145:
54:
484:. These cells are truly marine and have been isolated from both the coastal and the open ocean. All strains are obligate photoautrophs and are around 0.6–1.7 μm in diameter. This cluster is, however, further divided into a population that either contains (cluster 5.1) or does not contain (cluster 5.2) phycoerythrin. The reference strains are WH8103 for the phycoerythrin-containing strains and WH5701 for those strains that lack this pigment.
33:
611:, where cell abundances are often very low, population growth rates are often high and not drastically limited. Factors such as grazing, viral mortality, genetic variability, light adaptation, and temperature, as well as nutrients are certainly involved, but remain to be investigated on a rigorous and global scale. Despite the uncertainties, a relationship probably exists between ambient nitrogen concentrations and
686:' viruses co-exist with their host until stresses or nearing end of natural life span make them switch their host to virus production; if a mutation occurs that stops this final step, the host can carry the virus genes with no ill effects. And if a healthy host reproduces while infectious, its offspring can be infectious as well. It is likely such a process gave
417:
ranging from 39 to 71%, illustrating the large genetic diversity of this provisional taxon. Initially, attempts were made to divide the group into three subclusters, each with a specific range of genomic G+C content. The observation that open-ocean isolates alone nearly span the complete G+C
248:
oceans. The genus was first described in 1979, and was originally defined to include "small unicellular cyanobacteria with ovoid to cylindrical cells that reproduce by binary traverse fission in a single plane and lack sheaths". This definition of the genus
319:
is a mutant hybrid from UTEX 625 and is most closely related to S. elongatus PCC 7942 with 99.8% similarity. It has the shortest doubling time at “1.9 hours in a BG11 medium at 41°C under continuous 500 μmoles photons·m−2·s−1 white light with 3% CO2”.
583:
and exhibits an affinity for the higher-light areas. Below the mixed layer, cell concentrations rapidly decline. Vertical profiles are strongly influenced by hydrologic conditions and can be very variable both seasonally and spatially. Overall,
449:, respectively. Cluster 2 also is characterized by low salt tolerance. Cells are obligate photoautrotrophs, lack phycoerythrin, and are thermophilic. The reference strain PCC6715 was isolated from a hot spring in
681:
samples have been found to have viral proteins associated with photosynthesis. It is estimated 10% of all photosynthesis on earth is carried out with viral genes. Not all viruses immediately kill their hosts,
1157:
Palenik, B.; Brahamsha, B.; Larimer, F.W.; Land, M.; Hauser, L.; Chain, P.; Lamerdin, J.; Regala, W.; Allen, E.E.; McCarren, J.; Paulsen, I.; Dufresne, A.; Partensky, F.; Webb, E.A.; Waterbury, J. (2003).
2598:
L. Campbell; Liu, H.; Nolla, H.A.; Vaulot, D. (1997). "Annual variability of phytoplankton and bacteria in the subtropical North
Pacific Ocean at Station ALOHA during the 1991-1994 ENSO event".
600:
zone and in temperate open seas where stratification was recently established both profiles parallel each other and exhibit abundance maxima just about the subsurface chlorophyll maximum.
2666:
Partensky, F.; Blanchot, J.; Lantoine, F.; Neveux, J.; Marie, D. (1996). "Vertical
Structure of Picophytoplankton at Different Trophic Sites of the Tropical Northeastern Atlantic Ocean".
499:) found in these organisms. α-cyanobacteria were defined to contain a form IA, while β-cyanobacteria were defined to contain a form IB of this gene. In support for this division Badger
3019:
437:
Cluster 1 includes relatively large (1–1.5 μm) nonmotile obligate photoautotrophs that exhibit low salt tolerance. Reference strains for this cluster are PCC6301 (formerly
2302:
Landry, M.R.; Kirshtein, J.; Constantinou, J. (1996). "Abundances and distributions of picoplankton populations in the central equatorial
Pacific from 12°N to 12°S, 140°W".
2265:
Blanchot, J.; Rodier, M. (1996). "Picophytoplankton abundance and biomass in the western tropical
Pacific Ocean during the 1992 El Nino year: results from flow cytometry".
1869:
Waterbury, J.B.; Stanier, R.Y. (1981). "Isolation and growth of cyanobacteria from marine and hypersaline environments". In Starr; Stulp; Truper; Balows; Schleeper (eds.).
1614:
Yu, Jingjie; Liberton, Michelle; Cliften, Paul F.; Head, Richard D.; Jacobs, Jon M.; Smith, Richard D.; Koppenaal, David W.; Brand, Jerry J.; Pakrasi, Himadri B. (2015).
461:, i.e. capable of growth in both marine and freshwater environments. Several strains, including the reference strain PCC7003, are facultative heterotrophs and require
2993:
265:
cells with highly structured cell walls that may contain projections on their surface. Electron microscopy frequently reveals the presence of phosphate inclusions,
553:. Cells are generally much more abundant in nutrient-rich environments than in the oligotrophic ocean and prefer the upper, well-lit portion of the euphotic zone.
3045:
468:
for growth. Cluster 4 contains a single isolate, PCC7335. This strain is obligate marine. This strain contains phycoerthrin and was first isolated from the
144:
2722:
2806:
Waterbury, J.B.; Watson, S.W.; Valois, F.W.; Franks, D.G. (1986). "Biological and ecological characterization of the marine unicellular cyanobacterium
2228:
Campbell, L.; Vaulot, D. (1993). "Photosynthetic picoplankton community structure in the stubtropical North
Pacific Ocean new Hawaii (station ALOHA)".
1573:
Waterbury, J.B.; Watson, S.W.; Valois, F.W.; Franks, D.G. (1986). "Biological and ecological characterization of the marine unicellular cyanobacterium
2967:
2191:"Effect of El Niño Southern Oscillation events on the distribution and abundance of phytoplankton in the Western Pacific Tropical Ocean along 165°E"
1354:
Waterbury, J.B.; Watson, S.W.; Guillard, R.R.L.; Brand, L.E. (1979). "Wide-spread occurrence of a unicellular, marine planktonic, cyanobacterium".
3006:
296:
strains appear to be obligate photoautotrophs that are capable of supporting their nitrogen requirements using nitrate, ammonia, or in some cases
557:
has also been observed to occur at high abundances in environments with low salinities and/or low temperatures. It is usually far outnumbered by
2514:
1915:
1882:
1489:
253:
contained organisms of considerable genetic diversity and was later subdivided into subgroups based on the presence of the accessory pigment
1579:
2391:"Preferential uptake of ammonium in the presence of elevated nitrate concentrations by phytoplankton in the offshore Mississippi Plume"
2819:
2063:
3081:
1513:
Waterbury, J.B.; Willey, J.M.; Franks, D.G.; Valois, F.W.; Watson, S.W. (1985). "A cyanobacterium capable of swimming motility".
1445:
757:
537:
has been observed to occur at concentrations ranging between a few cells to 10 cells per ml in virtually all regions of oceanic
522:, but they have not derived from the common ancestor. Moreover, it has been estimated based on molecular dating that the first
3119:
2468:
2352:
2099:
597:
526:
lineage has appeared 3 billion years ago in thermal springs with subsequent radiation to marine and freshwater environments.
2699:"Recent advances in the use of molecular techniques to assess the genetic diversity of marine photosynthetic microorganisms"
3109:
3032:
1740:
3011:
2763:"Photoacclimation of Prochlorococcus sp.(Prochlorophyta) strains isolated from the North Atlantic and Mediterranean Sea"
575:
is apparently always present, although only at low concentrations, ranging from a few to 4×10³ cells per ml. Vertically
53:
1122:
563:
in all environments where they co-occur. Exceptions to this rule are areas of permanently enriched nutrients such as
975:
2767:
2635:
2149:
2058:. Bulletin de l'Institut Océanographique Monaco. Vol. NS 19. Monaco: Musée océanographique. pp. 457–475.
1316:
985:
661:
accounts for at least 22 times more carbon in these waters, thus may be of much greater significance to the global
3050:
2430:
2395:
2304:
1116:
875:
737:
450:
1976:
Dvořák, Petr; Casamatta, Dale A.; Poulíčková, Aloisie; Hašler, Petr; Ondřej, Vladan; Sanges, Remo (2014-11-01).
1088:
1068:
965:
925:
855:
795:
3114:
2668:
2600:
2267:
2230:
1941:
1058:
885:
865:
835:
703:
683:
2459:
Wawrik, B.; Paul, J.H.; Campbell, L.; Griffin, D.; Houchin, L.; Fuentes-Ortega, A.; Müller-Karger, F. (2003).
1035:
1025:
1015:
805:
657:
in warm oligotrophic waters. Assuming average cellular carbon concentrations, it has thus been estimated that
607:
still remain poorly understood, especially considering that even in the most nutrient-depleted regions of the
1078:
1048:
995:
895:
825:
815:
1781:
1223:
1098:
945:
935:
845:
781:
771:
503:
analyze the phylogeny of carboxysomal proteins, which appear to support this division. Also, two particular
1005:
955:
915:
905:
504:
2868:
1410:
636:
628:
2461:"Vertical structure of the phytoplankton community associated with a coastal plume in the Gulf of Mexico"
1311:
492:
568:
353:
2545:"Comparative Genomics of DNA Recombination and Repair in Cyanobacteria: Biotechnological Implications"
280:
by a gliding method and a novel uncharacterized, nonphototactic swimming method that does not involve
3086:
3060:
2941:
2677:
2644:
2609:
2477:
2361:
2313:
2276:
2239:
2158:
2108:
1993:
1827:
1775:
Rippka, R.; Cohen-Bazire, G. (1983). "The
Cyanobacteriales: a legitimate order based on type strains
1696:
1631:
1524:
1365:
1325:
1177:
410:
491:(2002) proposed the division of the cyanobacteria into a α- and a β-subcluster based on the type of
311:
have been produced in laboratory environments to include the fastest growing cyanobacteria to date,
2703:
2198:
632:
1286:
2906:
2190:
1851:
1720:
1548:
1381:
507:
systems appear to only be found in α-cyanobacteria, which lack carboxysomal carbonic anhydrases.
480:. The last cluster contains what had previously been referred to as ‘marine A and B clusters’ of
289:
48:
2920:
1898:
Waterbury, J.B.; Willey, J.M (1988). "Isolation and growth of marine planktonic cyanobacteria".
1314:(1979). "Chroococcoid cyanobacteria in the sea: a ubiquitous and diverse phototrophic biomass".
2845:
434:
into five clusters (equivalent to genera) based on morphology, physiology, and genetic traits.
3068:
2998:
2928:
2825:
2815:
2794:
2749:
2576:
2520:
2510:
2069:
2059:
2019:
2011:
1958:
1911:
1878:
1843:
1818:
1798:
1757:
1712:
1665:
1647:
1596:
1588:
1540:
1515:
1495:
1485:
1462:
1250:
1195:
511:
473:
2631:"The importance of Prochlorococcus to community structure in the central North Pacific Ocean"
2460:
3073:
2784:
2776:
2739:
2731:
2685:
2652:
2617:
2566:
2556:
2485:
2439:
2426:"Phytoplankton community structure and productivity along the axis of the Mississippi Plume"
2404:
2369:
2321:
2284:
2247:
2207:
2166:
2116:
2001:
1950:
1903:
1835:
1816:
Dyer, D.L.; Gafford, R.D. (1961). "Some characteristics of a thermophilic blue-green alga".
1790:
1749:
1704:
1655:
1639:
1532:
1454:
1419:
1373:
1356:
1333:
1240:
1232:
1185:
1168:
729:
337:
285:
105:
32:
2698:
1874:
1439:
Perkins, F.O.; Haas, L.W.; Phillips, D.E.; Webb, K.L. (1981). "Ultrastructure of a marine
1128:
711:
559:
469:
383:
370:
349:
178:
95:
2340:
2091:
1871:
The prokaryotes: a handbook on habitats, isolation, and identification of bacteria, Vol 1
1406:"Generic assignments, strains histories and properties of pure cultures of cyanobacteria"
2681:
2648:
2613:
2481:
2365:
2317:
2280:
2243:
2162:
2112:
1997:
1831:
1753:
1700:
1635:
1528:
1369:
1329:
1181:
631:
are coastally enriched with nutrients such as nitrate and phosphate, which drives large
2571:
2544:
1932:
1738:
Stanier, R.Y.; Cohen-Bazire, G. (1977). "Phototrophic prokaryotes: the cyanobacteria".
1660:
1615:
546:
186:
174:
122:
2789:
2762:
2621:
1794:
1620:
UTEX 2973, a fast growing cyanobacterial chassis for biosynthesis using light and CO2"
1245:
1214:
3103:
2744:
2717:
2689:
2325:
2288:
2251:
1907:
1236:
1135:
620:
542:
538:
402:
357:
333:
262:
254:
193:
85:
75:
2345:
sequences in ambient phytoplankton populations from the southeastern Gulf of Mexico"
2046:
Partensky F, Blanchot J, Vaulot D (1999). "Differential distribution and ecology of
1855:
1724:
1552:
1480:
Castenholz, R.W. (1982). "Motility and taxes". In N. G. Carr; B. A. Whitton (eds.).
2735:
1385:
749:
662:
593:
237:
2933:
368:
but no diagnostic pigment for this organism is known. Zeaxanthin is also found in
1839:
740:, and thus can take up extracellular DNA and recombine it into their own genome.
3037:
2980:
2900:
1536:
1268:
721:
580:
462:
391:
379:
345:
270:
197:
153:
1404:
Rippka, R.; Deruelles, J.; Waterbury, J.B.; Herdman, M.; Stanier, R.Y. (1979).
2657:
2630:
2211:
2171:
2140:
1708:
1337:
707:
608:
550:
458:
446:
414:
365:
341:
231:
196:
where it can be very abundant (generally 1,000 to 200,000 cells per ml). Many
182:
2891:
2561:
2524:
2073:
2015:
1716:
1651:
1600:
1592:
1424:
1405:
2954:
2915:
2854:
2829:
2718:"Prochlorococcus, a Marine Photosynthetic Prokaryote of Global Significance"
564:
387:
375:
281:
241:
234:
216:
3024:
2798:
2753:
2580:
2023:
1962:
1847:
1669:
1544:
1499:
1199:
571:. In the nutrient-depleted areas of the oceans, such as the central gyres,
1802:
1466:
1254:
2885:
2780:
1761:
677:
Free floating viruses have been found carrying photosynthetic genes, and
277:
266:
65:
2761:
Partensky, F.; Hoepffner, N.; Li, W.K.W.; Ulloa, O.; Vaulot, D. (1993).
1190:
1159:
148:
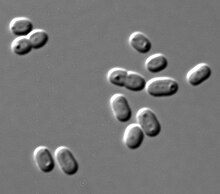
Transmission electron micrograph showing a species of the cyanobacteria
2972:
2814:. Ottawa, Canada: Department of Fisheries and Oceans. pp. 71–120.
2490:
2374:
2121:
496:
245:
219:, whereas the oceanic strain WH8102 has a genome of size 2.4 Mbp.
2985:
2444:
2425:
2409:
2390:
2006:
1977:
1643:
1377:
716:
477:
208:
189:
2862:
1954:
1458:
2959:
2139:
Olson, R.J.; Chisholm, S.W.; Zettler, E.R.; Armbrust, E.V. (1990).
1683:
Racharaks, Ratanachat; Peccia, Jordan (2019). "Cryopreservation of
627:
thrives particularly well is coastal plumes of major rivers. Such
442:
2092:"Composition of ultraphytoplankton in the central North Atlantic"
518:
revealed at least 12 groups, which morphologically correspond to
2389:
Wawrik, B.; John, D.; Gray, M.; Bronk, D.A.; Paul, J.H. (2004).
297:
261:
are coccoid cells between 0.6 and 1.6 μm in size. They are
2866:
2946:
2054:
in oceanic waters: a review". In Charpy L, Larkum AWD (eds.).
579:
is usually relatively equitably distributed throughout the
2697:
Partensky, F.; Guillou, L.; Simon, N.; Vaulot, D. (1997).
1287:"Picosynechococcus sp. PCC 7002 genome assembly ASM1948v1"
2509:(2nd ed.). University of Chicago Press. p. 45.
1902:. Methods in Enzymology. Vol. 167. pp. 100–5.
1215:"Physical genome map of the unicellular cyanobacterium
724:
V, functions in recombinational repair of DNA, and the
653:
is thought to be at least 100 times more abundant than
441:) and PCC6312, which were isolated from fresh water in
304:
species are traditionally not thought to fix nitrogen.
2538:
2536:
2534:
2543:
Cassier-Chauvat, C.; Veaudor, T.; Chauvat, F (2016).
623:, where light is not limiting. One environment where
1484:. University of California Press. pp. 413–439.
702:
species possess a set of genes that function in DNA
2875:
2189:Blanchot, J.; Rodier, M.; LeBouteiller, A. (1992).
1580:
Canadian
Bulletin of Fisheries and Aquatic Sciences
752:system, that is likely similar to the well-studied
453:. Cluster 3 includes phycoerythrin-lacking marine
269:granules, and more importantly, highly structured
732:, functions in error-prone DNA replication. Some
756:SOS system that is employed in the response to
497:ribulose 1,5-bisphosphate carboxylase/oxygenase
230:is one of the most important components of the
2339:Paul, J.H.; Wawrik, B.; Alfreider, A. (2000).
1931:Badger, M.R.; Hanson, D.; Price, G.D. (2002).
639:is often associated with large populations of
2716:F. Partensky; Hess, W.R.; Vaulot, D. (1999).
2629:L. Campbell; Nolla, H.A.; Vaulot, D. (1994).
8:
386:. Similarly, phycoerythrin is also found in
615:abundance, with an inverse relationship to
2863:
2723:Microbiology and Molecular Biology Reviews
284:. While some cyanobacteria are capable of
31:
20:
2788:
2743:
2656:
2570:
2560:
2489:
2443:
2408:
2373:
2170:
2145:in the North Atlantic and Pacific oceans"
2134:
2132:
2120:
2041:
2039:
2037:
2035:
2033:
2005:
1937:concentrating mechanism in cyanobacteria"
1659:
1568:
1566:
1564:
1562:
1423:
1244:
1189:
694:DNA recombination, repair and replication
603:The factors controlling the abundance of
422:is composed of at least several species.
336:, while its major accessory pigments are
2223:
2221:
2184:
2182:
143:
2844:Komárek, J.; Guiry, M.D. (2006-07-17).
2085:
2083:
1399:
1397:
1395:
1349:
1347:
1149:
1139:, another cyanobacterial model organism
307:In the last decade, several strains of
181:. Its size varies from 0.8 to 1.5
1982:: 3 billion years of global dominance"
643:and elevated form IA (cyanobacterial)
635:. High productivity in coastal river
192:are preferentially found in well–lit
156:appear as polyhedral dark structures.
7:
3061:9be56943-0e80-4925-b56f-7a5889c59f15
736:strains are naturally competent for
1754:10.1146/annurev.mi.31.100177.001301
830:A. E. Bailey-Watts & J. Komárek
328:The main photosynthetic pigment in
2141:"Pigment size and distribution of
714:. This set of genes includes the
588:abundance often parallels that of
418:spectrum, however, indicates that
300:as a sole nitrogen source. Marine
215:strain PCC7002 has a size of 3.4 M
14:
43:PCC 7002 cells in DIC microscopy
1446:Canadian Journal of Microbiology
1237:10.1128/jb.175.16.5106-5116.1993
52:
2424:Wawrik, B.; Paul, J.H. (2004).
1213:Chen, X.; Widger, W.R. (1993).
1160:"The genome of a motile marine
378:and as a minor pigment in some
340:. The four commonly recognized
177:that is very widespread in the
169:, in succession, and the Greek
2736:10.1128/MMBR.63.1.106-127.1999
2469:Marine Ecology Progress Series
2353:Marine Ecology Progress Series
2341:"Micro- and macrodiversity in
2100:Marine Ecology Progress Series
1933:"Evolution and diversity of CO
1273:sp. PCC 7002, complete genome"
976:Synechococcus roseo-persicinus
598:high-nutrient, low-chlorophyll
1:
2622:10.1016/S0967-0637(96)00102-1
1795:10.1016/s0769-2609(83)80094-5
1741:Annual Review of Microbiology
986:Synechococcus roseo-purpureus
744:strains also encode the gene
2690:10.1016/0967-0637(96)00056-8
2326:10.1016/0967-0645(96)00018-5
2289:10.1016/0967-0637(96)00026-X
2252:10.1016/0967-0637(93)90044-4
1908:10.1016/0076-6879(88)67009-1
1840:10.1126/science.134.3479.616
1689:Journal of Applied Phycology
1482:The biology of cyanobacteria
1030:(Moore & Carter) Komárek
728:gene complex whose product,
720:gene complex whose product,
413:very similar, yet exhibit a
173:, granule) is a unicellular
2812:Photosynthetic Picoplankton
1537:10.1126/science.230.4721.74
1123:Photosynthetic picoplankton
876:Synechococcus ferrunginosus
541:except in samples from the
409:is difficult. Isolates are
3138:
2636:Limnology and Oceanography
2150:Limnology and Oceanography
1877:, Berlin. pp. 221–3.
1317:Limnology and Oceanography
1089:Synechococcus viridissimus
1069:Synechococcus vantieghemii
970:Komárek & Anagnostidis
966:Synechococcus rhodobaktron
926:Synechococcus minutissimus
856:Synechococcus endogloeicus
796:Synechococcus bigranulatus
204:have also been described.
2658:10.4319/lo.1994.39.4.0954
2431:Aquatic Microbial Ecology
2396:Aquatic Microbial Ecology
2305:Deep-Sea Research Part II
2172:10.4319/lo.1990.35.1.0045
1709:10.1007/s10811-018-1714-9
1338:10.4319/lo.1979.24.5.0928
1117:Gloeomargarita lithophora
1059:Synechococcus sulphuricus
886:Synechococcus intermedius
866:Synechococcus epigloeicus
836:Synechococcus carcerarius
514:of 16S rRNA sequences of
451:Yellowstone National Park
135:
130:
49:Scientific classification
47:
39:
30:
23:
2669:Deep-Sea Research Part I
2601:Deep-Sea Research Part I
2562:10.3389/fmicb.2016.01809
2268:Deep-Sea Research Part I
2231:Deep-Sea Research Part I
1942:Functional Plant Biology
1425:10.1099/00221287-111-1-1
1036:Synechococcus spongiarum
1026:Synechococcus sigmoideus
1016:Synechococcus sciophilus
806:Synechococcus brunneolus
530:Ecology and distribution
2810:". In W.K.W. Li (ed.).
2212:10.1093/plankt/14.1.137
1777:Cyanobacterium stanieri
1685:Synechococcus elongatus
1618:Synechococcus elongatus
1275:. 2013-12-11. CP000951.
1224:Journal of Bacteriology
1079:Synechococcus violaceus
1049:Synechococcus subsalsus
996:Synechococcus salinarum
896:Synechococcus koidzumii
846:Synechococcus elongatus
826:Synechococcus capitatus
816:Synechococcus caldarius
313:Synechococcus elongatus
309:Synechococcus elongatus
1782:Annals of Microbiology
1099:Synechococcus vulcanus
1073:(Pringsheim) Bourrelly
946:Synechococcus nidulans
936:Synechococcus mundulus
782:Synechococcus arcuatus
772:Synechococcus ambiguus
738:genetic transformation
487:More recently, Badger
317:S. elongatus UTEX 2973
276:Cells are known to be
257:. The marine forms of
157:
3120:Marine microorganisms
2505:Zimmer, Carl (2015).
1006:Synechococcus salinus
956:Synechococcus rayssae
916:Synechococcus marinus
906:Synechococcus lividus
505:bicarbonate transport
147:
3110:Cyanobacteria genera
2781:10.1104/pp.101.1.285
2056:Marine cyanobacteria
1443:possessing spinae".
1219:sp. strain PCC 7002"
950:(Pringsheim) Komárek
673:Evolutionary history
633:phytoplankton blooms
2682:1996DSRI...43.1191P
2649:1994LimOc..39..954C
2614:1997DSRI...44..167C
2482:2003MEPS..251...87W
2366:2000MEPS..198....9P
2318:1996DSRII..43..871L
2281:1996DSRI...43..877B
2244:1993DSRI...40.2043C
2163:1990LimOc..35...45O
2113:1995MEPS..122....1L
2090:Li, W.K.W. (1995).
1998:2014MolEc..23.5538D
1832:1961Sci...134..616D
1701:2019JAPco..31.2267R
1636:2015NatSR...5E8132Y
1529:1985Sci...230...74W
1370:1979Natur.277..293W
1330:1979LimOc..24..928J
1191:10.1038/nature01943
1182:2003Natur.424.1037P
439:Anacycstis nidulans
292:growth, all marine
2491:10.3354/meps251087
2375:10.3354/meps198009
2122:10.3354/meps122001
1624:Scientific Reports
748:that regulates an
569:coastal watersheds
495:(large subunit of
430:2001) now divides
290:chemoheterotrophic
286:photoheterotrophic
179:marine environment
158:
3097:
3096:
3069:Open Tree of Life
2869:Taxon identifiers
2516:978-0-226-32026-7
2507:Planet of Viruses
2445:10.3354/ame035185
2410:10.3354/ame035185
2007:10.1111/mec.12948
1986:Molecular Ecology
1917:978-0-12-182068-8
1884:978-0-387-08871-6
1644:10.1038/srep08132
1491:978-0-520-04717-4
1176:(6952): 1037–42.
1104:
1094:
1084:
1074:
1064:
1054:
1044:
1031:
1021:
1011:
1001:
991:
981:
971:
961:
951:
941:
931:
921:
911:
901:
891:
881:
871:
861:
851:
841:
831:
821:
811:
801:
791:
777:
596:. In the Pacific
512:phylogenetic tree
457:species that are
354:allophycocyanin B
142:
141:
126:
16:Genus of bacteria
3127:
3090:
3089:
3077:
3076:
3064:
3063:
3054:
3053:
3041:
3040:
3038:NHMSYS0000607055
3028:
3027:
3015:
3014:
3002:
3001:
2989:
2988:
2976:
2975:
2963:
2962:
2950:
2949:
2937:
2936:
2924:
2923:
2911:
2910:
2909:
2896:
2895:
2894:
2864:
2859:
2850:Nägeli 1849: 56"
2833:
2802:
2792:
2768:Plant Physiology
2757:
2747:
2712:
2693:
2676:(8): 1191–1213.
2662:
2660:
2625:
2585:
2584:
2574:
2564:
2540:
2529:
2528:
2502:
2496:
2495:
2493:
2465:
2456:
2450:
2449:
2447:
2421:
2415:
2414:
2412:
2386:
2380:
2379:
2377:
2349:
2336:
2330:
2329:
2312:(4–6): 871–890.
2299:
2293:
2292:
2262:
2256:
2255:
2225:
2216:
2215:
2195:
2186:
2177:
2176:
2174:
2136:
2127:
2126:
2124:
2096:
2087:
2078:
2077:
2043:
2028:
2027:
2009:
1973:
1967:
1966:
1928:
1922:
1921:
1895:
1889:
1888:
1866:
1860:
1859:
1813:
1807:
1806:
1772:
1766:
1765:
1735:
1729:
1728:
1680:
1674:
1673:
1663:
1611:
1605:
1604:
1570:
1557:
1556:
1510:
1504:
1503:
1477:
1471:
1470:
1436:
1430:
1429:
1427:
1401:
1390:
1389:
1378:10.1038/277293a0
1351:
1342:
1341:
1307:
1301:
1300:
1298:
1297:
1283:
1277:
1276:
1265:
1259:
1258:
1248:
1210:
1204:
1203:
1193:
1154:
1102:
1092:
1082:
1072:
1062:
1052:
1039:
1029:
1019:
1009:
999:
989:
979:
969:
959:
949:
939:
929:
919:
909:
899:
889:
879:
869:
859:
849:
839:
829:
819:
809:
799:
789:
775:
730:DNA polymerase V
690:photosynthesis.
384:eustigmatophytes
338:phycobiliprotein
282:flagellar motion
165:(from the Greek
121:
106:Synechococcaceae
57:
56:
35:
21:
3137:
3136:
3130:
3129:
3128:
3126:
3125:
3124:
3115:Synechococcales
3100:
3099:
3098:
3093:
3085:
3080:
3072:
3067:
3059:
3057:
3049:
3044:
3036:
3031:
3023:
3018:
3010:
3005:
2997:
2992:
2984:
2979:
2971:
2966:
2958:
2953:
2945:
2940:
2932:
2927:
2919:
2914:
2905:
2904:
2899:
2890:
2889:
2884:
2871:
2843:
2840:
2822:
2805:
2760:
2715:
2696:
2665:
2628:
2597:
2594:
2592:Further reading
2589:
2588:
2549:Front Microbiol
2542:
2541:
2532:
2517:
2504:
2503:
2499:
2463:
2458:
2457:
2453:
2423:
2422:
2418:
2388:
2387:
2383:
2347:
2338:
2337:
2333:
2301:
2300:
2296:
2264:
2263:
2259:
2238:(10): 2043–60.
2227:
2226:
2219:
2194:(abstract page)
2193:
2188:
2187:
2180:
2138:
2137:
2130:
2094:
2089:
2088:
2081:
2066:
2048:Prochlorococcus
2045:
2044:
2031:
1992:(22): 5538–51.
1975:
1974:
1970:
1955:10.1071/PP01213
1936:
1930:
1929:
1925:
1918:
1897:
1896:
1892:
1885:
1875:Springer-Verlag
1868:
1867:
1863:
1826:(3479): 616–7.
1815:
1814:
1810:
1774:
1773:
1769:
1737:
1736:
1732:
1682:
1681:
1677:
1613:
1612:
1608:
1572:
1571:
1560:
1523:(4721): 74–76.
1512:
1511:
1507:
1492:
1479:
1478:
1474:
1459:10.1139/m81-049
1438:
1437:
1433:
1403:
1402:
1393:
1364:(5694): 293–4.
1353:
1352:
1345:
1310:Johnson, P.W.;
1309:
1308:
1304:
1295:
1293:
1285:
1284:
1280:
1267:
1266:
1262:
1231:(16): 5106–16.
1212:
1211:
1207:
1156:
1155:
1151:
1146:
1129:Prochlorococcus
1112:
1107:
850:(Nägeli) Nägeli
766:
696:
675:
659:Prochlorococcus
651:Prochlorococcus
617:Prochlorococcus
590:Prochlorococcus
560:Prochlorococcus
532:
470:intertidal zone
466:
424:Bergey's Manual
411:morphologically
405:description of
400:
371:Prochlorococcus
350:allophycocyanin
326:
225:
120:
96:Synechococcales
51:
17:
12:
11:
5:
3135:
3134:
3131:
3123:
3122:
3117:
3112:
3102:
3101:
3095:
3094:
3092:
3091:
3078:
3065:
3055:
3042:
3029:
3016:
3003:
2990:
2977:
2964:
2951:
2938:
2925:
2912:
2897:
2881:
2879:
2873:
2872:
2867:
2861:
2860:
2839:
2838:External links
2836:
2835:
2834:
2820:
2803:
2758:
2730:(1): 106–127.
2713:
2694:
2663:
2643:(4): 954–961.
2626:
2608:(2): 167–192.
2593:
2590:
2587:
2586:
2530:
2515:
2497:
2451:
2416:
2381:
2331:
2294:
2275:(6): 877–895.
2257:
2217:
2206:(1): 137–156.
2178:
2128:
2079:
2064:
2029:
1968:
1949:(3): 161–175.
1934:
1923:
1916:
1890:
1883:
1861:
1808:
1767:
1730:
1695:(4): 2267–76.
1675:
1606:
1558:
1505:
1490:
1472:
1453:(3): 318–329.
1431:
1391:
1343:
1324:(5): 928–935.
1312:Sieburth, J.M.
1302:
1278:
1260:
1205:
1148:
1147:
1145:
1142:
1141:
1140:
1132:
1125:
1120:
1111:
1108:
1106:
1105:
1095:
1085:
1075:
1065:
1055:
1045:
1032:
1022:
1012:
1002:
992:
982:
972:
962:
952:
942:
932:
922:
912:
902:
892:
882:
872:
862:
852:
842:
832:
822:
812:
802:
792:
778:
767:
765:
762:
695:
692:
674:
671:
547:Ross Ice Shelf
531:
528:
474:Puerto Peñasco
464:
399:
396:
364:also contains
360:. In addition
325:
322:
224:
221:
194:surface waters
187:photosynthetic
175:cyanobacterium
140:
139:
133:
132:
128:
127:
113:
109:
108:
103:
99:
98:
93:
89:
88:
83:
79:
78:
73:
69:
68:
63:
59:
58:
45:
44:
37:
36:
28:
27:
15:
13:
10:
9:
6:
4:
3:
2:
3133:
3132:
3121:
3118:
3116:
3113:
3111:
3108:
3107:
3105:
3088:
3083:
3079:
3075:
3070:
3066:
3062:
3056:
3052:
3047:
3043:
3039:
3034:
3030:
3026:
3025:synechococcus
3021:
3017:
3013:
3008:
3004:
3000:
2995:
2991:
2987:
2982:
2978:
2974:
2969:
2965:
2961:
2956:
2952:
2948:
2943:
2939:
2935:
2930:
2926:
2922:
2917:
2913:
2908:
2907:Synechococcus
2902:
2898:
2893:
2887:
2883:
2882:
2880:
2878:
2877:Synechococcus
2874:
2870:
2865:
2857:
2856:
2851:
2849:
2848:Synechococcus
2842:
2841:
2837:
2831:
2827:
2823:
2821:0-660-12243-X
2817:
2813:
2809:
2808:Synechococcus
2804:
2800:
2796:
2791:
2786:
2782:
2778:
2774:
2770:
2769:
2764:
2759:
2755:
2751:
2746:
2741:
2737:
2733:
2729:
2725:
2724:
2719:
2714:
2710:
2706:
2705:
2704:Vie et Milieu
2700:
2695:
2691:
2687:
2683:
2679:
2675:
2671:
2670:
2664:
2659:
2654:
2650:
2646:
2642:
2638:
2637:
2632:
2627:
2623:
2619:
2615:
2611:
2607:
2603:
2602:
2596:
2595:
2591:
2582:
2578:
2573:
2568:
2563:
2558:
2554:
2550:
2546:
2539:
2537:
2535:
2531:
2526:
2522:
2518:
2512:
2508:
2501:
2498:
2492:
2487:
2483:
2479:
2475:
2471:
2470:
2462:
2455:
2452:
2446:
2441:
2437:
2433:
2432:
2427:
2420:
2417:
2411:
2406:
2402:
2398:
2397:
2392:
2385:
2382:
2376:
2371:
2367:
2363:
2359:
2355:
2354:
2346:
2344:
2335:
2332:
2327:
2323:
2319:
2315:
2311:
2307:
2306:
2298:
2295:
2290:
2286:
2282:
2278:
2274:
2270:
2269:
2261:
2258:
2253:
2249:
2245:
2241:
2237:
2233:
2232:
2224:
2222:
2218:
2213:
2209:
2205:
2201:
2200:
2199:J. Plank. Res
2192:
2185:
2183:
2179:
2173:
2168:
2164:
2160:
2156:
2152:
2151:
2146:
2144:
2143:Synechococcus
2135:
2133:
2129:
2123:
2118:
2114:
2110:
2106:
2102:
2101:
2093:
2086:
2084:
2080:
2075:
2071:
2067:
2065:2-7260-0210-2
2061:
2057:
2053:
2052:Synechococcus
2049:
2042:
2040:
2038:
2036:
2034:
2030:
2025:
2021:
2017:
2013:
2008:
2003:
1999:
1995:
1991:
1987:
1983:
1981:
1980:Synechococcus
1972:
1969:
1964:
1960:
1956:
1952:
1948:
1944:
1943:
1938:
1927:
1924:
1919:
1913:
1909:
1905:
1901:
1900:Cyanobacteria
1894:
1891:
1886:
1880:
1876:
1872:
1865:
1862:
1857:
1853:
1849:
1845:
1841:
1837:
1833:
1829:
1825:
1821:
1820:
1812:
1809:
1804:
1800:
1796:
1792:
1788:
1784:
1783:
1778:
1771:
1768:
1763:
1759:
1755:
1751:
1747:
1743:
1742:
1734:
1731:
1726:
1722:
1718:
1714:
1710:
1706:
1702:
1698:
1694:
1690:
1686:
1679:
1676:
1671:
1667:
1662:
1657:
1653:
1649:
1645:
1641:
1637:
1633:
1629:
1625:
1621:
1619:
1610:
1607:
1602:
1598:
1594:
1590:
1586:
1582:
1581:
1576:
1575:Synechococcus
1569:
1567:
1565:
1563:
1559:
1554:
1550:
1546:
1542:
1538:
1534:
1530:
1526:
1522:
1518:
1517:
1509:
1506:
1501:
1497:
1493:
1487:
1483:
1476:
1473:
1468:
1464:
1460:
1456:
1452:
1448:
1447:
1442:
1441:Synechococcus
1435:
1432:
1426:
1421:
1417:
1413:
1412:
1407:
1400:
1398:
1396:
1392:
1387:
1383:
1379:
1375:
1371:
1367:
1363:
1359:
1358:
1350:
1348:
1344:
1339:
1335:
1331:
1327:
1323:
1319:
1318:
1313:
1306:
1303:
1292:
1288:
1282:
1279:
1274:
1272:
1271:Synechococcus
1264:
1261:
1256:
1252:
1247:
1242:
1238:
1234:
1230:
1226:
1225:
1220:
1218:
1217:Synechococcus
1209:
1206:
1201:
1197:
1192:
1187:
1183:
1179:
1175:
1171:
1170:
1165:
1163:
1162:Synechococcus
1153:
1150:
1143:
1138:
1137:
1136:Synechocystis
1133:
1131:
1130:
1126:
1124:
1121:
1119:
1118:
1114:
1113:
1109:
1101:
1100:
1096:
1091:
1090:
1086:
1081:
1080:
1076:
1071:
1070:
1066:
1061:
1060:
1056:
1051:
1050:
1046:
1043:
1038:
1037:
1033:
1028:
1027:
1023:
1018:
1017:
1013:
1008:
1007:
1003:
998:
997:
993:
988:
987:
983:
978:
977:
973:
968:
967:
963:
958:
957:
953:
948:
947:
943:
938:
937:
933:
928:
927:
923:
918:
917:
913:
908:
907:
903:
898:
897:
893:
888:
887:
883:
878:
877:
873:
868:
867:
863:
858:
857:
853:
848:
847:
843:
838:
837:
833:
828:
827:
823:
818:
817:
813:
808:
807:
803:
798:
797:
793:
788:
784:
783:
779:
774:
773:
769:
768:
763:
761:
759:
755:
751:
747:
743:
742:Synechococcus
739:
735:
734:Synechococcus
731:
727:
723:
719:
718:
713:
709:
705:
704:recombination
701:
700:Synechococcus
693:
691:
689:
688:Synechococcus
685:
680:
679:Synechococcus
672:
670:
668:
667:Synechococcus
664:
660:
656:
655:Synechococcus
652:
648:
646:
642:
641:Synechococcus
638:
634:
630:
626:
625:Synechococcus
622:
621:euphotic zone
619:in the upper
618:
614:
613:Synechococcus
610:
609:central gyres
606:
605:Synechococcus
601:
599:
595:
591:
587:
586:Synechococcus
582:
578:
577:Synechococcus
574:
573:Synechococcus
570:
566:
562:
561:
556:
555:Synechococcus
552:
548:
544:
543:McMurdo Sound
540:
539:euphotic zone
536:
535:Synechococcus
529:
527:
525:
524:Synechococcus
521:
520:Synechococcus
517:
516:Synechococcus
513:
510:The complete
508:
506:
502:
498:
494:
490:
485:
483:
482:Synechococcus
479:
475:
471:
467:
460:
456:
455:Synechococcus
452:
448:
444:
440:
435:
433:
432:Synechococcus
429:
425:
421:
420:Synechococcus
416:
412:
408:
407:Synechococcus
404:
397:
395:
393:
389:
385:
381:
377:
373:
372:
367:
363:
362:Synechococcus
359:
358:phycoerythrin
355:
351:
347:
343:
339:
335:
334:chlorophyll a
331:
330:Synechococcus
323:
321:
318:
314:
310:
305:
303:
302:Synechococcus
299:
295:
294:Synechococcus
291:
287:
283:
279:
274:
272:
268:
264:
263:Gram-negative
260:
259:Synechococcus
256:
255:phycoerythrin
252:
251:Synechococcus
247:
243:
239:
236:
233:
229:
228:Synechococcus
222:
220:
218:
214:
210:
205:
203:
202:Synechococcus
199:
195:
191:
190:coccoid cells
188:
184:
180:
176:
172:
168:
164:
163:
162:Synechococcus
155:
151:
150:Synechococcus
146:
138:
134:
129:
124:
119:
118:
117:Synechococcus
114:
111:
110:
107:
104:
101:
100:
97:
94:
91:
90:
87:
84:
81:
80:
77:
76:Cyanobacteria
74:
71:
70:
67:
64:
61:
60:
55:
50:
46:
42:
41:Synechococcus
38:
34:
29:
26:
25:Synechococcus
22:
19:
2876:
2853:
2847:
2811:
2807:
2775:(1): 295–6.
2772:
2766:
2727:
2721:
2708:
2702:
2673:
2667:
2640:
2634:
2605:
2599:
2552:
2548:
2506:
2500:
2473:
2467:
2454:
2435:
2429:
2419:
2400:
2394:
2384:
2357:
2351:
2342:
2334:
2309:
2303:
2297:
2272:
2266:
2260:
2235:
2229:
2203:
2197:
2157:(1): 45–58.
2154:
2148:
2142:
2104:
2098:
2055:
2051:
2047:
1989:
1985:
1979:
1971:
1946:
1940:
1926:
1899:
1893:
1870:
1864:
1823:
1817:
1811:
1789:(1): 21–36.
1786:
1780:
1776:
1770:
1745:
1739:
1733:
1692:
1688:
1687:UTEX 2973".
1684:
1678:
1627:
1623:
1617:
1609:
1584:
1578:
1574:
1520:
1514:
1508:
1481:
1475:
1450:
1444:
1440:
1434:
1415:
1411:Microbiology
1409:
1361:
1355:
1321:
1315:
1305:
1294:. Retrieved
1290:
1281:
1270:
1263:
1228:
1222:
1216:
1208:
1173:
1167:
1161:
1152:
1134:
1127:
1115:
1097:
1087:
1077:
1067:
1057:
1047:
1041:
1034:
1024:
1014:
1004:
994:
984:
974:
964:
954:
944:
934:
924:
914:
904:
894:
884:
874:
864:
854:
844:
834:
824:
814:
804:
794:
790:Fjerdingstad
786:
780:
770:
753:
750:SOS response
745:
741:
733:
725:
715:
699:
697:
687:
678:
676:
666:
663:carbon cycle
658:
654:
650:
649:
644:
640:
624:
616:
612:
604:
602:
594:water column
589:
585:
576:
572:
558:
554:
534:
533:
523:
519:
515:
509:
500:
488:
486:
481:
454:
438:
436:
431:
427:
423:
419:
406:
403:Phylogenetic
401:
392:cryptomonads
380:chlorophytes
369:
361:
329:
327:
316:
312:
308:
306:
301:
293:
275:
271:carboxysomes
258:
250:
238:picoplankton
227:
226:
223:Introduction
213:S. elongatus
212:
206:
201:
170:
166:
161:
160:
159:
154:carboxysomes
149:
136:
116:
115:
86:Cyanophyceae
40:
24:
18:
2981:iNaturalist
2901:Wikispecies
2438:: 175–184.
2403:: 185–196.
1748:: 255–274.
1630:(1): 8132.
722:exonuclease
712:replication
581:mixed layer
415:G+C content
388:rhodophytes
346:phycocyanin
342:phycobilins
315:UTEX 2973.
235:autotrophic
232:prokaryotic
200:species of
3104:Categories
2711:: 367–374.
2476:: 87–101.
1587:: 71–120.
1296:2024-06-09
1144:References
990:G. S. West
810:Rabenhorst
787:calcicolus
758:DNA damage
567:areas and
551:Antarctica
459:euryhaline
447:California
366:zeaxanthin
198:freshwater
2916:AlgaeBase
2855:AlgaeBase
2525:924717835
2074:606510917
2016:1365-294X
1717:0921-8971
1652:2045-2322
1601:565301343
1593:0706-6503
870:F. Hindák
860:F. Hindák
684:temperate
565:upwelling
463:vitamin B
426:(Herdman
398:Phylogeny
390:and some
376:red algae
242:temperate
137:See text
2886:Wikidata
2830:16576851
2799:12231684
2754:10066832
2581:27881980
2555:: 1809.
2360:: 9–18.
2024:25283338
1963:32689463
1856:45409007
1848:13725365
1725:59159665
1670:25633131
1553:29180516
1545:17817167
1418:: 1–61.
1200:12917641
1110:See also
1103:Copeland
1093:Copeland
910:Copeland
324:Pigments
288:or even
267:glycogen
246:tropical
167:synechos
131:Species
102:Family:
72:Phylum:
66:Bacteria
62:Domain:
2999:1073573
2973:3216715
2892:Q150962
2678:Bibcode
2645:Bibcode
2610:Bibcode
2572:5101192
2478:Bibcode
2362:Bibcode
2314:Bibcode
2277:Bibcode
2240:Bibcode
2159:Bibcode
2109:Bibcode
2107:: 1–8.
1994:Bibcode
1828:Bibcode
1819:Science
1803:6416126
1697:Bibcode
1661:5389031
1632:Bibcode
1525:Bibcode
1516:Science
1500:8169723
1467:6786719
1386:4270426
1366:Bibcode
1326:Bibcode
1255:8349551
1178:Bibcode
1000:Komárek
890:Gardner
764:Species
754:E. coli
698:Marine
592:in the
240:in the
112:Genus:
92:Order:
82:Class:
3087:160572
3074:269054
3058:NZOR:
2986:356581
2960:1SYNKG
2828:
2818:
2797:
2790:158675
2787:
2752:
2742:
2579:
2569:
2523:
2513:
2072:
2062:
2022:
2014:
1961:
1914:
1881:
1854:
1846:
1801:
1762:410354
1760:
1723:
1715:
1668:
1658:
1650:
1599:
1591:
1551:
1543:
1498:
1488:
1465:
1384:
1357:Nature
1253:
1246:204977
1243:
1198:
1169:Nature
1083:Grunow
1042:et al.
1040:Usher
980:Grunow
930:Negoro
900:Yoneda
880:Wawrik
840:Norris
717:recBCD
708:repair
647:mRNA.
637:plumes
629:plumes
501:et al.
489:et al.
478:Mexico
428:et al.
278:motile
209:genome
185:. The
171:kokkos
152:. The
125:, 1849
123:Nägeli
3082:WoRMS
2994:IRMNG
2947:12721
2921:43582
2745:98958
2464:(PDF)
2348:(PDF)
2095:(PDF)
1852:S2CID
1721:S2CID
1549:S2CID
1382:S2CID
1053:Skuja
1020:Skuja
1010:Frémy
940:Skuja
820:Okada
800:Skuja
785:var.
776:Skuja
726:umuCD
665:than
443:Texas
3051:1129
3046:NCBI
3020:LPSN
3007:ITIS
2968:GBIF
2955:EPPO
2934:7R7J
2826:OCLC
2816:ISBN
2795:PMID
2750:PMID
2577:PMID
2521:OCLC
2511:ISBN
2343:rbcL
2070:OCLC
2060:ISBN
2050:and
2020:PMID
2012:ISSN
1959:PMID
1912:ISBN
1879:ISBN
1844:PMID
1799:PMID
1787:134B
1779:?".
1758:PMID
1713:ISSN
1666:PMID
1648:ISSN
1597:OCLC
1589:ISSN
1541:PMID
1496:OCLC
1486:ISBN
1463:PMID
1291:NCBI
1251:PMID
1196:PMID
746:lexA
710:and
645:rbcL
545:and
493:rbcL
445:and
382:and
356:and
344:are
298:urea
207:The
3033:NBN
3012:773
2942:EoL
2929:CoL
2785:PMC
2777:doi
2773:101
2740:PMC
2732:doi
2686:doi
2653:doi
2618:doi
2567:PMC
2557:doi
2486:doi
2474:251
2440:doi
2405:doi
2370:doi
2358:198
2322:doi
2285:doi
2248:doi
2208:doi
2167:doi
2117:doi
2105:122
2002:doi
1951:doi
1904:doi
1836:doi
1824:134
1791:doi
1750:doi
1705:doi
1656:PMC
1640:doi
1585:214
1577:".
1533:doi
1521:230
1455:doi
1420:doi
1416:111
1374:doi
1362:277
1334:doi
1241:PMC
1233:doi
1229:175
1186:doi
1174:424
1063:Dor
960:Dor
920:Jao
706:,
549:in
472:in
332:is
244:to
211:of
3106::
3084::
3071::
3048::
3035::
3022::
3009::
2996::
2983::
2970::
2957::
2944::
2931::
2918::
2903::
2888::
2852:.
2824:.
2793:.
2783:.
2771:.
2765:.
2748:.
2738:.
2728:63
2726:.
2720:.
2709:47
2707:.
2701:.
2684:.
2674:43
2672:.
2651:.
2641:39
2639:.
2633:.
2616:.
2606:44
2604:.
2575:.
2565:.
2551:.
2547:.
2533:^
2519:.
2484:.
2472:.
2466:.
2436:35
2434:.
2428:.
2401:35
2399:.
2393:.
2368:.
2356:.
2350:.
2320:.
2310:43
2308:.
2283:.
2273:43
2271:.
2246:.
2236:40
2234:.
2220:^
2204:14
2202:.
2196:.
2181:^
2165:.
2155:35
2153:.
2147:.
2131:^
2115:.
2103:.
2097:.
2082:^
2068:.
2032:^
2018:.
2010:.
2000:.
1990:23
1988:.
1984:.
1957:.
1947:29
1945:.
1939:.
1910:.
1873:.
1850:.
1842:.
1834:.
1822:.
1797:.
1785:.
1756:.
1746:31
1744:.
1719:.
1711:.
1703:.
1693:31
1691:.
1664:.
1654:.
1646:.
1638:.
1626:.
1622:.
1595:.
1583:.
1561:^
1547:.
1539:.
1531:.
1519:.
1494:.
1461:.
1451:27
1449:.
1414:.
1408:.
1394:^
1380:.
1372:.
1360:.
1346:^
1332:.
1322:24
1320:.
1289:.
1249:.
1239:.
1227:.
1221:.
1194:.
1184:.
1172:.
1166:.
760:.
669:.
476:,
465:12
394:.
374:,
352:,
348:,
273:.
217:bp
183:μm
2858:.
2846:"
2832:.
2801:.
2779::
2756:.
2734::
2692:.
2688::
2680::
2661:.
2655::
2647::
2624:.
2620::
2612::
2583:.
2559::
2553:7
2527:.
2494:.
2488::
2480::
2448:.
2442::
2413:.
2407::
2378:.
2372::
2364::
2328:.
2324::
2316::
2291:.
2287::
2279::
2254:.
2250::
2242::
2214:.
2210::
2175:.
2169::
2161::
2125:.
2119::
2111::
2076:.
2026:.
2004::
1996::
1978:"
1965:.
1953::
1935:2
1920:.
1906::
1887:.
1858:.
1838::
1830::
1805:.
1793::
1764:.
1752::
1727:.
1707::
1699::
1672:.
1642::
1634::
1628:5
1616:"
1603:.
1555:.
1535::
1527::
1502:.
1469:.
1457::
1428:.
1422::
1388:.
1376::
1368::
1340:.
1336::
1328::
1299:.
1269:"
1257:.
1235::
1202:.
1188::
1180::
1164:"
682:'
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.