887:
546:
399:
882:{\displaystyle {\ce {H}}{-}{\overset {\displaystyle {\ce {H}} \atop |}{\underset {| \atop \displaystyle {\ce {H}}}{\ce {C}}}}{-}{\overset {\displaystyle {\ce {H}} \atop |}{\underset {| \atop \displaystyle {\ce {H}}}{\ce {C}}}}{\ce {-O-H->H}}{-}{\overset {\displaystyle {\ce {H}} \atop |}{\underset {| \atop \displaystyle {\ce {H}}}{\ce {C}}}}{-}{\overset {\displaystyle {\ce {H}} \atop |}{\underset {\| \atop \displaystyle {\ce {O}}}{\ce {C}}}}{\ce {->H}}{-}{\overset {\displaystyle {\ce {H}} \atop |}{\underset {| \atop \displaystyle {\ce {H}}}{\ce {C}}}}{-}{\overset {\color {white}{\displaystyle {\ce {H}} \atop |}}{\underset {\| \atop \displaystyle {\ce {O}}}{\ce {C}}}}{\ce {-O-H}}}
411:
468:
387:
54:
523:
2725:
910:), the intermediate structures can be toxic, and health problems arise when those intermediates cannot be cleared. When high levels of acetaldehyde occur in the blood, facial flushing, lightheadedness, palpitations, nausea, and general “hangover” symptoms occur. These symptoms are indicative of a medical condition known as the
377:, which functions as a cofactor. Cysteine and glutamate molecules interact with the aldehyde substrate. Many other residues will interact with NAD(P) to hold it in place. Magnesium may be used to help the enzyme function, although the degree to which magnesium assists the enzyme varies between different classes of aldehydes.
398:
499:. The enzyme's active site then goes through an isomorphic change whereby the NAD(P)H is moved, creating room for a water molecule to access the substrate. The water is primed by a glutamate in the active site, and the water makes a nucleophilic attack on the carbonyl carbon, kicking off the sulfur as a
368:
The active site of the aldehyde dehydrogenase enzyme is largely conserved throughout the different classes of the enzyme and, although the number of amino acids present in a subunit can change, the overall function of the site changes little. The active site binds to one molecule of an aldehyde and
944:
for NAD and has a higher maximum velocity than the wild-type allele. This mutation is common in Japan, where 41% of a non-alcoholic control group were ALDH2 deficient, where only 2–5% of an alcoholic group were ALDH2-deficient. In Taiwan, the numbers are similar, with 30% of the control group
1020:
explored aldehyde dehydrogenase inhibition as a pathogenic mechanism in
Parkinson disease. "This ALDH model for PD etiology may help explain the selective vulnerability of dopaminergic neurons in PD and provide a potential mechanism through which environmental toxicants contribute to PD
410:
305:). The oxygen comes from a water molecule. To date, nineteen ALDH genes have been identified within the human genome. These genes participate in a wide variety of biological processes including the detoxification of exogenously and endogenously generated aldehydes.
975:
in lab animals. ALDH2*2 is associated with increased odds of oropharyngolaryngeal, esophageal, gastric, colon, and lung cancer. However, they found no connection between increased levels of ALDH2*2 in the blood and an increased risk of liver cancer.
1708:
Kamino K, Nagasaka K, Imagawa M, Yamamoto H, Yoneda H, Ueki A, Kitamura S, Namekata K, Miki T, Ohta S (June 2000). "Deficiency in mitochondrial aldehyde dehydrogenase increases the risk for late-onset
Alzheimer's disease in the Japanese population".
1027:
models further confirm the involvement of ALDH family in neurodegeneration. Mice null for ALDH1a1 and ALDH2 exhibit
Parkinson's disease-like age-dependent deficits in motor performance and significant increase in biogenic aldehydes.
1327:
Liu ZJ, Sun YJ, Rose J, Chung YJ, Hsiao CD, Chang WR, Kuo I, Perozich J, Lindahl R, Hempel J, Wang BC (April 1997). "The first structure of an aldehyde dehydrogenase reveals novel interactions between NAD and the
Rossmann fold".
952:, which is why disulfiram is used to treat alcoholism. The patients show higher blood levels of acetaldehyde, and become violently ill upon consumption of even small amounts of alcohol. Several drugs (e.g.,
1031:
The ALDH2-/- mice display age-related memory deficits in various tasks, as well as endothelial dysfunction, brain atrophy, and other
Alzheimer's disease-associated pathologies, including marked increases in
945:
showing the deficiency and 6% of alcoholics displaying it. The deficiency is manifested by slow acetaldehyde removal, with low alcohol tolerance perhaps leading to a lower frequency of alcoholism.
404:
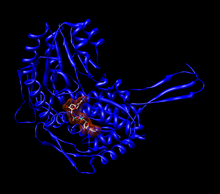
The active site of a human mitochondrial aldehyde dehydrogenase 2. Cys302 and Glu268 interact with the aldehyde substrate. The NAD is held in place by multiple residues (shown as wires or sticks).
386:
986:
found that ALDH1 is a potentially important, poor prognostic factor in breast cancer, associated with high histological grade, estrogen/progesteron receptor negativity and HER2 positivity.
2235:
997:
gene (the odds for LOAD in carriers of ALDH2*2 allele almost twice that of non-carriers). Moreover, ALDH gene, protein expression and activity are substantially decreased in the
2367:
1828:
Fitzmaurice AG, Rhodes SL, Lulla A, Murphy NP, Lam HA, O'Donnell KC, Barnhill L, Casida JE, Cockburn M, Sagasti A, Stahl MC, Maidment NT, Ritz B, Bronstein JM (January 2013).
1048:. These behavioral and biochemical Alzheimer's disease-like deficits were efficiently ameliorated when the ALDH2-/- mice were treated with isotope-reinforced, deuterated
2030:
231:
971:
and oropharyngolaryngeal cancers. The metabolized acetaldehyde in the blood, which is six times higher than in individuals without the mutation, has shown to be a
2291:
250:
2266:
2195:
374:
2228:
2185:
2170:
2165:
2160:
2129:
416:
The active site of the K487E mutant aldehyde dehydrogenase 2 with a space-filling model of NAD in the active site. The amino acid Glu349 is highlighted.
2202:
1889:"Neurodegeneration and motor dysfunction in mice lacking cytosolic and mitochondrial aldehyde dehydrogenases: implications for Parkinson's disease"
31:
467:
1557:
Yokoyama A, Muramatsu T, Ohmori T, Yokoyama T, Okuyama K, Takahashi H, Hasegawa Y, Higuchi S, Maruyama K, Shirakura K, Ishii H (August 1998).
2221:
2023:
1399:
1288:"Overview of the role of alcohol dehydrogenase and aldehyde dehydrogenase and their variants in the genesis of alcohol-related pathology"
979:
High expression of the genes that encode ALDH1A1 and ALDH2 is associated with a poor prognosis in patients with acute myeloid leukemia.
967:
found that decreased enzyme activity of aldehyde dehydrogenase-2, caused by the mutated ALDH2 allele, contributes to a higher chance of
514:
extract inhibited aldehyde dehydrogenase activity, lending credence to an Asian folklore warning against consuming durian with alcohol.
2357:
2313:
352:, such as those expressed in tumors, stomach, and cornea). In all three classes, constitutive and inducible forms exist. ALDH1 and
2444:
2284:
2262:
2134:
370:
67:
1371:
2016:
243:
2387:
170:
2600:
194:
2715:
898:
ALDH2 plays a crucial role in maintaining low blood levels of acetaldehyde during alcohol oxidation. In this pathway (
1948:"Deuterium-reinforced polyunsaturated fatty acids improve cognition in a mouse model of sporadic Alzheimer's disease"
1185:
Perez-Miller SJ, Hurley TD (June 2003). "Coenzyme isomerization is integral to catalysis in aldehyde dehydrogenase".
2279:
1049:
2585:
2701:
2688:
2675:
2662:
2649:
2636:
2623:
2399:
2345:
2323:
2301:
2258:
1744:
Grünblatt E, Riederer P (February 2016). "Aldehyde dehydrogenase (ALDH) in
Alzheimer's and Parkinson's disease".
360:
subunits. These enzymes are found in many tissues of the body but are at the highest concentration in the liver.
2595:
2549:
2492:
2249:
2043:
2002:
957:
271:
188:
81:
2403:
933:
for the mutation have reduced activity. Thus, the mutation is partially dominant. The ineffective homozygous
2497:
1475:
Thomasson HR, Edenberg HJ, Crabb DW, Mai XL, Jerome RE, Li TK, Wang SP, Lin YT, Lu RB, Yin SJ (April 1991).
175:
2362:
1002:
990:
911:
314:
2518:
2437:
1148:
531:
507:
255:
2590:
163:
53:
356:
are the most important enzymes for aldehyde oxidation, and both are tetrameric enzymes composed of 54
1900:
1841:
98:
2554:
484:
93:
63:
1998:
1946:
Elharram A, Czegledy NM, Golod M, Milne GL, Pollock E, Bennett BM, Shchepinov MS (December 2017).
948:
These symptoms are the same as those observed in people who drink while being treated by the drug
191:
2487:
1769:
1353:
1033:
330:
115:
1417:"Alcohol Dehydrogenases, Aldehyde Dehydrogenases, and Alcohol Use Disorders: A Critical Review"
2750:
2411:
1977:
1928:
1869:
1810:
1761:
1726:
1690:
1672:
1631:
1580:
1539:
1518:"Structure of mitochondrial aldehyde dehydrogenase: the genetic component of ethanol aversion"
1498:
1454:
1436:
1395:
1345:
1309:
1251:
1202:
1175:
1010:
968:
460:, and interactions between the cofactor and the fold allow for the action of the active site.
182:
1649:
Demir, Hale; Dulgar, Ozgecan; Gulle, Bugra Taygun; Turna, Hande; Ilvan, Sennur (2018-11-07).
2745:
2533:
2528:
2502:
2430:
2335:
1967:
1959:
1918:
1908:
1859:
1849:
1800:
1753:
1718:
1680:
1662:
1621:
1611:
1570:
1529:
1488:
1444:
1428:
1337:
1299:
1241:
1233:
1194:
1006:
998:
994:
326:
38:
989:
Some case-control studies claimed that carriage of ALDH2*2 allele was a risk of late-onset
392:
Tetramer of aldehyde dehydrogenase 2 with a space filling model of NAD in each active site.
2580:
2564:
2477:
1600:"Aldehyde Dehydrogenase Genes as Prospective Actionable Targets in Acute Myeloid Leukemia"
441:
298:
151:
2213:
1904:
1845:
1626:
1599:
522:
127:
2729:
2618:
2559:
2245:
1972:
1947:
1923:
1888:
1864:
1829:
1805:
1788:
1685:
1650:
1493:
1476:
1449:
1416:
1388:
1246:
1221:
1024:
527:
456:
through a channel extending from the surface of the enzyme. The active site contains a
226:
86:
1534:
1517:
206:
2739:
2523:
2482:
2104:
2008:
953:
535:
500:
457:
357:
201:
1773:
1651:"Prognostic value of aldehyde dehydrogenase 1 (ALDH1) in invasive breast carcinomas"
1357:
2472:
1037:
930:
903:
1830:"Aldehyde dehydrogenase inhibition as a pathogenic mechanism in Parkinson disease"
1913:
2696:
2631:
2467:
453:
210:
2724:
1834:
Proceedings of the
National Academy of Sciences of the United States of America
1575:
1558:
2382:
2305:
1757:
972:
949:
926:
329:. There are three different classes of these enzymes in mammals: class 1 (low
1887:
Wey MC, Fernandez E, Martinez PA, Sullivan P, Goldstein DS, Strong R (2012).
1676:
1667:
1598:
Dancik, Garrett; Varisli, Lokman; Tolan, Veysel; Vlahopoulos, Spiros (2023).
1559:"Alcohol-related cancers and aldehyde dehydrogenase-2 in Japanese alcoholics"
1440:
1237:
1222:"Non-P450 aldehyde oxidizing enzymes: the aldehyde dehydrogenase superfamily"
2670:
2644:
2349:
1854:
1477:"Alcohol and aldehyde dehydrogenase genotypes and alcoholism in Chinese men"
929:
individuals with the mutant allele have almost no ALDH2 activity, and those
922:
917:
There is a mutant form of aldehyde dehydrogenase, termed ALDH2*2, wherein a
322:
318:
286:
1981:
1932:
1873:
1814:
1765:
1730:
1722:
1694:
1635:
1458:
1313:
1255:
1206:
1616:
1584:
1543:
1502:
1349:
2190:
1136:
1132:
1045:
488:
433:
290:
282:
1386:
Rod Flower; Humphrey P. Rang; Maureen M. Dale; Ritter, James M. (2007).
1341:
937:
works at a rate of about 8% of the normal allele, for it shows a higher
2180:
2175:
2144:
2139:
2091:
2086:
2078:
2071:
2066:
2061:
1963:
1432:
1304:
1287:
1128:
1124:
1120:
1116:
1112:
1108:
1103:
1099:
1095:
1091:
1081:
1077:
1073:
1069:
1065:
1061:
907:
899:
496:
492:
274:
158:
139:
1198:
1179:
2683:
2453:
2377:
2372:
2327:
1005:
patients. These reports are in line with findings implementing toxic
934:
918:
511:
480:
278:
238:
134:
122:
110:
762:
661:
17:
1789:"Neurodegeneration and aldehyde load: from concept to therapeutics"
2657:
2113:
1086:
1041:
429:
The overall reaction catalysed by the aldehyde dehydrogenases is:
353:
59:
445:
146:
2426:
2217:
2012:
914:, also known as “Asian flush” or “Oriental flushing syndrome”.
466:
1220:
Marchitti SA, Brocker C, Stagos D, Vasiliou V (June 2008).
452:
In this NAD(P)-dependent reaction, the aldehyde enters the
2422:
891:
Metabolism of alcohol (ethanol) to acetaldehyde (ethanal)
491:
carbon of the aldehyde. The hydrogen is kicked off as a
1286:
Crabb DW, Matsumoto M, Chang D, You M (February 2004).
2713:
1516:
Steinmetz CG, Xie P, Weiner H, Hurley TD (May 1997).
1415:
Edenberg, Howard J.; McClintick, Jeanette N. (2018).
844:
832:
802:
792:
741:
731:
701:
691:
624:
614:
582:
572:
549:
526:
Role of aldehyde dehydrogenase (shown in red box) in
2368:
Branched-chain alpha-keto acid dehydrogenase complex
2609:
2573:
2542:
2511:
2460:
2398:
2344:
2322:
2300:
2257:
2153:
2122:
2102:
2051:
1711:
1372:"Durians and booze: Worse than a stinking hangover"
249:
237:
225:
220:
200:
181:
169:
157:
145:
133:
121:
109:
104:
92:
80:
75:
46:
1387:
1226:Expert Opinion on Drug Metabolism & Toxicology
881:
1470:
1468:
2292:Mycothiol-dependent formaldehyde dehydrogenase
1421:Alcoholism: Clinical and Experimental Research
1170:
1168:
1166:
1164:
925:in the active site at position 487 of ALDH2.
2438:
2229:
2024:
518:Pathology (aldehyde dehydrogenase deficiency)
8:
829:
728:
1281:
1279:
1277:
1275:
1273:
1271:
1269:
1267:
1265:
483:from a cysteine in the active site makes a
58:Monomer of human aldehyde dehydrogenase 2 (
2445:
2431:
2423:
2236:
2222:
2214:
2031:
2017:
2009:
217:
52:
2001:at the U.S. National Library of Medicine
1971:
1922:
1912:
1863:
1853:
1804:
1684:
1666:
1655:Bosnian Journal of Basic Medical Sciences
1625:
1615:
1574:
1533:
1492:
1448:
1303:
1245:
870:
862:
861:
851:
845:
842:
833:
822:
817:
809:
803:
793:
787:
780:
775:
763:
757:
756:
748:
742:
732:
721:
716:
708:
702:
692:
686:
679:
674:
662:
656:
648:
640:
639:
631:
625:
615:
609:
602:
597:
589:
583:
573:
567:
560:
555:
550:
548:
1793:Journal of Psychiatry & Neuroscience
1292:The Proceedings of the Nutrition Society
521:
2720:
1160:
893:and then to acetic acid (ethanoic acid)
382:
293:. They convert aldehydes (R–C(=O)
32:Aldehyde dehydrogenase (disambiguation)
2176:α-Aminoadipic semialdehyde — (ALDH7A1)
43:
7:
2203:Glyceraldehyde 3-phosphate — (GAPDH)
1394:. Edinburgh: Churchill Livingstone.
956:) cause a similar reaction known as
345:, mitochondrial), and class 3 (high
2130:Dimeric NADP-preferring — (ALDH3A1)
841:
471:Mechanism of Aldehyde Dehydrogenase
2358:Oxoglutarate dehydrogenase complex
2314:Formate dehydrogenase (cytochrome)
1481:American Journal of Human Genetics
843:
828:
801:
786:
740:
727:
700:
685:
623:
608:
581:
566:
25:
2285:Long-chain-aldehyde dehydrogenase
1007:lipid oxidation-derived aldehydes
2723:
409:
397:
385:
1746:Journal of Neural Transmission
1390:Rang & Dale's pharmacology
852:
810:
788:
749:
709:
687:
632:
610:
590:
568:
1:
1535:10.1016/S0969-2126(97)00224-4
47:Aldehyde dehydrogenase (NAD+)
1914:10.1371/journal.pone.0031522
313:Aldehyde dehydrogenase is a
1050:polyunsaturated fatty acids
495:and attacks NAD(P) to make
338:, cytosolic), class 2 (low
317:enzyme responsible for the
2767:
2280:Acetaldehyde dehydrogenase
2135:Fatty aldehyde — (ALDH3A2)
1787:Wood PL (September 2006).
36:
29:
2601:Michaelis–Menten kinetics
2083:Formyltetrahydrofolate —
1758:10.1007/s00702-014-1320-1
1330:Nature Structural Biology
1009:in these diseases and in
216:
51:
2493:Diffusion-limited enzyme
2003:Medical Subject Headings
1668:10.17305/bjbms.2018.3094
1576:10.1093/carcin/19.8.1383
1238:10.1517/17425255.4.6.697
958:disulfiram-like reaction
37:Not to be confused with
2040:Aldehyde dehydrogenases
1855:10.1073/pnas.1220399110
369:one molecule of either
268:Aldehyde dehydrogenases
2363:Pyruvate dehydrogenase
2275:Aldehyde dehydrogenase
1999:Aldehyde+dehydrogenase
1723:10.1006/bbrc.2000.2923
912:alcohol flush reaction
883:
539:
530:degradation, creating
472:
2586:Eadie–Hofstee diagram
2519:Allosteric regulation
1617:10.3390/genes14091807
1149:Alcohol dehydrogenase
884:
532:vanillylmandelic acid
525:
508:University of Tsukuba
470:
2596:Lineweaver–Burk plot
547:
30:For other uses, see
2404:iron–sulfur protein
1905:2012PLoSO...731522W
1846:2013PNAS..110..636F
1342:10.1038/nsb0497-317
1003:Parkinson's disease
993:independent of the
991:Alzheimer's disease
921:residue replaces a
768:
667:
506:Researchers at the
485:nucleophilic attack
70:in the active site.
64:space-filling model
2555:Enzyme superfamily
2488:Enzyme promiscuity
1964:10.1111/febs.14291
1433:10.1111/acer.13904
1305:10.1079/PNS2003327
1034:lipid peroxidation
879:
858:
850:
840:
838:
808:
800:
798:
747:
739:
737:
707:
699:
697:
630:
622:
620:
588:
580:
578:
540:
473:
2711:
2710:
2420:
2419:
2412:Pyruvate synthase
2211:
2210:
1958:(23): 4083–4095.
1427:(12): 2281–2297.
1401:978-0-443-06911-6
1199:10.1021/bi034182w
1011:neurodegeneration
877:
869:
859:
856:
848:
839:
836:
827:
824:
815:
814:
806:
799:
796:
785:
782:
773:
769:
766:
754:
753:
745:
738:
735:
726:
723:
714:
713:
705:
698:
695:
684:
681:
672:
668:
665:
655:
647:
637:
636:
628:
621:
618:
607:
604:
595:
594:
586:
579:
576:
565:
562:
553:
477:
476:
277:) are a group of
265:
264:
261:
260:
164:metabolic pathway
16:(Redirected from
2758:
2728:
2727:
2719:
2591:Hanes–Woolf plot
2534:Enzyme activator
2529:Enzyme inhibitor
2503:Enzyme catalysis
2447:
2440:
2433:
2424:
2336:Aldehyde oxidase
2238:
2231:
2224:
2215:
2058:Retinaldehyde —
2033:
2026:
2019:
2010:
1986:
1985:
1975:
1952:The FEBS Journal
1943:
1937:
1936:
1926:
1916:
1884:
1878:
1877:
1867:
1857:
1825:
1819:
1818:
1808:
1784:
1778:
1777:
1741:
1735:
1734:
1705:
1699:
1698:
1688:
1670:
1646:
1640:
1639:
1629:
1619:
1595:
1589:
1588:
1578:
1554:
1548:
1547:
1537:
1513:
1507:
1506:
1496:
1472:
1463:
1462:
1452:
1412:
1406:
1405:
1393:
1384:Figure 11-4 in:
1382:
1376:
1375:
1368:
1362:
1361:
1324:
1318:
1317:
1307:
1283:
1260:
1259:
1249:
1217:
1211:
1210:
1182:
1172:
999:substantia nigra
995:apolipoprotein E
888:
886:
885:
880:
878:
875:
874:
867:
866:
860:
857:
855:
849:
846:
837:
834:
825:
823:
821:
816:
813:
807:
804:
797:
794:
791:
783:
781:
779:
774:
771:
770:
767:
764:
758:
755:
752:
746:
743:
736:
733:
724:
722:
720:
715:
712:
706:
703:
696:
693:
690:
682:
680:
678:
673:
670:
669:
666:
663:
657:
653:
652:
645:
644:
638:
635:
629:
626:
619:
616:
613:
605:
603:
601:
596:
593:
587:
584:
577:
574:
571:
563:
561:
559:
554:
551:
463:
462:
413:
401:
389:
327:carboxylic acids
304:
303:–O–H
299:carboxylic acids
296:
218:
56:
44:
39:aldehyde oxidase
27:Group of enzymes
21:
2766:
2765:
2761:
2760:
2759:
2757:
2756:
2755:
2736:
2735:
2734:
2722:
2714:
2712:
2707:
2619:Oxidoreductases
2605:
2581:Enzyme kinetics
2569:
2565:List of enzymes
2538:
2507:
2478:Catalytic triad
2456:
2451:
2421:
2416:
2394:
2340:
2318:
2296:
2253:
2246:oxidoreductases
2242:
2212:
2207:
2149:
2118:
2098:
2047:
2037:
1995:
1990:
1989:
1945:
1944:
1940:
1886:
1885:
1881:
1827:
1826:
1822:
1786:
1785:
1781:
1743:
1742:
1738:
1707:
1706:
1702:
1648:
1647:
1643:
1597:
1596:
1592:
1556:
1555:
1551:
1515:
1514:
1510:
1474:
1473:
1466:
1414:
1413:
1409:
1402:
1385:
1383:
1379:
1370:
1369:
1365:
1326:
1325:
1321:
1285:
1284:
1263:
1219:
1218:
1214:
1184:
1174:
1173:
1162:
1157:
1145:
1058:
1021:pathogenesis."
943:
896:
895:
894:
892:
889:
545:
544:
520:
439:
427:
422:
421:
420:
417:
414:
405:
402:
393:
390:
366:
351:
344:
336:
311:
302:
294:
71:
42:
35:
28:
23:
22:
15:
12:
11:
5:
2764:
2762:
2754:
2753:
2748:
2738:
2737:
2733:
2732:
2709:
2708:
2706:
2705:
2692:
2679:
2666:
2653:
2640:
2627:
2613:
2611:
2607:
2606:
2604:
2603:
2598:
2593:
2588:
2583:
2577:
2575:
2571:
2570:
2568:
2567:
2562:
2557:
2552:
2546:
2544:
2543:Classification
2540:
2539:
2537:
2536:
2531:
2526:
2521:
2515:
2513:
2509:
2508:
2506:
2505:
2500:
2495:
2490:
2485:
2480:
2475:
2470:
2464:
2462:
2458:
2457:
2452:
2450:
2449:
2442:
2435:
2427:
2418:
2417:
2415:
2414:
2408:
2406:
2396:
2395:
2393:
2392:
2391:
2390:
2385:
2380:
2375:
2365:
2360:
2354:
2352:
2342:
2341:
2339:
2338:
2332:
2330:
2320:
2319:
2317:
2316:
2310:
2308:
2298:
2297:
2295:
2294:
2289:
2288:
2287:
2282:
2271:
2269:
2255:
2254:
2243:
2241:
2240:
2233:
2226:
2218:
2209:
2208:
2206:
2205:
2199:
2198:
2193:
2188:
2183:
2178:
2173:
2168:
2163:
2157:
2155:
2151:
2150:
2148:
2147:
2142:
2137:
2132:
2126:
2124:
2120:
2119:
2117:
2116:
2110:
2108:
2100:
2099:
2097:
2096:
2095:
2094:
2089:
2081:
2076:
2075:
2074:
2069:
2064:
2055:
2053:
2049:
2048:
2038:
2036:
2035:
2028:
2021:
2013:
2007:
2006:
1994:
1993:External links
1991:
1988:
1987:
1938:
1879:
1820:
1779:
1736:
1700:
1661:(4): 313–319.
1641:
1590:
1563:Carcinogenesis
1549:
1508:
1464:
1407:
1400:
1377:
1363:
1319:
1261:
1232:(6): 697–720.
1212:
1193:(23): 7100–9.
1159:
1158:
1156:
1153:
1152:
1151:
1144:
1141:
1140:
1139:
1106:
1089:
1084:
1057:
1054:
1044:and activated
1025:Knockout mouse
941:
890:
873:
865:
854:
831:
820:
812:
790:
778:
761:
751:
730:
719:
711:
689:
677:
660:
651:
643:
634:
612:
600:
592:
570:
558:
543:
542:
541:
528:norepinephrine
519:
516:
475:
474:
450:
449:
437:
426:
423:
419:
418:
415:
408:
406:
403:
396:
394:
391:
384:
381:
380:
379:
365:
362:
349:
342:
334:
310:
307:
301:(R–C(=O)
263:
262:
259:
258:
253:
247:
246:
241:
235:
234:
229:
223:
222:
214:
213:
204:
198:
197:
186:
179:
178:
173:
167:
166:
161:
155:
154:
149:
143:
142:
137:
131:
130:
125:
119:
118:
113:
107:
106:
102:
101:
96:
90:
89:
84:
78:
77:
73:
72:
57:
49:
48:
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
2763:
2752:
2749:
2747:
2744:
2743:
2741:
2731:
2726:
2721:
2717:
2703:
2699:
2698:
2693:
2690:
2686:
2685:
2680:
2677:
2673:
2672:
2667:
2664:
2660:
2659:
2654:
2651:
2647:
2646:
2641:
2638:
2634:
2633:
2628:
2625:
2621:
2620:
2615:
2614:
2612:
2608:
2602:
2599:
2597:
2594:
2592:
2589:
2587:
2584:
2582:
2579:
2578:
2576:
2572:
2566:
2563:
2561:
2560:Enzyme family
2558:
2556:
2553:
2551:
2548:
2547:
2545:
2541:
2535:
2532:
2530:
2527:
2525:
2524:Cooperativity
2522:
2520:
2517:
2516:
2514:
2510:
2504:
2501:
2499:
2496:
2494:
2491:
2489:
2486:
2484:
2483:Oxyanion hole
2481:
2479:
2476:
2474:
2471:
2469:
2466:
2465:
2463:
2459:
2455:
2448:
2443:
2441:
2436:
2434:
2429:
2428:
2425:
2413:
2410:
2409:
2407:
2405:
2401:
2397:
2389:
2386:
2384:
2381:
2379:
2376:
2374:
2371:
2370:
2369:
2366:
2364:
2361:
2359:
2356:
2355:
2353:
2351:
2347:
2343:
2337:
2334:
2333:
2331:
2329:
2325:
2321:
2315:
2312:
2311:
2309:
2307:
2303:
2299:
2293:
2290:
2286:
2283:
2281:
2278:
2277:
2276:
2273:
2272:
2270:
2268:
2264:
2260:
2256:
2251:
2247:
2244:Aldehyde/oxo
2239:
2234:
2232:
2227:
2225:
2220:
2219:
2216:
2204:
2201:
2200:
2197:
2194:
2192:
2189:
2187:
2184:
2182:
2179:
2177:
2174:
2172:
2169:
2167:
2164:
2162:
2159:
2158:
2156:
2152:
2146:
2143:
2141:
2138:
2136:
2133:
2131:
2128:
2127:
2125:
2121:
2115:
2112:
2111:
2109:
2106:
2105:mitochondrial
2101:
2093:
2090:
2088:
2085:
2084:
2082:
2080:
2077:
2073:
2070:
2068:
2065:
2063:
2060:
2059:
2057:
2056:
2054:
2050:
2045:
2041:
2034:
2029:
2027:
2022:
2020:
2015:
2014:
2011:
2004:
2000:
1997:
1996:
1992:
1983:
1979:
1974:
1969:
1965:
1961:
1957:
1953:
1949:
1942:
1939:
1934:
1930:
1925:
1920:
1915:
1910:
1906:
1902:
1899:(2): e31522.
1898:
1894:
1890:
1883:
1880:
1875:
1871:
1866:
1861:
1856:
1851:
1847:
1843:
1840:(2): 636–41.
1839:
1835:
1831:
1824:
1821:
1816:
1812:
1807:
1802:
1798:
1794:
1790:
1783:
1780:
1775:
1771:
1767:
1763:
1759:
1755:
1751:
1747:
1740:
1737:
1732:
1728:
1724:
1720:
1716:
1712:
1704:
1701:
1696:
1692:
1687:
1682:
1678:
1674:
1669:
1664:
1660:
1656:
1652:
1645:
1642:
1637:
1633:
1628:
1623:
1618:
1613:
1609:
1605:
1604:Genes (Basel)
1601:
1594:
1591:
1586:
1582:
1577:
1572:
1569:(8): 1383–7.
1568:
1564:
1560:
1553:
1550:
1545:
1541:
1536:
1531:
1528:(5): 701–11.
1527:
1523:
1519:
1512:
1509:
1504:
1500:
1495:
1490:
1487:(4): 677–81.
1486:
1482:
1478:
1471:
1469:
1465:
1460:
1456:
1451:
1446:
1442:
1438:
1434:
1430:
1426:
1422:
1418:
1411:
1408:
1403:
1397:
1392:
1391:
1381:
1378:
1373:
1367:
1364:
1359:
1355:
1351:
1347:
1343:
1339:
1336:(4): 317–26.
1335:
1331:
1323:
1320:
1315:
1311:
1306:
1301:
1297:
1293:
1289:
1282:
1280:
1278:
1276:
1274:
1272:
1270:
1268:
1266:
1262:
1257:
1253:
1248:
1243:
1239:
1235:
1231:
1227:
1223:
1216:
1213:
1208:
1204:
1200:
1196:
1192:
1188:
1181:
1177:
1171:
1169:
1167:
1165:
1161:
1154:
1150:
1147:
1146:
1142:
1138:
1134:
1130:
1126:
1122:
1118:
1114:
1110:
1107:
1105:
1101:
1097:
1093:
1090:
1088:
1085:
1083:
1079:
1075:
1071:
1067:
1063:
1060:
1059:
1055:
1053:
1051:
1047:
1043:
1039:
1035:
1029:
1026:
1022:
1019:
1014:
1012:
1008:
1004:
1000:
996:
992:
987:
985:
980:
977:
974:
970:
966:
961:
959:
955:
954:metronidazole
951:
946:
940:
936:
932:
928:
924:
920:
915:
913:
909:
905:
901:
871:
863:
818:
776:
759:
717:
675:
658:
649:
641:
598:
556:
537:
536:catecholamine
533:
529:
524:
517:
515:
513:
509:
504:
502:
501:leaving group
498:
494:
490:
486:
482:
469:
465:
464:
461:
459:
458:Rossmann fold
455:
447:
443:
435:
432:
431:
430:
424:
412:
407:
400:
395:
388:
383:
378:
376:
372:
363:
361:
359:
355:
348:
341:
337:
333:
328:
324:
320:
316:
308:
306:
300:
292:
288:
284:
280:
276:
273:
269:
257:
254:
252:
248:
245:
242:
240:
236:
233:
230:
228:
224:
219:
215:
212:
208:
205:
203:
202:Gene Ontology
199:
196:
193:
190:
187:
184:
180:
177:
174:
172:
168:
165:
162:
160:
156:
153:
150:
148:
144:
141:
140:NiceZyme view
138:
136:
132:
129:
126:
124:
120:
117:
114:
112:
108:
103:
100:
97:
95:
91:
88:
85:
83:
79:
74:
69:
65:
61:
55:
50:
45:
40:
33:
19:
2697:Translocases
2694:
2681:
2668:
2655:
2642:
2632:Transferases
2629:
2616:
2473:Binding site
2274:
2039:
1955:
1951:
1941:
1896:
1892:
1882:
1837:
1833:
1823:
1799:(5): 296–7.
1796:
1792:
1782:
1752:(2): 83–90.
1749:
1745:
1739:
1717:(1): 192–6.
1714:
1710:
1703:
1658:
1654:
1644:
1607:
1603:
1593:
1566:
1562:
1552:
1525:
1521:
1511:
1484:
1480:
1424:
1420:
1410:
1389:
1380:
1366:
1333:
1329:
1322:
1298:(1): 49–63.
1295:
1291:
1229:
1225:
1215:
1190:
1187:Biochemistry
1186:
1038:amyloid-beta
1030:
1023:
1017:
1016:Fitzmaurice
1015:
1013:in general.
988:
983:
981:
978:
964:
962:
947:
938:
931:heterozygous
916:
904:acetaldehyde
897:
534:, the major
505:
478:
451:
428:
367:
346:
339:
331:
312:
267:
266:
128:BRENDA entry
2468:Active site
1610:(9): 1807.
538:metabolite.
510:found that
454:active site
364:Active site
315:polymorphic
116:IntEnz view
76:Identifiers
2740:Categories
2671:Isomerases
2645:Hydrolases
2512:Regulation
2306:cytochrome
2103:Family 2 (
1155:References
1052:(D-PUFA).
1036:products,
973:carcinogen
969:esophageal
950:disulfiram
927:Homozygous
185:structures
152:KEGG entry
99:9028-86-8
2550:EC number
2350:disulfide
1677:1840-4812
1522:Structure
1441:1530-0277
1183:;
963:Yokoyama
923:glutamate
872:−
864:−
830:‖
819:−
777:−
729:‖
718:−
676:−
650:−
642:−
599:−
557:−
436:+ NAD + H
425:Mechanism
323:aldehydes
319:oxidation
291:aldehydes
287:oxidation
105:Databases
62:) with a
2751:EC 1.2.1
2574:Kinetics
2498:Cofactor
2461:Activity
2196:ALDH18A1
2191:ALDH16A1
2123:Family 3
2052:Family 1
1982:29024570
1933:22384032
1893:PLOS ONE
1874:23267077
1815:16951732
1774:24270982
1766:25298080
1731:10873585
1695:29924962
1636:37761947
1627:10531322
1459:30320893
1358:21436007
1314:15099407
1256:18611112
1207:12795606
1143:See also
1137:ALDH18A1
1133:ALDH16A1
1046:caspases
760:→
659:→
489:carbonyl
309:Function
295:–H
283:catalyse
256:proteins
244:articles
232:articles
189:RCSB PDB
2746:Enzymes
2730:Biology
2684:Ligases
2454:Enzymes
2186:ALDH9A1
2181:ALDH8A1
2171:ALDH6A1
2166:ALDH5A1
2161:ALDH4A1
2145:ALDH3B2
2140:ALDH3B1
2092:ALDH1L2
2087:ALDH1L1
2079:ALDH1B1
2072:ALDH1A3
2067:ALDH1A2
2062:ALDH1A1
1973:5716852
1924:3284575
1901:Bibcode
1865:3545765
1842:Bibcode
1806:1557683
1686:6252102
1585:9744533
1544:9195888
1503:2014795
1494:1682953
1450:6286250
1350:9095201
1247:2658643
1129:ALDH9A1
1125:ALDH8A1
1121:ALDH7A1
1117:ALDH6A1
1113:ALDH5A1
1109:ALDH4A1
1104:ALDH3B2
1100:ALDH3B1
1096:ALDH3A2
1092:ALDH3A1
1082:ALDH1L2
1078:ALDH1L1
1074:ALDH1B1
1070:ALDH1A3
1066:ALDH1A2
1062:ALDH1A1
908:acetate
900:ethanol
497:NAD(P)H
493:hydride
487:on the
279:enzymes
275:1.2.1.3
211:QuickGO
176:profile
159:MetaCyc
94:CAS no.
87:1.2.1.3
2716:Portal
2658:Lyases
2378:BCKDHB
2373:BCKDHA
2328:oxygen
2046:1.2.1)
2005:(MeSH)
1980:
1970:
1931:
1921:
1872:
1862:
1813:
1803:
1772:
1764:
1729:
1693:
1683:
1675:
1634:
1624:
1583:
1542:
1501:
1491:
1457:
1447:
1439:
1398:
1356:
1348:
1312:
1254:
1244:
1205:
1018:et al.
984:et al.
982:Demir
965:et al.
935:allele
919:lysine
512:durian
481:sulfur
239:PubMed
221:Search
207:AmiGO
195:PDBsum
135:ExPASy
123:BRENDA
111:IntEnz
82:EC no.
2610:Types
2400:1.2.7
2346:1.2.4
2324:1.2.3
2302:1.2.2
2259:1.2.1
2154:Other
2114:ALDH2
1770:S2CID
1354:S2CID
1087:ALDH2
1056:Genes
1042:p-tau
442:RCOOH
354:ALDH2
297:) to
281:that
171:PRIAM
60:ALDH2
2702:list
2695:EC7
2689:list
2682:EC6
2676:list
2669:EC5
2663:list
2656:EC4
2650:list
2643:EC3
2637:list
2630:EC2
2624:list
2617:EC1
2267:NADP
2252:1.2)
1978:PMID
1929:PMID
1870:PMID
1811:PMID
1762:PMID
1727:PMID
1691:PMID
1673:ISSN
1632:PMID
1581:PMID
1540:PMID
1499:PMID
1455:PMID
1437:ISSN
1396:ISBN
1346:PMID
1310:PMID
1252:PMID
1203:PMID
1180:1o02
765:ALDH
446:NADH
440:O →
434:RCHO
375:NADP
285:the
251:NCBI
192:PDBe
147:KEGG
68:NAD+
18:ALDH
2388:DLD
2383:DBT
2265:or
2263:NAD
1968:PMC
1960:doi
1956:284
1919:PMC
1909:doi
1860:PMC
1850:doi
1838:110
1801:PMC
1754:doi
1750:123
1719:doi
1715:273
1681:PMC
1663:doi
1622:PMC
1612:doi
1571:doi
1530:doi
1489:PMC
1445:PMC
1429:doi
1338:doi
1300:doi
1242:PMC
1234:doi
1195:doi
1176:PDB
1001:of
906:to
902:to
664:ADH
448:+ H
373:or
371:NAD
358:kDa
325:to
321:of
289:of
227:PMC
183:PDB
66:of
2742::
2402::
2348::
2326::
2304::
2261::
2250:EC
2044:EC
1976:.
1966:.
1954:.
1950:.
1927:.
1917:.
1907:.
1895:.
1891:.
1868:.
1858:.
1848:.
1836:.
1832:.
1809:.
1797:31
1795:.
1791:.
1768:.
1760:.
1748:.
1725:.
1713:.
1689:.
1679:.
1671:.
1659:18
1657:.
1653:.
1630:.
1620:.
1608:14
1606:.
1602:.
1579:.
1567:19
1565:.
1561:.
1538:.
1524:.
1520:.
1497:.
1485:48
1483:.
1479:.
1467:^
1453:.
1443:.
1435:.
1425:42
1423:.
1419:.
1352:.
1344:.
1332:.
1308:.
1296:63
1294:.
1290:.
1264:^
1250:.
1240:.
1228:.
1224:.
1201:.
1191:42
1189:.
1178::
1163:^
1135:,
1131:,
1127:,
1123:,
1119:,
1115:,
1111:,
1102:,
1098:,
1094:,
1080:,
1076:,
1072:,
1068:,
1064:,
1040:,
960:.
503:.
479:A
444:+
272:EC
209:/
2718::
2704:)
2700:(
2691:)
2687:(
2678:)
2674:(
2665:)
2661:(
2652:)
2648:(
2639:)
2635:(
2626:)
2622:(
2446:e
2439:t
2432:v
2248:(
2237:e
2230:t
2223:v
2107:)
2042:(
2032:e
2025:t
2018:v
1984:.
1962::
1935:.
1911::
1903::
1897:7
1876:.
1852::
1844::
1817:.
1776:.
1756::
1733:.
1721::
1697:.
1665::
1638:.
1614::
1587:.
1573::
1546:.
1532::
1526:5
1505:.
1461:.
1431::
1404:.
1374:.
1360:.
1340::
1334:4
1316:.
1302::
1258:.
1236::
1230:4
1209:.
1197::
942:m
939:K
876:H
868:O
853:|
847:H
835:O
826:C
811:|
805:H
795:H
789:|
784:C
772:H
750:|
744:H
734:O
725:C
710:|
704:H
694:H
688:|
683:C
671:H
654:H
646:O
633:|
627:H
617:H
611:|
606:C
591:|
585:H
575:H
569:|
564:C
552:H
438:2
350:m
347:K
343:m
340:K
335:m
332:K
270:(
41:.
34:.
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.