110:
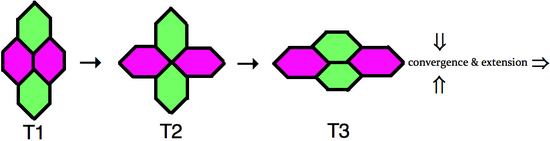
anterior-posterior neighbors selectively shrinks, resulting in an obligatory intermediate type 2 junction, where the four cells share a vertex. Upon resolution of the type 2 junction, a new type 3 junction forms perpendicular to the original type 1 configuration. During this process the two dorsal-ventral cells have become neighbors. When multiple clusters of cells intercalate in the dorsal-ventral axis, through junctional neighbor exchange, the outcome is an extension of germ-band in the anterior-posterior axis.
65:
embryo, effectively folding over onto the dorsal side of the egg. Multiple individual cells intercalating mediolateral to the anterior-posterior axis drive the resulting global elongation of the embryo. In addition, cell shape changes, and oriented cell divisions in the posterior germ-band are in part required for full elongation. However, elongation of the body axis seems to be primarily linked to changes in cell neighbor relations.
20:
123:
89:
113:
In addition to the simple neighbor exchange, higher-ordered rosette formations have been observed in which five or more cells meet at a vertex. Multicellular rosettes form and resolve in a directional fashion that promotes germ-band elongation. Neighbor exchange and multicellular rosette formation
64:
and is divided into two phases. The fast phase, in which most of the extension occurs, takes about 25 minutes. The remaining extension continues during the slow phase and is completed in the following 70 minutes. During this process the ventral germ-band extends around the posterior end of the
183:
provides an anterior-posterior pulling force that contributes to germ-band extension through passive cell shape changes. Although anterior-posterior patterning mutants fail to fully extend their germ-bands, during the fast phase the elongation length is normal despite defects in polarized cell
27:
embryo at the start of germ-band elongation and 30 minutes later. The germ-band (grey) is posterior to the cephalic furrow (curved-line) and folds dorsally (red arrow) upon cell intercalation. Each rectangle represents a field of cells before (0 min) and after (30 min) convergent extension.
109:
has captured this process of cell neighbor exchange, which is schematically represented to the right. In the type 1 configuration, two cells contact each other along the anterior-posterior axis, whereas two dorsal-ventral cells do not directly contact. Next, the cell boundary between the two
143:
mutants are defective in germ-band extension, which supports the idea that polarized protein localization is critical for cell rearrangements. One mechanism by which myosin II might promote polarized cell remodeling is through contractile activity that creates tension orienting junctional
147:
The source that establishes planar polarity during germ-band extension remains elusive. Polarized intercalation is largely unaffected in mutant embryos that lack dorsal-ventral cell types. Yet, mutations that disrupt segmental patterning along the anterior-posterior axis, such as
191:
In addition, there is evidence that mechanical tension is necessary and sufficient for the cortical localization of myosin II. Thus, not only can myosin II generate tension but it may also be up-regulated by tensile forces, creating a
184:
intercalation. Time-lapse analysis revealed that an increase in cell shape stretching in the anterior-posterior axis was compensating for aberrant cell intercalation, independent of anterior-posterior patterning. Furthermore, during
92:
Schematic of neighbor exchange or an elementary T1 process involving four cells. T1-magenta cells are in direct contact. T2-all cells share a common vertex. T3-resolution results in green cells sharing a common
138:
localizes to the anterior-posterior boundaries of cells, destabilizing adherens junctions, whereas the
Bazooka/Par-3 complex localizes to dorsal-ventral boundaries, stabilizing adherens junctions. Moreover,
308:
Butler, L.C.; Blanchard, G.B.; Kabla, A.J.; Lawrence, N.J.; Welchman, D.P.; Mahadevan, L.; Adams, R.J.; Sanson, B. (2009). "Cell shape changes indicate a role for extrinsic tensile forces in
164:
of eve or runt is sufficient to locally reorient the polarity of nearby cells. This evidence argues that planar polarity is established by cell-cell interactions, and not by a
188:
development, it has been suggested that intercalary cell behavior relaxes the stress imposed on the germ-band, allowing stretched cells to restore to isometric shapes.
130:
The dorsal-ventral intercalation of cells during germ-band extension ultimately arises from the asymmetric localization of proteins within individual cells.
114:
involve oriented junctional remodeling, which indicates that the intercalating cells are intrinsically polarized within the plane of the epithelium.
144:
disassembly. However, the precise mechanism in which asymmetrically localized protein complexes encourage directed intercalation remains disputed.
388:
Bertet, C.; Sulak, L.; Lecuit, T. (2004). "Myosin-dependent junction remodelling controls planar cell intercalation and axis elongation".
168:. Thus, polarizing information can spread from one cell to the next, downstream of an Eve-dependent signal that remains to be identified.
621:
40:
126:
Asymmetric localization of proteins to reciprocal cell borders in the apical plane of a polarized epithelial.
45:
106:
53:, which develops into the segmented trunk of the embryo, approximately doubles in length along the
161:
131:
98:
196:
loop that allows cells to dynamically respond to fluctuations in their mechanical environment.
72:. However, the basis of germ-band elongation is applicable to many organisms including other
586:
535:
494:
446:
405:
370:
329:
287:
242:
193:
576:
566:
525:
484:
436:
397:
360:
321:
277:
232:
553:
Fernandez-Gonzalez, R.; de Matos Simoes, S.; Röper J.-C.; Eaton, S.; Zallen, J.A. (2009).
28:
Colored nuclei arbitrarily mark rows of cells in order to visualize tissue morphogenesis.
581:
554:
156:
150:
489:
468:
615:
73:
36:
425:"Multicellular rosette formation links planar cell polarity to tissue morphogenesis"
180:
61:
19:
571:
441:
424:
423:
Blankenship, J.T.; Backovic, S.T.; Sanny, J.S.; Weitz, O.; Zallen, J.A. (2006).
160:, decrease cell intercalation and subsequent germ-band elongation. Furthermore,
77:
530:
513:
102:
50:
606:
185:
165:
135:
122:
590:
539:
498:
450:
409:
374:
333:
291:
246:
237:
216:
177:
54:
401:
282:
261:
221:
germ-band extension and its regulation by pair-rule segmentation genes"
88:
365:
348:
555:"Myosin II dynamics are regulated by tension in intercalating cells"
325:
57:
axis while subsequently narrowing along the dorsal-ventral axis.
469:"Patterned gene expression directs bipolar planar polarity in
97:
In order for cells to intercalate between one another the
262:"Oriented cell divisions in the extending germband of
303:
301:
8:
349:"Mechanisms of elongation in embryogenesis"
68:This article describes axis elongation in
580:
570:
529:
488:
440:
364:
281:
236:
60:Germ-band extension begins shortly after
121:
87:
18:
16:Morphogenic process during embryogenesis
607:Jennifer Zallen's Lab - Sloan-Kettering
462:
460:
260:da Silva, S.M.; Vincent, J.-P. (2007).
210:
208:
204:
105:tissue must be dynamically remodeled.
7:
467:Zallen, J.A.; Wieschaus, E. (2004).
514:"Animal development: Crowd control"
101:that maintain the integrity of the
215:Irvine, K.; Wieschaus, E. (1994).
14:
1:
490:10.1016/s1534-5807(04)00060-7
572:10.1016/j.devcel.2009.09.003
442:10.1016/j.devcel.2006.09.007
217:"Cell intercalation during
638:
39:process widely studied in
531:10.1016/j.cub.2004.08.047
176:Researchers suggest that
166:long-range polarizing cue
134:reveals that non-muscle
46:Drosophila melanogaster
312:germ-band extension".
127:
94:
29:
622:Developmental biology
238:10.1242/dev.120.4.827
125:
107:Time-lapse microscopy
91:
22:
402:10.1038/nature02590
347:Keller, R. (2006).
314:Nature Cell Biology
33:Germ-band extension
559:Developmental Cell
477:Developmental Cell
429:Developmental Cell
283:10.1242/dev.004911
162:ectopic expression
132:Immunofluorescence
128:
99:adherens junctions
95:
55:anterior-posterior
30:
524:(17): R716–R718.
512:Baum, B. (2004).
396:(6992): 667–671.
366:10.1242/dev.02406
359:(12): 2291–2302.
276:(17): 3049–3054.
194:positive feedback
629:
595:
594:
584:
574:
550:
544:
543:
533:
509:
503:
502:
492:
464:
455:
454:
444:
420:
414:
413:
385:
379:
378:
368:
344:
338:
337:
305:
296:
295:
285:
257:
251:
250:
240:
212:
637:
636:
632:
631:
630:
628:
627:
626:
612:
611:
603:
598:
552:
551:
547:
518:Current Biology
511:
510:
506:
466:
465:
458:
422:
421:
417:
387:
386:
382:
346:
345:
341:
326:10.1038/ncb1894
307:
306:
299:
259:
258:
254:
214:
213:
206:
202:
174:
120:
118:Molecular basis
86:
41:the development
17:
12:
11:
5:
635:
633:
625:
624:
614:
613:
610:
609:
602:
601:External links
599:
597:
596:
565:(5): 736–743.
545:
504:
483:(3): 343–355.
456:
435:(4): 459–470.
415:
380:
339:
320:(7): 859–864.
297:
252:
231:(4): 827–841.
203:
201:
198:
173:
172:Tensile forces
170:
119:
116:
85:
84:Cellular basis
82:
15:
13:
10:
9:
6:
4:
3:
2:
634:
623:
620:
619:
617:
608:
605:
604:
600:
592:
588:
583:
578:
573:
568:
564:
560:
556:
549:
546:
541:
537:
532:
527:
523:
519:
515:
508:
505:
500:
496:
491:
486:
482:
478:
474:
472:
463:
461:
457:
452:
448:
443:
438:
434:
430:
426:
419:
416:
411:
407:
403:
399:
395:
391:
384:
381:
376:
372:
367:
362:
358:
354:
350:
343:
340:
335:
331:
327:
323:
319:
315:
311:
304:
302:
298:
293:
289:
284:
279:
275:
271:
267:
265:
256:
253:
248:
244:
239:
234:
230:
226:
222:
220:
211:
209:
205:
199:
197:
195:
189:
187:
182:
179:
171:
169:
167:
163:
159:
158:
153:
152:
145:
142:
137:
133:
124:
117:
115:
111:
108:
104:
100:
90:
83:
81:
79:
75:
74:invertebrates
71:
66:
63:
58:
56:
52:
49:in which the
48:
47:
42:
38:
34:
26:
23:Schematic of
21:
562:
558:
548:
521:
517:
507:
480:
476:
470:
432:
428:
418:
393:
389:
383:
356:
352:
342:
317:
313:
309:
273:
269:
263:
255:
228:
224:
218:
190:
181:invagination
175:
155:
149:
146:
140:
129:
112:
96:
69:
67:
62:gastrulation
59:
44:
32:
31:
24:
353:Development
270:Development
225:Development
78:vertebrates
37:morphogenic
471:Drosophila
310:Drosophila
264:Drosophila
219:Drosophila
200:References
103:epithelial
70:Drosophila
25:Drosophila
186:wild type
136:myosin II
93:boundary.
51:germ-band
616:Category
591:19879198
540:15341764
499:15030758
451:17011486
410:15190355
375:16720874
334:19503074
292:17652351
178:mesoderm
582:2854079
247:7600960
141:bazooka
80:alike.
589:
579:
538:
497:
449:
408:
390:Nature
373:
332:
290:
245:
35:is a
587:PMID
536:PMID
495:PMID
447:PMID
406:PMID
371:PMID
330:PMID
288:PMID
243:PMID
157:runt
154:and
76:and
577:PMC
567:doi
526:doi
485:doi
437:doi
398:doi
394:429
361:doi
357:133
322:doi
278:doi
274:134
233:doi
229:120
151:eve
43:of
618::
585:.
575:.
563:17
561:.
557:.
534:.
522:14
520:.
516:.
493:.
479:.
475:.
459:^
445:.
433:11
431:.
427:.
404:.
392:.
369:.
355:.
351:.
328:.
318:11
316:.
300:^
286:.
272:.
268:.
241:.
227:.
223:.
207:^
593:.
569::
542:.
528::
501:.
487::
481:6
473:"
453:.
439::
412:.
400::
377:.
363::
336:.
324::
294:.
280::
266:"
249:.
235::
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.