57:
44:
411:
The ability to produce penicillin appears to have evolved over millions of years, and is shared with several other related fungi. It is believed to confer a selective advantage during competition with bacteria for food sources. Some bacteria have consequently developed the counter-ability to survive
310:
cannot be identified based on colour alone. Observations of morphology and microscopic features are needed to confirm its identity and DNA sequencing is essential to distinguish it from closely related species such as
202:
regions and can be found on salted food products, but it is mostly found in indoor environments, especially in damp or water-damaged buildings. It has been recognised as a species complex that includes
1293:
Martín JF, Gutiérrez S, Fernández FJ, Velasco J, Fierro F, Marcos AT, Kosalkova K (1994). "Expression of genes and processing of enzymes for the biosynthesis of penicillins and cephalosporins".
1231:
Shen HD, Chou H, Tam MF, Chang CY, Lai HY, Wang SR (October 2003). "Molecular and immunological characterization of Pen ch 18, the vacuolar serine protease major allergen of
1593:
1632:
981:
Ali H, Ries MI, Lankhorst PP, van der Hoeven RA, Schouten OL, Noga M, Hankemeier T, van Peij NN, Bovenberg RA, Vreeken RJ, Driessen AJ (2014-06-02).
679:
Lyratzopoulos, G.; Ellis, M.; Nerringer, R.; Denning, D. W. (October 2002). "Invasive infection due to penicillium species other than P. marneffei".
1567:
1655:
1724:
439:
1541:
215:
Molecular phylogeny has established that
Alexander Fleming's first discovered penicillin producing strain is of a distinct species,
1709:
1681:
416:, enzymes that degrade penicillin. Penicillinase production is one mechanism by which bacteria can become penicillin resistant.
1176:
Böhm J, Hoff B, O'Gorman CM, Wolfers S, Klix V, Binger D, Zadra I, Kürnsteiner H, Pöggeler S, Dyer PS, Kück U (January 2013).
388:. As the mold grows, it uses up the sugar and starts to make penicillin only after using up most of the nutrients for growth.
353:
1619:
435:
strains used for the industrial production of penicillin contain multiple tandem copies of the penicillin gene cluster.
56:
372:
ushered in a new age of antibiotics derived from microorganisms. Penicillin is an antibiotic isolated from growing
983:"A non-canonical NRPS is involved in the synthesis of fungisporin and related hydrophobic cyclic tetrapeptides in
630:
Houbraken, J.; Frisvad, J.C.; Seifert, K.A.; Overy, D.P.; Tuthill, D.M.; Valdez, J.G.; Samson, R.A. (2012-12-31).
1637:
954:
Matsuda Y, Awakawa T, Abe I (September 2013). "Reconstituted biosynthesis of fungal meroterpenoid andrastin A".
1719:
1396:
1729:
722:
Ali H, Ries MI, Nijland JG, Lankhorst PP, Hankemeier T, Bovenberg RA, Vreeken RJ, Driessen AJ (2013-06-12).
901:
Guzman-Chavez F, Salo O, Samol M, Ries M, Kuipers J, Bovenberg RA, Vreeken RJ, Driessen AJ (October 2018).
785:
Viggiano A, Salo O, Ali H, Szymanski W, Lankhorst PP, Nygård Y, Bovenberg RA, Driessen AJ (February 2018).
1470:
258:
230:
1338:"The penicillin gene cluster is amplified in tandem repeats linked by conserved hexanucleotide sequences"
401:
323:, after the mating types (MAT1-1 or MAT1-2) of the strains had been determined using PCR amplification.
151:
1714:
1686:
1647:
1559:
1515:
1349:
1061:
998:
802:
739:
490:
1178:"Sexual reproduction and mating-type-mediated strain development in the penicillin-producing fungus
724:"A branched biosynthetic pathway is involved in production of roquefortine and related compounds in
1044:
Salo O, Guzmán-Chávez F, Ries MI, Lankhorst PP, Bovenberg RA, Vreeken RJ, Driessen AJ (July 2016).
1428:
1318:
1260:
563:
270:
217:
51:
254:
1598:
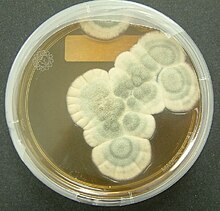
306:, the conidia are blue to blue-green, and the mold sometimes exudes a yellow pigment. However,
1668:
1502:
1420:
1377:
1310:
1252:
1213:
1140:
1087:
1026:
936:
883:
828:
767:
704:
696:
661:
612:
555:
516:
349:
1673:
1412:
1367:
1357:
1302:
1244:
1203:
1193:
1130:
1122:
1077:
1069:
1016:
1006:
963:
926:
918:
873:
863:
818:
810:
757:
747:
688:
651:
643:
602:
594:
547:
506:
498:
357:
331:
1353:
1105:
Guzmán-Chávez F, Salo O, Nygård Y, Lankhorst PP, Bovenberg RA, Driessen AJ (July 2017).
1065:
1002:
806:
743:
494:
319:
was discovered in 2013 by mating cultures in the dark on oatmeal agar supplemented with
1554:
1449:
1208:
1177:
1135:
1106:
1082:
1045:
1021:
982:
931:
902:
878:
847:
823:
786:
762:
723:
656:
631:
607:
582:
511:
478:
413:
341:
266:
246:
118:
98:
1703:
1432:
1372:
1337:
1248:
299:
1322:
1264:
632:"New penicillin-producing Penicillium species and an overview of section Chrysogena"
598:
583:"Fleming's penicillin producing strain is not Penicillium chrysogenum but P. rubens"
567:
1507:
167:
43:
17:
302:. The conidia are typically carried by air currents to new colonisation sites. In
1046:"Identification of a Polyketide Synthase Involved in Sorbicillin Biosynthesis by
1011:
752:
466:. Utrecht, the Netherlands: CBS-KNAW- Fungal Biodiversity Centre. pp. 1–398.
1624:
1580:
1336:
Fierro F, Barredo JL, Díez B, Gutierrez S, Fernández FJ, Martín JF (June 1995).
282:
262:
199:
190:
128:
1342:
Proceedings of the
National Academy of Sciences of the United States of America
1186:
Proceedings of the
National Academy of Sciences of the United States of America
1416:
967:
397:
369:
238:
234:
108:
88:
1493:
868:
700:
647:
1546:
1528:
1362:
1198:
1126:
477:
Andersen B, Frisvad JC, Søndergaard I, Rasmussen IS, Larsen LS (June 2011).
377:
345:
250:
195:
68:
1424:
1256:
1217:
1144:
1091:
1030:
940:
887:
832:
771:
708:
692:
665:
616:
520:
431:
are closely linked, forming a cluster on chromosome I. Some high-producing
1381:
1314:
479:"Associations between fungal species and water-damaged building materials"
1606:
1487:
1073:
903:"Deregulation of secondary metabolism in a histone deacetylase mutant of
814:
559:
502:
385:
295:
274:
1572:
1306:
551:
226:
1585:
1611:
922:
320:
291:
78:
1464:
1533:
1395:
Pohl C, Kiel JA, Driessen AJ, Bovenberg RA, Nygård Y (July 2016).
381:
462:
Samson RA, Houbraken J, Thrane U, Frisvad JC, Andersen B (2010).
1660:
1520:
534:
Samson RA, Hadlok R, Stolk AC (1977). "A taxonomic study of the
1468:
846:
Guzmán-Chávez F, Zwahlen RD, Bovenberg RA, Driessen AJ (2018).
581:
Houbraken, Jos; Frisvad, Jens C.; Samson, Robert A. (2011).
787:"Pathway for the Biosynthesis of the Pigment Chrysogine by
419:
The principal genes responsible for producing penicillin,
1107:"Mechanism and regulation of sorbicillin biosynthesis by
334:
have been implicated as the major allergenic proteins.
636:
Persoonia - Molecular
Phylogeny and Evolution of Fungi
340:
has been used industrially to produce penicillin and
330:
are important human allergens. Vacuolar and alkaline
1477:
442:techniques are available for editing the genome of
380:. The mold is grown in a liquid culture containing
1161:de Hoog GS, Guarro J, Gené J, Figueras F (2000),
1165:, Centraalbureau voor Schimmelcultures (Utrecht)
225:It has rarely been reported as a cause of human
1282:. Williams & Wilkins Company (Baltimore).
8:
290:usually reproduces by forming dry chains of
1156:
1154:
1465:
384:and other nutrients including a source of
42:
31:
1371:
1361:
1207:
1197:
1134:
1081:
1020:
1010:
930:
877:
867:
822:
761:
751:
655:
606:
510:
280:Like the many other species of the genus
454:
188:) is a species of fungus in the genus
1054:Applied and Environmental Microbiology
795:Applied and Environmental Microbiology
1397:"CRISPR/Cas9 Based Genome Editing of
1163:Atlas of Clinical Fungi - 2nd Edition
852:as Cell Factory for Natural Products"
7:
1648:a27c03ee-91df-44cd-a1ac-d5e4960c681e
438:Similar to other filamentous fungi,
440:CRISPR/Cas9-mediated genome editing
348:waste, and to produce the enzymes
25:
412:penicillin exposure by producing
1249:10.1034/j.1398-9995.2003.00107.x
55:
599:10.5598/imafungus.2011.02.01.12
326:The airborne asexual spores of
1458:Tom Volk's fungus of the month
354:phosphogluconate dehydrogenase
229:. It is the source of several
1:
261:YWA1/melanin, andrastatin A,
1012:10.1371/journal.pone.0098212
753:10.1371/journal.pone.0065328
1746:
1725:Taxa named by Charles Thom
1280:A manual of the Penicillia
395:
1417:10.1021/acssynbio.6b00082
1278:Raper KB, Thom C (1949).
968:10.1016/j.tet.2013.07.029
856:Frontiers in Microbiology
398:Penicillin § History
157:
150:
52:Scientific classification
50:
41:
34:
869:10.3389/fmicb.2018.02768
681:The Journal of Infection
648:10.3767/003158512X660571
483:Appl. Environ. Microbiol
1710:Fungi described in 1910
1479:Penicillium chrysogenum
1452:Penicillium chrysogenum
1399:Penicillium chrysogenum
1363:10.1073/pnas.92.13.6200
1295:Antonie van Leeuwenhoek
1233:Penicillium chrysogenum
1199:10.1073/pnas.1217943110
1180:Penicillium chrysogenum
1127:10.1111/1751-7915.12736
1115:Microbial Biotechnology
1109:Penicillium chrysogenum
1048:Penicillium chrysogenum
985:Penicillium chrysogenum
905:Penicillium chrysogenum
850:Penicillium chrysogenum
789:Penicillium chrysogenum
726:Penicillium chrysogenum
540:Antonie van Leeuwenhoek
536:Penicillium chrysogenum
444:Penicillium chrysogenum
433:Penicillium chrysogenum
179:Penicillium chrysogenum
161:Penicillium chrysogenum
36:Penicillium chrysogenum
693:10.1053/jinf.2002.1056
407:Genetics and evolution
315:. The sexual stage of
1405:ACS Synthetic Biology
464:Food and Indoor Fungi
402:History of penicillin
396:Further information:
233:, most significantly
1074:10.1128/AEM.00350-16
815:10.1128/AEM.02246-17
503:10.1128/AEM.02513-10
298:) from brush-shaped
231:β-lactam antibiotics
1354:1995PNAS...92.6200F
1066:2016ApEnM..82.3971S
1003:2014PLoSO...998212A
807:2018ApEnM..84E2246V
744:2013PLoSO...865328A
495:2011ApEnM..77.4180A
328:P. chrysogenum
317:P. chrysogenum
308:P. chrysogenum
304:P. chrysogenum
288:P. chrysogenum
243:P. chrysogenum
185:Penicillium notatum
182:(formerly known as
143:P. chrysogenum
18:Penicillium notatum
1307:10.1007/BF00871951
552:10.1007/BF00395671
237:. Other secondary
194:. It is common in
1697:
1696:
1669:Open Tree of Life
1471:Taxon identifiers
1060:(13): 3971–3978.
962:(38): 8199–8204.
368:The discovery of
350:polyamine oxidase
175:
174:
27:Species of fungus
16:(Redirected from
1737:
1690:
1689:
1677:
1676:
1664:
1663:
1651:
1650:
1641:
1640:
1628:
1627:
1625:BMSSYS0000012947
1615:
1614:
1602:
1601:
1589:
1588:
1576:
1575:
1563:
1562:
1550:
1549:
1537:
1536:
1524:
1523:
1511:
1510:
1498:
1497:
1496:
1466:
1461:
1437:
1436:
1392:
1386:
1385:
1375:
1365:
1333:
1327:
1326:
1290:
1284:
1283:
1275:
1269:
1268:
1243:(10): 993–1002.
1228:
1222:
1221:
1211:
1201:
1173:
1167:
1166:
1158:
1149:
1148:
1138:
1102:
1096:
1095:
1085:
1041:
1035:
1034:
1024:
1014:
978:
972:
971:
951:
945:
944:
934:
923:10.1002/mbo3.598
911:MicrobiologyOpen
898:
892:
891:
881:
871:
843:
837:
836:
826:
782:
776:
775:
765:
755:
719:
713:
712:
676:
670:
669:
659:
627:
621:
620:
610:
578:
572:
571:
531:
525:
524:
514:
474:
468:
467:
459:
332:serine proteases
213:P. cyaneofulvum.
163:
60:
59:
46:
32:
21:
1745:
1744:
1740:
1739:
1738:
1736:
1735:
1734:
1720:Medicinal fungi
1700:
1699:
1698:
1693:
1685:
1680:
1672:
1667:
1659:
1654:
1646:
1644:
1636:
1631:
1623:
1618:
1610:
1605:
1597:
1592:
1584:
1579:
1571:
1566:
1558:
1553:
1545:
1540:
1532:
1527:
1519:
1514:
1506:
1501:
1492:
1491:
1486:
1473:
1448:
1445:
1440:
1394:
1393:
1389:
1335:
1334:
1330:
1292:
1291:
1287:
1277:
1276:
1272:
1230:
1229:
1225:
1175:
1174:
1170:
1160:
1159:
1152:
1104:
1103:
1099:
1043:
1042:
1038:
980:
979:
975:
953:
952:
948:
900:
899:
895:
845:
844:
840:
784:
783:
779:
721:
720:
716:
678:
677:
673:
629:
628:
624:
580:
579:
575:
533:
532:
528:
476:
475:
471:
461:
460:
456:
452:
409:
404:
394:
366:
358:glucose oxidase
267:secalonic acids
209:P. meleagrinum,
171:
165:
159:
146:
54:
28:
23:
22:
15:
12:
11:
5:
1743:
1741:
1733:
1732:
1730:Fungus species
1727:
1722:
1717:
1712:
1702:
1701:
1695:
1694:
1692:
1691:
1678:
1665:
1652:
1642:
1629:
1616:
1603:
1590:
1577:
1564:
1551:
1538:
1525:
1512:
1499:
1483:
1481:
1475:
1474:
1469:
1463:
1462:
1444:
1443:External links
1441:
1439:
1438:
1387:
1348:(13): 6200–4.
1328:
1285:
1270:
1223:
1192:(4): 1476–81.
1168:
1150:
1121:(4): 958–968.
1097:
1036:
973:
946:
893:
838:
777:
714:
687:(3): 184–195.
671:
622:
573:
526:
489:(12): 4180–8.
469:
453:
451:
448:
414:penicillinases
408:
405:
393:
390:
365:
362:
342:xanthocillin X
338:P. chrysogenum
247:roquefortine C
173:
172:
166:
155:
154:
148:
147:
140:
138:
134:
133:
126:
122:
121:
119:Aspergillaceae
116:
112:
111:
106:
102:
101:
99:Eurotiomycetes
96:
92:
91:
86:
82:
81:
76:
72:
71:
66:
62:
61:
48:
47:
39:
38:
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
1742:
1731:
1728:
1726:
1723:
1721:
1718:
1716:
1713:
1711:
1708:
1707:
1705:
1688:
1683:
1679:
1675:
1670:
1666:
1662:
1657:
1653:
1649:
1643:
1639:
1634:
1630:
1626:
1621:
1617:
1613:
1608:
1604:
1600:
1595:
1591:
1587:
1582:
1578:
1574:
1569:
1565:
1561:
1556:
1552:
1548:
1543:
1539:
1535:
1530:
1526:
1522:
1517:
1513:
1509:
1504:
1500:
1495:
1489:
1485:
1484:
1482:
1480:
1476:
1472:
1467:
1459:
1455:
1453:
1447:
1446:
1442:
1434:
1430:
1426:
1422:
1418:
1414:
1411:(7): 754–64.
1410:
1406:
1402:
1400:
1391:
1388:
1383:
1379:
1374:
1369:
1364:
1359:
1355:
1351:
1347:
1343:
1339:
1332:
1329:
1324:
1320:
1316:
1312:
1308:
1304:
1301:(3): 227–43.
1300:
1296:
1289:
1286:
1281:
1274:
1271:
1266:
1262:
1258:
1254:
1250:
1246:
1242:
1238:
1234:
1227:
1224:
1219:
1215:
1210:
1205:
1200:
1195:
1191:
1187:
1183:
1181:
1172:
1169:
1164:
1157:
1155:
1151:
1146:
1142:
1137:
1132:
1128:
1124:
1120:
1116:
1112:
1110:
1101:
1098:
1093:
1089:
1084:
1079:
1075:
1071:
1067:
1063:
1059:
1055:
1051:
1049:
1040:
1037:
1032:
1028:
1023:
1018:
1013:
1008:
1004:
1000:
997:(6): e98212.
996:
992:
988:
986:
977:
974:
969:
965:
961:
957:
950:
947:
942:
938:
933:
928:
924:
920:
917:(5): e00598.
916:
912:
908:
906:
897:
894:
889:
885:
880:
875:
870:
865:
861:
857:
853:
851:
842:
839:
834:
830:
825:
820:
816:
812:
808:
804:
800:
796:
792:
790:
781:
778:
773:
769:
764:
759:
754:
749:
745:
741:
738:(6): e65328.
737:
733:
729:
727:
718:
715:
710:
706:
702:
698:
694:
690:
686:
682:
675:
672:
667:
663:
658:
653:
649:
645:
642:(1): 78–100.
641:
637:
633:
626:
623:
618:
614:
609:
604:
600:
596:
592:
588:
584:
577:
574:
569:
565:
561:
557:
553:
549:
546:(2): 169–75.
545:
541:
537:
530:
527:
522:
518:
513:
508:
504:
500:
496:
492:
488:
484:
480:
473:
470:
465:
458:
455:
449:
447:
445:
441:
436:
434:
430:
426:
422:
417:
415:
406:
403:
399:
391:
389:
387:
383:
379:
375:
371:
363:
361:
359:
355:
351:
347:
343:
339:
335:
333:
329:
324:
322:
318:
314:
309:
305:
301:
300:conidiophores
297:
293:
289:
285:
284:
278:
276:
272:
268:
264:
260:
256:
252:
248:
244:
240:
236:
232:
228:
224:
221:, and not of
220:
219:
214:
210:
206:
201:
197:
193:
192:
187:
186:
181:
180:
169:
164:
162:
156:
153:
152:Binomial name
149:
145:
144:
139:
136:
135:
132:
131:
127:
124:
123:
120:
117:
114:
113:
110:
107:
104:
103:
100:
97:
94:
93:
90:
87:
84:
83:
80:
77:
74:
73:
70:
67:
64:
63:
58:
53:
49:
45:
40:
37:
33:
30:
19:
1478:
1457:
1451:
1408:
1404:
1398:
1390:
1345:
1341:
1331:
1298:
1294:
1288:
1279:
1273:
1240:
1236:
1232:
1226:
1189:
1185:
1179:
1171:
1162:
1118:
1114:
1108:
1100:
1057:
1053:
1047:
1039:
994:
990:
984:
976:
959:
955:
949:
914:
910:
904:
896:
859:
855:
849:
841:
798:
794:
788:
780:
735:
731:
725:
717:
684:
680:
674:
639:
635:
625:
593:(1): 87–95.
590:
586:
576:
543:
539:
535:
529:
486:
482:
472:
463:
457:
443:
437:
432:
428:
424:
420:
418:
410:
373:
367:
337:
336:
327:
325:
316:
312:
307:
303:
287:
281:
279:
242:
222:
216:
212:
208:
204:
189:
184:
183:
178:
177:
176:
160:
158:
142:
141:
129:
35:
29:
1715:Penicillium
1581:iNaturalist
956:Tetrahedron
374:Penicillium
344:, to treat
283:Penicillium
271:sorbicillin
263:fungisporin
239:metabolites
223:P. notatum.
200:subtropical
191:Penicillium
130:Penicillium
1704:Categories
587:IMA Fungus
450:References
376:mold in a
370:penicillin
255:chrysogine
235:penicillin
205:P. notatum
109:Eurotiales
89:Ascomycota
85:Division:
1433:206764457
701:0163-4453
538:series".
378:fermenter
346:pulp mill
313:P. rubens
251:meleagrin
218:P. rubens
196:temperate
137:Species:
75:Kingdom:
69:Eukaryota
1607:MycoBank
1599:11315450
1555:Fungorum
1488:Wikidata
1425:27072635
1323:25327312
1265:28229046
1257:14510716
1218:23307807
1145:28618182
1092:27107123
1031:24887561
991:PLOS ONE
941:29575742
888:30524395
862:: 2768.
833:29196288
772:23776469
732:PLOS ONE
709:12387776
666:23606767
617:22679592
568:41843432
521:21531835
386:nitrogen
275:PR-toxin
245:include
115:Family:
65:Domain:
1573:3466349
1521:2944750
1494:Q137155
1382:7597101
1350:Bibcode
1315:7847890
1237:Allergy
1209:3557024
1136:5481523
1083:4907180
1062:Bibcode
1022:4041764
999:Bibcode
932:6182556
879:6262359
824:5795073
803:Bibcode
763:3680398
740:Bibcode
657:3589797
608:3317369
512:3131638
491:Bibcode
392:History
364:Science
296:conidia
227:disease
125:Genus:
105:Order:
95:Class:
1687:100537
1674:226485
1661:100537
1645:NZOR:
1612:165757
1586:383409
1560:165757
1534:PENICH
1431:
1423:
1380:
1370:
1321:
1313:
1263:
1255:
1216:
1206:
1143:
1133:
1090:
1080:
1029:
1019:
939:
929:
886:
876:
831:
821:
770:
760:
707:
699:
664:
654:
615:
605:
566:
560:413477
558:
519:
509:
427:, and
400:, and
356:, and
321:biotin
292:spores
273:, and
170:(1910)
1682:WoRMS
1594:IRMNG
1547:20537
1542:EUNIS
1508:76HV7
1429:S2CID
1373:41670
1319:S2CID
1261:S2CID
801:(4).
564:S2CID
429:penDE
421:pcbAB
382:sugar
259:6-MSA
79:Fungi
1656:OBIS
1638:5076
1633:NCBI
1568:GBIF
1529:EPPO
1421:PMID
1378:PMID
1311:PMID
1253:PMID
1214:PMID
1141:PMID
1088:PMID
1027:PMID
937:PMID
884:PMID
829:PMID
768:PMID
705:PMID
697:ISSN
662:PMID
613:PMID
556:PMID
517:PMID
425:pcbC
294:(or
211:and
198:and
168:Thom
1620:NBN
1516:EoL
1503:CoL
1413:doi
1368:PMC
1358:doi
1303:doi
1245:doi
1235:".
1204:PMC
1194:doi
1190:110
1131:PMC
1123:doi
1078:PMC
1070:doi
1017:PMC
1007:doi
964:doi
927:PMC
919:doi
874:PMC
864:doi
819:PMC
811:doi
758:PMC
748:doi
689:doi
652:PMC
644:doi
603:PMC
595:doi
548:doi
507:PMC
499:doi
241:of
1706::
1684::
1671::
1658::
1635::
1622::
1609::
1596::
1583::
1570::
1557::
1544::
1531::
1518::
1505::
1490::
1456:.
1427:.
1419:.
1407:.
1403:.
1376:.
1366:.
1356:.
1346:92
1344:.
1340:.
1317:.
1309:.
1299:65
1297:.
1259:.
1251:.
1241:58
1239:.
1212:.
1202:.
1188:.
1184:.
1153:^
1139:.
1129:.
1119:10
1117:.
1113:.
1086:.
1076:.
1068:.
1058:82
1056:.
1052:.
1025:.
1015:.
1005:.
993:.
989:.
960:69
958:.
935:.
925:.
913:.
909:.
882:.
872:.
858:.
854:.
827:.
817:.
809:.
799:84
797:.
793:.
766:.
756:.
746:.
734:.
730:.
703:.
695:.
685:45
683:.
660:.
650:.
640:29
638:.
634:.
611:.
601:.
589:.
585:.
562:.
554:.
544:43
542:.
515:.
505:.
497:.
487:77
485:.
481:.
446:.
423:,
360:.
352:,
286:,
277:.
269:,
265:,
257:,
253:,
249:,
207:,
1460:.
1454:"
1450:"
1435:.
1415::
1409:5
1401:"
1384:.
1360::
1352::
1325:.
1305::
1267:.
1247::
1220:.
1196::
1182:"
1147:.
1125::
1111:"
1094:.
1072::
1064::
1050:"
1033:.
1009::
1001::
995:9
987:"
970:.
966::
943:.
921::
915:7
907:"
890:.
866::
860:9
848:"
835:.
813::
805::
791:"
774:.
750::
742::
736:8
728:"
711:.
691::
668:.
646::
619:.
597::
591:2
570:.
550::
523:.
501::
493::
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.