neutral theory was proposed by
47:events and so slower molecular evolution.
1635:
1625:
1584:
1574:
1533:
1523:
1475:
1465:
1401:
1352:
1260:
1057:
1051:
1020:
999:
993:
968:
962:
939:
914:
894:
874:
851:
845:
822:
813:
798:
792:
771:
765:
740:
734:
711:
702:
691:
670:
664:
637:
631:
608:
599:
588:
565:
556:
541:
535:
523:{\displaystyle |s|\ll {\frac {1}{N_{e}}}}
512:
503:
495:
487:
485:
461:
431:
420:
399:
386:
371:
365:
341:
335:
315:
295:
270:
264:
244:
224:
193:
182:
181:
174:
156:
133:
1015:and the variance in the distribution of
619:{\displaystyle -s\gg {\frac {1}{N_{e}}}}
27:that accounts for the fact that not all
1129:
310:is constant between species, and that
1665:- Perspectives on Molecular Evolution
25:neutral theory of molecular evolution
7:
1327:Ohta T, Tachida H (September 1990).
659:decreases almost exponentially with
35:are greater than one divided by the
1427:The origins of genome architecture
16:Variant of one theory of evolution
14:
1440:Rajon E, Masel J (January 2011).
1429:. Sunderland: Sinauer Associates.
1510:(29): E4257–E4258. July 2016.
1249:Theoretical Population Biology
1118:History of molecular evolution
496:
488:
187:
1:
1614:Genome Biology and Evolution
1300:(8): 673–7, discussion 683.
290:. Early models assumed that
259:is the generation time, and
1576:10.1534/genetics.116.192567
128:The rate of substitution,
1705:
1394:10.1093/genetics/128.1.183
1345:10.1093/genetics/126.1.219
1039:The "drift barrier" theory
930:function, but allows the
626:(extremely deleterious),
37:effective population size
23:is a modification of the
1086:Saccharomyces cerevisiae
1525:10.1073/pnas.1609677113
1467:10.1073/pnas.1012918108
652:{\displaystyle P_{fix}}
1376:Tachida H (May 1991).
1306:10.1002/bies.950180811
1271:10.1006/tpbi.1996.0007
1067:
1029:
1009:
978:
948:
923:
903:
883:
861:
834:
781:
750:
723:
680:
653:
620:
577:
530:(completely neutral),
524:
470:
446:
351:
324:
304:
280:
253:
239:is the mutation rate,
233:
209:
142:
125:
117:
1068:
1066:{\displaystyle N_{e}}
1030:
1010:
1008:{\displaystyle N_{e}}
979:
977:{\displaystyle N_{e}}
949:
924:
904:
884:
862:
860:{\displaystyle N_{e}}
835:
782:
780:{\displaystyle N_{e}}
751:
749:{\displaystyle N_{e}}
724:
681:
679:{\displaystyle N_{e}}
654:
621:
578:
525:
478:selection coefficient
471:
447:
352:
350:{\displaystyle N_{e}}
325:
305:
281:
279:{\displaystyle N_{e}}
254:
234:
210:
143:
141:{\displaystyle \rho }
123:
114:
97:molecular systematics
33:selection coefficient
1050:
1019:
992:
961:
938:
913:
893:
873:
844:
791:
764:
756:populations through
733:
690:
663:
630:
587:
534:
484:
480:of a mutation. When
460:
364:
334:
314:
294:
263:
243:
223:
155:
132:
1684:Population genetics
1679:Molecular evolution
1516:2016PNAS..113E4257.
1458:2011PNAS..108.1082R
1205:1973Natur.246...96O
1151:1968Natur.217..624K
1627:10.1093/gbe/evz275
1063:
1025:
1005:
974:
944:
919:
899:
879:
857:
830:
777:
746:
719:
676:
649:
616:
573:
520:
466:
442:
347:
320:
300:
276:
249:
229:
205:
138:
126:
118:
1145:(5129): 624–626.
1028:{\displaystyle s}
947:{\displaystyle s}
922:{\displaystyle s}
902:{\displaystyle s}
882:{\displaystyle s}
828:
717:
686:. Mutations with
614:
571:
518:
469:{\displaystyle s}
440:
323:{\displaystyle g}
303:{\displaystyle u}
252:{\displaystyle g}
232:{\displaystyle u}
190:
1696:
1650:
1649:
1639:
1629:
1620:(1): 3754–3761.
1605:
1599:
1598:
1588:
1578:
1554:
1548:
1547:
1537:
1527:
1496:
1490:
1489:
1479:
1469:
1452:(3): 1082–1087.
1437:
1431:
1430:
1425:Lynch M (2007).
1422:
1416:
1415:
1405:
1373:
1367:
1366:
1356:
1324:
1318:
1317:
1289:
1283:
1282:
1264:
1244:
1233:
1232:
1213:10.1038/246096a0
1188:
1179:
1178:
1159:10.1038/217624a0
1134:
1080:Escherichia coli
1072:
1070:
1069:
1064:
1062:
1061:
1034:
1032:
1031:
1026:
1014:
1012:
1011:
1006:
1004:
1003:
983:
981:
980:
975:
973:
972:
953:
951:
950:
945:
928:
926:
925:
920:
908:
906:
905:
900:
888:
886:
885:
880:
866:
864:
863:
858:
856:
855:
840:is dependent on
839:
837:
836:
831:
829:
827:
826:
814:
809:
808:
786:
784:
783:
778:
776:
775:
755:
753:
752:
747:
745:
744:
728:
726:
725:
720:
718:
716:
715:
703:
685:
683:
682:
677:
675:
674:
658:
656:
655:
650:
648:
647:
625:
623:
622:
617:
615:
613:
612:
600:
582:
580:
579:
574:
572:
570:
569:
557:
552:
551:
529:
527:
526:
521:
519:
517:
516:
504:
499:
491:
475:
473:
472:
467:
451:
449:
448:
443:
441:
439:
438:
437:
436:
435:
408:
407:
406:
387:
382:
381:
356:
354:
353:
348:
346:
345:
329:
327:
326:
321:
309:
307:
306:
301:
285:
283:
282:
277:
275:
274:
258:
256:
255:
250:
238:
236:
235:
230:
214:
212:
211:
206:
204:
203:
192:
191:
183:
179:
178:
147:
145:
144:
139:
69:generation times
1704:
1703:
1699:
1698:
1697:
1695:
1694:
1693:
1669:
1668:
1659:
1654:
1653:
1607:
1606:
1602:
1556:
1555:
1551:
1498:
1497:
1493:
1439:
1438:
1434:
1424:
1423:
1419:
1375:
1374:
1370:
1326:
1325:
1321:
1291:
1290:
1286:
1262:10.1.1.332.2080
1246:
1245:
1236:
1199:(5428): 96–98.
1190:
1189:
1182:
1136:
1135:
1131:
1126:
1114:
1053:
1048:
1047:
1041:
1017:
1016:
995:
990:
989:
964:
959:
958:
936:
935:
911:
910:
891:
890:
871:
870:
847:
842:
841:
818:
794:
789:
788:
767:
762:
761:
736:
731:
730:
707:
688:
687:
666:
661:
660:
633:
628:
627:
604:
585:
584:
561:
537:
532:
531:
508:
482:
481:
458:
457:
427:
416:
409:
395:
388:
367:
362:
361:
337:
332:
331:
330:increases with
312:
311:
292:
291:
266:
261:
260:
241:
240:
221:
220:
180:
170:
153:
152:
130:
129:
109:
64:
17:
12:
11:
5:
1702:
1700:
1692:
1691:
1689:Neutral theory
1686:
1681:
1671:
1670:
1667:
1666:
1658:
1657:External links
1655:
1652:
1651:
1600:
1569:(1): 397–407.
1549:
1491:
1432:
1417:
1388:(1): 183–192.
1368:
1339:(1): 219–229.
1319:
1284:
1255:(2): 128–142.
1234:
1180:
1128:
1127:
1125:
1122:
1121:
1120:
1113:
1110:
1060:
1056:
1040:
1037:
1024:
1002:
998:
971:
967:
943:
918:
898:
878:
854:
850:
825:
821:
817:
812:
807:
804:
801:
797:
774:
770:
743:
739:
714:
710:
706:
701:
698:
695:
673:
669:
646:
643:
640:
636:
611:
607:
603:
598:
595:
592:
568:
564:
560:
555:
550:
547:
544:
540:
515:
511:
507:
502:
498:
494:
490:
465:
454:
453:
434:
430:
426:
423:
419:
415:
412:
405:
402:
398:
394:
391:
385:
380:
377:
374:
370:
344:
340:
319:
299:
273:
269:
248:
228:
217:
216:
202:
199:
196:
189:
186:
177:
173:
169:
166:
163:
160:
137:
108:
105:
101:DNA sequencing
63:
60:
15:
13:
10:
9:
6:
4:
3:
2:
1701:
1690:
1687:
1685:
1682:
1680:
1677:
1676:
1674:
1664:
1661:
1660:
1656:
1647:
1643:
1638:
1633:
1628:
1623:
1619:
1615:
1611:
1604:
1601:
1596:
1592:
1587:
1582:
1577:
1572:
1568:
1564:
1560:
1553:
1550:
1545:
1541:
1536:
1531:
1526:
1521:
1517:
1513:
1509:
1505:
1501:
1495:
1492:
1487:
1483:
1478:
1473:
1468:
1463:
1459:
1455:
1451:
1447:
1443:
1436:
1433:
1428:
1421:
1418:
1413:
1409:
1404:
1399:
1395:
1391:
1387:
1383:
1379:
1372:
1369:
1364:
1360:
1355:
1350:
1346:
1342:
1338:
1334:
1330:
1323:
1320:
1315:
1311:
1307:
1303:
1299:
1295:
1288:
1285:
1280:
1276:
1272:
1268:
1263:
1258:
1254:
1250:
1243:
1241:
1239:
1235:
1230:
1226:
1222:
1218:
1214:
1210:
1206:
1202:
1198:
1194:
1187:
1185:
1181:
1176:
1172:
1168:
1164:
1160:
1156:
1152:
1148:
1144:
1140:
1133:
1130:
1123:
1119:
1116:
1115:
1111:
1109:
1107:
1106:S. cerevisiae
1103:
1100:
1096:
1092:
1088:
1087:
1082:
1081:
1076:
1058:
1054:
1045:
1044:Michael Lynch
1038:
1036:
1022:
1000:
996:
986:
969:
965:
955:
941:
933:
916:
896:
876:
867:
852:
848:
823:
819:
815:
810:
805:
802:
799:
795:
772:
768:
759:
758:genetic drift
741:
737:
712:
708:
704:
699:
696:
693:
671:
667:
644:
641:
638:
634:
609:
605:
601:
596:
593:
590:
566:
562:
558:
553:
548:
545:
542:
538:
513:
509:
505:
500:
492:
479:
463:
432:
428:
424:
421:
417:
413:
410:
403:
400:
396:
392:
389:
383:
378:
375:
372:
368:
360:
359:
358:
342:
338:
317:
297:
289:
271:
267:
246:
226:
200:
197:
194:
184:
175:
171:
167:
164:
161:
158:
151:
150:
149:
135:
122:
116:illustrated).
113:
106:
104:
102:
98:
92:
90:
84:
81:
80:noncoding DNA
77:
72:
70:
61:
59:
57:
56:Michael Lynch
53:
48:
46:
42:
41:genetic drift
38:
34:
30:
26:
22:
1617:
1613:
1603:
1566:
1562:
1552:
1507:
1503:
1494:
1449:
1445:
1435:
1426:
1420:
1385:
1381:
1371:
1336:
1332:
1322:
1297:
1293:
1287:
1252:
1248:
1196:
1192:
1142:
1138:
1132:
1105:
1090:
1084:
1078:
1075:proofreading
1042:
987:
956:
932:mean fitness
868:
455:
218:
127:
93:
88:
85:
73:
65:
49:
20:
18:
1102:deamination
760:. In large-
583:, and when
52:Tomoko Ohta
1673:Categories
1124:References
1104:errors in
76:amino acid
1294:BioEssays
1257:CiteSeerX
811:≪
700:≃
694:−
597:≫
591:−
501:≪
422:−
414:−
401:−
393:−
188:¯
159:ρ
136:ρ
29:mutations
1646:31841128
1595:27838629
1563:Genetics
1544:27402746
1486:21199946
1382:Genetics
1333:Genetics
1112:See also
45:fixation
1637:6988749
1586:5223517
1535:4961203
1512:Bibcode
1477:3024668
1454:Bibcode
1412:2060776
1403:1204447
1363:2227381
1354:1204126
1314:8779656
1279:8813019
1229:4226804
1221:4585855
1201:Bibcode
1175:4161261
1167:5637732
1147:Bibcode
1091:E. coli
476:is the
62:Origins
1644:
1634:
1593:
1583:
1542:
1532:
1484:
1474:
1410:
1400:
1361:
1351:
1312:
1277:
1259:
1227:
1219:
1193:Nature
1173:
1165:
1139:Nature
456:where
219:where
107:Theory
89:Nature
1225:S2CID
1171:S2CID
288:fixed
1642:PMID
1591:PMID
1540:PMID
1482:PMID
1408:PMID
1359:PMID
1310:PMID
1275:PMID
1217:PMID
1163:PMID
19:The
1632:PMC
1622:doi
1581:PMC
1571:doi
1567:205
1530:PMC
1520:doi
1508:113
1472:PMC
1462:doi
1450:108
1398:PMC
1390:doi
1386:128
1349:PMC
1341:doi
1337:126
1302:doi
1267:doi
1209:doi
1197:246
1155:doi
1143:217
1097:to
148:is
1675::
1640:.
1630:.
1618:12
1616:.
1612:.
1589:.
1579:.
1565:.
1561:.
1538:.
1528:.
1518:.
1506:.
1502:.
1480:.
1470:.
1460:.
1448:.
1444:.
1406:.
1396:.
1384:.
1380:.
1357:.
1347:.
1335:.
1331:.
1308:.
1298:18
1296:.
1273:.
1265:.
1253:49
1251:.
1237:^
1223:.
1215:.
1207:.
1195:.
1183:^
1169:.
1161:.
1153:.
1141:.
1108:.
1648:.
1624::
1597:.
1573::
1546:.
1522::
1514::
1488:.
1464::
1456::
1414:.
1392::
1365:.
1343::
1316:.
1304::
1281:.
1269::
1231:.
1211::
1203::
1177:.
1157::
1149::
1099:U
1095:C
1059:e
1055:N
1023:s
1001:e
997:N
970:e
966:N
942:s
917:s
897:s
877:s
853:e
849:N
824:e
820:N
816:1
806:x
803:i
800:f
796:P
773:e
769:N
742:e
738:N
713:e
709:N
705:1
697:s
672:e
668:N
645:x
642:i
639:f
635:P
610:e
606:N
602:1
594:s
567:e
563:N
559:1
554:=
549:x
546:i
543:f
539:P
514:e
510:N
506:1
497:|
493:s
489:|
464:s
452:,
433:e
429:N
425:s
418:e
411:1
404:s
397:e
390:1
384:=
379:x
376:i
373:f
369:P
343:e
339:N
318:g
298:u
272:e
268:N
247:g
227:u
215:,
201:x
198:i
195:f
185:P
176:e
172:N
168:g
165:u
162:=
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.