20:
121:
found in low densities, such as some cell surface antigens. As the resolution of scanning electron microscopy (SEM) increased, so too did the need for nanoparticle-sized labels such as immunogold. In 1975, Horisberger and coworkers successfully visualised gold nanoparticles with a diameter of less
141:
The prepared sample is then incubated with a specific antibody designed to bind the molecule of interest. Next, a secondary antibody which has gold particles attached is added, and it binds to the primary antibody. Gold can also be attached to
677:
Van Laere O, De Wael L, De Mey J (1985). "Immuno gold staining (IGS) and immuno gold silver staining (IGSS) for the identification of the plant pathogenic bacterium
Erwinia amylovora (Burrill) Winslow et al".
192:. The silver enhancement increases the particle size, also making scanning electron microscopy possible. In order to produce the silver-enhanced gold particles, colloidal gold particles are placed in an
255:
used to embed samples for imaging. Thus, only accessible molecules can be targeted and visualized. Labeling prior to embedding the sample can reduce the negative impact of this limitation.
100:
Immunogold labeling can introduce artifacts, as the gold particles reside some distance from the labelled object and very thin sectioning is required during sample preparation.
225:
An inherent limitation to the immunogold technique is that the gold particle is around 15-30 nm away from the site to which the primary antibody is bound (when using a
244:
may appear as a 'spike' depending on which plane the sectioning occurred. To overcome this limitation serial sections can be taken, which can then be compiled into a
229:
labeling strategy). The precise location of the targeted molecule can therefore not be accurately calculated. Gold particles can be created with a diameter of 1
236:
Thin sections are required for immunogold labeling and these can produce misleading images; a thin slice of a cell component may not give an accurate view of its
208:
site and silver is deposited onto the particle. An example of the application of silver-enhanced immunogold labeling (IGSS) was in the identification of the
172:
by using two different-sized gold particles. An extension of this method used three different sized gold particles to track the localisation of regulatory
385:
188:
Although immunogold labeling is typically used for transmission electron microscopy, when the gold is 'silver-enhanced' it can be seen using
160:
The electron-dense gold particle can now be seen under an electron microscope as a black dot, indirectly labeling the molecule of interest.
19:
728:
233:(or lower) but another limitation is then realized—at these sizes the gold label becomes hard to distinguish from tissue structure.
226:
86:
63:
340:
90:
51:
97:. The labeling technique can be adapted to distinguish multiple objects by using differently-sized gold particles.
23:
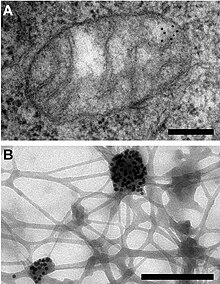
Two images produced using immunogold labeling and transmission electron microscopy: (A) Gold particles are marking
559:"Coloidal gold, ferritin and peroxidase as markers for electron microscopic double labeling lectin techniques"
600:"Double immunogold staining method for the simultaneous ultrastructural localization of regulatory peptides"
245:
117:. It was first applied in transmission electron microscopy (TEM) and was especially useful in highlighting
189:
168:
Immunogold labeling can be used to visualize more than one target simultaneously. This can be achieved in
94:
28:
718:
639:
Bendayan M (January 1982). "Double immunocytochemical labeling applying the protein A-gold technique".
264:
197:
723:
169:
135:
47:
237:
180:
separately, the immunogold particles attached to both sides can then be viewed simultaneously.
695:
656:
621:
580:
539:
504:
460:
443:
Hermann R, Walther P, Müller M (1996). "Immunogold labeling in scanning electron microscopy".
418:
381:
317:
213:
483:"Ultrastructural localization of intracellular antigens by the use of protein A-gold complex"
687:
648:
611:
570:
531:
494:
452:
410:
401:
Faulk WP, Taylor GM (November 1971). "An immunocolloid method for the electron microscope".
307:
297:
151:
75:
71:
55:
312:
284:
176:. A more complex method of multi-site labeling involves labeling opposite sides of an
712:
414:
251:
A further limitation is that antibodies and gold particles cannot penetrate the
241:
344:
205:
110:
59:
289:
230:
154:
147:
143:
131:
543:
522:
Porter, K; Blum, J (1953). "A study in
Microtomy for Electron Microscopy".
321:
108:
Immunogold labeling was first used in 1971 by Faulk and Taylor to identify
699:
660:
625:
535:
464:
422:
302:
652:
616:
599:
584:
508:
499:
482:
209:
173:
114:
79:
575:
558:
691:
456:
177:
118:
67:
285:"The functional organization of mitochondrial genomes in human cells"
193:
122:
than 30 nm and this soon became an established SEM technique.
598:
Tapia FJ, Varndell IM, Probert L, De Mey J, Polak JM (July 1983).
252:
150:
instead of a secondary antibody, as these proteins bind mammalian
24:
18:
85:
First used in 1971, immunogold labeling has been applied to both
201:
130:
First, a thin section of the sample is cut, often using a
31:(B) mtDNA marked with gold particles after extraction.
341:"Immunogold Labeling in Scanning Electron Microscopy"
50:. This staining technique is an equivalent of the
672:
670:
58:particles are most often attached to secondary
16:Staining technique used in electron microscopy
8:
481:Roth J, Bendayan M, Orci L (December 1978).
82:scatter to give high contrast 'dark spots'.
380:(5th ed.). New York: Garland Science.
615:
574:
498:
311:
301:
371:
369:
367:
365:
363:
361:
275:
476:
474:
335:
333:
331:
283:Iborra FJ, Kimura H, Cook PR (2004).
74:component. Gold is used for its high
7:
438:
436:
434:
432:
376:Alberts, Bruce; et al. (2008).
46:) is a staining technique used in
14:
227:primary and secondary antibodies
87:transmission electron microscopy
557:Roth J, Binder M (March 1978).
445:Histochemistry and Cell Biology
204:. Gold particles then act as a
184:Uses in brightfield microscopy
62:which are in turn attached to
1:
378:Molecular biology of the cell
54:technique for visible light.
415:10.1016/0019-2791(71)90496-4
91:scanning electron microscopy
66:designed to bind a specific
238:three-dimensional structure
52:indirect immunofluorescence
745:
729:Electron microscopy stains
134:. Various other stages of
164:Labeling multiple objects
157:in a non-specific way.
641:J. Histochem. Cytochem
604:J. Histochem. Cytochem
563:J. Histochem. Cytochem
487:J. Histochem. Cytochem
190:brightfield microscopy
95:brightfield microscopy
32:
536:10.1002/ar.1091170403
524:The Anatomical Record
303:10.1186/1741-7007-2-9
138:may then take place.
22:
653:10.1177/30.1.6172469
617:10.1177/31.7.6189888
500:10.1177/26.12.366014
265:Immunohistochemistry
576:10.1177/26.3.632554
170:electron microscopy
48:electron microscopy
40:immunogold staining
36:Immunogold labeling
692:10.1007/BF00509198
457:10.1007/BF02473200
200:containing silver
136:sample preparation
64:primary antibodies
33:
387:978-0-8153-4106-2
246:three-dimensional
240:. For example, a
214:Erwinia amylovora
736:
704:
703:
674:
665:
664:
636:
630:
629:
619:
595:
589:
588:
578:
554:
548:
547:
519:
513:
512:
502:
478:
469:
468:
440:
427:
426:
398:
392:
391:
373:
356:
355:
353:
352:
343:. Archived from
337:
326:
325:
315:
305:
280:
78:which increases
76:electron density
744:
743:
739:
738:
737:
735:
734:
733:
709:
708:
707:
676:
675:
668:
638:
637:
633:
597:
596:
592:
556:
555:
551:
521:
520:
516:
493:(12): 1074–81.
480:
479:
472:
442:
441:
430:
403:Immunochemistry
400:
399:
395:
388:
375:
374:
359:
350:
348:
339:
338:
329:
282:
281:
277:
273:
261:
223:
186:
166:
128:
106:
17:
12:
11:
5:
742:
740:
732:
731:
726:
721:
711:
710:
706:
705:
680:Histochemistry
666:
631:
590:
549:
530:(4): 685–710.
514:
470:
428:
409:(11): 1081–3.
393:
386:
357:
327:
274:
272:
269:
268:
267:
260:
257:
222:
219:
185:
182:
178:antigenic site
165:
162:
127:
124:
105:
102:
56:Colloidal gold
15:
13:
10:
9:
6:
4:
3:
2:
741:
730:
727:
725:
722:
720:
717:
716:
714:
701:
697:
693:
689:
685:
681:
673:
671:
667:
662:
658:
654:
650:
646:
642:
635:
632:
627:
623:
618:
613:
610:(7): 977–81.
609:
605:
601:
594:
591:
586:
582:
577:
572:
568:
564:
560:
553:
550:
545:
541:
537:
533:
529:
525:
518:
515:
510:
506:
501:
496:
492:
488:
484:
477:
475:
471:
466:
462:
458:
454:
450:
446:
439:
437:
435:
433:
429:
424:
420:
416:
412:
408:
404:
397:
394:
389:
383:
379:
372:
370:
368:
366:
364:
362:
358:
347:on 2014-02-06
346:
342:
336:
334:
332:
328:
323:
319:
314:
309:
304:
299:
295:
292:
291:
286:
279:
276:
270:
266:
263:
262:
258:
256:
254:
249:
247:
243:
239:
234:
232:
228:
220:
218:
216:
215:
211:
207:
203:
199:
195:
191:
183:
181:
179:
175:
171:
163:
161:
158:
156:
153:
149:
145:
139:
137:
133:
125:
123:
120:
116:
113:
112:
103:
101:
98:
96:
93:, as well as
92:
88:
83:
81:
77:
73:
69:
65:
61:
57:
53:
49:
45:
41:
37:
30:
26:
21:
719:Biochemistry
686:(5): 397–9.
683:
679:
644:
640:
634:
607:
603:
593:
569:(3): 163–9.
566:
562:
552:
527:
523:
517:
490:
486:
451:(1): 31–39.
448:
444:
406:
402:
396:
377:
349:. Retrieved
345:the original
293:
288:
278:
250:
235:
224:
212:
187:
167:
159:
140:
129:
109:
107:
99:
84:
43:
39:
35:
34:
29:mitochondria
647:(1): 81–5.
242:microtubule
221:Limitations
724:Microscopy
713:Categories
351:2010-07-08
271:References
206:nucleation
196:enhancing
155:Fc regions
111:Salmonella
60:antibodies
290:BMC Biol.
148:protein G
144:protein A
132:microtome
126:Technique
70:or other
27:near the
544:13124776
322:15157274
259:See also
210:pathogen
198:solution
174:peptides
119:proteins
115:antigens
80:electron
700:2416717
661:6172469
626:6189888
465:8858365
423:4110101
248:image.
104:History
68:antigen
698:
659:
624:
585:632554
583:
542:
509:366014
507:
463:
421:
384:
320:
313:425603
310:
194:acidic
296:: 9.
253:resin
25:mtDNA
696:PMID
657:PMID
622:PMID
581:PMID
540:PMID
505:PMID
461:PMID
419:PMID
382:ISBN
318:PMID
202:ions
89:and
72:cell
688:doi
649:doi
612:doi
571:doi
532:doi
528:117
495:doi
453:doi
449:106
411:doi
308:PMC
298:doi
152:IgG
146:or
44:IGS
38:or
715::
694:.
684:83
682:.
669:^
655:.
645:30
643:.
620:.
608:31
606:.
602:.
579:.
567:26
565:.
561:.
538:.
526:.
503:.
491:26
489:.
485:.
473:^
459:.
447:.
431:^
417:.
405:.
360:^
330:^
316:.
306:.
287:.
231:nm
217:.
702:.
690::
663:.
651::
628:.
614::
587:.
573::
546:.
534::
511:.
497::
467:.
455::
425:.
413::
407:8
390:.
354:.
324:.
300::
294:2
42:(
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.