84:
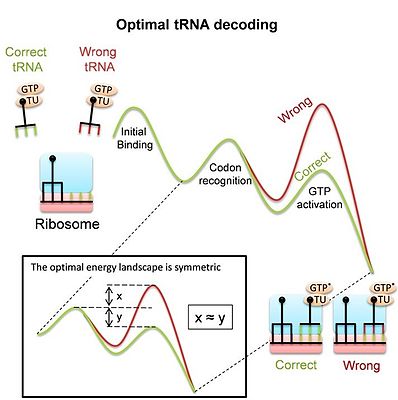
200:: The curves show the free energy landscape of codon recognition as suggested by experiments. In the stages that are sensitive to codon identity, the pathways of correct (green) and wrong (red) tRNAs split. The multistage kinetics include: Initial binding and codon recognition: a complex of elongation factor (EF-Tu) and aminoacyl-tRNA binds to the ribosome. Codon is recognized by pairing with the anticodon, and by additional interaction with the "decoding center" of the ribosome. As a result, correct (cognate) tRNAs are more stable than non-cognate ones. GTP activation and hydrolysis: Codon recognition leads to global conformational changes of the ribosome and tRNA, which are different for cognate or non-cognate tRNAs and affect GTP activation and hydrolysis by EF-Tu.
34:- such as a structural mismatch between a molecular recognizer and its target - enhances the recognition specificity and quality. Conformational proofreading does not require the consumption of energy and may therefore be used in any molecular recognition system. Conformational proofreading is especially useful in scenarios where the recognizer has to select the appropriate target among many similar competitors. Proteins evolve the capacity for conformational proofreading through fine-tuning their geometry, flexibility and chemical interactions with the target.
190:
173:. Decoding is a major determinant of fitness and requires accurate and fast selection of correct tRNAs among many similar competitors. One must have in mind that most binding events are by non-matching (“non-cognate”) tRNAs and the ribosome needs to reject those as fast as possible in order to vacate the binding site. At the same time, the ribosome should keep the matching tRNAs bound long enough to allow the protein synthesis process ensue. Despite the importance of tRNA decoding, it was unclear until recently whether the modern ribosome, and in particular its
274:(equivalently, an irreversible intermediate stage) is introduced during the formation of the correct or incorrect complexes. This time delay reduces the production rates of both complexes but enhances the fidelity beyond the equilibrium limit. The irreversibility of the scheme requires an energy source. The time delay in kinetic proofreading is analogous to the spatial difference in conformational proofreading. However, the conformational proofreading can be an equilibrium scheme that does not consume energy.
144:
the energy barrier predicted by the conformational proofreading model. The data indicate a physical picture for homology recognition in which the fidelity of the search process is governed by the distance between the DNA-binding sites. The authors conclude that their interpretation of the measurements "is akin to a
249:
at sites of damage. Analysis of the rates of dissociation for the transient binding molecules on both undamaged and damaged DNA show multiple dwell times over three orders of magnitude... These intermediate states are believed to represent discrete UV-DDB conformers on the trajectory to stable damage
129:
in E. coli, or members of its superfamily in other organisms. RecA first polymerizes along a stretch of single-stranded DNA, and then this protein-DNA filament searches for homology along double-stranded DNA. In the RecA-DNA filament, the distance between bases increases significantly with respect to
143:
The group of C. Dekker (Delft
University) directly probed the interactions involved in homology search by combining magnetic and optical tweezers. They have found that homology search and recognition requires opening of the helix and can therefore be accelerated by unwinding the DNA. This is exactly
148:
scheme ... where the dsDNA, and not the RecA filament, is the active, recognizing search entity. A large conformational mismatch exists between the target-bound and unbound states of the dsDNA. The target-bound state is accessed via energetically unfavorable intermediate states, as discussed above.
134:
on the search, since the double-stranded DNA has to stretch by the same magnitude to check for homology. By formulating the DNA recognition process as a signal detection problem, it was shown that the experimentally observed RecA-induced DNA deformation and the binding energetics are fine-tuned to
177:
during decoding, are the outcome of adaptation to its task as a decoder or the result of other constraints. Recent study derived the energy landscape that provides optimal discrimination between competing tRNA substrates, and thereby optimal tRNA decoding. The optimal landscape is a symmetric one
70:
between the recognizer (lock) and the key, reduces the binding probability to the correct target but reduces even more the binding probability to a similar wrong target and thus improves the specificity. Yet, introducing too much deformation drastically reduces binding probability to the correct
106:
which is the difference, Fitness = Prob(Correct) − Prob(Wrong), is maximal at a non-zero deformation energy. This barrier is optimal in the sense that it significantly reduces the binding probability while keeping the correct binding probability roughly the same. Biochemical measurements of
135:
ensure optimal sequence detection. The amount of deformation is such that binding to homologous DNA sequences only slightly decreases, while binding to wrong sequences decreases significantly. This is exactly the conformational proofreading mechanism.
149:
The conformational mismatch improves the selectivity of the recognition reaction." In other words, they identified the energetic barrier and have shown that indeed the double-stranded DNA is the active participant, since it has to pass this barrier.
79:
mechanism, are advantageous for enhancing the specificity of recognition. Such conformational changes may be fine-tuned by mutations that affect the mechanical response of the recognizer, also at positions far from the binding site.
124:
facilitates the exchange of genetic material between homologous DNA molecules. This crucial process requires detecting a specific homologous DNA sequence within a huge variety of heterologous sequences. The detection is mediated by
59:, then the binding probability will be high since no deformation is required upon binding. At the same time, the recognizer might also bind to a competitor with a similar structure with high probability. Introducing an
182:. This model suggests that conformational changes of the ribosome and tRNA during decoding are means to obtain such an optimal tRNA decoder. The fact that both homologous recombination and tRNA decoding utilize
42:
Molecular recognition takes place in a noisy, crowded biological environment and the recognizer often has to cope with the task of selecting its target among a variety of similar competitors. For example, the
225:
A recent study shows that conformational proofreading is used by human DNA repair mechanisms. The research focused on the question of how DNA-repair proteins scan the human genome for
71:
target. Therefore, the optimal balance between maximizing the correct binding probability and minimizing the incorrect binding probability is achieved when the recognizer is slightly
1002:
237:(UV-DDB) performs a 3D search. The authors find that "UV-DDB examines sites on DNA in discrete steps before forming long-lived, nonmotile UV-DDB dimers (
356:
Savir Y, Tlusty T (2008). "Optimal Design of a
Molecular Recognizer : Molecular Recognition as a Bayesian Signal Detection Problem".
189:
833:"Single-molecule analysis reveals human UV-damaged DNA-binding protein (UV-DDB) dimerizes on DNA via multiple kinetic intermediates"
211:
adapted landscape implies that the ratio of forward and backward rates is inverted between the correct and wrong energy landscapes.
83:
94:: The binding probability to homologous (correct) and non-homologous (wrong) DNA sequences decrease with the deformation
739:
Chen Z, Yang H, Pavletich NP (May 2008). "Mechanism of homologous recombination from the RecA-ssDNA/dsDNA structures".
476:"Achieving fidelity in homologous recombination despite extreme complexity: informed decisions by molecular profiling"
234:
230:
202:
The conformational proofreading model explains these conformational changes as a means to enhance tRNA recognition.
892:"Kinetic proofreading: a new mechanism for reducing errors in biosynthetic processes requiring high specificity"
416:
Savir Y, Tlusty T (November 2010). "RecA-mediated homology search as a nearly optimal signal detection system".
296:"Conformational proofreading: the impact of conformational changes on the specificity of molecular recognition"
121:
55:
codon among many structurally similar tRNAs. If the recognizer and its correct target match perfectly like a
186:
suggests that this is a generic mechanism that may be utilized broadly by molecular recognition systems.
1007:
27:
790:
De
Vlaminck I, van Loenhout MT, Zweifel L, den Blanken J, Hooning K, Hage S, et al. (June 2012).
903:
844:
748:
693:
590:
541:
375:
307:
267:
252:
UV-DDB recognizes lesions using a conformational proofreading mechanism via multiple intermediates.
65:
56:
772:
608:
451:
425:
391:
365:
120:
is utilized in the system of homologous recombination to discern between similar DNA sequences.
75:. This suggests that conformational changes during molecular recognition processes, such as the
831:
Ghodke H, Wang H, Hsieh CL, Woldemeskel S, Watkins SC, Rapić-Otrin V, Van Houten B (May 2014).
178:(see image). The study shows that the measured landscape of the prokaryotic ribosome is indeed
966:
931:
872:
813:
764:
721:
662:
559:
505:
443:
335:
95:
60:
31:
1012:
958:
921:
911:
862:
852:
803:
756:
711:
701:
652:
644:
633:"General Theory of Specific Binding: Insights from a Genetic-Mechano-Chemical Protein Model"
598:
549:
495:
487:
435:
383:
325:
315:
76:
987:
907:
848:
752:
697:
594:
545:
379:
311:
867:
832:
792:"Mechanism of homology recognition in DNA recombination from dual-molecule experiments"
657:
632:
500:
475:
330:
295:
962:
926:
891:
716:
681:
996:
612:
776:
455:
395:
107:
RecA-induced recombination suggest that the observed deformation is nearly optimal.
130:
the bare 3.4 Å in the double-strand (by 50% on average). This sets a significant
808:
791:
491:
439:
320:
139:
Experimental evidence for conformational proofreading by homologous recombination
226:
162:
161:
is a complex molecular machine that, in order to synthesize proteins during the
896:
Proceedings of the
National Academy of Sciences of the United States of America
837:
Proceedings of the
National Academy of Sciences of the United States of America
686:
Proceedings of the
National Academy of Sciences of the United States of America
554:
529:
250:
detection." The authors conclude from their detailed kinetic measurements that
387:
648:
30:
systems, suggested by
Yonatan Savir and Tsvi Tlusty, in which introducing an
916:
857:
876:
817:
768:
725:
666:
563:
509:
447:
339:
970:
935:
706:
158:
44:
760:
530:"The ribosome as an optimal decoder: a lesson in molecular recognition"
682:"Application of a Theory of Enzyme Specificity to Protein Synthesis"
603:
578:
233:(NER). Detailed single-molecule measurements revealed how the human
949:
Ninio J (1975). "Kinetic amplification of enzyme discrimination".
631:
McBride JM, Eckmann JP, Tlusty T (November 2022). Echave J (ed.).
430:
370:
98:. The wrong binding probability decreases before the correct one.
242:
238:
170:
166:
126:
52:
48:
194:
Ribosome uses conformational proofreading for tRNA decoding
88:
Conformational proofreading in homologous recombination
112:
Use by homologous recombination for homology search
474:Rambo RP, Williams GJ, Tainer JA (November 2010).
523:
521:
519:
469:
467:
465:
411:
409:
407:
405:
294:Savir Y, Tlusty T (May 2007). Scalas E (ed.).
8:
229:-induced damage during the initial step of
925:
915:
866:
856:
807:
715:
705:
656:
602:
553:
499:
429:
369:
329:
319:
188:
82:
283:
38:Balancing correct and incorrect binding
169:by pairing their codons with matching
7:
1003:Mathematical and theoretical biology
626:
624:
622:
351:
349:
289:
287:
16:Mechanism in molecular recognition
14:
153:Use by ribosome for tRNA decoding
528:Savir Y, Tlusty T (April 2013).
358:IEEE J Sel Topics Signal Process
262:Relation to kinetic proofreading
637:Molecular Biology and Evolution
235:UV-damaged DNA-binding protein
1:
963:10.1016/S0300-9084(75)80139-8
680:Koshland DE (February 1958).
890:Hopfield JJ (October 1974).
809:10.1016/j.molcel.2012.03.029
492:10.1016/j.molcel.2010.10.032
440:10.1016/j.molcel.2010.10.020
321:10.1371/journal.pone.0000468
175:large conformational changes
216:In other biological systems
184:conformational proofreading
146:conformational proofreading
118:conformational proofreading
20:Conformational proofreading
1029:
555:10.1016/j.cell.2013.03.032
231:nucleotide excision repair
47:has to select the correct
26:is a general mechanism of
388:10.1109/JSTSP.2008.923859
257:Other recognition schemes
122:Homologous recombination
24:conformational selection
917:10.1073/pnas.71.10.4135
858:10.1073/pnas.1323856111
165:process, has to decode
649:10.1093/molbev/msac217
221:Human UV-damage repair
212:
108:
192:
86:
28:molecular recognition
707:10.1073/pnas.44.2.98
268:kinetic proofreading
908:1974PNAS...71.4135H
849:2014PNAS..111E1862G
843:(18): E1862–E1871.
761:10.1038/nature06971
753:2008Natur.453..489C
698:1958PNAS...44...98K
595:2008Natur.453..701A
546:2013APS..MARY46006T
380:2008ISTSP...2..390S
312:2007PLoSO...2..468S
102:: As a result, the
213:
109:
902:(10): 4135–4139.
747:(7194): 489–484.
132:energetic barrier
116:The mechanism of
63:, in particular,
51:that matches the
32:energetic barrier
1020:
988:Related articles
975:
974:
946:
940:
939:
929:
919:
887:
881:
880:
870:
860:
828:
822:
821:
811:
787:
781:
780:
736:
730:
729:
719:
709:
677:
671:
670:
660:
628:
617:
616:
606:
574:
568:
567:
557:
525:
514:
513:
503:
471:
460:
459:
433:
413:
400:
399:
373:
353:
344:
343:
333:
323:
291:
1028:
1027:
1023:
1022:
1021:
1019:
1018:
1017:
993:
992:
984:
979:
978:
948:
947:
943:
889:
888:
884:
830:
829:
825:
789:
788:
784:
738:
737:
733:
679:
678:
674:
643:(11): msac217.
630:
629:
620:
604:10.1038/453701e
577:Alon U (2008).
576:
575:
571:
527:
526:
517:
473:
472:
463:
415:
414:
403:
355:
354:
347:
293:
292:
285:
280:
264:
259:
248:
223:
218:
155:
141:
114:
40:
17:
12:
11:
5:
1026:
1024:
1016:
1015:
1010:
1005:
995:
994:
991:
990:
983:
982:External links
980:
977:
976:
957:(5): 587–595.
941:
882:
823:
802:(5): 616–624.
796:Molecular Cell
782:
731:
672:
618:
579:"Journal Club"
569:
540:(2): 471–479.
515:
486:(3): 347–348.
480:Molecular Cell
461:
424:(3): 388–396.
418:Molecular Cell
401:
364:(3): 390–399.
345:
282:
281:
279:
276:
263:
260:
258:
255:
246:
222:
219:
217:
214:
154:
151:
140:
137:
113:
110:
96:energy barrier
61:energy barrier
57:lock and a key
39:
36:
15:
13:
10:
9:
6:
4:
3:
2:
1025:
1014:
1011:
1009:
1006:
1004:
1001:
1000:
998:
989:
986:
985:
981:
972:
968:
964:
960:
956:
952:
945:
942:
937:
933:
928:
923:
918:
913:
909:
905:
901:
897:
893:
886:
883:
878:
874:
869:
864:
859:
854:
850:
846:
842:
838:
834:
827:
824:
819:
815:
810:
805:
801:
797:
793:
786:
783:
778:
774:
770:
766:
762:
758:
754:
750:
746:
742:
735:
732:
727:
723:
718:
713:
708:
703:
699:
695:
692:(2): 98–104.
691:
687:
683:
676:
673:
668:
664:
659:
654:
650:
646:
642:
638:
634:
627:
625:
623:
619:
614:
610:
605:
600:
596:
592:
589:(7196): 701.
588:
584:
580:
573:
570:
565:
561:
556:
551:
547:
543:
539:
535:
531:
524:
522:
520:
516:
511:
507:
502:
497:
493:
489:
485:
481:
477:
470:
468:
466:
462:
457:
453:
449:
445:
441:
437:
432:
427:
423:
419:
412:
410:
408:
406:
402:
397:
393:
389:
385:
381:
377:
372:
367:
363:
359:
352:
350:
346:
341:
337:
332:
327:
322:
317:
313:
309:
305:
301:
297:
290:
288:
284:
277:
275:
273:
269:
261:
256:
254:
253:
244:
240:
236:
232:
228:
220:
215:
210:
206:
203:
199:
195:
191:
187:
185:
181:
176:
172:
168:
164:
160:
152:
150:
147:
138:
136:
133:
128:
123:
119:
111:
105:
101:
97:
93:
89:
85:
81:
78:
74:
69:
67:
62:
58:
54:
50:
46:
37:
35:
33:
29:
25:
21:
1008:Biomolecules
954:
950:
944:
899:
895:
885:
840:
836:
826:
799:
795:
785:
744:
740:
734:
689:
685:
675:
640:
636:
586:
582:
572:
537:
533:
483:
479:
421:
417:
361:
357:
303:
299:
271:
265:
251:
224:
208:
204:
201:
197:
193:
183:
179:
174:
156:
145:
142:
131:
117:
115:
103:
99:
91:
87:
72:
64:
41:
23:
19:
18:
306:(5): e468.
163:translation
77:induced fit
997:Categories
278:References
272:time delay
270:schema, a
209:symmetric
73:off target
66:structural
951:Biochimie
431:1011.4382
371:1007.4527
180:symmetric
877:24760829
818:22560720
769:18497818
726:16590179
667:36208205
613:29639642
564:23582332
510:21070960
448:21070965
340:17520027
300:PLOS ONE
159:ribosome
68:mismatch
45:ribosome
1013:Enzymes
971:1182215
936:4530290
904:Bibcode
868:4020048
845:Bibcode
777:4416531
749:Bibcode
694:Bibcode
658:9641994
591:Bibcode
542:Bibcode
501:3003302
456:1682936
396:7210763
376:Bibcode
331:1868595
308:Bibcode
266:In the
104:Fitness
969:
934:
927:434344
924:
875:
865:
816:
775:
767:
741:Nature
724:
717:335371
714:
665:
655:
611:
583:Nature
562:
508:
498:
454:
446:
394:
338:
328:
207:: The
100:Bottom
773:S2CID
609:S2CID
452:S2CID
426:arXiv
392:S2CID
366:arXiv
205:inset
171:tRNAs
167:mRNAs
967:PMID
932:PMID
873:PMID
814:PMID
765:PMID
722:PMID
663:PMID
560:PMID
534:Cell
506:PMID
444:PMID
336:PMID
243:DDB2
239:DDB1
198:main
157:The
127:RecA
53:mRNA
49:tRNA
959:doi
922:PMC
912:doi
863:PMC
853:doi
841:111
804:doi
757:doi
745:453
712:PMC
702:doi
653:PMC
645:doi
599:doi
587:453
550:doi
538:153
496:PMC
488:doi
436:doi
384:doi
326:PMC
316:doi
92:Top
22:or
999::
965:.
955:57
953:.
930:.
920:.
910:.
900:71
898:.
894:.
871:.
861:.
851:.
839:.
835:.
812:.
800:46
798:.
794:.
771:.
763:.
755:.
743:.
720:.
710:.
700:.
690:44
688:.
684:.
661:.
651:.
641:39
639:.
635:.
621:^
607:.
597:.
585:.
581:.
558:.
548:.
536:.
532:.
518:^
504:.
494:.
484:40
482:.
478:.
464:^
450:.
442:.
434:.
422:40
420:.
404:^
390:.
382:.
374:.
360:.
348:^
334:.
324:.
314:.
302:.
298:.
286:^
227:UV
196:.
90:.
973:.
961::
938:.
914::
906::
879:.
855::
847::
820:.
806::
779:.
759::
751::
728:.
704::
696::
669:.
647::
615:.
601::
593::
566:.
552::
544::
512:.
490::
458:.
438::
428::
398:.
386::
378::
368::
362:2
342:.
318::
310::
304:2
247:2
245:)
241:-
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.