123:(SEF). LGFMS tumors present with many clinical and pathological features that are similar to those in SEF. Indeed, current studies suggest that LGFMS may be an early form of SEF. For example, a tumor may present with features typical of LGFMS but over time progress to features typical of sclerosing epithelioid fibrosarcomas. This progression may be particularly evident in recurrent or metastatic "LGFMS" tumors. Since the World Health Organization has classified LGFMS as one of the malignant fibroblastic and myofibroblastic tumors that is distinctly different than SEF, SEF and LGFMS are here regarded as separate tumor forms.
29:
443:
surgically resected LGFMS tumors have tended to recur at the site or resection in up to 75% of cases. These recurrences can develop as long as 15 years (median: 3.5 years) after the initial diagnosis of the disease. Following initial surgical resection, metastases have been reported to develop in up to 50% of individuals. These metastases have been diagnosed up to 45 years (median: 5 years) after the initial diagnosis of LGFMS. They were found mostly in the lungs, pleura, and chest wall, occasionally in the
447:(i.e. the sac containing the heart), and rarely in a bone, the liver, or the heart. Recurrences of the tumors at the site of their resection as well as many of their metastases have been treated by surgical resections; this treatment may be repeated multiple times in individuals who develop repeated local recurrences and/or metastases.
156:, or abdominal cavity. Tumors occurring in subcutaneous and subfacial tissues are often 3–11 cm in size (largest diameter) but in sites where the tumor can go undetected for long periods such as the buttocks, mediastinum, or abdominal cavity have been reported to be as large as 25, 23.5, and 50 cm, respectively.
442:
The generally recommended treatment for LGFMS is aggressive, wide surgical excision of the tumor including even those located in difficult to access sites such as the mediastinum. Complete removal of all tumor tissue is critical in lowering the rates of recurrences and metastases. Over the long term,
1311:
Krystel-Whittemore M, Taylor MS, Rivera M, Lennerz JK, Le LP, Dias-Santagata D, Iafrate AJ, Deshpande V, Chebib I, Nielsen GP, Wu CL, Nardi V (November 2019). "Novel and established EWSR1 gene fusions and associations identified by next-generation sequencing and fluorescence in-situ hybridization".
687:
Martínez-Trufero J, Cruz Jurado J, Gómez-Mateo MC, Bernabeu D, Floría LJ, Lavernia J, Sebio A, García Del Muro X, Álvarez R, Correa R, Hernández-León CN, Marquina G, Hindi N, Redondo A, Martínez V, Asencio JM, Mata C, Valverde
Morales CM, Martin-Broto J (September 2021). "Uncommon and peculiar soft
254:
protein is express by the tumor cells of almost all LGFMS cases and therefore is commonly used as a sensitive and specific marker for LGFMS. (Since a small subset of LGFMS tumors have been reported to consist of cells that are MUC4-negative, the diagnosis of LGFMS should not be absolutely dependent
320:
fusion gene is expressed in a minority of LGMFS cases. Fusion genes containing part of an AFET gene family gene occur in a wide range of tumors and are thought to contribute to the development of at least some of these tumors. While the role of the cited fusion genes in the development of LGFMS is
131:
Individuals initially diagnosed with LGFMS had been characterized as young adults (mean age: 33 years) or children >5 years old (13% to 19% of all cases), although a more recent study reported that individuals first diagnosed with the disease ranged in age between 6 and 67 years (mean age: 45.6
456:
to treat LGFMS are limited and this tumor does not appear very sensitive to either treatment modality. In one long-term study of 33 treated individuals with LGFMS, 14 died of their tumors from 3 to 42 years (median: 15 years) after their initial diagnosis and 19 were alive after 5½ to 70 years
107:
LGFMS tumors occur in individuals of almost any age but up to 20% are less than 18 years/old. The tumors typically involve the proximal extremities but can occur virtually anywhere in the body. The progression of these tumors commonly takes an indolent, prolonged course involving many years or
200:
and other elements that resemble, or are indistinguishable from, sclerosing epithelioid fibrosarcoma. This hybrid pattern is more often seen in recurrent and metastatic LGGMS tumors, may progress to dominate the tissues, and may therefore warrant changing the diagnosis of LGFMD to sclerosing
636:
Porrino J, Al-Dasuqi K, Irshaid L, Wang A, Kani K, Haims A, Maloney E (June 2021). "Update of pediatric soft tissue tumors with review of conventional MRI appearance-part 1: tumor-like lesions, adipocytic tumors, fibroblastic and myofibroblastic tumors, and perivascular tumors".
1506:
Porteus C, Gan Q, Gong Y, Pantanowitz L, Henderson-Jackson E, Saeed-Vafa D, Mela N, Peterson D, Ahmad N, Ahmed A, Bui M (2020). "Sclerosing epithelioid fibrosarcoma: cytologic characterization with histologic, immunohistologic, molecular, and clinical correlation of 8 cases".
451:
combined with partial surgical resection or radiation therapy alone have been used in cases where the primary, recurrent, or metastatic tumor cannot be safely or totally removed. However, the experiences with radiation therapy and
201:
epithelioid fibrosarcoma. While not definitive, LGMFS tumors that are dominated with cells that have an undifferentiated or immature round-cell morphology may be more aggressive than tumors with the typical cell morphology.
382:. However, sclerosing epithelioid fibrosarcoma tumors may contain areas that are histopathologically indistinguishable from LGFMS and contain neoplastic cells that expression MUC4 protein in most cases, and a
353:
findings ma also help in the diagnosis. These findings can differentiate LGFMS from various spindle-shaped cell and myxoid neoplasms including benign soft tissue tumors, malignant soft tissue tumors (e.g.
172:
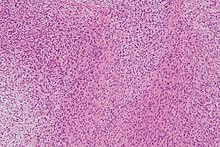
LGMFS tumors typically contain one or multiple nodules embedded in a grey-white whorled cut surface. They appear to be infiltrating adjacent structures/tissues in a minority of cases. Microscopic
329:
The diagnosis of LGFMS rest primary on its tumors' histology consisting of spindle-shaped fibroblastic cells in the appropriate background that express the MUC4 protein and either a
1217:
Sambri A, Righi A, Tuzzato G, Donati D, Bianchi G (2018). "Low-grade fibromyxoid sarcoma of the extremities: a clinicopathologic study of 24 cases and review of the literature".
192:. The spindle-shaped cells typically occur in an alternating fibrous and myxoid (i.e. more blue or purple compared to normal connective tissue because of excessive uptake of the
263:
The neoplastic cells in LGFMS tumors express one of various fusion genes. Fusion genes are abnormal genes consisting of parts of two different genes that merge as a result of
688:
tissue sarcomas: Multidisciplinary review and practical recommendations for diagnosis and treatment. Spanish group for
Sarcoma research (GEIS - GROUP). Part I".
1586:
1262:
Lindén M, Thomsen C, Grundevik P, Jonasson E, Andersson D, Runnberg R, Dolatabadi S, Vannas C, Luna
Santamaria M, Fagman H, Ståhlberg A, Åman P (May 2019).
1581:
101:
474:
Evans HL (November 1987). "Low-grade fibromyxoid sarcoma. A report of two metastasizing neoplasms having a deceptively benign appearance".
394:
fusion gene in rare cases. Based on the relative rates of expression in the two tumor types, the presence of tumor neoplastic cells that:
371:
120:
1053:"Low-grade fibromyxoid sarcoma incidentally discovered as an asymptomatic mediastinal mass: a case report and review of the literature"
180:
tumor tissue show whorled and bundled, uniform, bland-appearing, slender fibroblastic spindle-shaped cells with elongated, tapered
848:
Mustafa S, VandenBussche CJ, Ali SZ, Siddiqui MT, Wakely PE (2020). "Cytomorphologic findings of low-grade fibromyxoid sarcoma".
1456:
Chew W, Benson C, Thway K, Hayes A, Miah A, Zaidi S, Lee AT, Messiou C, Fisher C, van der Graaf WT, Jones RL (September 2018).
1005:
Saab-Chalhoub MW, Al-Rohil RN (April 2019). "Low-grade fibromyxoid sarcoma of acral sites: Case report and literature review".
367:
524:"EWSR1-The Most Common Rearranged Gene in Soft Tissue Lesions, Which Also Occurs in Different Bone Lesions: An Updated Review"
359:
116:
tissues surrounding the lungs. These metastasis can develop decades after the tumor's initial presentation and diagnosis.
734:
Evans, Harry L (2011). "Low-Grade
Fibromyxoid Sarcoma: A Clinicopathologic Study of 33 Cases With Long-Term Follow-Up".
28:
346:
157:
1156:"MRI and CT of Low-Grade Fibromyxoid Sarcoma in Children: A Report From Children's Oncology Group Study ARST0332"
268:
100:. The World Health Organization (2020) classified LGFMS as a specific type of tumor in the category of malignant
410:
fusion gene suggest that the tumor is a SEF but in any case is unlikely to be a low-grade fibromyxoid sarcoma.
350:
161:
953:"Low-Grade Fibromyxoid Sarcoma of the Head and Neck: A Clinicopathologic Series and Review of the Literature"
281:
457:(median: 13 years] of their diagnosis with 6 individuals having persistent tumors and 13 being tumor-free.
422:
fusion gene is present in >90% of low-grade fibromyxoid sarcomas but only ~20% of SEF tumor cells}; or
145:
276:
1204:"CREB3L2 cAMP responsive element binding protein 3 like 2 [Homo sapiens (Human)] - Gene - NCBI"
522:
Flucke U, van Noesel MM, Siozopoulou V, Creytens D, Tops BB, van Gorp JM, Hiemcke-Jiwa LS (June 2021).
108:
decades. Over this time, however, the tumors often recur at the site of their surgical removal and/or
355:
204:
272:
133:
84:; on microscopic examination, they are found to be composed of spindle-shaped cells that resemble
1532:
1435:
1383:
1337:
1030:
922:
873:
759:
662:
379:
292:
1524:
1487:
1458:"Clinical Characteristics and efficacy of chemotherapy in sclerosing epithelioid fibrosarcoma"
1427:
1375:
1329:
1293:
1244:
1185:
1136:
1084:
1022:
982:
914:
865:
813:
751:
705:
654:
607:
555:
491:
448:
81:
48:
1596:
1516:
1477:
1469:
1417:
1367:
1321:
1283:
1275:
1234:
1226:
1175:
1167:
1126:
1118:
1074:
1064:
1014:
972:
964:
904:
857:
803:
793:
743:
697:
646:
597:
589:
545:
535:
483:
375:
363:
197:
1107:"Huge mesenteric low-grade fibromyxoid sarcoma: A case report and review of the literature"
893:"Sclerosing epithelioid fibrosarcoma: in-depth review of a genetically heterogeneous tumor"
321:
not known, the presence of one of these fusion genes is helpful in diagnosing the disease.
1591:
1154:
Sargar K, Kao SC, Spunt SL, Hawkins DS, Parham DM, Coffin C, McCarville MB (August 2015).
173:
149:
141:
1482:
1457:
1288:
1263:
1180:
1155:
1131:
1106:
1079:
1052:
977:
952:
808:
781:
602:
577:
550:
523:
119:
LGFMS can be difficult to distinguish from other mesenchymal tumors, particularly from
418:
fusion gene is more likely to be a low-grade fibromyxoid sarcoma than a SEF (i.e. the
1575:
1536:
1439:
1387:
1341:
926:
877:
666:
189:
1034:
782:"MRI findings of low-grade fibromyxoid sarcoma: a case report and literature review"
763:
453:
231:
181:
1406:"Sclerosing Epithelioid Fibrosarcoma: A Distinct Sarcoma With Aggressive Features"
1325:
177:
40:
1422:
1405:
747:
1264:"FET family fusion oncoproteins target the SWI/SNF chromatin remodeling complex"
540:
444:
243:
193:
153:
109:
93:
73:
1520:
1371:
1069:
861:
701:
650:
96:, i.e. genes composed of parts of two different genes that form as a result of
1473:
968:
798:
402:
or an otherwise rearranged FUS gene strongly suggest that the tumor is a SEF;
188:
and do not appear to be rapidly proliferating as defined by their low rate of
89:
85:
77:
36:
1122:
1279:
487:
239:
185:
53:
1528:
1491:
1431:
1379:
1358:
Agaimy A (January 2020). "What is new in epithelioid soft tissue tumors?".
1333:
1297:
1248:
1230:
1189:
1140:
1088:
1026:
986:
918:
869:
817:
755:
709:
658:
611:
593:
578:"The 2020 WHO Classification of Soft Tissue Tumours: news and perspectives"
559:
1203:
495:
1171:
264:
97:
1561:
1239:
1105:
Geramizadeh B, Zare Z, Dehghanian AR, Bolandparvaz S, Marzban M (2018).
1051:
Sajid MI, Arshad S, Abdul-Ghafar J, Fatimi SH, Din NU (February 2021).
301:
219:
196:
stain) background. However, the tissues may contain sites dominated by
69:
1018:
909:
892:
235:
137:
113:
287:
247:
227:
216:
132:
year). LGFMS generally presents as a painless mass located in the
299:(i.e. cAMP responsive element binding protein 3 like 2 gene) or
309:
fusion gene is expressed in the majority of LGMFS cases while a
305:(i.e. CAMP responsive element binding protein 3 like 1 gene). A
251:
223:
212:
208:
780:
Yue Y, Liu Y, Song L, Chen X, Wang Y, Wang Z (February 2018).
140:(i.e. beneath the skin) tissues of the upper or lower limbs,
144:, or, less commonly, the head and neck areas or within the
951:
Cowan ML, Thompson LD, Leon ME, Bishop JA (June 2016).
164:
often give results suggesting that a tumor is a LGFMS.
1551:
1555:
207:analyses of LGFMS tumors shows cells that express:
47:
21:
72:first described by H. L. Evans in 1987. LGFMS are
1509:Journal of the American Society of Cytopathology
850:Journal of the American Society of Cytopathology
576:Sbaraglia M, Bellan E, Dei Tos AP (April 2021).
1451:
1449:
1399:
1397:
1353:
1351:
1100:
1098:
1046:
1044:
1000:
998:
996:
946:
944:
942:
940:
938:
936:
891:Murshed KA, Al-Bozom I, Ammar A (August 2021).
88:. These fibroblastic, spindle-shaped cells are
843:
841:
839:
837:
835:
833:
831:
829:
827:
775:
773:
682:
680:
678:
676:
631:
629:
627:
625:
623:
621:
571:
569:
517:
515:
513:
511:
509:
507:
505:
279:. The fusion genes in LGFMS merge part of the
1462:Medical Oncology (Northwood, London, England)
8:
729:
727:
725:
723:
721:
719:
378:that have a prominent myxoid background, or
1552:
1410:The American Journal of Surgical Pathology
736:The American Journal of Surgical Pathology
92:cells that in most cases of LGFMS express
27:
18:
1481:
1421:
1287:
1238:
1179:
1130:
1078:
1068:
976:
908:
807:
797:
601:
549:
539:
434:fusion gene is likely to be a SEF tumor.
250:proteins in virtually no cases. However,
466:
255:on finding MUC4-positive tumor cells.)
102:fibroblastic and myofibroblastic tumors
1160:AJR. American Journal of Roentgenology
476:American Journal of Clinical Pathology
398:express the MUC4 marker protein and a
368:myxoid dermatofibrosarcoma protuberans
7:
1587:Connective and soft tissue neoplasms
386:fusion gene in >20% of cases, a
39:of a low-grade fibromyxoid sarcoma.
390:fusion gene in ~10% of cases, or a
372:extraskeletal myxoid chondrosarcoma
121:sclerosing epithelioid fibrosarcoma
14:
1404:Warmke LM, Meis JM (March 2021).
358:), and sarcomas (e.g., malignant
528:Diagnostics (Basel, Switzerland)
1582:Dermal and subcutaneous growths
1057:Journal of Medical Case Reports
1007:Journal of Cutaneous Pathology
360:peripheral nerve sheath tumora
68:) is a rare type of low-grade
1:
1326:10.1016/j.humpath.2019.08.006
786:BMC Musculoskeletal Disorders
178:hematoxylin and eosin stained
62:Low-grade fibromyxoid sarcoma
22:Low-grade fibromyxoid sarcoma
1423:10.1097/PAS.0000000000001559
748:10.1097/PAS.0b013e31822b3687
230:proteins in rare cases; and
1219:Polish Journal of Pathology
541:10.3390/diagnostics11061093
345:fusion gene in most cases.
1613:
1521:10.1016/j.jasc.2020.05.005
1372:10.1007/s00428-019-02677-8
1070:10.1186/s13256-020-02605-4
862:10.1016/j.jasc.2020.01.006
702:10.1016/j.ctrv.2021.102259
651:10.1007/s00256-021-03836-2
347:Magnetic resonance imaging
291:gene (which is one of the
269:chromosomal translocations
265:large scale gene mutations
158:Magnetic resonance imaging
1474:10.1007/s12032-018-1192-6
969:10.1007/s12105-015-0647-8
799:10.1186/s12891-018-1976-z
162:computed tomography scans
35:
26:
1123:10.1177/2036361318777031
690:Cancer Treatment Reviews
351:computed tomography scan
293:FET protein family genes
148:, heart, kidney, brain,
1280:10.15252/embr.201845766
957:Head and Neck Pathology
438:Treatment and prognosis
400:FUS-CREB3L2 fusion gene
184:. The cells have scant
112:usually to the lung or
1231:10.5114/pjp.2018.79541
594:10.32074/1591-951X-213
273:interstitial deletions
146:gastrointestinal tract
488:10.1093/ajcp/88.5.615
1172:10.2214/AJR.14.13972
1117:: 2036361318777031.
370:, or in rare cases,
356:desmoid fibromatosis
211:(also termed EMA),
205:Immunohistochemical
639:Skeletal Radiology
380:myxoid liposarcoma
259:Gene abnormalities
82:connective tissues
1569:
1568:
1019:10.1111/cup.13413
910:10.1111/apm.13157
742:(10): 1450–1462.
449:Radiation therapy
376:synovial sarcomas
295:) with part of a
198:epithelioid cells
59:
58:
16:Medical condition
1604:
1553:
1541:
1540:
1503:
1497:
1495:
1485:
1453:
1444:
1443:
1425:
1401:
1392:
1391:
1355:
1346:
1345:
1308:
1302:
1301:
1291:
1259:
1253:
1252:
1242:
1214:
1208:
1207:
1200:
1194:
1193:
1183:
1151:
1145:
1144:
1134:
1102:
1093:
1092:
1082:
1072:
1048:
1039:
1038:
1002:
991:
990:
980:
948:
931:
930:
912:
888:
882:
881:
845:
822:
821:
811:
801:
777:
768:
767:
731:
714:
713:
684:
671:
670:
633:
616:
615:
605:
573:
564:
563:
553:
543:
519:
500:
499:
471:
364:myxofibrosarcoma
31:
19:
1612:
1611:
1607:
1606:
1605:
1603:
1602:
1601:
1572:
1571:
1570:
1565:
1564:
1550:
1545:
1544:
1505:
1504:
1500:
1455:
1454:
1447:
1403:
1402:
1395:
1360:Virchows Archiv
1357:
1356:
1349:
1314:Human Pathology
1310:
1309:
1305:
1261:
1260:
1256:
1216:
1215:
1211:
1202:
1201:
1197:
1153:
1152:
1148:
1104:
1103:
1096:
1050:
1049:
1042:
1004:
1003:
994:
950:
949:
934:
890:
889:
885:
847:
846:
825:
779:
778:
771:
733:
732:
717:
686:
685:
674:
635:
634:
619:
575:
574:
567:
521:
520:
503:
473:
472:
468:
463:
440:
327:
261:
222:in some cases;
174:histopathologic
170:
150:retroperitoneum
129:
17:
12:
11:
5:
1610:
1608:
1600:
1599:
1594:
1589:
1584:
1574:
1573:
1567:
1566:
1560:
1559:
1557:
1556:Classification
1549:
1548:External links
1546:
1543:
1542:
1515:(6): 513–519.
1498:
1445:
1416:(3): 317–328.
1393:
1347:
1303:
1254:
1225:(3): 219–225.
1209:
1195:
1146:
1094:
1040:
1013:(4): 271–276.
992:
932:
903:(8): 455–460.
883:
856:(3): 191–201.
823:
769:
715:
672:
645:(3): 477–504.
617:
565:
501:
465:
464:
462:
459:
439:
436:
326:
323:
314:EWSR1-CREB3L2,
260:
257:
169:
166:
128:
125:
76:tumors of the
57:
56:
51:
45:
44:
33:
32:
24:
23:
15:
13:
10:
9:
6:
4:
3:
2:
1609:
1598:
1595:
1593:
1590:
1588:
1585:
1583:
1580:
1579:
1577:
1563:
1558:
1554:
1547:
1538:
1534:
1530:
1526:
1522:
1518:
1514:
1510:
1502:
1499:
1493:
1489:
1484:
1479:
1475:
1471:
1467:
1463:
1459:
1452:
1450:
1446:
1441:
1437:
1433:
1429:
1424:
1419:
1415:
1411:
1407:
1400:
1398:
1394:
1389:
1385:
1381:
1377:
1373:
1369:
1365:
1361:
1354:
1352:
1348:
1343:
1339:
1335:
1331:
1327:
1323:
1319:
1315:
1307:
1304:
1299:
1295:
1290:
1285:
1281:
1277:
1273:
1269:
1265:
1258:
1255:
1250:
1246:
1241:
1236:
1232:
1228:
1224:
1220:
1213:
1210:
1205:
1199:
1196:
1191:
1187:
1182:
1177:
1173:
1169:
1166:(2): 414–20.
1165:
1161:
1157:
1150:
1147:
1142:
1138:
1133:
1128:
1124:
1120:
1116:
1112:
1108:
1101:
1099:
1095:
1090:
1086:
1081:
1076:
1071:
1066:
1062:
1058:
1054:
1047:
1045:
1041:
1036:
1032:
1028:
1024:
1020:
1016:
1012:
1008:
1001:
999:
997:
993:
988:
984:
979:
974:
970:
966:
962:
958:
954:
947:
945:
943:
941:
939:
937:
933:
928:
924:
920:
916:
911:
906:
902:
898:
894:
887:
884:
879:
875:
871:
867:
863:
859:
855:
851:
844:
842:
840:
838:
836:
834:
832:
830:
828:
824:
819:
815:
810:
805:
800:
795:
791:
787:
783:
776:
774:
770:
765:
761:
757:
753:
749:
745:
741:
737:
730:
728:
726:
724:
722:
720:
716:
711:
707:
703:
699:
695:
691:
683:
681:
679:
677:
673:
668:
664:
660:
656:
652:
648:
644:
640:
632:
630:
628:
626:
624:
622:
618:
613:
609:
604:
599:
595:
591:
587:
583:
579:
572:
570:
566:
561:
557:
552:
547:
542:
537:
533:
529:
525:
518:
516:
514:
512:
510:
508:
506:
502:
497:
493:
489:
485:
481:
477:
470:
467:
460:
458:
455:
450:
446:
437:
435:
433:
432:EWSR1-CREB3L2
429:
428:EWSR1-CREB3L1
425:
421:
417:
413:
409:
408:EWSR1-CREB3L1
405:
401:
397:
393:
392:EWSR1-CREB3L2
389:
385:
384:EWSR1-CREB3L1
381:
377:
373:
369:
365:
361:
357:
352:
348:
344:
340:
339:EWSR1-CREB3L2
336:
332:
324:
322:
319:
315:
312:
308:
304:
303:
298:
294:
290:
289:
284:
283:
278:
274:
270:
266:
258:
256:
253:
249:
245:
241:
237:
233:
229:
225:
221:
218:
214:
210:
206:
202:
199:
195:
191:
187:
183:
179:
175:
167:
165:
163:
159:
155:
151:
147:
143:
139:
135:
126:
124:
122:
117:
115:
111:
105:
103:
99:
95:
91:
87:
83:
79:
75:
71:
67:
63:
55:
52:
50:
46:
42:
41:H&E stain
38:
34:
30:
25:
20:
1512:
1508:
1501:
1465:
1461:
1413:
1409:
1366:(1): 81–96.
1363:
1359:
1317:
1313:
1306:
1271:
1268:EMBO Reports
1267:
1257:
1240:11585/667908
1222:
1218:
1212:
1198:
1163:
1159:
1149:
1114:
1110:
1060:
1056:
1010:
1006:
963:(2): 161–6.
960:
956:
900:
896:
886:
853:
849:
789:
785:
739:
735:
693:
689:
642:
638:
588:(2): 70–84.
585:
581:
531:
527:
482:(5): 615–9.
479:
475:
469:
454:chemotherapy
441:
431:
427:
423:
419:
415:
411:
407:
406:express the
403:
399:
395:
391:
387:
383:
362:, low-grade
343:EWSR2-CREBL1
342:
338:
334:
330:
328:
318:EWSR2-CREBL1
317:
313:
311:FUS-CREB3L1,
310:
306:
300:
296:
286:
280:
262:
203:
171:
134:subcutaneous
130:
127:Presentation
118:
106:
94:fusion genes
65:
61:
60:
1468:(11): 138.
1111:Rare Tumors
582:Pathologica
534:(6): 1093.
445:pericardium
426:express an
420:FUS-CREB3L2
416:FUS-CREB3L2
388:FUS-CREB3L2
335:FUS-CREB3L1
331:FUS-CREB3L2
307:FUS-CREB3L2
244:cytokeratin
194:hematoxylin
154:mediastinum
110:metastasize
86:fibroblasts
74:soft tissue
1576:Categories
696:: 102259.
461:References
414:express a
277:inversions
90:neoplastic
78:mesenchyme
37:Micrograph
1537:220369922
1440:221084763
1388:207893952
1342:201117873
1320:: 65–73.
1063:(1): 50.
927:235232220
878:212810533
792:(1): 65.
667:235678096
325:Diagnosis
240:caldesmon
215:, and/or
186:cytoplasm
176:views of
168:Pathology
138:subfacial
98:mutations
80:-derived
54:Pathology
49:Specialty
1529:32624384
1492:30187231
1432:32769431
1380:31686193
1334:31430493
1298:30962207
1249:30509048
1190:26204295
1141:29854356
1089:33526082
1035:58592480
1027:30632203
987:26276044
919:34048081
870:32197967
818:29482535
764:21592343
756:21921785
710:34311246
659:34191084
612:33179614
560:34203801
267:such as
220:proteins
1597:Sarcoma
1483:6132781
1289:6500973
1181:4570741
1132:5971385
1080:7851906
978:4838961
809:6389061
603:8167394
551:8232650
496:3673943
302:CREB3L1
297:CREB3L2
190:mitosis
114:pleural
70:sarcoma
1592:Cancer
1535:
1527:
1490:
1480:
1438:
1430:
1386:
1378:
1340:
1332:
1296:
1286:
1247:
1188:
1178:
1139:
1129:
1087:
1077:
1033:
1025:
985:
975:
925:
917:
876:
868:
816:
806:
762:
754:
708:
665:
657:
610:
600:
558:
548:
494:
246:, and
236:desmin
182:nuclei
1533:S2CID
1436:S2CID
1384:S2CID
1338:S2CID
1274:(5).
1031:S2CID
923:S2CID
897:APMIS
874:S2CID
760:S2CID
663:S2CID
341:, or
288:EWSR2
275:, or
248:CD117
217:bcl-2
142:trunk
66:LGFMS
1525:PMID
1488:PMID
1428:PMID
1376:PMID
1330:PMID
1294:PMID
1245:PMID
1186:PMID
1137:PMID
1085:PMID
1023:PMID
983:PMID
915:PMID
866:PMID
814:PMID
752:PMID
706:PMID
655:PMID
608:PMID
556:PMID
492:PMID
349:and
252:MUC4
232:S100
226:and
224:CD34
213:CD99
209:MUC1
160:and
1517:doi
1478:PMC
1470:doi
1418:doi
1368:doi
1364:476
1322:doi
1284:PMC
1276:doi
1235:hdl
1227:doi
1176:PMC
1168:doi
1164:205
1127:PMC
1119:doi
1075:PMC
1065:doi
1015:doi
973:PMC
965:doi
905:doi
901:129
858:doi
804:PMC
794:doi
744:doi
698:doi
647:doi
598:PMC
590:doi
586:113
546:PMC
536:doi
484:doi
430:or
316:or
285:or
282:FUS
228:SMA
136:or
1578::
1531:.
1523:.
1511:.
1486:.
1476:.
1466:35
1464:.
1460:.
1448:^
1434:.
1426:.
1414:45
1412:.
1408:.
1396:^
1382:.
1374:.
1362:.
1350:^
1336:.
1328:.
1318:93
1316:.
1292:.
1282:.
1272:20
1270:.
1266:.
1243:.
1233:.
1223:69
1221:.
1184:.
1174:.
1162:.
1158:.
1135:.
1125:.
1115:10
1113:.
1109:.
1097:^
1083:.
1073:.
1061:15
1059:.
1055:.
1043:^
1029:.
1021:.
1011:46
1009:.
995:^
981:.
971:.
961:10
959:.
955:.
935:^
921:.
913:.
899:.
895:.
872:.
864:.
852:.
826:^
812:.
802:.
790:19
788:.
784:.
772:^
758:.
750:.
740:35
738:.
718:^
704:.
694:99
692:.
675:^
661:.
653:.
643:51
641:.
620:^
606:.
596:.
584:.
580:.
568:^
554:.
544:.
532:11
530:.
526:.
504:^
490:.
480:88
478:.
424:4)
412:3)
404:2)
396:1)
374:,
366:,
337:,
333:,
271:,
242:,
238:,
234:,
152:,
104:.
1562:D
1539:.
1519::
1513:9
1496:)
1494:.
1472::
1442:.
1420::
1390:.
1370::
1344:.
1324::
1300:.
1278::
1251:.
1237::
1229::
1206:.
1192:.
1170::
1143:.
1121::
1091:.
1067::
1037:.
1017::
989:.
967::
929:.
907::
880:.
860::
854:9
820:.
796::
766:.
746::
712:.
700::
669:.
649::
614:.
592::
562:.
538::
498:.
486::
64:(
43:.
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.