222:
1265:, and higher plants are not the same, but there are many analogous functions and similar structures. Three main features are similar between the different photosystems. First, redox potential is negative enough to reduce ferredoxin. Next, the electron-accepting reaction centers include iron–sulfur proteins. Last, redox centres in complexes of both photosystems are constructed upon a protein subunit dimer. The photosystem of green sulfur bacteria even contains all of the same cofactors of the
31:
1282:
337:
discoveries was not yet recognized at the time. Louis
Duysens first proposed the concepts of Photosystems I and II in 1960, and, in the same year, a proposal by Fay Bendall and Robert Hill assembled earlier discoveries into a coherent theory of serial photosynthetic reactions. Hill and Bendall's hypothesis was later confirmed in experiments conducted in 1961 by the Duysens and Witt groups.
48:
336:
This photosystem is known as PSI because it was discovered before
Photosystem II, although future experiments showed that Photosystem II is actually the first enzyme of the photosynthetic electron transport chain. Aspects of PSI were discovered in the 1950s, but the significance of these
1269:
in PSI. The number and degree of similarities between the two photosystems strongly indicates that PSI and the analogous photosystem of green sulfur bacteria evolved from a common ancestral photosystem.
1244:
found on the thylakoid membrane is vital to photosystem I. This thylakoid transmembrane protein helps assemble the components of photosystem I. Without it, photosynthesis would be inefficient.
992:. P700 receives energy from antenna molecules and uses the energy from each photon to raise an electron to a higher energy level (P700*). These electrons are moved in pairs in an
3078:
967:
reaction centers. The energy passed around by antenna molecules is directed to the reaction center. There may be as many as 120 or as few as 25 chlorophyll molecules per P700.
2619:
2482:
2624:
2331:
1031:
The two modified chlorophyll molecules are early electron acceptors in PSI. They are present one per PsaA/PsaB side, forming two branches electrons can take to reach F
185:
2970:
2372:
1119:
is tied to the PSI complex. Various experiments have shown some disparity between theories of iron–sulfur cofactor orientation and operation order. In one model, F
1943:
1172:. Thylakoid membranes have one binding site for each function of Fd. The main function of Fd is to carry an electron from the iron-sulfur complex to the enzyme
441:
204:
1043:
of the same side, which then passes the electron to the quinone on the same side. Different species seems to have different preferences for either A/B branch.
1662:
Plöchinger, Magdalena; Torabi, Salar; Rantala, Marjaana; Tikkanen, Mikko; Suorsa, Marjaana; Jensen, Poul-Erik; Aro, Eva Mari; Meurer, Jörg (September 2016).
506:
2487:
2243:"The chloroplast ycf3 and ycf4 open reading frames of Chlamydomonas reinhardtii are required for the accumulation of the photosystem I complex"
1927:
1731:
1592:
Jagannathan B, Golbeck JH (June 2009). "Understanding of the binding interface between PsaC and the PsaA/PsaB heterodimer in photosystem I".
1484:
Webber AN, Malkin R (May 1990). "Photosystem I reaction-centre proteins contain leucine zipper motifs. A proposed role in dimer formation".
963:
chloroplasts have around 200 chlorophylls and 50 carotenoids. Located within the antenna complex of PSI are molecules of chlorophyll called
2636:
937:
3163:
2963:
2365:
1537:"Breaking biological symmetry in membrane proteins: the asymmetrical orientation of PsaC on the pseudo-C2 symmetric Photosystem I core"
3056:
2679:
1763:
1782:
197:
2646:
3121:
2453:
2328:
1232:
is an electron carrier that transfers the electron from cytochrome b6f to the P700 cofactor of PSI in its ionized state P700.
164:
3126:
3098:
3071:
3049:
3034:
2999:
2956:
2358:
2162:
1803:
Shubin VV, Karapetyan NV, Krasnovsky AA (January 1986). "Molecular arrangement of pigment-protein complex of photosystem 1".
140:
2835:
2206:
Hope AB (January 2000). "Electron transfers amongst cytochrome f, plastocyanin and photosystem I: kinetics and mechanisms".
308:. Ultimately, the electrons that are transferred by Photosystem I are used to produce the moderate-energy hydrogen carrier
405:
1351:"Physiological Functions of Cyclic Electron Transport Around Photosystem I in Sustaining Photosynthesis and Plant Growth"
3044:
2641:
1218:
1173:
853:
3231:
366:
2820:
1433:
Fromme P, Mathis P (2004). "Unraveling the photosystem I reaction center: a history, or the sum of many efforts".
2936:
2923:
2910:
2897:
2884:
2871:
2858:
2592:
2543:
2520:
2497:
2463:
2435:
2395:
279:
259:
158:
2830:
1664:"The Low Molecular Weight Protein PsaI Stabilizes the Light-Harvesting Complex II Docking Site of Photosystem I"
416:, also iron-sulfur clusters, are located in a 9-kDa protein called PsaC that binds to the PsaA/PsaB core near F
3221:
3148:
2784:
2727:
2601:
2533:
2477:
2386:
1627:
Saenger W, Jordan P, Krauss N (April 2002). "The assembly of protein subunits and cofactors in photosystem I".
1266:
631:
63:
2501:
1088:
145:
3216:
3180:
3158:
3138:
2732:
393:; two cysteines are provided each by PsaA and PsaB. The two cysteines in each are proximal and located in a
321:
3143:
3039:
2067:
Vassiliev IR, Antonkine ML, Golbeck JH (October 2001). "Iron-sulfur clusters in type I reaction centers".
603:
317:
209:
133:
2753:
2672:
1314:
Golbeck JH (1987). "Structure, function and organization of the
Photosystem I reaction center complex".
1254:
1016:′ molecule. However, if P700 forms a complex with other antenna molecules, it can no longer be a dimer.
2825:
221:
2408:
2122:
2018:
1863:
1812:
1493:
1442:
1295:
702:
408:
of the cysteines and could contribute to dimerisation of PsaA/PsaB. The terminal electron acceptors F
386:
374:
235:
20:
161:
3195:
3185:
3153:
2789:
2631:
2606:
1241:
394:
346:
85:
440:
Related large transmembrane proteins involved in the binding of P700, A0, A1, and Fx. Part of the
3226:
3190:
2722:
2611:
2140:
2044:
1987:
1937:
1887:
1836:
1574:
1517:
1466:
997:
230:
1723:
996:
process from P700* to electron acceptors, leaving behind P700. The pair of P700* - P700 has an
2272:
2223:
2084:
2036:
1966:(Phylloquinone) Restores the Turnover of FeS centers of Ether-extracted Spinach PSI Particles"
1923:
1879:
1828:
1759:
1727:
1693:
1644:
1609:
1566:
1509:
1458:
1410:
1372:
1331:
1005:
293:
152:
1406:
1004:. The reaction center is made of two chlorophyll molecules and is therefore referred to as a
2768:
2763:
2737:
2665:
2472:
2308:
2262:
2254:
2215:
2188:
2130:
2076:
2026:
1977:
1915:
1871:
1820:
1683:
1675:
1636:
1601:
1556:
1548:
1501:
1450:
1402:
1362:
1323:
948:. The number of these pigment molecules varies from organism to organism. For instance, the
945:
121:
2983:
2815:
2799:
2712:
2335:
1367:
1350:
1112:
917:
282:
97:
1454:
68:
2126:
2022:
1867:
1816:
1786:
1561:
1536:
1497:
1446:
3131:
3014:
2991:
2853:
2794:
2382:
2350:
2312:
2267:
2242:
1919:
1854:
Rutherford AW, Heathcote P (December 1985). "Primary photochemistry in photosystem-I".
1688:
1663:
1287:
398:
325:
180:
2219:
2192:
2080:
1716:
1640:
1327:
920:
of the pigment molecules in the antenna complex induces electron and energy transfer.
345:
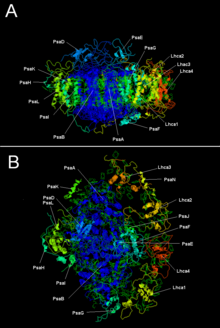
Two main subunits of PSI, PsaA and PsaB, are closely related proteins involved in the
239:. The 21 protein-coding genes involved in photosynthesis are displayed as green boxes.
3210:
3026:
2758:
2717:
2448:
2340:
2135:
2110:
2031:
2006:
1982:
1961:
1751:
1505:
1258:
1052:
982:
949:
826:
790:
401:
271:
24:
2144:
2048:
1991:
1891:
1840:
1578:
1521:
1470:
30:
2707:
2560:
1229:
705:
671:
313:
301:
424:
Components of PSI (protein subunits, lipids, pigments, coenzymes, and cofactors).
2948:
595:
579:
563:
547:
531:
515:
494:
478:
3004:
2931:
2866:
2702:
2510:
929:
736:
275:
255:
2258:
1281:
3116:
3093:
2547:
2524:
2399:
1552:
1393:
Nelson N, Yocum CF (2006). "Structure and function of photosystems I and II".
1277:
1143:
1139:
941:
933:
820:
796:
661:
370:
305:
2299:
Lockau W, Nitschke W (1993). "Photosystem I and its
Bacterial Counterparts".
940:
from photons when they become photoexcited. Antenna molecules can absorb all
2905:
2879:
1157:
to NADPH. Fd moves to carry an electron either to a lone thylakoid or to an
989:
903:
297:
2345:
2227:
2088:
1883:
1832:
1714:
Raven PH, Evert RF, Eichhorn SE (2005). "Photosynthesis, Light, and Life".
1697:
1648:
1613:
1570:
1462:
1414:
1376:
2276:
2161:
Madoz J, Fernández Recio J, Gómez Moreno C, Fernández VM (November 1998).
2040:
1513:
1335:
2241:
Boudreau E, Takahashi Y, Lemieux C, Turmel M, Rochaix JD (October 1997).
590:
574:
558:
542:
526:
510:
489:
473:
390:
109:
1679:
2551:
1875:
1824:
960:
883:
745:
617:
128:
1605:
47:
3088:
3083:
2979:
2918:
2688:
1253:
Molecular data show that PSI likely evolved from the photosystems of
1158:
378:
289:
192:
104:
92:
80:
34:
Light-dependent reactions of photosynthesis at the thylakoid membrane
3173:
3168:
2892:
2580:
2575:
2570:
2439:
993:
681:
309:
286:
267:
263:
220:
29:
2007:"Is phylloquinone an obligate electron carrier in photosystem I?"
3066:
3061:
2565:
2423:
2418:
2413:
1262:
1063:
in order to receive the electron and in turn is re-oxidized by F
1001:
976:
964:
116:
2952:
2661:
2354:
1316:
Biochimica et
Biophysica Acta (BBA) - Reviews on Bioenergetics
1206:
This enzyme transfers the electron from reduced ferredoxin to
936:
mounted on two proteins. These pigment molecules transmit the
312:. The photon energy absorbed by Photosystem I also produces a
2329:
Photosystem I: Molecule of the Month in the
Protein Data Bank
1059:, is the next early electron acceptor in PSI. It oxidizes A
2657:
1257:. The photosystems of green sulfur bacteria and those of
1221:
may also accept an electron from NADPH by binding to it.
1008:. The dimer is thought to be composed of one chlorophyll
959:) has about 100 chlorophylls and 20 carotenoids, whereas
2163:"Investigation of the Diaphorase Reaction of Ferredoxin–
2111:"Two Sites of Interaction of Ferredoxin with thylakoids"
16:
Second protein complex in photosynthetic light reactions
773:
Early electron acceptor of modified chlorophyll in ETC
397:
between the ninth and tenth transmembrane segments. A
3079:
Branched-chain alpha-keto acid dehydrogenase complex
1758:(4th ed.). Sunderland, MA: Sinauer Associates.
988:
that best absorbs light at a wavelength of 700
3181:
Phosphoenolpyruvate sugar phosphotransferase system
3109:
3025:
2990:
2844:
2808:
2777:
2746:
2695:
2591:
2542:
2519:
2496:
2462:
2434:
2394:
2208:
Biochimica et
Biophysica Acta (BBA) - Bioenergetics
2069:
Biochimica et
Biophysica Acta (BBA) - Bioenergetics
203:
191:
179:
174:
151:
139:
127:
115:
103:
91:
79:
74:
62:
57:
40:
2483:4-Hydroxy-3-methylbut-2-enyl diphosphate reductase
1910:Grotjohann, I; Fromme, P (2013). "Photosystem I".
1722:(7th ed.). New York: W. H. Freeman. pp.
1715:
981:The P700 reaction center is composed of modified
2341:Photosystem I in A Companion to Plant Physiology
928:The antenna complex is composed of molecules of
3000:Photosynthetic reaction center complex proteins
472:Required for assembly, helps bind ferredoxin.
2964:
2673:
2366:
1914:(Second ed.). London. pp. 503–507.
1905:
1903:
1901:
1388:
1386:
442:photosynthetic reaction centre protein family
8:
2062:
2060:
2058:
2005:Palace GP, Franke JE, Warden JT (May 1987).
1091:reaction centers are found in PSI. Labeled F
1039:accepts electrons from P700*, passes it to A
2595:: Acting on X-H and Y-H to form an X-Y bond
19:"PS I" redirects here. For other uses, see
2971:
2957:
2949:
2680:
2666:
2658:
2373:
2359:
2351:
1942:: CS1 maint: location missing publisher (
349:of the vital electron transfer cofactors P
171:
46:
2294:
2292:
2290:
2288:
2286:
2266:
2134:
2030:
1981:
1798:
1796:
1687:
1560:
1366:
2346:James Barber FRS Photosystems I & II
2156:
2154:
2104:
2102:
2100:
2098:
1709:
1707:
1535:Jagannathan B, Golbeck JH (April 2009).
1428:
1426:
1424:
1407:10.1146/annurev.arplant.57.032905.105350
1067:, from which the electron is passed to F
422:
2488:7-Hydroxymethyl chlorophyll a reductase
1955:
1953:
1777:
1775:
1745:
1743:
1306:
1935:
1146:protein that facilitates reduction of
1079:appears to be the rate-limiting step.
252:plastocyanin–ferredoxin oxidoreductase
37:
2181:Bioelectrochemistry and Bioenergetics
2174:Reductase by Electrochemical Methods"
1629:Current Opinion in Structural Biology
1368:10.1146/annurev-arplant-043015-112002
7:
2637:Tetrahydrocannabinolic acid synthase
1912:Encyclopedia of biological chemistry
1541:Cellular and Molecular Life Sciences
1217:to complete the reduction to NADPH.
457:Iron-sulfur center; apoprotein for F
3164:Mitochondrial trifunctional protein
2109:Forti G, Maria P, Grubas G (1985).
1349:Yamori W, Shikanai T (April 2016).
320:. PSI is composed of more than 110
52:Plant photosystem I with LHC I
2313:10.1111/j.1399-3054.1993.tb05512.x
1920:10.1016/B978-0-12-378630-2.00287-5
1455:10.1023/B:PRES.0000030657.88242.e1
1103:, they serve as electron relays. F
723:Phosphatidylglycerol phospholipid
715:Phosphatidylglycerol phospholipid
14:
1752:"Ch. 7: Topic 7.8: Photosystem I"
841:Early electron acceptor vitamin K
2647:Dichlorochromopyrrolate synthase
1280:
694:Monogalactosyldiglyceride lipid
3122:Carbamoyl phosphate synthase II
2454:(Methionine synthase) reductase
505:May stabilize PsaL. Stabilizes
3127:Aspartate carbamoyltransferase
3035:Pyruvate dehydrogenase complex
1395:Annual Review of Plant Biology
1355:Annual Review of Plant Biology
260:photosynthetic light reactions
1:
3159:Glycine decarboxylase complex
3154:Fatty acid synthetase complex
2625:(-)-bisdechlorogeodin-forming
2620:(+)-bisdechlorogeodin-forming
2220:10.1016/S0005-2728(99)00101-2
2193:10.1016/S0302-4598(98)00175-5
2081:10.1016/S0005-2728(01)00197-9
1641:10.1016/S0959-440X(02)00317-2
1328:10.1016/s0304-4173(87)80002-2
1012:molecule and one chlorophyll
957:Thermosynechococcus elongatus
2466:: Acting on CH or CH2 groups
2136:10.1016/0014-5793(85)80698-0
2032:10.1016/0014-5793(87)80113-8
1983:10.1016/0014-5793(89)81215-3
1783:"The Photosynthetic Process"
1506:10.1016/0014-5793(90)80749-9
1055:, sometimes called vitamin K
748:molecules in antenna system
2642:Cannabidiolic acid synthase
759:5 pigment molecules in ETC
3248:
3191:Sucrase-isomaltase complex
3057:Oxoglutarate dehydrogenase
974:
785:1 pigment molecule in ETC
367:integral membrane proteins
324:, significantly more than
18:
2836:Michaelis–Menten kinetics
1750:Zeiger E, Taiz L (2006).
1553:10.1007/s00018-009-8673-x
1131:to reach the ferredoxin.
1127:, which passes it on to F
439:
365:. PsaA and PsaB are both
316:that is used to generate
280:integral membrane protein
170:
45:
3149:Electron transport chain
2728:Diffusion-limited enzyme
2602:Isopenicillin N synthase
2534:Nitrogenase (flavodoxin)
2478:Ribonucleotide reductase
2259:10.1093/emboj/16.20.6095
1960:Itoh S, Iwaki M (1989).
1267:electron transport chain
1115:of the PSI complex and F
819:Early electron acceptor
804:Coenzymes and cofactors
666:Electron carrier in ETC
632:electron transport chain
507:light-harvesting complex
3139:P450-containing systems
1856:Photosynthesis Research
1805:Photosynthesis Research
1435:Photosynthesis Research
1123:passes an electron to F
953:Synechococcus elongatus
3144:Cytochrome b6f complex
1020:Modified chlorophyll A
240:
35:
2984:multienzyme complexes
2821:Eadie–Hofstee diagram
2754:Allosteric regulation
2301:Physiologia Plantarum
1255:green sulfur bacteria
865:oxidoreductase enzyme
845:phylloquinone in ETC
404:seems to be present
341:Components and action
294:transfer of electrons
224:
33:
2831:Lineweaver–Burk plot
2523:: Acting on reduced
2502:iron–sulfur proteins
2409:Superoxide dismutase
1296:Biohybrid solar cell
1087:Three proteinaceous
1075:. The reduction of F
971:P700 reaction center
944:of light within the
703:Phosphatidylglycerol
236:Arabidopsis thaliana
21:PSI (disambiguation)
3196:Tryptophan synthase
3186:Polyketide synthase
2632:Aureusidin synthase
2607:Columbamine oxidase
2127:1985FEBSL.186..149F
2023:1987FEBSL.215...58P
1868:1985PhoRe...6..295R
1817:1986PhoRe...9....3S
1680:10.1104/pp.16.00647
1498:1990FEBSL.264....1W
1447:2004PhoRe..80..109F
1242:Ycf4 protein domain
1236:Ycf4 protein domain
1083:Iron–sulfur complex
994:oxidation/reduction
656:From PsaAB; In ETC
425:
379:iron-sulfur cluster
314:proton-motive force
2790:Enzyme superfamily
2723:Enzyme promiscuity
2616:Sulochrin oxidase
2612:Reticuline oxidase
2334:2011-04-11 at the
1876:10.1007/BF00054105
1825:10.1007/BF00029726
998:electric potential
799:pigment molecules
645:From PsaC; In ETC
423:
244:Photosystem I
241:
231:chloroplast genome
36:
3232:Protein complexes
3204:
3203:
2946:
2945:
2655:
2654:
1929:978-0-12-378630-2
1733:978-0-7167-1007-3
1718:Biology of Plants
1606:10.1021/bi900243f
910:
909:
429:Protein subunits
219:
218:
215:
214:
134:metabolic pathway
3239:
2973:
2966:
2959:
2950:
2826:Hanes–Woolf plot
2769:Enzyme activator
2764:Enzyme inhibitor
2738:Enzyme catalysis
2682:
2675:
2668:
2659:
2473:Xanthine oxidase
2375:
2368:
2361:
2352:
2317:
2316:
2296:
2281:
2280:
2270:
2253:(20): 6095–104.
2247:The EMBO Journal
2238:
2232:
2231:
2203:
2197:
2196:
2178:
2173:
2172:
2171:
2158:
2149:
2148:
2138:
2106:
2093:
2092:
2064:
2053:
2052:
2034:
2002:
1996:
1995:
1985:
1957:
1948:
1947:
1941:
1933:
1907:
1896:
1895:
1851:
1845:
1844:
1800:
1791:
1790:
1785:. Archived from
1779:
1770:
1769:
1756:Plant Physiology
1747:
1738:
1737:
1721:
1711:
1702:
1701:
1691:
1668:Plant Physiology
1659:
1653:
1652:
1624:
1618:
1617:
1589:
1583:
1582:
1564:
1532:
1526:
1525:
1481:
1475:
1474:
1430:
1419:
1418:
1390:
1381:
1380:
1370:
1346:
1340:
1339:
1311:
1290:
1285:
1284:
1216:
1215:
1214:
1201:
1200:
1199:
1184:
1183:
1182:
1171:
1170:
1169:
1156:
1155:
1154:
1113:protein subunits
946:visible spectrum
938:resonance energy
900:
899:
898:
880:
879:
878:
864:
863:
862:
676:Soluble protein
426:
373:that contain 11
292:to catalyze the
254:) is one of two
225:Location of the
172:
50:
38:
3247:
3246:
3242:
3241:
3240:
3238:
3237:
3236:
3222:Light reactions
3207:
3206:
3205:
3200:
3105:
3021:
2986:
2977:
2947:
2942:
2854:Oxidoreductases
2840:
2816:Enzyme kinetics
2804:
2800:List of enzymes
2773:
2742:
2713:Catalytic triad
2691:
2686:
2656:
2651:
2587:
2538:
2515:
2492:
2458:
2430:
2390:
2383:oxidoreductases
2379:
2336:Wayback Machine
2325:
2320:
2298:
2297:
2284:
2240:
2239:
2235:
2205:
2204:
2200:
2176:
2170:
2168:
2167:
2166:
2164:
2160:
2159:
2152:
2108:
2107:
2096:
2075:(1–3): 139–60.
2066:
2065:
2056:
2004:
2003:
1999:
1965:
1959:
1958:
1951:
1934:
1930:
1909:
1908:
1899:
1853:
1852:
1848:
1802:
1801:
1794:
1781:
1780:
1773:
1766:
1749:
1748:
1741:
1734:
1713:
1712:
1705:
1661:
1660:
1656:
1626:
1625:
1621:
1600:(23): 5405–16.
1591:
1590:
1586:
1534:
1533:
1529:
1483:
1482:
1478:
1441:(1–3): 109–24.
1432:
1431:
1422:
1392:
1391:
1384:
1348:
1347:
1343:
1313:
1312:
1308:
1304:
1286:
1279:
1276:
1251:
1238:
1227:
1213:
1211:
1210:
1209:
1207:
1204:
1202:reductase (FNR)
1198:
1196:
1195:
1194:
1192:
1181:
1179:
1178:
1177:
1175:
1168:
1166:
1165:
1164:
1162:
1153:
1151:
1150:
1149:
1147:
1137:
1130:
1126:
1122:
1118:
1110:
1106:
1102:
1098:
1094:
1085:
1078:
1074:
1070:
1066:
1062:
1058:
1049:
1042:
1038:
1034:
1029:
1027:
1023:
979:
973:
926:
924:Antenna complex
918:Photoexcitation
915:
897:
895:
894:
893:
891:
877:
875:
874:
873:
871:
861:
859:
858:
857:
855:
844:
837:
824:
815:
769:
653:
642:
627:
609:
464:
460:
419:
415:
411:
384:
364:
360:
356:
352:
343:
334:
278: I is an
53:
28:
23:. For PS1, see
17:
12:
11:
5:
3245:
3243:
3235:
3234:
3229:
3224:
3219:
3217:Photosynthesis
3209:
3208:
3202:
3201:
3199:
3198:
3193:
3188:
3183:
3178:
3177:
3176:
3171:
3161:
3156:
3151:
3146:
3141:
3136:
3135:
3134:
3132:Dihydroorotase
3129:
3124:
3113:
3111:
3107:
3106:
3104:
3103:
3102:
3101:
3096:
3091:
3086:
3076:
3075:
3074:
3069:
3064:
3054:
3053:
3052:
3047:
3042:
3031:
3029:
3023:
3022:
3020:
3019:
3018:
3017:
3012:
3002:
2996:
2994:
2992:Photosynthesis
2988:
2987:
2978:
2976:
2975:
2968:
2961:
2953:
2944:
2943:
2941:
2940:
2927:
2914:
2901:
2888:
2875:
2862:
2848:
2846:
2842:
2841:
2839:
2838:
2833:
2828:
2823:
2818:
2812:
2810:
2806:
2805:
2803:
2802:
2797:
2792:
2787:
2781:
2779:
2778:Classification
2775:
2774:
2772:
2771:
2766:
2761:
2756:
2750:
2748:
2744:
2743:
2741:
2740:
2735:
2730:
2725:
2720:
2715:
2710:
2705:
2699:
2697:
2693:
2692:
2687:
2685:
2684:
2677:
2670:
2662:
2653:
2652:
2650:
2649:
2644:
2639:
2634:
2629:
2628:
2627:
2622:
2614:
2609:
2604:
2598:
2596:
2589:
2588:
2586:
2585:
2584:
2583:
2578:
2573:
2568:
2557:
2555:
2540:
2539:
2537:
2536:
2530:
2528:
2517:
2516:
2514:
2513:
2507:
2505:
2494:
2493:
2491:
2490:
2485:
2480:
2475:
2469:
2467:
2460:
2459:
2457:
2456:
2451:
2445:
2443:
2432:
2431:
2429:
2428:
2427:
2426:
2421:
2416:
2405:
2403:
2392:
2391:
2380:
2378:
2377:
2370:
2363:
2355:
2349:
2348:
2343:
2338:
2324:
2323:External links
2321:
2319:
2318:
2307:(2): 372–381.
2282:
2233:
2198:
2187:(1): 179–183.
2169:
2150:
2121:(2): 149–152.
2094:
2054:
1997:
1963:
1949:
1928:
1897:
1862:(4): 295–316.
1846:
1792:
1789:on 2009-02-19.
1771:
1764:
1739:
1732:
1703:
1674:(1): 450–463.
1654:
1619:
1584:
1547:(7): 1257–70.
1527:
1476:
1420:
1382:
1341:
1322:(3): 167–204.
1305:
1303:
1300:
1299:
1298:
1292:
1291:
1288:Biology portal
1275:
1272:
1250:
1247:
1237:
1234:
1226:
1223:
1212:
1203:
1197:
1189:
1180:
1167:
1152:
1136:
1133:
1128:
1124:
1120:
1116:
1108:
1104:
1100:
1096:
1092:
1084:
1081:
1076:
1072:
1068:
1064:
1060:
1056:
1048:
1045:
1040:
1036:
1032:
1028:
1025:
1021:
1018:
1000:of about −1.2
975:Main article:
972:
969:
950:cyanobacterium
925:
922:
914:
911:
908:
907:
901:
896:
888:
887:
881:
876:
868:
867:
860:
851:
847:
846:
842:
839:
835:
831:
830:
822:
817:
813:
809:
808:
805:
801:
800:
793:
787:
786:
783:
775:
774:
771:
767:
761:
760:
757:
750:
749:
742:
733:
732:
729:
725:
724:
721:
717:
716:
713:
709:
708:
700:
696:
695:
692:
688:
687:
684:
678:
677:
674:
668:
667:
664:
658:
657:
654:
651:
647:
646:
643:
640:
636:
635:
630:From PsaC; In
628:
625:
621:
620:
614:
607:
600:
599:
588:
584:
583:
572:
568:
567:
556:
552:
551:
540:
536:
535:
524:
520:
519:
503:
499:
498:
487:
483:
482:
470:
466:
465:
462:
458:
455:
451:
450:
446:
445:
438:
434:
433:
430:
417:
413:
409:
399:leucine zipper
382:
369:of 730 to 750
362:
358:
354:
350:
342:
339:
333:
330:
326:Photosystem II
300:membrane from
217:
216:
213:
212:
207:
201:
200:
195:
189:
188:
183:
177:
176:
168:
167:
156:
149:
148:
143:
137:
136:
131:
125:
124:
119:
113:
112:
107:
101:
100:
95:
89:
88:
83:
77:
76:
72:
71:
66:
60:
59:
55:
54:
51:
43:
42:
15:
13:
10:
9:
6:
4:
3:
2:
3244:
3233:
3230:
3228:
3225:
3223:
3220:
3218:
3215:
3214:
3212:
3197:
3194:
3192:
3189:
3187:
3184:
3182:
3179:
3175:
3172:
3170:
3167:
3166:
3165:
3162:
3160:
3157:
3155:
3152:
3150:
3147:
3145:
3142:
3140:
3137:
3133:
3130:
3128:
3125:
3123:
3120:
3119:
3118:
3115:
3114:
3112:
3108:
3100:
3097:
3095:
3092:
3090:
3087:
3085:
3082:
3081:
3080:
3077:
3073:
3070:
3068:
3065:
3063:
3060:
3059:
3058:
3055:
3051:
3048:
3046:
3043:
3041:
3038:
3037:
3036:
3033:
3032:
3030:
3028:
3027:Dehydrogenase
3024:
3016:
3013:
3011:
3008:
3007:
3006:
3003:
3001:
2998:
2997:
2995:
2993:
2989:
2985:
2981:
2974:
2969:
2967:
2962:
2960:
2955:
2954:
2951:
2938:
2934:
2933:
2928:
2925:
2921:
2920:
2915:
2912:
2908:
2907:
2902:
2899:
2895:
2894:
2889:
2886:
2882:
2881:
2876:
2873:
2869:
2868:
2863:
2860:
2856:
2855:
2850:
2849:
2847:
2843:
2837:
2834:
2832:
2829:
2827:
2824:
2822:
2819:
2817:
2814:
2813:
2811:
2807:
2801:
2798:
2796:
2795:Enzyme family
2793:
2791:
2788:
2786:
2783:
2782:
2780:
2776:
2770:
2767:
2765:
2762:
2760:
2759:Cooperativity
2757:
2755:
2752:
2751:
2749:
2745:
2739:
2736:
2734:
2731:
2729:
2726:
2724:
2721:
2719:
2718:Oxyanion hole
2716:
2714:
2711:
2709:
2706:
2704:
2701:
2700:
2698:
2694:
2690:
2683:
2678:
2676:
2671:
2669:
2664:
2663:
2660:
2648:
2645:
2643:
2640:
2638:
2635:
2633:
2630:
2626:
2623:
2621:
2618:
2617:
2615:
2613:
2610:
2608:
2605:
2603:
2600:
2599:
2597:
2594:
2590:
2582:
2579:
2577:
2574:
2572:
2569:
2567:
2564:
2563:
2562:
2559:
2558:
2556:
2553:
2549:
2545:
2541:
2535:
2532:
2531:
2529:
2526:
2522:
2518:
2512:
2509:
2508:
2506:
2503:
2499:
2495:
2489:
2486:
2484:
2481:
2479:
2476:
2474:
2471:
2470:
2468:
2465:
2461:
2455:
2452:
2450:
2449:Ceruloplasmin
2447:
2446:
2444:
2441:
2437:
2433:
2425:
2422:
2420:
2417:
2415:
2412:
2411:
2410:
2407:
2406:
2404:
2401:
2397:
2393:
2388:
2384:
2376:
2371:
2369:
2364:
2362:
2357:
2356:
2353:
2347:
2344:
2342:
2339:
2337:
2333:
2330:
2327:
2326:
2322:
2314:
2310:
2306:
2302:
2295:
2293:
2291:
2289:
2287:
2283:
2278:
2274:
2269:
2264:
2260:
2256:
2252:
2248:
2244:
2237:
2234:
2229:
2225:
2221:
2217:
2213:
2209:
2202:
2199:
2194:
2190:
2186:
2182:
2175:
2157:
2155:
2151:
2146:
2142:
2137:
2132:
2128:
2124:
2120:
2116:
2112:
2105:
2103:
2101:
2099:
2095:
2090:
2086:
2082:
2078:
2074:
2070:
2063:
2061:
2059:
2055:
2050:
2046:
2042:
2038:
2033:
2028:
2024:
2020:
2016:
2012:
2008:
2001:
1998:
1993:
1989:
1984:
1979:
1975:
1971:
1967:
1956:
1954:
1950:
1945:
1939:
1931:
1925:
1921:
1917:
1913:
1906:
1904:
1902:
1898:
1893:
1889:
1885:
1881:
1877:
1873:
1869:
1865:
1861:
1857:
1850:
1847:
1842:
1838:
1834:
1830:
1826:
1822:
1818:
1814:
1811:(1–2): 3–12.
1810:
1806:
1799:
1797:
1793:
1788:
1784:
1778:
1776:
1772:
1767:
1765:0-87893-856-7
1761:
1757:
1753:
1746:
1744:
1740:
1735:
1729:
1725:
1720:
1719:
1710:
1708:
1704:
1699:
1695:
1690:
1685:
1681:
1677:
1673:
1669:
1665:
1658:
1655:
1650:
1646:
1642:
1638:
1635:(2): 244–54.
1634:
1630:
1623:
1620:
1615:
1611:
1607:
1603:
1599:
1595:
1588:
1585:
1580:
1576:
1572:
1568:
1563:
1558:
1554:
1550:
1546:
1542:
1538:
1531:
1528:
1523:
1519:
1515:
1511:
1507:
1503:
1499:
1495:
1491:
1487:
1480:
1477:
1472:
1468:
1464:
1460:
1456:
1452:
1448:
1444:
1440:
1436:
1429:
1427:
1425:
1421:
1416:
1412:
1408:
1404:
1400:
1396:
1389:
1387:
1383:
1378:
1374:
1369:
1364:
1360:
1356:
1352:
1345:
1342:
1337:
1333:
1329:
1325:
1321:
1317:
1310:
1307:
1301:
1297:
1294:
1293:
1289:
1283:
1278:
1273:
1271:
1268:
1264:
1260:
1259:cyanobacteria
1256:
1248:
1246:
1243:
1235:
1233:
1231:
1224:
1222:
1220:
1190:
1188:
1186:
1161:that reduces
1160:
1145:
1141:
1134:
1132:
1114:
1111:are bound to
1090:
1082:
1080:
1054:
1053:phylloquinone
1047:Phylloquinone
1046:
1044:
1019:
1017:
1015:
1011:
1007:
1003:
999:
995:
991:
987:
986:
978:
970:
968:
966:
962:
958:
954:
951:
947:
943:
939:
935:
931:
923:
921:
919:
912:
905:
902:
890:
889:
885:
882:
870:
869:
866:
852:
849:
848:
840:
833:
832:
828:
827:phylloquinone
825:
818:
811:
810:
806:
803:
802:
798:
794:
792:
789:
788:
784:
781:
777:
776:
772:
770:
763:
762:
758:
756:
752:
751:
747:
743:
741:
738:
735:
734:
730:
727:
726:
722:
719:
718:
714:
711:
710:
707:
704:
701:
698:
697:
693:
690:
689:
685:
683:
680:
679:
675:
673:
670:
669:
665:
663:
660:
659:
655:
649:
648:
644:
638:
637:
633:
629:
623:
622:
619:
615:
613:
611:
602:
601:
598:
597:
592:
589:
586:
585:
582:
581:
576:
573:
570:
569:
566:
565:
560:
557:
554:
553:
550:
549:
544:
541:
538:
537:
534:
533:
528:
525:
522:
521:
518:
517:
512:
508:
504:
501:
500:
497:
496:
491:
488:
485:
484:
481:
480:
475:
471:
468:
467:
456:
453:
452:
448:
447:
443:
436:
435:
431:
428:
427:
421:
407:
403:
400:
396:
392:
388:
380:
376:
375:transmembrane
372:
368:
348:
340:
338:
331:
329:
327:
323:
319:
315:
311:
307:
303:
299:
295:
291:
288:
284:
281:
277:
273:
272:cyanobacteria
269:
265:
261:
257:
253:
249:
245:
238:
237:
232:
229:genes in the
228:
223:
211:
208:
206:
202:
199:
196:
194:
190:
187:
184:
182:
178:
173:
169:
166:
163:
160:
157:
154:
150:
147:
144:
142:
138:
135:
132:
130:
126:
123:
120:
118:
114:
111:
110:NiceZyme view
108:
106:
102:
99:
96:
94:
90:
87:
84:
82:
78:
73:
70:
67:
65:
61:
56:
49:
44:
41:Photosystem I
39:
32:
26:
25:PlayStation 1
22:
3009:
2932:Translocases
2929:
2916:
2903:
2890:
2877:
2867:Transferases
2864:
2851:
2708:Binding site
2561:Glutaredoxin
2546:: Acting on
2500:: Acting on
2438:: Oxidizing
2398:: Acting on
2304:
2300:
2250:
2246:
2236:
2211:
2207:
2201:
2184:
2180:
2118:
2115:FEBS Letters
2114:
2072:
2068:
2017:(1): 58–62.
2014:
2011:FEBS Letters
2010:
2000:
1976:(1): 47–52.
1973:
1970:FEBS Letters
1969:
1911:
1859:
1855:
1849:
1808:
1804:
1787:the original
1755:
1717:
1671:
1667:
1657:
1632:
1628:
1622:
1597:
1594:Biochemistry
1593:
1587:
1544:
1540:
1530:
1489:
1486:FEBS Letters
1485:
1479:
1438:
1434:
1398:
1394:
1358:
1354:
1344:
1319:
1315:
1309:
1252:
1239:
1230:Plastocyanin
1228:
1225:Plastocyanin
1205:
1138:
1086:
1050:
1030:
1013:
1009:
984:
983:chlorophyll
980:
956:
952:
927:
916:
807:Description
779:
778:Chlorophyll
765:
764:Chlorophyll
754:
753:Chlorophyll
739:
731:Description
706:phospholipid
686:Description
672:Plastocyanin
605:
594:
578:
562:
546:
530:
514:
509:II binding.
493:
477:
432:Description
377:segments. A
344:
335:
302:plastocyanin
256:photosystems
251:
247:
243:
242:
234:
226:
98:BRENDA entry
3005:Photosystem
2703:Active site
2511:Nitrogenase
2402:as acceptor
2214:(1): 5–26.
1191:Ferredoxin–
1174:ferredoxin–
1089:iron–sulfur
942:wavelengths
934:carotenoids
930:chlorophyll
854:Ferredoxin-
737:Chlorophyll
604:cytochrome
387:coordinated
371:amino acids
296:across the
276:Photosystem
86:IntEnz view
58:Identifiers
3211:Categories
2906:Isomerases
2880:Hydrolases
2747:Regulation
2548:phosphorus
2525:flavodoxin
2400:superoxide
2389:1.15–1.21)
1962:"Vitamin K
1492:(1): 1–4.
1401:: 521–65.
1361:: 81–106.
1302:References
1142:(Fd) is a
1140:Ferredoxin
1135:Ferredoxin
797:carotenoid
791:β-Carotene
662:Ferredoxin
406:downstream
306:ferredoxin
285:that uses
155:structures
122:KEGG entry
3227:EC 1.97.1
2785:EC number
2554:in donors
2504:as donors
1938:cite book
1249:Evolution
1185:reductase
904:Magnesium
821:vitamin K
728:Pigments
596:IPR012986
580:IPR010010
564:IPR036592
548:IPR035982
532:IPR002615
516:IPR001302
495:IPR003375
479:IPR003685
391:cysteines
322:cofactors
298:thylakoid
75:Databases
69:1.97.1.12
2809:Kinetics
2733:Cofactor
2696:Activity
2527:as donor
2332:Archived
2228:10611452
2145:83495051
2089:11687212
2049:42983611
1992:84602152
1892:21845584
1884:24442951
1841:26158482
1833:24442279
1698:27406169
1649:11959504
1614:19432395
1579:32418758
1571:19132290
1562:11131447
1522:42294700
1471:13832448
1463:16328814
1415:16669773
1377:26927905
1274:See also
691:MGDG II
616:Soluble
591:InterPro
575:InterPro
559:InterPro
543:InterPro
527:InterPro
511:InterPro
490:InterPro
474:InterPro
389:by four
381:called F
353:, Acc, A
210:proteins
198:articles
186:articles
159:RCSB PDB
2980:Enzymes
2919:Ligases
2689:Enzymes
2552:arsenic
2277:9321389
2268:1326293
2123:Bibcode
2041:3552735
2019:Bibcode
1864:Bibcode
1813:Bibcode
1724:121–127
1689:5074619
1514:2186925
1494:Bibcode
1443:Bibcode
1336:3333014
1144:soluble
1099:, and F
961:spinach
884:Calcium
829:in ETC
746:pigment
712:PG III
618:protein
612:complex
593::
577::
561::
545::
529::
513::
492::
476::
361:, and F
347:binding
332:History
283:complex
258:in the
146:profile
129:MetaCyc
3089:BCKDHB
3084:BCKDHA
2893:Lyases
2381:Other
2275:
2265:
2226:
2143:
2087:
2047:
2039:
1990:
1926:
1890:
1882:
1839:
1831:
1762:
1730:
1696:
1686:
1647:
1612:
1577:
1569:
1559:
1520:
1512:
1469:
1461:
1413:
1375:
1334:
1159:enzyme
913:Photon
720:PG IV
682:Lipids
634:(ETC)
290:energy
270:, and
268:plants
193:PubMed
175:Search
165:PDBsum
105:ExPASy
93:BRENDA
81:IntEnz
64:EC no.
3174:HADHB
3169:HADHA
3110:Other
2845:Types
2581:GLRX5
2576:GLRX3
2571:GLRX2
2440:metal
2177:(PDF)
2141:S2CID
2045:S2CID
1988:S2CID
1888:S2CID
1837:S2CID
1575:S2CID
1518:S2CID
1467:S2CID
1263:algae
1107:and F
1071:and F
1024:and A
1006:dimer
1002:volts
699:PG I
587:PsaX
571:PsaM
555:PsaL
539:PsaK
523:PsaJ
502:PsaI
486:PsaE
469:PsaD
461:and F
454:PsaC
449:PsaB
437:PsaA
412:and F
402:motif
310:NADPH
287:light
264:algae
250:, or
141:PRIAM
3067:DLST
3062:OGDH
2937:list
2930:EC7
2924:list
2917:EC6
2911:list
2904:EC5
2898:list
2891:EC4
2885:list
2878:EC3
2872:list
2865:EC2
2859:list
2852:EC1
2593:1.21
2566:GLRX
2544:1.20
2521:1.19
2498:1.18
2464:1.17
2442:ions
2436:1.16
2424:SOD3
2419:SOD2
2414:SOD1
2396:1.15
2273:PMID
2224:PMID
2212:1456
2165:NADP
2085:PMID
2073:1507
2037:PMID
1944:link
1924:ISBN
1880:PMID
1829:PMID
1760:ISBN
1728:ISBN
1694:PMID
1645:PMID
1610:PMID
1567:PMID
1510:PMID
1459:PMID
1411:PMID
1373:PMID
1332:PMID
1240:The
1208:NADP
1193:NADP
1176:NADP
1163:NADP
1148:NADP
977:P700
965:P700
932:and
906:ion
886:ion
856:NADP
850:FNR
395:loop
205:NCBI
162:PDBe
117:KEGG
3117:CAD
3099:DLD
3094:DBT
3072:DLD
2550:or
2309:doi
2263:PMC
2255:doi
2216:doi
2189:doi
2131:doi
2119:186
2077:doi
2027:doi
2015:215
1978:doi
1974:243
1916:doi
1872:doi
1821:doi
1684:PMC
1676:doi
1672:172
1637:doi
1602:doi
1557:PMC
1549:doi
1502:doi
1490:264
1451:doi
1403:doi
1363:doi
1324:doi
1320:895
1219:FNR
1095:, F
1035:. A
838:-B
816:-A
795:22
744:90
385:is
357:, A
351:700
318:ATP
304:to
262:of
248:PSI
233:of
227:psa
181:PMC
153:PDB
3213::
3050:E3
3045:E2
3040:E1
3015:II
2982::
2387:EC
2305:88
2303:.
2285:^
2271:.
2261:.
2251:16
2249:.
2245:.
2222:.
2210:.
2185:47
2183:.
2179:.
2153:^
2139:.
2129:.
2117:.
2113:.
2097:^
2083:.
2071:.
2057:^
2043:.
2035:.
2025:.
2013:.
2009:.
1986:.
1972:.
1968:.
1952:^
1940:}}
1936:{{
1922:.
1900:^
1886:.
1878:.
1870:.
1858:.
1835:.
1827:.
1819:.
1807:.
1795:^
1774:^
1754:.
1742:^
1726:.
1706:^
1692:.
1682:.
1670:.
1666:.
1643:.
1633:12
1631:.
1608:.
1598:48
1596:.
1573:.
1565:.
1555:.
1545:66
1543:.
1539:.
1516:.
1508:.
1500:.
1488:.
1465:.
1457:.
1449:.
1439:80
1437:.
1423:^
1409:.
1399:57
1397:.
1385:^
1371:.
1359:67
1357:.
1353:.
1330:.
1318:.
1261:,
1187:.
1051:A
990:nm
892:Mg
872:Ca
782:′
444:.
420:.
328:.
274:.
266:,
3010:I
2972:e
2965:t
2958:v
2939:)
2935:(
2926:)
2922:(
2913:)
2909:(
2900:)
2896:(
2887:)
2883:(
2874:)
2870:(
2861:)
2857:(
2681:e
2674:t
2667:v
2385:(
2374:e
2367:t
2360:v
2315:.
2311::
2279:.
2257::
2230:.
2218::
2195:.
2191::
2147:.
2133::
2125::
2091:.
2079::
2051:.
2029::
2021::
1994:.
1980::
1964:1
1946:)
1932:.
1918::
1894:.
1874::
1866::
1860:6
1843:.
1823::
1815::
1809:9
1768:.
1736:.
1700:.
1678::
1651:.
1639::
1616:.
1604::
1581:.
1551::
1524:.
1504::
1496::
1473:.
1453::
1445::
1417:.
1405::
1379:.
1365::
1338:.
1326::
1129:b
1125:a
1121:x
1117:x
1109:b
1105:a
1101:b
1097:a
1093:x
1077:x
1073:a
1069:b
1065:x
1061:1
1057:1
1041:1
1037:0
1033:x
1026:1
1022:0
1014:a
1010:a
985:a
955:(
843:1
836:K
834:Q
823:1
814:K
812:Q
780:a
768:0
766:a
755:a
740:a
652:x
650:F
641:b
639:F
626:a
624:F
610:f
608:6
606:b
463:b
459:a
418:X
414:B
410:A
383:x
363:x
359:1
355:0
246:(
27:.
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.