44:
136:
is formed with five NH groups, and so on. The main chain atoms form part of an incomplete ring with the NH groups all pointing roughly towards the centre of the ring. Because their concavities are often wider than simple nests, compound nests are commonly employed by proteins for binding multi-atom anions such as
135:
If two nests overlap such that residue i+1 of the first nest is residue i of the second nest, a compound nest is formed. This has four NH groups instead of three. If three nests overlap such that residues i+1 and i+2 of the first nest are residue i of the second and third nest, a wider compound nest
60:
to a negatively charged, or partially negatively charged, atom, often an oxygen atom. The NH of the second residue may also be hydrogen bonded to the same atom but usually points somewhat away. These main chain atoms form a concavity called a nest into which an anionic atom fits. Such anionic atoms
223:
Watson, JD; Milner-White (2002). "A novel main-chain anion-binding site in proteins: The nest. A particular combination of phi,psi values in successive residues gives rise to anion-binding sites that occur commonly and are found often at functionally important regions".
164:
Simple nests are of two kinds called RL and LR depending on the sign of the phi angles of the first two nest residues. R residues have negative phi values (as in right-handed alpha-helices) and L residues have positive phi values (as in the left-handed
111:
Nests vary in their degree of concavity. A few have so little that the concavity is lost; these peptides often bind cations via their main chain CO groups, instead of anions via their NH groups. The specificity filter of the
601:
Watson, JD; Milner-White (2002). "The conformations of polypeptide chains where the main-chain parts of successive residues are enantiomeric. Their occurrence in cation and anion-binding regions of proteins".
721:
Hayward, S; Milner-White (2011). "Simulation of the β- to α-sheet transition results in a twisted sheet for antiparallel and an α-nanotube for parallel strands: implications for amyloid formation".
206:
have an RL nest at the beginning of the first beta-strand, with the function of recognizing the carboxylate group at the C-terminus of the domain's peptide or protein ligand.
370:
Berkessel, A; Koch (2006). "Asymmetric enone epoxidation by solid-phase bound peptides: further evidence for catalyst helicity and catalytic activity of individual strands".
764:
Bianchi, A; Giorgi A; Ruzza P; Toniolo C (2013). "A synthetic hexapeptide designed to resemble a proteinaceous P-loop nest is shown to bind inorganic phosphate".
191:) bound to a carboxylate side chain. These have been engineered to give rise to monoclonal nest-containing antibodies specific for proteins with phosphorylated
169:). Eighty percent of nests are RL and 20% are LR. When two nests overlap they may be RLR or LRL. When three nests overlap they may be RLRL or LRLR, and so on.
47:
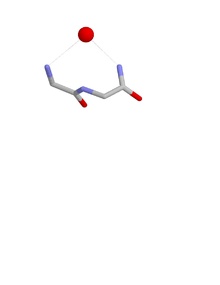
RL nest bound to an egg oxygen. carbons grey, oxygens red and nitrogens blue. Hydrogen atoms omitted. Hydrogen bonds are grey dotted lines.
335:
Pajewski, R; Ferdani (2005). "Cation
Dependence of Chloride Ion Complexation by Open-Chained Receptor Molecules in Chloroform Solution".
188:
1038:
678:
Hayward, S; Milner-White (2008). "The geometry of α-sheet: Implications for its possible function as amyloid precursor in proteins".
124:
are employed by proteins to transport molecules across membranes. This near-linear conformation is also that found in a strand of
40:
residues lack NH groups so are rare in nests. About one in 12 of amino acid residues in proteins, on average, belongs to a nest.
56:
The conformation of a nest is such that the NH groups of the first and third amino acid residues are liable to be
43:
36:
residues. The main chain NH groups bind the anions while the side chain atoms are often not involved.
1043:
149:
507:"Anion recognition in water: Recent advances from a supramolecular and macromolecular perspective"
789:
746:
703:
583:
639:"Amyloid formation may involve alpha- to beta sheet interconversion via peptide plane flipping"
1015:
964:
889:
838:
781:
738:
695:
660:
619:
575:
536:
487:
438:
387:
352:
317:
276:
241:
113:
1005:
995:
954:
944:
879:
869:
828:
820:
773:
730:
687:
650:
611:
567:
526:
518:
477:
469:
428:
418:
379:
344:
307:
268:
233:
21:
933:"Motivated Proteins: A web application for studying small three-dimensional protein motifs"
259:
Pal, D; Suhnel (2002). "New principles of protein structure: nests, eggs and what next?".
66:
556:"Collaborative routes to clarifying the murky waters of aquaeous supramolcular chemistry"
69:
is a functional example of a nest. Another occurs at the bottom of a deep cavity in the
1010:
983:
959:
932:
884:
857:
833:
808:
531:
506:
482:
457:
433:
406:
173:
121:
89:
1032:
587:
145:
57:
707:
793:
750:
88:
Nests are defined by the conformation of the main chain atoms, namely the phi, psi
152:. The synthesized peptide Ser-Gly-Ala-Gly-Lys-Thr, designed as a minimal peptide
858:"PDZ Domains and their binding Partners: Structure Specificity and Modification"
166:
125:
78:
61:
are sometimes called eggs and more than one egg may occur bound to a nest. The
655:
638:
312:
295:
203:
176:
incorporates an RL nest in the last three of its six residues. The nest binds
74:
70:
33:
1000:
949:
917:
196:
137:
117:
82:
1019:
968:
893:
842:
785:
742:
699:
664:
623:
615:
579:
540:
522:
491:
442:
391:
356:
321:
280:
272:
245:
237:
911:
874:
423:
407:"Predicting the conformations of proteins and peptides in early evolution"
473:
184:
177:
62:
296:"Recurring main-chain anion-binding motifs in short polypeptides: nests"
187:
proteins have RLR nests within the hairpin loops of their H-chain CDRs (
777:
734:
691:
571:
37:
29:
25:
383:
348:
824:
458:"ProFunc: a server for predicting protein function from 3D structure"
192:
153:
141:
92:
of the first two amino acids in the nest. For a typical (RL) nest phi
807:
Koerber, JT; Thomsen ND; Hannigan BT; DeGrado WF; Wells JA (2013).
42:
555:
156:, was shown to bind inorganic phosphate strongly at neutral pH.
85:
synthesis, thereby preventing bacterial cells from multiplying.
809:"Nature-inspired design of motif-specific antibody scaffolds"
32:. Each consists of the main chain atoms of three consecutive
24:. It is a small recurring anion-binding feature of both
554:Cremer, P; Flood AS; Gibb BC; Mobley DL (2018).
120:exhibit this more linear conformation in which
984:"MSDmotif: exploring protein sites and motifs"
8:
81:group utilized during the final stages of
1009:
999:
958:
948:
883:
873:
832:
654:
530:
505:Langton, MJ; Serpell CJ; Beer PD (2016).
481:
432:
422:
311:
337:Journal of the American Chemical Society
923:
511:Angewandte Chemie International Edition
215:
180:oxygen atoms preceding it in sequence.
7:
189:complementarity determining regions
405:Milner-White, EJ; Russell (2006).
294:Milner-White, EJ; Nissink (2004).
14:
931:Leader, DP; Milner-White (2009).
637:Milner-White, EJ; Watson (2006).
862:Cell Communication and Signaling
300:Acta Crystallographica Section D
456:Watson, JD; Laskowski (2005).
1:
982:Golovin, A; Henrick (2008).
604:Journal of Molecular Biology
226:Journal of Molecular Biology
856:Lee, H-J; Zheng JJ (2010).
1060:
1039:Protein structural motifs
656:10.1016/j.str.2006.06.016
313:10.1107/s0907444904021390
116:and the water channel of
22:protein structural motif
1001:10.1186/1471-2105-9-312
950:10.1186/1471-2105-10-60
468:(Web Server): W89–W93.
65:hole of the intestinal
616:10.1006/jmbi.2001.5228
523:10.1002/anie.201506589
462:Nucleic Acids Research
273:10.1002/anie.200290009
238:10.1006/jmbi.2001.5227
48:
910:Motivated Proteins:
875:10.1186/1478-811x-8-8
424:10.1186/1745-6150-3-3
150:iron-sulphur clusters
46:
813:Nature Biotechnology
343:(51): 18281–18295.
83:bacterial cell wall
988:BMC Bioinformatics
937:BMC Bioinformatics
778:10.1002/prot.24038
735:10.1002/prot.23154
692:10.1002/prot.21717
572:10.1038/nchem.2894
474:10.1093/nar/gki414
77:which binds a key
52:Nest conformations
49:
729:(11): 3193–3207.
384:10.1002/bip.20413
349:10.1021/ja0558894
306:(11): 1935–1942.
267:(24): 4663–4665.
261:Angew Chem Int Ed
114:potassium channel
1051:
1024:
1023:
1013:
1003:
979:
973:
972:
962:
952:
928:
898:
897:
887:
877:
853:
847:
846:
836:
825:10.1038/nbt.2672
804:
798:
797:
772:(5): 1418–1424.
761:
755:
754:
718:
712:
711:
675:
669:
668:
658:
649:(9): 1369–1376.
634:
628:
627:
598:
592:
591:
560:Nature Chemistry
551:
545:
544:
534:
517:(6): 1974–1987.
502:
496:
495:
485:
453:
447:
446:
436:
426:
402:
396:
395:
367:
361:
360:
332:
326:
325:
315:
291:
285:
284:
256:
250:
249:
220:
67:serine proteases
1059:
1058:
1054:
1053:
1052:
1050:
1049:
1048:
1029:
1028:
1027:
981:
980:
976:
930:
929:
925:
907:
902:
901:
855:
854:
850:
819:(10): 916–921.
806:
805:
801:
763:
762:
758:
720:
719:
715:
677:
676:
672:
636:
635:
631:
610:(15): 183–191.
600:
599:
595:
553:
552:
548:
504:
503:
499:
455:
454:
450:
404:
403:
399:
369:
368:
364:
334:
333:
329:
293:
292:
288:
258:
257:
253:
222:
221:
217:
212:
162:
133:
122:carbonyl groups
107:
103:
99:
95:
90:dihedral angles
58:hydrogen bonded
54:
12:
11:
5:
1057:
1055:
1047:
1046:
1041:
1031:
1030:
1026:
1025:
974:
922:
921:
920:
914:
906:
905:External links
903:
900:
899:
848:
799:
756:
713:
686:(1): 415–425.
670:
629:
593:
546:
497:
448:
411:Biology Direct
397:
362:
327:
286:
251:
232:(2): 171–182.
214:
213:
211:
208:
174:Schellman loop
161:
158:
132:
131:Compound nests
129:
105:
101:
97:
93:
53:
50:
20:is a type of
13:
10:
9:
6:
4:
3:
2:
1056:
1045:
1042:
1040:
1037:
1036:
1034:
1021:
1017:
1012:
1007:
1002:
997:
993:
989:
985:
978:
975:
970:
966:
961:
956:
951:
946:
942:
938:
934:
927:
924:
918:
915:
912:
909:
908:
904:
895:
891:
886:
881:
876:
871:
867:
863:
859:
852:
849:
844:
840:
835:
830:
826:
822:
818:
814:
810:
803:
800:
795:
791:
787:
783:
779:
775:
771:
767:
760:
757:
752:
748:
744:
740:
736:
732:
728:
724:
717:
714:
709:
705:
701:
697:
693:
689:
685:
681:
674:
671:
666:
662:
657:
652:
648:
644:
640:
633:
630:
625:
621:
617:
613:
609:
605:
597:
594:
589:
585:
581:
577:
573:
569:
565:
561:
557:
550:
547:
542:
538:
533:
528:
524:
520:
516:
512:
508:
501:
498:
493:
489:
484:
479:
475:
471:
467:
463:
459:
452:
449:
444:
440:
435:
430:
425:
420:
416:
412:
408:
401:
398:
393:
389:
385:
381:
377:
373:
366:
363:
358:
354:
350:
346:
342:
338:
331:
328:
323:
319:
314:
309:
305:
301:
297:
290:
287:
282:
278:
274:
270:
266:
262:
255:
252:
247:
243:
239:
235:
231:
227:
219:
216:
209:
207:
205:
200:
198:
194:
190:
186:
181:
179:
175:
170:
168:
160:Types of nest
159:
157:
155:
151:
147:
146:Walker motifs
143:
139:
130:
128:
127:
123:
119:
115:
109:
91:
86:
84:
80:
76:
72:
68:
64:
59:
51:
45:
41:
39:
35:
31:
27:
23:
19:
991:
987:
977:
940:
936:
926:
865:
861:
851:
816:
812:
802:
769:
765:
759:
726:
722:
716:
683:
679:
673:
646:
642:
632:
607:
603:
596:
563:
559:
549:
514:
510:
500:
465:
461:
451:
414:
410:
400:
378:(1): 90–96.
375:
371:
365:
340:
336:
330:
303:
299:
289:
264:
260:
254:
229:
225:
218:
201:
183:A number of
182:
171:
163:
140:, as in the
134:
110:
87:
55:
17:
15:
1044:Tripeptides
916:PDBeMotif:
566:(1): 8–16.
372:Biopolymers
204:PDZ domains
167:alpha helix
126:alpha sheet
79:carboxylate
1033:Categories
994:(1): 312.
210:References
197:threonines
138:phosphates
96:=-90°; psi
75:vancomycin
71:antibiotic
34:amino acid
943:(1): 60.
643:Structure
588:205298633
148:, and in
118:aquaporin
104:=80°; psi
1020:18637174
969:19210785
894:20509869
843:23955275
786:22275093
766:Proteins
743:21989939
723:Proteins
708:43848293
700:17957773
680:Proteins
665:16962968
624:11779238
580:29256514
541:26612067
492:15980588
443:18226248
392:16283656
357:16366583
322:15502299
281:12481319
246:11779237
185:antibody
178:carbonyl
100:=0°; phi
73:peptide
63:oxyanion
30:peptides
26:proteins
1011:2491636
960:2651126
885:2891790
834:3795957
794:5401588
751:8761012
532:4755225
483:1160175
434:2241844
193:serines
38:Proline
1018:
1008:
967:
957:
892:
882:
841:
831:
792:
784:
749:
741:
706:
698:
663:
622:
586:
578:
539:
529:
490:
480:
441:
431:
390:
355:
320:
279:
244:
172:Every
154:P-loop
142:P-loop
108:=20°.
868:: 8.
790:S2CID
747:S2CID
704:S2CID
584:S2CID
417:: 3.
202:Most
1016:PMID
965:PMID
890:PMID
839:PMID
782:PMID
739:PMID
696:PMID
661:PMID
620:PMID
576:PMID
537:PMID
488:PMID
439:PMID
388:PMID
353:PMID
318:PMID
277:PMID
242:PMID
195:and
28:and
18:Nest
16:The
1006:PMC
996:doi
955:PMC
945:doi
880:PMC
870:doi
829:PMC
821:doi
774:doi
731:doi
688:doi
651:doi
612:doi
608:315
568:doi
527:PMC
519:doi
478:PMC
470:doi
429:PMC
419:doi
380:doi
345:doi
341:127
308:doi
304:D60
269:doi
234:doi
230:315
144:or
106:i+1
102:i+1
1035::
1014:.
1004:.
990:.
986:.
963:.
953:.
941:10
939:.
935:.
888:.
878:.
864:.
860:.
837:.
827:.
817:31
815:.
811:.
788:.
780:.
770:80
768:.
745:.
737:.
727:79
725:.
702:.
694:.
684:71
682:.
659:.
647:14
645:.
641:.
618:.
606:.
582:.
574:.
564:10
562:.
558:.
535:.
525:.
515:55
513:.
509:.
486:.
476:.
466:33
464:.
460:.
437:.
427:.
413:.
409:.
386:.
376:84
374:.
351:.
339:.
316:.
302:.
298:.
275:.
265:41
263:.
240:.
228:.
199:.
1022:.
998::
992:9
971:.
947::
919:.
913:;
896:.
872::
866:8
845:.
823::
796:.
776::
753:.
733::
710:.
690::
667:.
653::
626:.
614::
590:.
570::
543:.
521::
494:.
472::
445:.
421::
415:3
394:.
382::
359:.
347::
324:.
310::
283:.
271::
248:.
236::
98:i
94:i
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.