624:
57:
44:
227:
283:(the time it takes a culture of microbes to double in cells per unit volume) of approximately 32 hours. By using the more optimal medium MA2, this can be reduced to 24 hours. Culturing is done at 42 °C (108 °F) under white fluorescent light with an approximate intensity of 50 μmol photons m s (μE). However, under a higher light intensity of 90 μE with 5% CO
594:
would for transformation purposes. To transform, cells are briefly exposed to 30% PEG with the DNA of interest, resulting in transient transformation. In this method, the DNA is taken up as a circular element and is not integrated into the genome of the organism because no homologous regions exist
1012:
Matsuzaki M; Misumi O; Shin-i T; Maruyama S; Takahara M; Miyagishima S; Mori T; Nishida K; Yagisawa F; Nishida K; Yoshida Y; Nishimura Y; Nakao S; Kobayashi T; Momoyama Y; Higashiyama T; Minoda A; Sano M; Nomoto H; Oishi K; Hayashi H; Ohta F; Nishizaka S; Haga S; Miura S; Morishita T; Kabeya Y;
652:
makes it the perfect organism for studying mechanisms of eukaryotic cell and organelle division. Synchronization of the division of organelles in cultured cells can be very simple and usually involves the use of light and dark cycles. The chemical agent
291:
can be further increased, with a doubling time of approximately 9.2 hours. Higher light is not necessarily beneficial, as above 90 μE the growth rate begins to decrease. This may be due to photodamage occurring to the photosynthetic apparatus.
704:, as might be expected, has a particularly unusual pH range in which it can function. Despite the fact that the mechanism of PSII requires protons to be quickly released, and lower pH solutions should alter the ability to do this,
1484:
Imamura S; Yoshihara S; Nakano S; Shiozaki N; Yamada A; Tanaka K; Takahashi H; Asayama M; Shirai M (2003). "Purification, characterization, and gene expression of all sigma factors of RNA polymerase in a cyanobacterium".
619:
machinery can be used to insert the gene at these regions. The same transformation procedure as is used for transient expression can be used here, except with the homologous DNA segments to allow for genome integration.
429:(CTAB) method may be employed. In this method, a high-salt extraction buffer is first applied and cells are disrupted, after which a chloroform-phenol mixture is used to extract the DNA at room temperature.
550:, which allowed cells to survive in the presence of 5-FOA as long as uracil was provided. By transforming this mutant with a PCR fragment carrying both a gene of interest and a functional copy of
279:
in the laboratory in
Modified Allen's medium (MA) or a modified form with twice the concentration of some elements called MA2. Using MA medium, growth rates are not particularly fast, with a
223:
in 2004; its plastid was sequenced in 2000 and 2003, and its mitochondrion in 1998. The organism has been considered the simplest of eukaryotic cells for its minimalist cellular organization.
449:
As is the case for DNA and RNA, the protocol for protein extraction is also an adaptation of the protocol used in cyanobacteria. Cells are disrupted using glass beads and vortexing in a 10%
868:
1013:
Terasawa K; Suzuki Y; Ishii Y; Asakawa S; Takano H; Ohta N; Kuroiwa H; Tanaka K; Shimizu N; Sugano S; Sato N; Nozaki H; Ogasawara N; Kohara Y; Kuroiwa T (2004).
881:
1284:, Reveals the Molecular Basis of the Metabolic Flexibility of Galdieria sulphuraria and Significant Differences in Carbohydrate Metabolism of Both Algae"
1757:
Imoto Y; Kuroiwa H; Yoshida Y; Ohnuma M; Fujiwara T; Yoshida M; Nishida K; Yagisawa F; Hirooka S; Miyagishima S; Misumi O; Kawano S; Kuroiwa T (2013).
1352:
Kobayashi Y; Ohnuma M; Kuroiwa T; Tanaka K; Hanaoka M (2010). "The basics of cultivation and molecular genetic analysis of the unicellular red alga
855:
611:. By including regions of DNA several hundred base pairs long on the ends of the gene of interest that are complementary to a sequence within the
308:, meaning it is not capable of taking up fixed carbon from its environment and must rely on oxygenic photosynthesis to fix carbon from CO
1146:
398:
for transient or stable transformation, and methods for selection including a uracil auxotroph that can be used as a selection marker.
1965:
1098:
355:. The reduced nature of the genome has led to several other unusual features. While most eukaryotes contain 10 or so copies of the
1378:
Castenholz RW; McDermott TR (2010). "The
Cyanidiales: Ecology, Biodiversity, and Biogeography". In Seckbach J; Chapman DJ (eds.).
917:
1715:
Terui S; Suzuki K; Takahiashi H; Itoh R; Kuroiwa T (1995). "High synchronization of chloroplast division in the ultramicro-alga
56:
1077:
Kuroiwa T; Kuroiwa H; Sakai A; Takahashi H; Toda K; Itoh R (1998). "The division apparatus of plastids and mitochondria".
603:
To create a stable mutant line, gene targeting can be used to insert a gene of interest into a particular location of the
527:
1901:"A reaction center-dependent photoprotection mechanism in a highly robust photosystem II from an extremophilic red alga,
488:
commonly used for selection of successfully transformed individuals in the laboratory, but it resistant to some, notably
370:
contains many genes not present in the chloroplast genomes of other algae and plants. Most of its genes are intronless.
425:(PCR), wherein phenol is added to whole cells and incubated at 65 °C to extract DNA. If purer DNA is required, the
1181:"Improvement of culture conditions and evidence for nuclear transformation by homologous recombination in a red alga,
426:
1899:
Krupnik T; Kotabova E; van
Bezouwen LS; Mazur R; Garstka M; Nixon PJ; Barber J; Kana R; Boekema EJ; Kargul J (2013).
1443:
Ohta, N; Matsuzaki, M; Misumi, O; Miyagishima, S. Y.; Nozaki, H; Tanaka, K; Shin-i, T; Kohara, Y; Kuroiwa, T (2003).
807:
949:«Cyanidioschyzon merolae»: a new alga of thermal acidic environments. P De Luca, R Taddei and L Varano, Webbia, 1978
886:
518:
in the presence of a compound, 5-FOA, which in and of itself is non-toxic but is converted to the toxic compound
422:
1960:
608:
470:
352:
543:
207:
is considered an excellent model system for study of cellular and organelle division processes, as well as
1522:"Tetrapyrrole signal as a cell-cycle coordinator from organelle to nuclear DNA replication in plant cells"
769:
179:
adapted to high sulfur acidic hot spring environments (pH 1.5, 45 °C). The cellular architecture of
1573:"The plant‐specific TFIIB related protein, PBRP, is a general transcription factor for RNA polymerase I"
718:
144:
1445:"Complete sequenced analysis of the plastid genome of the unicellular red alga Cyanidioschyzon merolae"
922:
829:
567:
363:
only contains two, a fact that researchers have taken advantage of when studying organelle division.
821:
747:
623:
335:
and was found to contain 5,331 genes, of which 86.3% were found to be expressed and only 26 contain
628:
575:
1736:
648:
The extremely simple divisome, simple cell architecture, and ability to synchronize divisions in
212:
51:
873:
1936:
1881:
1840:
1790:
1759:"Single-membrane–bounded peroxisome division revealed by isolation of dynamin-based machinery"
1697:
1602:
1553:
1502:
1466:
1425:
1313:
1256:
1206:
1104:
1094:
1040:
983:
894:
636:
458:
419:
43:
909:
708:
PSII is capable of exchanging and splitting water at the same rate as other related species.
1926:
1916:
1871:
1830:
1780:
1770:
1728:
1687:
1677:
1637:
1592:
1584:
1543:
1533:
1494:
1456:
1415:
1303:
1295:
1246:
1196:
1142:
1086:
1030:
975:
500:
899:
539:
91:
1815:"Substrate water exchange in photosystem II core complexes of the extremophilic red alga
1133:
as model system for investigating the dividing apparatus of mitochondria and plastids".
1931:
1900:
1785:
1758:
1692:
1661:
1597:
1572:
1548:
1521:
1308:
1276:"Comparative Genomics of Two Closely Related Unicellular Thermo-Acidophilic Red Algae,
1275:
697:
685:
519:
454:
331:
was sequenced in 2004. The reduced, extremely simple, compact genome is made up of 20
71:
1498:
1090:
17:
1954:
1732:
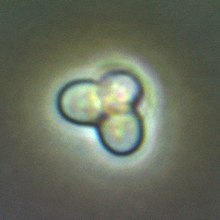
635:, showing two cells, one in which the plastid has begun to divide. Courtesy of Prof.
531:
418:
extraction, which is a quick extraction that can be used to isolate DNA suitable for
407:
348:
280:
268:
243:
188:
1740:
979:
554:, researchers can confirm that the gene of interest has been incorporated into the
466:
276:
208:
111:
1666:: single- and multi-copy insertion using authentic and chimeric selection markers"
300:
plates for purposes of colony selection or strain maintenance in the laboratory.
1876:
1859:
1835:
1814:
1682:
792:
801:
689:
666:
654:
579:
485:
251:
247:
226:
184:
101:
1520:
Kobayashia Y; Kanesakia Y; Tanakab A; Kuroiwac H; Kuroiwac T; Tanaka K (2009).
378:
As is the case with most model organisms, genetic tools have been developed in
1060:
658:
616:
591:
489:
332:
305:
297:
220:
176:
1461:
1444:
1400:"Polyethylene Glycol (PEG)-Mediated Transient Gene Expression in a Red Alga,
987:
657:
can be added to easily and effectively synchronize chloroplast division. The
1921:
1775:
1642:
1621:
1538:
842:
816:
760:. World-wide electronic publication, National University of Ireland, Galway.
756:
724:
583:
570:
allows short-term experiments to be done using labeled or modified genes in
511:
493:
321:
264:
200:
196:
81:
1940:
1885:
1844:
1794:
1701:
1606:
1588:
1557:
1506:
1470:
1429:
1317:
1260:
1210:
1044:
1299:
1147:
10.1002/(sici)1521-1878(199804)20:4<344::aid-bies11>3.0.co;2-2
1108:
1420:
1399:
1201:
1180:
786:
670:
462:
450:
1251:
1230:
1035:
1014:
441:
cells using a variant of the hot phenol method described above for DNA.
339:, which contained strict consensus sequences. Strikingly, the genome of
860:
736:
696:
has some significant differences from that of other related organisms.
356:
234:
in flasks and a 10-litre (2.2 imp gal; 2.6 US gal)
192:
1620:
Yagisawa F; Nishida K; Okano Y; Minoda A; Tanaka K; Kuroiwa T (2004).
566:
While chromosomal integration of genes creates a stable transformant,
1858:
Bricker TM; Roose JL; Fagerlund RD; Frankel LK; Eaton-Rye JJ (2012).
665:
as a system, where peroxisome division can be synchronized using the
523:
508:
415:
336:
324:
235:
216:
173:
763:
359:
required for pinching membranes to separate dividing compartments,
847:
622:
514:(requiring exogenous uracil). The mutant was developed by growing
225:
1373:
1371:
344:
834:
767:
1179:
Minoda A; Sakagami R; Yagisawa F; Kuroiwa T; Tanaka K (2004).
387:
383:
366:
Although possessing a small genome, the chloroplast genome of
1660:
Fujiwara T; Ohnuma M; Yoshida M; Kuroiwa T; Hirano T (2013).
1398:
Ohnuma M; Yokoyama T; Inouye T; Sekine Y; Tanaka K (2008).
263:
Originally isolated by De Luca in 1978 from the solfatane
242:
is blue-green: it makes little if any of the red pigment
1347:
1231:"Genome sequence of the ultrasmall unicellular red alga
1015:"Genome sequence of the ultrasmall unicellular red alga
246:, and hence only displays the second red-algal pigment,
1345:
1343:
1341:
1339:
1337:
1335:
1333:
1331:
1329:
1327:
562:
Polyethylene glycol (PEG) mediated transient expression
1655:
1653:
1007:
1005:
1003:
1001:
999:
997:
1808:
1806:
1804:
966: »: a new alga of thermal acidic environments".
1813:
Nilsson H; Krupnik T; Kargul J; Messinger J (2014).
1393:
1391:
1389:
957:
955:
1823:
Biochimica et
Biophysica Acta (BBA) - Bioenergetics
1752:
1750:
1174:
1172:
1170:
1168:
1166:
1164:
1162:
1160:
1158:
1156:
776:
1059:
477:Transformant selection and uracil auxotrophic line
453:buffer containing the reducing agent DTT to break
410:protocols, are used for the isolation of DNA from
203:divisions can be synchronized. For these reasons,
1622:"Isolation of cycloheximide-resistant mutants of
574:. Transient expression can be achieved using a
558:genome if it can grow without exogenous uracil.
457:within proteins. This extraction will result in
1719:by treatment with both light and aphidicolin".
1224:
1222:
1220:
661:division mechanism was first ascertained using
1120:
1118:
382:. These include methods for the isolation of
183:is extremely simple, containing only a single
287:applied through bubbling, the growth rate of
8:
1358:Journal of Endocytobiosis and Cell Research
688:. Notably, the subunit composition of the
172:is a small (2μm), club-shaped, unicellular
1860:"The extrinsic proteins of photosystem II"
764:
42:
31:
1930:
1920:
1875:
1834:
1784:
1774:
1691:
1681:
1641:
1596:
1547:
1537:
1460:
1419:
1307:
1250:
1200:
1125:Kuroiwa (1998). "The primitive red algae
1034:
347:genes and an extremely minimal number of
1274:Barbier, Guillaume; et al. (2005).
590:lacks a cell wall, it behaves much as a
160:P.De Luca, R.Taddei & L.Varano, 1978
1571:Imamura S; Hanaoka M; Tanaka K (2008).
968:Journal of Plant Taxonomy and Geography
962:De Luca P; Taddei R; Varano L (1978). "
942:
586:enzymatically eliminated), and because
238:. Although classified as a red alga,
219:was the first full algal genome to be
644:Studying cell and organelle divisions
7:
1081:. International Review of Cytology.
526:in the uracil biosynthetic pathway,
1229:Matsuzaki, M.; et al. (2004).
673:in addition to light-dark cycles.
25:
1733:10.1111/j.0022-3646.1995.00958.x
1662:"Gene targeting in the red alga
437:Total RNA may be extracted from
427:Cetyl trimethyl ammonium bromide
199:. In addition, the cellular and
55:
394:, the introduction of DNA into
259:Isolation and growth in culture
980:10.1080/00837792.1978.10670110
406:Several methods, derived from
1:
1499:10.1016/s0022-2836(02)01242-1
1091:10.1016/s0074-7696(08)60415-5
1066:. Cambridge University Press.
351:gene copies, as shown in the
1877:10.1016/j.bbabio.2011.07.006
1836:10.1016/j.bbabio.2014.04.001
1683:10.1371/journal.pone.0073608
1380:Red Algae in the Genomic Age
684:is also used in researching
582:(plant cells with the rigid
615:genome, the organism's own
1982:
1058:Robert Edward Lee (1999).
627:Freeze fracture deep etch
528:orotidine 5'-monophosphate
1966:Species described in 1978
746:Guiry, M.D.; Guiry, G.M.
423:polymerase chain reaction
150:
143:
52:Scientific classification
50:
41:
34:
609:homologous recombination
544:loss-of-function mutants
304:is an obligate oxygenic
250:, and the green pigment
1922:10.1074/jbc.m113.484659
1903:Cyanidioschyzon merolae
1817:Cyanidioschyzon merolae
1776:10.1073/pnas.1303483110
1717:Cyanidioschyzon merolae
1664:Cyanidioschyzon merolae
1643:10.1508/cytologia.69.97
1624:Cyanidioschyzon merolae
1539:10.1073/pnas.0804270105
1402:Cyanidioschyzon merolae
1354:Cyanidioschyzon merolae
1282:Cyanidioschyzon merolae
1233:Cyanidioschyzon merolae
1183:Cyanidioschyzon merolae
1131:Cyanidioschyzon merolae
1017:Cyanidioschyzon merolae
964:Cyanidioschyzon merolae
808:Cyanidioschyzon merolae
778:Cyanidioschyzon merolae
750:Cyanidioschyzon merolae
738:Cyanidioschyzon merolae
677:Photosynthesis research
461:, which can be used in
353:genome comparison table
169:Cyanidioschyzon merolae
154:Cyanidioschyzon merolae
1864:Biochim. Biophys. Acta
1589:10.1038/emboj.2008.151
1462:10.1093/dnares/10.2.67
640:
578:(PEG) based method in
503:for transformation in
255:
18:Cyanidioschyzon merolæ
1763:Proc. Natl. Acad. Sci
1526:Proc. Natl. Acad. Sci
1300:10.1104/pp.104.051169
1278:Galdieria sulphuraria
719:Galdieria sulphuraria
626:
484:is sensitive to many
414:. The first is a hot
296:can also be grown on
230:Growing the red alga
229:
568:transient expression
1915:(32): 23529–23542.
1382:. pp. 357–371.
1252:10.1038/nature02398
1127:Cyanidium caldarium
1036:10.1038/nature02398
629:electron microscopy
576:polyethylene glycol
1421:10.1093/pcp/pcm157
1408:Plant Cell Physiol
1202:10.1093/pcp/pch087
1189:Plant Cell Physiol
641:
459:denatured proteins
445:Protein extraction
267:of Campi Flegrei (
256:
213:structural biology
1769:(23): 9583–9588.
1583:(17): 2317–2327.
1245:(6983): 653–657.
1029:(6983): 653–657.
933:
932:
895:Open Tree of Life
770:Taxon identifiers
669:-disrupting drug
637:Ursula Goodenough
595:for integration.
534:, encoded by the
420:DNA amplification
374:Molecular biology
343:contains only 30
215:. The organism's
165:
164:
16:(Redirected from
1973:
1945:
1944:
1934:
1924:
1896:
1890:
1889:
1879:
1855:
1849:
1848:
1838:
1829:(8): 1257–1262.
1810:
1799:
1798:
1788:
1778:
1754:
1745:
1744:
1712:
1706:
1705:
1695:
1685:
1657:
1648:
1647:
1645:
1617:
1611:
1610:
1600:
1568:
1562:
1561:
1551:
1541:
1517:
1511:
1510:
1481:
1475:
1474:
1464:
1440:
1434:
1433:
1423:
1395:
1384:
1383:
1375:
1366:
1365:
1349:
1322:
1321:
1311:
1288:Plant Physiology
1271:
1265:
1264:
1254:
1226:
1215:
1214:
1204:
1176:
1151:
1150:
1122:
1113:
1112:
1074:
1068:
1067:
1065:
1055:
1049:
1048:
1038:
1009:
992:
991:
959:
950:
947:
926:
925:
913:
912:
903:
902:
890:
889:
877:
876:
864:
863:
851:
850:
838:
837:
825:
824:
812:
811:
810:
797:
796:
795:
765:
761:
501:selection marker
499:A commonly used
467:Western blotting
275:can be grown in
156:
60:
59:
46:
32:
21:
1981:
1980:
1976:
1975:
1974:
1972:
1971:
1970:
1961:Cyanidiophyceae
1951:
1950:
1949:
1948:
1898:
1897:
1893:
1857:
1856:
1852:
1812:
1811:
1802:
1756:
1755:
1748:
1714:
1713:
1709:
1659:
1658:
1651:
1619:
1618:
1614:
1570:
1569:
1565:
1519:
1518:
1514:
1483:
1482:
1478:
1442:
1441:
1437:
1397:
1396:
1387:
1377:
1376:
1369:
1351:
1350:
1325:
1273:
1272:
1268:
1228:
1227:
1218:
1178:
1177:
1154:
1124:
1123:
1116:
1101:
1079:Int. Rev. Cytol
1076:
1075:
1071:
1057:
1056:
1052:
1011:
1010:
995:
961:
960:
953:
948:
944:
939:
934:
929:
921:
916:
908:
906:
898:
893:
885:
880:
872:
867:
859:
854:
846:
841:
833:
828:
820:
815:
806:
805:
800:
791:
790:
785:
772:
745:
733:
714:
679:
646:
601:
564:
542:led to several
540:Random mutation
479:
455:disulfide bonds
447:
435:
404:
376:
318:
311:
286:
261:
161:
158:
152:
139:
136:C. merolae
123:Cyanidioschyzon
92:Cyanidiophyceae
54:
36:Cyanidioschyzon
28:
27:Species of alga
23:
22:
15:
12:
11:
5:
1979:
1977:
1969:
1968:
1963:
1953:
1952:
1947:
1946:
1891:
1870:(1): 121–142.
1850:
1800:
1746:
1707:
1649:
1612:
1563:
1532:(3): 803–807.
1512:
1493:(5): 857–872.
1476:
1435:
1414:(1): 117–120.
1385:
1367:
1323:
1294:(2): 460–474.
1266:
1216:
1195:(6): 667–671.
1152:
1141:(4): 344–354.
1114:
1099:
1069:
1050:
993:
951:
941:
940:
938:
935:
931:
930:
928:
927:
914:
904:
891:
878:
865:
852:
839:
826:
813:
798:
782:
780:
774:
773:
768:
743:
742:
740:Genome Project
732:
731:External links
729:
728:
727:
722:
713:
710:
698:Photosystem II
686:photosynthesis
678:
675:
645:
642:
600:
599:Gene targeting
597:
563:
560:
520:5-Fluorouracil
478:
475:
446:
443:
434:
431:
408:cyanobacterial
403:
400:
375:
372:
317:
314:
309:
284:
260:
257:
191:and lacking a
163:
162:
159:
148:
147:
141:
140:
133:
131:
127:
126:
119:
115:
114:
109:
105:
104:
99:
95:
94:
89:
85:
84:
79:
75:
74:
72:Archaeplastida
69:
62:
61:
48:
47:
39:
38:
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
1978:
1967:
1964:
1962:
1959:
1958:
1956:
1942:
1938:
1933:
1928:
1923:
1918:
1914:
1910:
1909:J. Biol. Chem
1906:
1904:
1895:
1892:
1887:
1883:
1878:
1873:
1869:
1865:
1861:
1854:
1851:
1846:
1842:
1837:
1832:
1828:
1824:
1820:
1818:
1809:
1807:
1805:
1801:
1796:
1792:
1787:
1782:
1777:
1772:
1768:
1764:
1760:
1753:
1751:
1747:
1742:
1738:
1734:
1730:
1726:
1722:
1718:
1711:
1708:
1703:
1699:
1694:
1689:
1684:
1679:
1676:(9): e73608.
1675:
1671:
1667:
1665:
1656:
1654:
1650:
1644:
1639:
1635:
1631:
1627:
1625:
1616:
1613:
1608:
1604:
1599:
1594:
1590:
1586:
1582:
1578:
1574:
1567:
1564:
1559:
1555:
1550:
1545:
1540:
1535:
1531:
1527:
1523:
1516:
1513:
1508:
1504:
1500:
1496:
1492:
1488:
1480:
1477:
1472:
1468:
1463:
1458:
1454:
1450:
1446:
1439:
1436:
1431:
1427:
1422:
1417:
1413:
1409:
1405:
1403:
1394:
1392:
1390:
1386:
1381:
1374:
1372:
1368:
1363:
1359:
1355:
1348:
1346:
1344:
1342:
1340:
1338:
1336:
1334:
1332:
1330:
1328:
1324:
1319:
1315:
1310:
1305:
1301:
1297:
1293:
1289:
1285:
1283:
1279:
1270:
1267:
1262:
1258:
1253:
1248:
1244:
1240:
1236:
1234:
1225:
1223:
1221:
1217:
1212:
1208:
1203:
1198:
1194:
1190:
1186:
1184:
1175:
1173:
1171:
1169:
1167:
1165:
1163:
1161:
1159:
1157:
1153:
1148:
1144:
1140:
1136:
1132:
1128:
1121:
1119:
1115:
1110:
1106:
1102:
1100:9780123645852
1096:
1092:
1088:
1084:
1080:
1073:
1070:
1064:
1063:
1054:
1051:
1046:
1042:
1037:
1032:
1028:
1024:
1020:
1018:
1008:
1006:
1004:
1002:
1000:
998:
994:
989:
985:
981:
977:
973:
969:
965:
958:
956:
952:
946:
943:
936:
924:
919:
915:
911:
905:
901:
896:
892:
888:
883:
879:
875:
870:
866:
862:
857:
853:
849:
844:
840:
836:
831:
827:
823:
818:
814:
809:
803:
799:
794:
788:
784:
783:
781:
779:
775:
771:
766:
762:
759:
758:
753:
751:
741:
739:
735:
734:
730:
726:
723:
721:
720:
716:
715:
711:
709:
707:
703:
699:
695:
691:
687:
683:
676:
674:
672:
668:
664:
660:
656:
651:
643:
638:
634:
630:
625:
621:
618:
614:
610:
606:
598:
596:
593:
589:
585:
581:
577:
573:
569:
561:
559:
557:
553:
549:
545:
541:
537:
533:
532:decarboxylase
529:
525:
521:
517:
513:
510:
506:
502:
497:
495:
491:
487:
483:
476:
474:
472:
468:
464:
460:
456:
452:
444:
442:
440:
433:RNA isolation
432:
430:
428:
424:
421:
417:
413:
409:
402:DNA isolation
401:
399:
397:
393:
389:
385:
381:
373:
371:
369:
364:
362:
358:
354:
350:
349:ribosomal RNA
346:
342:
338:
334:
330:
326:
323:
322:megabase pair
315:
313:
307:
303:
299:
295:
290:
282:
281:doubling time
278:
274:
270:
269:Naples, Italy
266:
258:
253:
249:
245:
244:phycoerythrin
241:
237:
233:
228:
224:
222:
218:
214:
210:
206:
202:
198:
194:
190:
189:mitochondrion
187:and a single
186:
182:
178:
175:
171:
170:
157:
155:
149:
146:
145:Binomial name
142:
138:
137:
132:
129:
128:
125:
124:
120:
117:
116:
113:
110:
107:
106:
103:
100:
97:
96:
93:
90:
87:
86:
83:
80:
77:
76:
73:
70:
67:
64:
63:
58:
53:
49:
45:
40:
37:
33:
30:
19:
1912:
1908:
1902:
1894:
1867:
1863:
1853:
1826:
1822:
1816:
1766:
1762:
1724:
1720:
1716:
1710:
1673:
1669:
1663:
1633:
1629:
1623:
1615:
1580:
1576:
1566:
1529:
1525:
1515:
1490:
1487:J. Mol. Biol
1486:
1479:
1455:(2): 67–77.
1452:
1449:DNA Research
1448:
1438:
1411:
1407:
1401:
1379:
1361:
1357:
1353:
1291:
1287:
1281:
1277:
1269:
1242:
1238:
1232:
1192:
1188:
1182:
1138:
1134:
1130:
1126:
1082:
1078:
1072:
1061:
1053:
1026:
1022:
1016:
974:(1): 37–44.
971:
967:
963:
945:
777:
755:
749:
744:
737:
717:
705:
701:
693:
690:photosystems
681:
680:
662:
649:
647:
632:
612:
604:
602:
587:
571:
565:
555:
551:
547:
535:
515:
504:
498:
481:
480:
448:
438:
436:
411:
405:
395:
391:
379:
377:
367:
365:
360:
340:
328:
319:
301:
293:
288:
272:
262:
239:
231:
209:biochemistry
204:
180:
168:
167:
166:
153:
151:
135:
134:
122:
121:
112:Cyanidiaceae
65:
35:
29:
1727:: 958–961.
802:Wikispecies
667:microtubule
655:aphidicolin
607:genome via
580:protoplasts
507:involves a
486:antibiotics
333:chromosomes
252:chlorophyll
248:phycocyanin
185:chloroplast
102:Cyanidiales
1955:Categories
1636:: 97–100.
937:References
706:C. merolae
702:C. merolae
700:(PSII) of
694:C. merolae
682:C. merolae
663:C. merolae
659:peroxisome
650:C. merolae
633:C. merolae
617:DNA repair
613:C. merolae
605:C. merolae
592:protoplast
588:C. merolae
572:C. merolae
556:C. merolae
516:C. merolae
505:C. merolae
490:ampicillin
482:C. merolae
473:staining.
439:C. merolae
412:C. merolae
396:C. merolae
392:C. merolae
380:C. merolae
368:C. merolae
361:C. merolae
341:C. merolae
329:C. merolae
306:phototroph
302:C. merolae
298:gellan gum
294:C. merolae
289:C. merolae
273:C. merolae
240:C. merolae
232:C. merolae
205:C. merolae
181:C. merolae
82:Rhodophyta
78:Division:
1721:J. Phycol
1630:Cytologia
1135:BioEssays
1062:Phycology
988:0083-7792
817:AlgaeBase
793:Q16034656
757:AlgaeBase
725:Red algae
631:image of
584:cell wall
512:auxotroph
494:kanamycin
471:Coomassie
465:gels for
320:The 16.5
265:fumaroles
221:sequenced
201:organelle
197:cell wall
130:Species:
1941:23775073
1886:21801710
1845:24726350
1795:23696667
1741:84124611
1702:24039997
1670:PLOS ONE
1607:18668124
1558:19141634
1507:12527296
1471:12755171
1430:18003671
1364:: 53–61.
1318:15710685
1261:15071595
1211:15215501
1085:: 1–41.
1045:15071595
874:10006752
787:Wikidata
712:See also
671:oryzalin
463:SDS-PAGE
451:glycerol
357:dynamins
177:red alga
108:Family:
1932:5395030
1786:3677435
1693:3764038
1598:2529366
1549:2625283
1309:1065348
1109:9522454
910:1469579
861:2668940
337:introns
277:culture
193:vacuole
174:haploid
118:Genus:
98:Order:
88:Class:
1939:
1929:
1884:
1843:
1793:
1783:
1739:
1700:
1690:
1605:
1595:
1577:EMBO J
1556:
1546:
1505:
1469:
1428:
1316:
1306:
1259:
1239:Nature
1209:
1107:
1097:
1043:
1023:Nature
986:
923:611455
907:uBio:
835:912450
552:Ura5.3
548:Ura5.3
538:gene.
536:Ura5.3
530:(OMP)
524:enzyme
522:by an
509:uracil
416:phenol
325:genome
316:Genome
236:carboy
217:genome
1737:S2CID
918:WoRMS
900:85503
887:45157
869:IRMNG
848:KYAME
822:36733
390:from
66:Clade
1937:PMID
1882:PMID
1868:1817
1841:PMID
1827:1837
1791:PMID
1698:PMID
1603:PMID
1554:PMID
1503:PMID
1467:PMID
1426:PMID
1404:10D"
1314:PMID
1280:and
1257:PMID
1235:10D"
1207:PMID
1185:10D"
1129:and
1105:PMID
1095:ISBN
1041:PMID
1019:10D"
984:ISSN
882:NCBI
856:GBIF
843:EPPO
492:and
469:and
386:and
345:tRNA
211:and
195:and
1927:PMC
1917:doi
1913:288
1872:doi
1831:doi
1781:PMC
1771:doi
1767:110
1729:doi
1688:PMC
1678:doi
1638:doi
1593:PMC
1585:doi
1544:PMC
1534:doi
1530:106
1495:doi
1491:325
1457:doi
1416:doi
1356:".
1304:PMC
1296:doi
1292:137
1247:doi
1243:428
1197:doi
1143:doi
1087:doi
1083:181
1031:doi
1027:428
976:doi
830:EoL
692:in
546:in
388:RNA
384:DNA
327:of
271:),
1957::
1935:.
1925:.
1911:.
1907:.
1880:.
1866:.
1862:.
1839:.
1825:.
1821:.
1803:^
1789:.
1779:.
1765:.
1761:.
1749:^
1735:.
1725:31
1723:.
1696:.
1686:.
1672:.
1668:.
1652:^
1634:69
1632:.
1628:.
1601:.
1591:.
1581:27
1579:.
1575:.
1552:.
1542:.
1528:.
1524:.
1501:.
1489:.
1465:.
1453:10
1451:.
1447:.
1424:.
1412:49
1410:.
1406:.
1388:^
1370:^
1362:20
1360:.
1326:^
1312:.
1302:.
1290:.
1286:.
1255:.
1241:.
1237:.
1219:^
1205:.
1193:45
1191:.
1187:.
1155:^
1139:20
1137:.
1117:^
1103:.
1093:.
1039:.
1025:.
1021:.
996:^
982:.
972:33
970:.
954:^
920::
897::
884::
871::
858::
845::
832::
819::
804::
789::
754:.
496:.
312:.
68::
1943:.
1919::
1905:"
1888:.
1874::
1847:.
1833::
1819:"
1797:.
1773::
1743:.
1731::
1704:.
1680::
1674:8
1646:.
1640::
1626:"
1609:.
1587::
1560:.
1536::
1509:.
1497::
1473:.
1459::
1432:.
1418::
1320:.
1298::
1263:.
1249::
1213:.
1199::
1149:.
1145::
1111:.
1089::
1047:.
1033::
990:.
978::
752:"
748:"
639:.
310:2
285:2
254:.
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.