1495:(Pol III). It is important to note, however, that the ligands of RIG-I are still being investigated and are controversial. Also notable, is that RIG-I can work together with MDA5 against viruses that RIG-I itself may not create a significant enough response. In addition, for many viruses, effective RIG-I-mediated antiviral responses are dependent on functionally active LGP2. Cells are synthesizing multiple types of RNA at all times, so it is important that RIG-I is not going to bind to those RNAs. Native RNA from inside the cell contains an N
393:
370:
267:
292:
644:
1392:
399:
298:
40:
1370:. Other studies have shown that in different microenvironments, such as in cancerous cells, RIG-I has more functions other than viral recognition. RIG-I orthologs are found in mammals, geese, ducks, some fish, and some reptiles. RIG-I is in most cells, including various innate immune system cells, and is usually in an inactive state.
1467:
enters a cell, and it takes over the cell's machinery to self replicate. Once a virus has begun replication, the infected cell is no longer useful and potentially harmful to its host, and the host's immune system must be notified. RIG-I functions as a pattern recognition receptor and PRR's are the
1485:
RIG-I is located in the cytoplasm where its function is to recognize its PAMP, which are ideally short (<300 base pairs) dsRNA with a 5′ triphosphate (5′ ppp). However, it has been noted that while not ideal, and response is weakened, RIG-I can recognize 5′ diphosphate (5′pp). This ability is
1490:
opens up more doors for recognition. An example of viruses evolving to evade RIG-I is in the case of certain retroviruses, such as HIV-1, encode a protease that directs RIG-I to the lysosome for degradation, and thereby evade RIG-I mediated signaling. The dsRNA can come from single-stranded RNA
3343:
Note: RARRES3 (Gene ID: 5920) and DDX58 (Gene ID: 23586) share the RIG1/RIG-1 alias in common. RIG1 is a widely used alternative name for DExD/H-box helicase 58 (DDX58), which can be confused with the retinoic acid receptor responder 3 (RARRES3) gene, since they share the same alias.
1528:, which create an antiviral environment. Once the IFN1s leave the cell, they can bind to IFN1 receptors on the cell surface from which they came from, or other cells close by. This will upregulate the production of more IFN1s, boosting an antiviral environment. IFN1 also activates the
1516:) CARD domain and bind. RIG-I CARD interactions have their own regulatory system. Although RIG-I expresses a CARD at all times, it must be activated by the ligand before it will allow both CARDs to interact with the MAVS CARD. This interaction will start the pathway to making
1395:
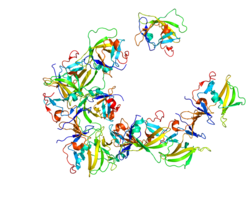
This image shows the basic structure of RIG-I while it is inactive and active. Shown are the N-terminus CARDs, the DExD/H helicase domain, and the C-terminus repressor domain (RD). Included on the active structure is the most common RIG-I PAMP, dsRNA with a 5' triphosphate
66:
1491:(ssRNA) viruses or from dsRNA viruses. The ssRNA viruses are not typically recognized as ssRNA, but through intermittent replication products in the form of dsRNA. RIG-I is also able to detect non-self 5′-triphosphorylated dsRNA transcribed from AT-rich dsDNA by
1507:
RIG-I is a signaling molecule and is usually in a condensed resting state until it is activated. Once RIG-I is bound to its PAMP, molecules such as PACT and zinc antiviral protein short isoform (ZAPs), help keep RIG-I in an activated state which then keeps the
1476:
cells. In addition, they are located in many different parts of those cells, such as the cell membrane, endosomal membrane, and in the cytosol, to provide the most protection against many types of invaders (i.e., extracellular and intracellular microbes).
1580:(ATRA) in leukemia cells. RIG-I and the other genes were assigned the temporary name of RIG (retinoic acid–induced gene) in the format of RIG-A, RIG-B etc by the group, however they performed no additional characterization on RIG-I.
1374:
that have been designed to have a deleted or non-functioning RIG-I gene are not healthy and typically die embryonically. If they survive, the mice have serious developmental dysfunction. The stimulator of interferon genes
1472:(PAMP). Once the PAMP is recognized, it can then lead to a signaling cascade producing an inflammatory response or an interferon response. PRRs are located in many different cell types, however most notably active in the
1355:(cap-0), but not RNA with a 5' m7G cap having a ribose 2′-O-methyl modification (cap-1). These are often generated during a viral infection but can also be host-derived. Once activated by the dsRNA, the N-terminus
2682:
Imaizumi T, Yagihashi N, Hatakeyama M, Yamashita K, Ishikawa A, Taima K, et al. (July 2004). "Expression of retinoic acid-inducible gene-I in vascular smooth muscle cells stimulated with interferon-gamma".
2614:
Cui XF, Imaizumi T, Yoshida H, Borden EC, Satoh K (June 2004). "Retinoic acid-inducible gene-I is induced by interferon-gamma and regulates the expression of interferon-gamma stimulated gene 15 in MCF-7 cells".
2644:
Yoneyama M, Kikuchi M, Natsukawa T, Shinobu N, Imaizumi T, Miyagishi M, et al. (July 2004). "The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses".
2776:
Sakaki H, Imaizumi T, Matsumiya T, Kusumi A, Nakagawa H, Kubota K, et al. (February 2005). "Retinoic acid-inducible gene-I is induced by interleukin-1beta in cultured human gingival fibroblasts".
2585:
Imaizumi T, Aratani S, Nakajima T, Carlson M, Matsumiya T, Tanji K, et al. (March 2002). "Retinoic acid-inducible gene-I is induced in endothelial cells by LPS and regulates expression of COX-2".
3203:
1366:
Type-I IFNs have three main functions: to limit the virus from spreading to nearby cells, promote an innate immune response, including inflammatory responses, and help activate the
2747:
Imaizumi T, Hatakeyama M, Yamashita K, Yoshida H, Ishikawa A, Taima K, et al. (2004). "Interferon-gamma induces retinoic acid-inducible gene-I in endothelial cells".
3057:
Kawai T, Takahashi K, Sato S, Coban C, Kumar H, Kato H, et al. (October 2005). "IPS-1, an adaptor triggering RIG-I- and Mda5-mediated type I interferon induction".
406:
305:
3196:
1032:
1013:
228:
1544:
cells (BrCa) to resist treatments and grow because of an IFN response to noncoding RNA. In contrast, RIG-I in other types of cancer, such as acute
2885:"Inhibition of RIG-I-dependent signaling to the interferon pathway during hepatitis C virus expression and restoration of signaling by IKKepsilon"
3189:
2225:"cGAS-STING effectively restricts murine norovirus infection but antagonizes the antiviral action of N-terminus of RIG-I in mouse macrophages"
1657:
1639:
1469:
3356:
2807:"Regulating intracellular antiviral defense and permissiveness to hepatitis C virus RNA replication through a cellular RNA helicase, RIG-I"
1552:, can act as a tumor suppressor. If cancer causing viruses infect a cell, however, RIG-I can lead to cell death. Cell death can occur via
392:
1232:
1239:
369:
3371:
2928:"Human ISG15 conjugation targets both IFN-induced and constitutively expressed proteins functioning in diverse cellular pathways"
3247:
1626:
1605:
3016:"Identification and characterization of MAVS, a mitochondrial antiviral signaling protein that activates NF-kappaB and IRF 3"
1572:
In 2000, RIG-I was named by researchers from the
Shanghai Institute of Hematology who identified novel genes that respond to
1622:
1426:(CARDs) that are important for interactions with mitochondrial antiviral signaling protein (MAVS). RIG-I is a member of the
291:
266:
63:
3366:
2272:
1601:
3302:
3262:
3224:
3216:
2714:"Upregulation of retinoic acid-inducible gene-I in T24 urinary bladder carcinoma cells stimulated with interferon-gamma"
1864:"Structural basis for m7G recognition and 2'-O-methyl discrimination in capped RNAs by the innate immune receptor RIG-I"
1460:
1302:
208:
1540:
Usually, RIG-I recognizes foreign RNA. However, it can sometimes recognize "self" RNAs. RIG-I has been shown to enable
1379:
antagonizes RIG-I by binding its N-terminus, probably as to avoid overactivation of RIG-I signaling and the associated
405:
304:
1529:
1758:"RIG-I-mediated antiviral signaling is inhibited in HIV-1 infection by a protease-mediated sequestration of RIG-I"
1347:
that is activated by immunostimulatory RNAs from viruses as well as RNAs of other origins. RIG-I recognizes short
398:
297:
2512:"Gene expression networks underlying retinoic acid-induced differentiation of acute promyelocytic leukemia cells"
2850:"Distinct poly(I-C) and virus-activated signaling pathways leading to interferon-beta production in hepatocytes"
1549:
1077:
216:
3242:
2981:"Shared and unique functions of the DExD/H-box helicases RIG-I, MDA5, and LGP2 in antiviral innate immunity"
1058:
3130:
2116:"RIG-I like receptor sensing of host RNAs facilitates the cell-intrinsic immune response to KSHV infection"
1367:
2298:"MAVS forms functional prion-like aggregates to activate and propagate antiviral innate immune response"
1517:
1318:
2461:Żeromski J, Kaczmarek M, Boruczkowski M, Kierepa A, Kowala-Piaskowska A, Mozer-Lisewska I (June 2019).
1512:(CARDs) ready for binding. The molecule will migrate to the mitochondrial antiviral signaling protein (
3181:
2556:
Bowie AG, Fitzgerald KA (April 2007). "RIG-I: tri-ing to discriminate between self and non-self RNA".
3212:
3145:
2939:
2415:
2127:
1875:
1818:
1473:
1463:(PRRs) are a part of the innate immune system used for recognizing invaders. In a viral infection, a
1310:
280:
199:, DEAD (Asp-Glu-Ala-Asp) box polypeptide 58, RIGI, RLR-1, SGMRT2, RIG-I, DEXD/H-box helicase 58, RIG1
195:
2979:
Yoneyama M, Kikuchi M, Matsumoto K, Imaizumi T, Miyagishi M, Taira K, et al. (September 2005).
2712:
Imaizumi T, Yagihashi N, Hatakeyama M, Yamashita K, Ishikawa A, Taima K, et al. (August 2004).
1805:
Goubau D, Schlee M, Deddouche S, Pruijssers AJ, Zillinger T, Goldeck M, et al. (October 2014).
3275:
3252:
1561:
1427:
1408:
1348:
1341:
3131:"Cardif is an adaptor protein in the RIG-I antiviral pathway and is targeted by hepatitis C virus"
3129:
Meylan E, Curran J, Hofmann K, Moradpour D, Binder M, Bartenschlager R, Tschopp J (October 2005).
3361:
3232:
3169:
3082:
3045:
2670:
2065:
Chiang JJ, Sparrer KM, van Gent M, Lässig C, Huang T, Osterrieder N, et al. (January 2018).
1756:
Solis M, Nakhaei P, Jalalirad M, Lacoste J, Douville R, Arguello M, et al. (February 2011).
1492:
240:
1215:
1194:
1168:
1147:
3270:
3161:
3117:
3074:
3037:
3002:
2967:
2914:
2871:
2836:
2793:
2764:
2735:
2700:
2662:
2632:
2602:
2573:
2533:
2492:
2443:
2381:
2327:
2254:
2205:
2153:
2096:
2047:
1991:
1903:
1844:
1787:
1735:
1306:
643:
188:
56:
17:
3237:
3153:
3107:
3066:
3027:
2992:
2957:
2947:
2904:
2896:
2861:
2826:
2818:
2785:
2756:
2725:
2692:
2654:
2624:
2594:
2565:
2523:
2482:
2474:
2433:
2423:
2371:
2361:
2317:
2309:
2244:
2236:
2195:
2187:
2143:
2135:
2086:
2078:
2067:"Viral unmasking of cellular 5S rRNA pseudogene transcripts induces RIG-I-mediated immunity"
2037:
2029:
1981:
1971:
1893:
1883:
1862:
Devarkar SC, Wang C, Miller MT, Ramanathan A, Jiang F, Khan AG, et al. (January 2016).
1834:
1826:
1777:
1769:
1725:
1717:
1545:
485:
416:
360:
315:
1391:
1359:(CARDs) migrate and bind with CARDs attached to mitochondrial antiviral signaling protein (
236:
2402:
Satoh T, Kato H, Kumagai Y, Yoneyama M, Sato S, Matsushita K, et al. (January 2010).
1314:
460:
1960:"Comparative Structure and Function Analysis of the RIG-I-Like Receptors: RIG-I and MDA5"
3149:
2943:
2883:
Breiman A, Grandvaux N, Lin R, Ottone C, Akira S, Yoneyama M, et al. (April 2005).
2419:
2348:
Amarante-Mendes GP, Adjemian S, Branco LM, Zanetti LC, Weinlich R, Bortoluci KR (2018).
2131:
1879:
1822:
2962:
2927:
2909:
2884:
2487:
2462:
2438:
2403:
2376:
2349:
2322:
2297:
2249:
2224:
2200:
2175:
2148:
2115:
2091:
2066:
2042:
2017:
1986:
1959:
1898:
1863:
1839:
1806:
1782:
1757:
1730:
1709:
1371:
1352:
39:
2831:
2806:
1313:
for recognizing cells that have been infected with a virus. These viruses can include
947:
942:
937:
932:
927:
922:
917:
912:
907:
902:
897:
892:
887:
882:
877:
872:
867:
862:
857:
852:
847:
842:
837:
832:
827:
811:
806:
801:
796:
791:
786:
781:
776:
771:
766:
750:
745:
740:
735:
730:
725:
720:
715:
710:
705:
700:
695:
690:
685:
3350:
3326:
2789:
1541:
672:
3086:
3049:
2900:
2822:
2674:
220:
3173:
1411:
1380:
1344:
1326:
478:
257:
2240:
1807:"Antiviral immunity via RIG-I-mediated recognition of RNA bearing 5'-diphosphates"
863:
positive regulation of granulocyte macrophage colony-stimulating factor production
3112:
3095:
2510:
Liu TX, Zhang JW, Tao J, Zhang RB, Zhang QH, Zhao CJ, et al. (August 2000).
244:
3330:
2997:
2980:
2805:
Sumpter R, Loo YM, Foy E, Li K, Yoneyama M, Fujita T, et al. (March 2005).
1721:
1662:
National Center for
Biotechnology Information, U.S. National Library of Medicine
1644:
National Center for
Biotechnology Information, U.S. National Library of Medicine
1513:
1509:
1439:
1423:
1360:
1356:
1334:
1322:
3032:
3015:
2932:
Proceedings of the
National Academy of Sciences of the United States of America
2478:
2408:
Proceedings of the
National Academy of Sciences of the United States of America
2313:
2139:
1868:
Proceedings of the
National Academy of Sciences of the United States of America
853:
cytoplasmic pattern recognition receptor signaling pathway in response to virus
561:
2760:
2696:
2404:"LGP2 is a positive regulator of RIG-I- and MDA5-mediated antiviral responses"
2082:
2033:
1419:
1415:
1330:
377:
274:
224:
2569:
2528:
2511:
2366:
1976:
3310:
3096:"VISA is an adapter protein required for virus-triggered IFN-beta signaling"
2952:
2428:
1888:
1573:
1557:
1553:
977:
621:
499:
444:
431:
343:
330:
232:
3165:
3121:
3078:
3041:
3006:
2971:
2918:
2875:
2866:
2849:
2840:
2797:
2768:
2739:
2704:
2666:
2636:
2606:
2598:
2577:
2537:
2496:
2447:
2385:
2350:"Pattern Recognition Receptors and the Host Cell Death Molecular Machinery"
2331:
2273:"DDX58 DExD/H-box helicase 58 [Homo sapiens (human)] - Gene - NCBI"
2258:
2209:
2157:
2100:
2051:
1995:
1907:
1848:
1791:
1739:
1486:
important as many viruses have evolved to evade RIG-I, so having the dual
1279:
2730:
2713:
1773:
1525:
1263:
1122:
1103:
3157:
2018:"RIG-I: a multifunctional protein beyond a pattern recognition receptor"
1830:
1521:
1468:
molecules that start the notification process. PRRs recognize specific
1299:
1089:
1044:
2463:"Significance and Role of Pattern Recognition Receptors in Malignancy"
2223:
Yu P, Miao Z, Li Y, Bansal R, Peppelenbosch MP, Pan Q (January 2021).
1274:
165:
161:
157:
153:
149:
145:
141:
137:
133:
129:
125:
121:
117:
113:
109:
105:
101:
97:
93:
89:
85:
1487:
1247:
999:
2628:
2191:
3070:
2658:
1532:
pathway, leading to the production of IFN-stimulated genes (ISGs).
1464:
1376:
2296:
Hou F, Sun L, Zheng H, Skaug B, Jiang QX, Chen ZJ (August 2011).
962:
958:
948:
positive regulation of myeloid dendritic cell cytokine production
3290:
3285:
3094:
Xu LG, Wang YY, Han KJ, Li LY, Zhai Z, Shu HB (September 2005).
1435:
1431:
1404:
943:
positive regulation of DNA-binding transcription factor activity
652:
212:
3185:
2926:
Zhao C, Denison C, Huibregtse JM, Gygi S, Krug RM (July 2005).
1351:(dsRNA) in the cytosol with a 5' tri- or di-phosphate end or a
2114:
Zhao Y, Ye X, Dunker W, Song Y, Karijolich J (November 2018).
2016:
Xu XX, Wan H, Nie L, Shao T, Xiang LX, Shao JZ (March 2018).
1499:
2'O-Methyl self RNA marker that deters RIG-I from binding.
898:
positive regulation of transcription by RNA polymerase II
913:
positive regulation of defense response to virus by host
468:
2343:
2341:
1309:(IFN1) response. RIG-I is an essential molecule in the
2848:
Li K, Chen Z, Kato N, Gale M, Lemon SM (April 2005).
633:
928:
positive regulation of response to cytokine stimulus
3319:
3301:
3261:
3223:
2587:
1208:
1187:
1161:
1140:
878:
negative regulation of type I interferon production
1618:
1616:
1614:
1597:
1595:
1593:
1438:). RIG-I and MDA5 are both involved in activating
838:positive regulation of interferon-alpha production
3014:Seth RB, Sun L, Ea CK, Chen ZJ (September 2005).
2467:Archivum Immunologiae et Therapiae Experimentalis
918:positive regulation of interferon-beta production
415:
314:
2169:
2167:
1434:(MDA5) and Laboratory of genetics physiology 2 (
873:positive regulation of interleukin-6 production
868:positive regulation of interleukin-8 production
1953:
1951:
1949:
1947:
1945:
1943:
1941:
1939:
1937:
1623:GRCm38: Ensembl release 89: ENSMUSG00000040296
1560:pathway, or through IFN-dependent T cells and
1418:that binds to the target RNA. Included on the
1363:) to activate the signaling pathway for IFN1.
3197:
2397:
2395:
1935:
1933:
1931:
1929:
1927:
1925:
1923:
1921:
1919:
1917:
1432:Melanoma Differentiation-Associated protein 5
8:
1751:
1749:
1703:
1701:
1699:
1697:
1695:
1693:
1691:
1407:in humans. RIG-I is a helical ATP-dependent
908:regulation of type III interferon production
2718:The Tohoku Journal of Experimental Medicine
2176:"Regulation of type I interferon responses"
1689:
1687:
1685:
1683:
1681:
1679:
1677:
1675:
1673:
1671:
1602:GRCh38: Ensembl release 89: ENSG00000107201
3204:
3190:
3182:
2011:
2009:
2007:
2005:
1510:caspase activation and recruitment domains
1424:caspase activation and recruitment domains
1357:caspase activation and recruitment domains
973:
668:
456:
355:
252:
74:
3111:
3031:
2996:
2961:
2951:
2908:
2865:
2830:
2729:
2527:
2486:
2437:
2427:
2375:
2365:
2321:
2248:
2199:
2147:
2090:
2041:
1985:
1975:
1897:
1887:
1838:
1781:
1729:
1390:
2174:Ivashkiv LB, Donlin LT (January 2014).
1589:
1305:(PRR) that can mediate induction of a
1470:Pathogen-Associated Molecular Patterns
1442:and triggering an antiviral response.
858:positive regulation of gene expression
29:
420:
381:
376:
319:
278:
273:
7:
1414:with a repressor domain (RD) on the
933:cellular response to exogenous dsRNA
2854:The Journal of Biological Chemistry
1205:
1184:
1158:
1137:
1113:
1094:
1068:
1049:
1023:
1004:
638:
556:
494:
473:
25:
1451:As a Pattern Recognition Receptor
27:Mammalian protein found in humans
2790:10.1111/j.1399-302X.2005.00181.x
2778:Oral Microbiology and Immunology
1710:"RIG-I in RNA virus recognition"
1493:DNA-dependent RNA polymerase III
751:ubiquitin protein ligase binding
642:
404:
397:
391:
368:
303:
296:
290:
265:
38:
2901:10.1128/JVI.79.7.3969-3978.2005
2823:10.1128/JVI.79.5.2689-2699.2005
1353:5' 7-methyl guanosine (m7G) cap
1296:retinoic acid-inducible gene I
653:More reference expression data
622:More reference expression data
18:Retinoic acid-inducible gene I
1:
3217:Pattern recognition receptors
2617:Biochemistry and Cell Biology
2241:10.1080/19490976.2021.1959839
1461:Pattern Recognition Receptors
1456:Pattern Recognition Receptors
604:epithelium of small intestine
389:
288:
3113:10.1016/j.molcel.2005.08.014
1708:Kell AM, Gale M (May 2015).
1520:and type-1 Interferon (IFN1;
1303:pattern recognition receptor
828:regulation of cell migration
3357:Genes on human chromosome 9
2998:10.4049/jimmunol.175.5.2851
1722:10.1016/j.virol.2015.02.017
1319:Japanese Encephalitis virus
833:response to exogenous dsRNA
731:double-stranded DNA binding
726:double-stranded RNA binding
711:single-stranded RNA binding
546:superficial temporal artery
3388:
3033:10.1016/j.cell.2005.08.012
2479:10.1007/s00005-019-00540-x
2314:10.1016/j.cell.2011.06.041
2180:Nature Reviews. Immunology
2140:10.1038/s41467-018-07314-7
1430:(RLRs) that also includes
1340:RIG-I is an ATP-dependent
2761:10.1080/10623320490512156
2697:10.1016/j.lfs.2004.01.030
2083:10.1038/s41590-017-0005-y
2034:10.1007/s13238-017-0431-5
1658:"Mouse PubMed Reference:"
1640:"Human PubMed Reference:"
1568:Identification and Naming
1518:proinflammatory cytokines
1503:Type-1 Interferon Pathway
1278:
1273:
1269:
1262:
1246:
1227:
1212:
1191:
1180:
1165:
1144:
1133:
1120:
1116:
1101:
1097:
1088:
1075:
1071:
1056:
1052:
1043:
1030:
1026:
1011:
1007:
998:
983:
976:
972:
956:
938:defense response to virus
782:bicellular tight junction
716:identical protein binding
671:
667:
650:
641:
632:
619:
568:
559:
506:
497:
467:
459:
455:
438:
425:
388:
367:
358:
354:
337:
324:
287:
264:
255:
251:
206:
203:
193:
186:
181:
82:
77:
60:
55:
50:
46:
37:
32:
2570:10.1016/j.it.2007.02.002
2529:10.1182/blood.V96.4.1496
2367:10.3389/fimmu.2018.02379
1977:10.3389/fimmu.2019.01586
1550:hepatocellular carcinoma
1400:RIG-I is encoded by the
923:protein deubiquitination
542:internal globus pallidus
518:tendon of biceps brachii
3372:Intracellular receptors
3243:Formyl peptide receptor
2953:10.1073/pnas.0504754102
2429:10.1073/pnas.0912986107
2354:Frontiers in Immunology
1964:Frontiers in Immunology
1958:Brisse M, Ly H (2019).
1889:10.1073/pnas.1515152113
1233:Chr 9: 32.46 – 32.53 Mb
903:RIG-I signaling pathway
2867:10.1074/jbc.M414139200
2599:10.1006/bbrc.2002.6650
1397:
1368:adaptive immune system
1240:Chr 4: 40.2 – 40.24 Mb
893:innate immune response
580:lumbar spinal ganglion
576:mesenteric lymph nodes
2985:Journal of Immunology
2120:Nature Communications
1394:
843:immune system process
3367:RIG-I-like receptors
3213:Innate immune system
2731:10.1620/tjem.203.313
2558:Trends in Immunology
2277:www.ncbi.nlm.nih.gov
1774:10.1128/JVI.01635-10
1716:. 479–480: 110–121.
1562:natural killer cells
1474:innate immune system
1428:RIG-I like receptors
1311:innate immune system
746:nucleic acid binding
383:Chromosome 4 (mouse)
281:Chromosome 9 (human)
78:List of PDB id codes
51:Available structures
3276:RIG-I-like receptor
3253:Complement receptor
3225:Membrane-bound_PRRs
3158:10.1038/nature04193
3150:2005Natur.437.1167M
3144:(7062): 1167–1172.
2944:2005PNAS..10210200Z
2938:(29): 10200–10205.
2889:Journal of Virology
2860:(17): 16739–16747.
2811:Journal of Virology
2420:2010PNAS..107.1512S
2132:2018NatCo...9.4841Z
1880:2016PNAS..113..596D
1831:10.1038/nature13590
1823:2014Natur.514..372G
1762:Journal of Virology
1349:double-stranded RNA
3248:Scavenger receptor
3233:Toll-like receptor
2022:Protein & Cell
1398:
1078:ENSMUSG00000040296
883:detection of virus
821:Biological process
802:actin cytoskeleton
760:Cellular component
736:hydrolase activity
686:nucleotide binding
679:Molecular function
510:buccal mucosa cell
3340:
3339:
3271:NOD-like receptor
3059:Nature Immunology
2691:(10): 1171–1180.
2647:Nature Immunology
2071:Nature Immunology
1817:(7522): 372–375.
1307:type-I interferon
1289:
1288:
1285:
1284:
1258:
1257:
1223:
1222:
1202:
1201:
1176:
1175:
1155:
1154:
1129:
1128:
1110:
1109:
1084:
1083:
1065:
1064:
1039:
1038:
1020:
1019:
968:
967:
848:response to virus
701:metal ion binding
691:helicase activity
663:
662:
659:
658:
628:
627:
615:
614:
553:
552:
451:
450:
350:
349:
245:DDX58 - orthologs
177:
176:
173:
172:
61:Ortholog search:
16:(Redirected from
3379:
3263:Cytoplasmic_PRRs
3238:Mannose receptor
3206:
3199:
3192:
3183:
3177:
3135:
3125:
3115:
3090:
3053:
3035:
3010:
3000:
2991:(5): 2851–2858.
2975:
2965:
2955:
2922:
2912:
2895:(7): 3969–3978.
2879:
2869:
2844:
2834:
2817:(5): 2689–2699.
2801:
2772:
2755:(3–4): 169–173.
2743:
2733:
2708:
2678:
2640:
2610:
2581:
2542:
2541:
2531:
2522:(4): 1496–1504.
2507:
2501:
2500:
2490:
2458:
2452:
2451:
2441:
2431:
2414:(4): 1512–1517.
2399:
2390:
2389:
2379:
2369:
2345:
2336:
2335:
2325:
2293:
2287:
2286:
2284:
2283:
2269:
2263:
2262:
2252:
2220:
2214:
2213:
2203:
2171:
2162:
2161:
2151:
2111:
2105:
2104:
2094:
2062:
2056:
2055:
2045:
2013:
2000:
1999:
1989:
1979:
1955:
1912:
1911:
1901:
1891:
1859:
1853:
1852:
1842:
1802:
1796:
1795:
1785:
1768:(3): 1224–1236.
1753:
1744:
1743:
1733:
1705:
1666:
1665:
1654:
1648:
1647:
1636:
1630:
1620:
1609:
1599:
1546:myeloid leukemia
1271:
1270:
1242:
1235:
1218:
1206:
1197:
1185:
1181:RefSeq (protein)
1171:
1159:
1150:
1138:
1114:
1095:
1069:
1050:
1024:
1005:
974:
696:zinc ion binding
669:
655:
646:
639:
624:
564:
562:Top expressed in
557:
522:white blood cell
502:
500:Top expressed in
495:
474:
457:
447:
434:
423:
408:
401:
395:
384:
372:
356:
346:
333:
322:
307:
300:
294:
283:
269:
253:
247:
198:
191:
168:
75:
69:
48:
47:
42:
30:
21:
3387:
3386:
3382:
3381:
3380:
3378:
3377:
3376:
3347:
3346:
3341:
3336:
3320:Other/ungrouped
3315:
3297:
3257:
3219:
3210:
3180:
3133:
3128:
3093:
3065:(10): 981–988.
3056:
3013:
2978:
2925:
2882:
2847:
2804:
2775:
2746:
2711:
2681:
2643:
2629:10.1139/o04-041
2613:
2584:
2555:
2551:
2549:Further reading
2546:
2545:
2509:
2508:
2504:
2460:
2459:
2455:
2401:
2400:
2393:
2347:
2346:
2339:
2295:
2294:
2290:
2281:
2279:
2271:
2270:
2266:
2222:
2221:
2217:
2192:10.1038/nri3581
2173:
2172:
2165:
2113:
2112:
2108:
2064:
2063:
2059:
2015:
2014:
2003:
1957:
1956:
1915:
1861:
1860:
1856:
1804:
1803:
1799:
1755:
1754:
1747:
1707:
1706:
1669:
1656:
1655:
1651:
1638:
1637:
1633:
1621:
1612:
1600:
1591:
1586:
1570:
1538:
1536:In Cancer Cells
1505:
1498:
1483:
1458:
1453:
1448:
1389:
1315:West Nile virus
1280:View/Edit Mouse
1275:View/Edit Human
1238:
1231:
1228:Location (UCSC)
1214:
1193:
1167:
1146:
1059:ENSG00000107201
952:
816:
792:ruffle membrane
787:plasma membrane
772:cell projection
755:
706:protein binding
651:
620:
611:
606:
602:
598:
594:
590:
586:
582:
578:
574:
560:
549:
544:
540:
536:
534:Achilles tendon
532:
528:
524:
520:
516:
512:
498:
442:
429:
421:
411:
410:
409:
402:
382:
359:Gene location (
341:
328:
320:
310:
309:
308:
301:
279:
256:Gene location (
207:
194:
187:
84:
62:
28:
23:
22:
15:
12:
11:
5:
3385:
3383:
3375:
3374:
3369:
3364:
3359:
3349:
3348:
3338:
3337:
3335:
3334:
3323:
3321:
3317:
3316:
3314:
3313:
3307:
3305:
3299:
3298:
3296:
3295:
3294:
3293:
3288:
3283:
3273:
3267:
3265:
3259:
3258:
3256:
3255:
3250:
3245:
3240:
3235:
3229:
3227:
3221:
3220:
3211:
3209:
3208:
3201:
3194:
3186:
3179:
3178:
3126:
3106:(6): 727–740.
3100:Molecular Cell
3091:
3071:10.1038/ni1243
3054:
3026:(5): 669–682.
3011:
2976:
2923:
2880:
2845:
2802:
2773:
2744:
2724:(4): 313–318.
2709:
2679:
2659:10.1038/ni1087
2653:(7): 730–737.
2641:
2623:(3): 401–405.
2611:
2593:(1): 274–279.
2582:
2564:(4): 147–150.
2552:
2550:
2547:
2544:
2543:
2502:
2473:(3): 133–141.
2453:
2391:
2337:
2308:(3): 448–461.
2288:
2264:
2235:(1): 1959839.
2215:
2163:
2106:
2057:
2028:(3): 246–253.
2001:
1913:
1874:(3): 596–601.
1854:
1797:
1745:
1667:
1649:
1631:
1610:
1588:
1587:
1585:
1582:
1569:
1566:
1537:
1534:
1504:
1501:
1496:
1482:
1479:
1457:
1454:
1452:
1449:
1447:
1444:
1388:
1385:
1287:
1286:
1283:
1282:
1277:
1267:
1266:
1260:
1259:
1256:
1255:
1253:
1251:
1244:
1243:
1236:
1229:
1225:
1224:
1221:
1220:
1210:
1209:
1203:
1200:
1199:
1189:
1188:
1182:
1178:
1177:
1174:
1173:
1163:
1162:
1156:
1153:
1152:
1142:
1141:
1135:
1131:
1130:
1127:
1126:
1118:
1117:
1111:
1108:
1107:
1099:
1098:
1092:
1086:
1085:
1082:
1081:
1073:
1072:
1066:
1063:
1062:
1054:
1053:
1047:
1041:
1040:
1037:
1036:
1028:
1027:
1021:
1018:
1017:
1009:
1008:
1002:
996:
995:
990:
985:
981:
980:
970:
969:
966:
965:
954:
953:
951:
950:
945:
940:
935:
930:
925:
920:
915:
910:
905:
900:
895:
890:
885:
880:
875:
870:
865:
860:
855:
850:
845:
840:
835:
830:
824:
822:
818:
817:
815:
814:
809:
804:
799:
794:
789:
784:
779:
774:
769:
763:
761:
757:
756:
754:
753:
748:
743:
738:
733:
728:
723:
718:
713:
708:
703:
698:
693:
688:
682:
680:
676:
675:
665:
664:
661:
660:
657:
656:
648:
647:
636:
630:
629:
626:
625:
617:
616:
613:
612:
610:
609:
605:
601:
597:
593:
589:
585:
581:
577:
573:
569:
566:
565:
554:
551:
550:
548:
547:
543:
539:
535:
531:
527:
523:
519:
515:
511:
507:
504:
503:
491:
490:
482:
471:
465:
464:
461:RNA expression
453:
452:
449:
448:
440:
436:
435:
427:
424:
419:
413:
412:
403:
396:
390:
386:
385:
380:
374:
373:
365:
364:
352:
351:
348:
347:
339:
335:
334:
326:
323:
318:
312:
311:
302:
295:
289:
285:
284:
277:
271:
270:
262:
261:
249:
248:
205:
201:
200:
192:
184:
183:
179:
178:
175:
174:
171:
170:
80:
79:
71:
70:
59:
53:
52:
44:
43:
35:
34:
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
3384:
3373:
3370:
3368:
3365:
3363:
3360:
3358:
3355:
3354:
3352:
3345:
3332:
3328:
3327:Immunophilins
3325:
3324:
3322:
3318:
3312:
3309:
3308:
3306:
3304:
3303:Secreted_PRRs
3300:
3292:
3289:
3287:
3284:
3282:
3279:
3278:
3277:
3274:
3272:
3269:
3268:
3266:
3264:
3260:
3254:
3251:
3249:
3246:
3244:
3241:
3239:
3236:
3234:
3231:
3230:
3228:
3226:
3222:
3218:
3214:
3207:
3202:
3200:
3195:
3193:
3188:
3187:
3184:
3175:
3171:
3167:
3163:
3159:
3155:
3151:
3147:
3143:
3139:
3132:
3127:
3123:
3119:
3114:
3109:
3105:
3101:
3097:
3092:
3088:
3084:
3080:
3076:
3072:
3068:
3064:
3060:
3055:
3051:
3047:
3043:
3039:
3034:
3029:
3025:
3021:
3017:
3012:
3008:
3004:
2999:
2994:
2990:
2986:
2982:
2977:
2973:
2969:
2964:
2959:
2954:
2949:
2945:
2941:
2937:
2933:
2929:
2924:
2920:
2916:
2911:
2906:
2902:
2898:
2894:
2890:
2886:
2881:
2877:
2873:
2868:
2863:
2859:
2855:
2851:
2846:
2842:
2838:
2833:
2828:
2824:
2820:
2816:
2812:
2808:
2803:
2799:
2795:
2791:
2787:
2783:
2779:
2774:
2770:
2766:
2762:
2758:
2754:
2750:
2745:
2741:
2737:
2732:
2727:
2723:
2719:
2715:
2710:
2706:
2702:
2698:
2694:
2690:
2686:
2685:Life Sciences
2680:
2676:
2672:
2668:
2664:
2660:
2656:
2652:
2648:
2642:
2638:
2634:
2630:
2626:
2622:
2618:
2612:
2608:
2604:
2600:
2596:
2592:
2588:
2583:
2579:
2575:
2571:
2567:
2563:
2559:
2554:
2553:
2548:
2539:
2535:
2530:
2525:
2521:
2517:
2513:
2506:
2503:
2498:
2494:
2489:
2484:
2480:
2476:
2472:
2468:
2464:
2457:
2454:
2449:
2445:
2440:
2435:
2430:
2425:
2421:
2417:
2413:
2409:
2405:
2398:
2396:
2392:
2387:
2383:
2378:
2373:
2368:
2363:
2359:
2355:
2351:
2344:
2342:
2338:
2333:
2329:
2324:
2319:
2315:
2311:
2307:
2303:
2299:
2292:
2289:
2278:
2274:
2268:
2265:
2260:
2256:
2251:
2246:
2242:
2238:
2234:
2230:
2226:
2219:
2216:
2211:
2207:
2202:
2197:
2193:
2189:
2185:
2181:
2177:
2170:
2168:
2164:
2159:
2155:
2150:
2145:
2141:
2137:
2133:
2129:
2125:
2121:
2117:
2110:
2107:
2102:
2098:
2093:
2088:
2084:
2080:
2076:
2072:
2068:
2061:
2058:
2053:
2049:
2044:
2039:
2035:
2031:
2027:
2023:
2019:
2012:
2010:
2008:
2006:
2002:
1997:
1993:
1988:
1983:
1978:
1973:
1969:
1965:
1961:
1954:
1952:
1950:
1948:
1946:
1944:
1942:
1940:
1938:
1936:
1934:
1932:
1930:
1928:
1926:
1924:
1922:
1920:
1918:
1914:
1909:
1905:
1900:
1895:
1890:
1885:
1881:
1877:
1873:
1869:
1865:
1858:
1855:
1850:
1846:
1841:
1836:
1832:
1828:
1824:
1820:
1816:
1812:
1808:
1801:
1798:
1793:
1789:
1784:
1779:
1775:
1771:
1767:
1763:
1759:
1752:
1750:
1746:
1741:
1737:
1732:
1727:
1723:
1719:
1715:
1711:
1704:
1702:
1700:
1698:
1696:
1694:
1692:
1690:
1688:
1686:
1684:
1682:
1680:
1678:
1676:
1674:
1672:
1668:
1663:
1659:
1653:
1650:
1645:
1641:
1635:
1632:
1628:
1624:
1619:
1617:
1615:
1611:
1607:
1603:
1598:
1596:
1594:
1590:
1583:
1581:
1579:
1578:retinoic acid
1577:
1567:
1565:
1563:
1559:
1555:
1551:
1547:
1543:
1542:breast cancer
1535:
1533:
1531:
1527:
1523:
1519:
1515:
1511:
1502:
1500:
1494:
1489:
1480:
1478:
1475:
1471:
1466:
1462:
1455:
1450:
1445:
1443:
1441:
1437:
1433:
1429:
1425:
1421:
1417:
1413:
1410:
1406:
1403:
1393:
1386:
1384:
1382:
1378:
1373:
1372:Knockout mice
1369:
1364:
1362:
1358:
1354:
1350:
1346:
1343:
1338:
1336:
1335:coronaviruses
1332:
1328:
1324:
1320:
1316:
1312:
1308:
1304:
1301:
1297:
1293:
1281:
1276:
1272:
1268:
1265:
1261:
1254:
1252:
1249:
1245:
1241:
1237:
1234:
1230:
1226:
1219:
1217:
1211:
1207:
1204:
1198:
1196:
1190:
1186:
1183:
1179:
1172:
1170:
1164:
1160:
1157:
1151:
1149:
1143:
1139:
1136:
1134:RefSeq (mRNA)
1132:
1125:
1124:
1119:
1115:
1112:
1106:
1105:
1100:
1096:
1093:
1091:
1087:
1080:
1079:
1074:
1070:
1067:
1061:
1060:
1055:
1051:
1048:
1046:
1042:
1035:
1034:
1029:
1025:
1022:
1016:
1015:
1010:
1006:
1003:
1001:
997:
994:
991:
989:
986:
982:
979:
975:
971:
964:
960:
955:
949:
946:
944:
941:
939:
936:
934:
931:
929:
926:
924:
921:
919:
916:
914:
911:
909:
906:
904:
901:
899:
896:
894:
891:
889:
888:viral process
886:
884:
881:
879:
876:
874:
871:
869:
866:
864:
861:
859:
856:
854:
851:
849:
846:
844:
841:
839:
836:
834:
831:
829:
826:
825:
823:
820:
819:
813:
810:
808:
805:
803:
800:
798:
797:cell junction
795:
793:
790:
788:
785:
783:
780:
778:
775:
773:
770:
768:
765:
764:
762:
759:
758:
752:
749:
747:
744:
742:
739:
737:
734:
732:
729:
727:
724:
722:
719:
717:
714:
712:
709:
707:
704:
702:
699:
697:
694:
692:
689:
687:
684:
683:
681:
678:
677:
674:
673:Gene ontology
670:
666:
654:
649:
645:
640:
637:
635:
631:
623:
618:
607:
603:
599:
595:
591:
587:
584:otolith organ
583:
579:
575:
571:
570:
567:
563:
558:
555:
545:
541:
537:
533:
529:
525:
521:
517:
514:skin of thigh
513:
509:
508:
505:
501:
496:
493:
492:
489:
487:
483:
481:
480:
476:
475:
472:
470:
466:
462:
458:
454:
446:
441:
437:
433:
428:
418:
414:
407:
400:
394:
387:
379:
375:
371:
366:
362:
357:
353:
345:
340:
336:
332:
327:
317:
313:
306:
299:
293:
286:
282:
276:
272:
268:
263:
259:
254:
250:
246:
242:
238:
234:
230:
226:
222:
218:
214:
210:
202:
197:
190:
185:
180:
169:
167:
163:
159:
155:
151:
147:
143:
139:
135:
131:
127:
123:
119:
115:
111:
107:
103:
99:
95:
91:
87:
81:
76:
73:
72:
68:
65:
58:
54:
49:
45:
41:
36:
31:
19:
3342:
3280:
3141:
3137:
3103:
3099:
3062:
3058:
3023:
3019:
2988:
2984:
2935:
2931:
2892:
2888:
2857:
2853:
2814:
2810:
2784:(1): 47–50.
2781:
2777:
2752:
2748:
2721:
2717:
2688:
2684:
2650:
2646:
2620:
2616:
2590:
2586:
2561:
2557:
2519:
2515:
2505:
2470:
2466:
2456:
2411:
2407:
2357:
2353:
2305:
2301:
2291:
2280:. Retrieved
2276:
2267:
2232:
2229:Gut Microbes
2228:
2218:
2186:(1): 36–49.
2183:
2179:
2123:
2119:
2109:
2077:(1): 53–62.
2074:
2070:
2060:
2025:
2021:
1967:
1963:
1871:
1867:
1857:
1814:
1810:
1800:
1765:
1761:
1713:
1661:
1652:
1643:
1634:
1575:
1571:
1539:
1506:
1484:
1459:
1412:RNA helicase
1401:
1399:
1381:autoimmunity
1365:
1345:RNA helicase
1339:
1327:Sendai virus
1295:
1291:
1290:
1213:
1192:
1166:
1145:
1121:
1102:
1076:
1057:
1031:
1012:
992:
987:
807:cytoskeleton
484:
477:
204:External IDs
83:
3331:Cyclophilin
2749:Endothelium
2126:(1): 4841.
1481:RIG-I PAMPs
1323:influenza A
741:ATP binding
721:RNA binding
572:granulocyte
538:granulocyte
443:40,239,828
430:40,203,773
422:4|4 A5
342:32,526,208
329:32,455,302
182:Identifiers
3351:Categories
2282:2020-02-29
1629:, May 2017
1608:, May 2017
1584:References
1420:N-terminus
1416:C-terminus
1409:DExD/H box
1342:DExD/H box
1331:flavivirus
488:(ortholog)
225:HomoloGene
3362:Helicases
3311:Collectin
1558:caspase-3
1554:apoptosis
1446:Functions
1387:Structure
1300:cytosolic
1216:NP_766277
1195:NP_055129
1169:NM_172689
1148:NM_014314
978:Orthologs
767:cytoplasm
233:GeneCards
3166:16177806
3122:16153868
3087:31479259
3079:16127453
3050:11104354
3042:16125763
3007:16116171
2972:16009940
2919:15767399
2876:15737993
2841:15708988
2798:15612946
2769:15370293
2740:15297736
2705:15219805
2675:34876422
2667:15208624
2637:15181474
2607:11890704
2578:17307033
2538:10942397
2497:30976817
2448:20080593
2386:30459758
2360:: 2379.
2332:21782231
2259:34347572
2210:24362405
2158:30451863
2101:29180807
2052:28593618
1996:31379819
1970:: 1586.
1908:26733676
1849:25119032
1792:21084468
1740:25749629
1714:Virology
1625:–
1604:–
1556:via the
1530:JAK-STAT
1422:are two
1396:(5'ppp).
1264:Wikidata
957:Sources:
777:membrane
600:calvaria
526:monocyte
3174:4391603
3146:Bibcode
2963:1177427
2940:Bibcode
2910:1061556
2488:6509067
2439:2824407
2416:Bibcode
2377:6232773
2323:3179916
2250:8344765
2201:4084561
2149:6242832
2128:Bibcode
2092:5815369
2043:5829270
1987:6652118
1899:4725518
1876:Bibcode
1840:4201573
1819:Bibcode
1783:3020501
1731:4424084
1627:Ensembl
1606:Ensembl
1298:) is a
1090:UniProt
1045:Ensembl
984:Species
963:QuickGO
812:cytosol
592:jejunum
588:utricle
463:pattern
221:2442858
189:Aliases
3172:
3164:
3138:Nature
3120:
3085:
3077:
3048:
3040:
3005:
2970:
2960:
2917:
2907:
2874:
2839:
2832:548482
2829:
2796:
2767:
2738:
2703:
2673:
2665:
2635:
2605:
2576:
2536:
2495:
2485:
2446:
2436:
2384:
2374:
2330:
2320:
2257:
2247:
2208:
2198:
2156:
2146:
2099:
2089:
2050:
2040:
1994:
1984:
1906:
1896:
1847:
1837:
1811:Nature
1790:
1780:
1738:
1728:
1488:ligand
1333:, and
1250:search
1248:PubMed
1123:Q6Q899
1104:O95786
1033:230073
1000:Entrez
634:BioGPS
608:zygote
596:spleen
321:9p21.1
213:609631
3281:RIG-I
3170:S2CID
3134:(PDF)
3083:S2CID
3046:S2CID
2671:S2CID
2516:Blood
1576:trans
1526:IFNβ)
1465:virus
1402:DDX58
1377:STING
1292:RIG-I
1014:23586
993:Mouse
988:Human
959:Amigo
530:blood
486:Mouse
479:Human
426:Start
361:Mouse
325:Start
258:Human
237:DDX58
229:32215
196:DDX58
33:DDX58
3291:LGP2
3286:MDA5
3162:PMID
3118:PMID
3075:PMID
3038:PMID
3020:Cell
3003:PMID
2968:PMID
2915:PMID
2872:PMID
2837:PMID
2794:PMID
2765:PMID
2736:PMID
2701:PMID
2663:PMID
2633:PMID
2603:PMID
2574:PMID
2534:PMID
2493:PMID
2444:PMID
2382:PMID
2328:PMID
2302:Cell
2255:PMID
2206:PMID
2154:PMID
2097:PMID
2048:PMID
1992:PMID
1904:PMID
1845:PMID
1788:PMID
1736:PMID
1574:all-
1548:and
1524:and
1522:IFNα
1514:MAVS
1440:MAVS
1436:LGP2
1405:gene
1361:MAVS
469:Bgee
417:Band
378:Chr.
316:Band
275:Chr.
209:OMIM
166:5F9H
162:5F9F
158:5F98
154:5E3H
150:4P4H
146:4ON9
142:4NQK
138:4BPB
134:4AY2
130:3ZD7
126:3ZD6
122:3OG8
118:3NCU
114:3LRR
110:3LRN
106:2YKG
102:2RMJ
98:2QFD
94:2QFB
90:2LWE
86:2LWD
67:RCSB
64:PDBe
3154:doi
3142:437
3108:doi
3067:doi
3028:doi
3024:122
2993:doi
2989:175
2958:PMC
2948:doi
2936:102
2905:PMC
2897:doi
2862:doi
2858:280
2827:PMC
2819:doi
2786:doi
2757:doi
2726:doi
2722:203
2693:doi
2655:doi
2625:doi
2595:doi
2591:292
2566:doi
2524:doi
2483:PMC
2475:doi
2434:PMC
2424:doi
2412:107
2372:PMC
2362:doi
2318:PMC
2310:doi
2306:146
2245:PMC
2237:doi
2196:PMC
2188:doi
2144:PMC
2136:doi
2087:PMC
2079:doi
2038:PMC
2030:doi
1982:PMC
1972:doi
1894:PMC
1884:doi
1872:113
1835:PMC
1827:doi
1815:514
1778:PMC
1770:doi
1726:PMC
1718:doi
439:End
338:End
241:OMA
217:MGI
57:PDB
3353::
3215::
3168:.
3160:.
3152:.
3140:.
3136:.
3116:.
3104:19
3102:.
3098:.
3081:.
3073:.
3061:.
3044:.
3036:.
3022:.
3018:.
3001:.
2987:.
2983:.
2966:.
2956:.
2946:.
2934:.
2930:.
2913:.
2903:.
2893:79
2891:.
2887:.
2870:.
2856:.
2852:.
2835:.
2825:.
2815:79
2813:.
2809:.
2792:.
2782:20
2780:.
2763:.
2753:11
2751:.
2734:.
2720:.
2716:.
2699:.
2689:75
2687:.
2669:.
2661:.
2649:.
2631:.
2621:82
2619:.
2601:.
2589:.
2572:.
2562:28
2560:.
2532:.
2520:96
2518:.
2514:.
2491:.
2481:.
2471:67
2469:.
2465:.
2442:.
2432:.
2422:.
2410:.
2406:.
2394:^
2380:.
2370:.
2356:.
2352:.
2340:^
2326:.
2316:.
2304:.
2300:.
2275:.
2253:.
2243:.
2233:13
2231:.
2227:.
2204:.
2194:.
2184:14
2182:.
2178:.
2166:^
2152:.
2142:.
2134:.
2122:.
2118:.
2095:.
2085:.
2075:19
2073:.
2069:.
2046:.
2036:.
2024:.
2020:.
2004:^
1990:.
1980:.
1968:10
1966:.
1962:.
1916:^
1902:.
1892:.
1882:.
1870:.
1866:.
1843:.
1833:.
1825:.
1813:.
1809:.
1786:.
1776:.
1766:85
1764:.
1760:.
1748:^
1734:.
1724:.
1712:.
1670:^
1660:.
1642:.
1613:^
1592:^
1564:.
1383:.
1337:.
1329:,
1325:,
1321:,
1317:,
961:/
445:bp
432:bp
344:bp
331:bp
239:;
235::
231:;
227::
223:;
219::
215:;
211::
164:,
160:,
156:,
152:,
148:,
144:,
140:,
136:,
132:,
128:,
124:,
120:,
116:,
112:,
108:,
104:,
100:,
96:,
92:,
88:,
3333:)
3329:(
3205:e
3198:t
3191:v
3176:.
3156::
3148::
3124:.
3110::
3089:.
3069::
3063:6
3052:.
3030::
3009:.
2995::
2974:.
2950::
2942::
2921:.
2899::
2878:.
2864::
2843:.
2821::
2800:.
2788::
2771:.
2759::
2742:.
2728::
2707:.
2695::
2677:.
2657::
2651:5
2639:.
2627::
2609:.
2597::
2580:.
2568::
2540:.
2526::
2499:.
2477::
2450:.
2426::
2418::
2388:.
2364::
2358:9
2334:.
2312::
2285:.
2261:.
2239::
2212:.
2190::
2160:.
2138::
2130::
2124:9
2103:.
2081::
2054:.
2032::
2026:9
1998:.
1974::
1910:.
1886::
1878::
1851:.
1829::
1821::
1794:.
1772::
1742:.
1720::
1664:.
1646:.
1497:1
1294:(
363:)
260:)
243::
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.