114:
141:
158:
25:
39:
197:
Differences in the biosynthetic pathways of enediynes are due to the different origins of the -yne carbons as well as differences in isotope incorporation patterns. More differentiation comes from the attachment of various functional groups at different positions to the enediyne warheads during their
169:
that acts as a nucleophile and attacks electrophiles in order to achieve a more stable form. In biological systems, once the diradical is positioned in the minor groove of double-stranded DNA, it abstracts two hydrogen atoms from the sugars opposite strands at either the C1, C4, or C5 positions. The
205:
Due to the cytotoxicity of the enediyne chromophores, their biosynthesis is tightly regulated, although the regulatory mechanisms are still largely unclear. Organisms that produce enediynes have been shown to protect themselves with a self-resistance mechanism that uses a self-sacrificing protein.
109:
Bergman cyclization restructures the enediyne ring into two smaller rings. One electron from each of the enediyne triple bonds is pushed to the adjacent single bonds, generating two new double bonds. Meanwhile, another pair of electrons (one from each alkyne) is used form a new covalent bond. The
144:
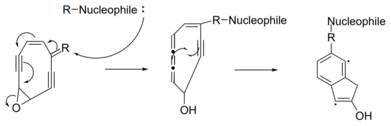
A generic enediyne molecule is pictured above. During Myers-Saito cyclization, electrons pushing begins at the diene and generates an unstable cumulene. This cumulene and the nearby alkyne donate one electron to split the enediyne ring into two fused
105:
or Myers-Saito cyclization. The triggering mechanism can be attributed to an intramolecular nucleophilic attack initiated by one of the variable regions of the molecule. Triggering can also occur via attack by an external nucleophile.
248:
protein H1. Neocarzinostatin is an example of an enediyne that undergoes triggering via Myers-Saito cyclization. An analog of neocarzinostatin, SMANCS, has been approved for use in Japan as an antitumor drug for liver cancer.
132:
with a double bond in a variable group. A nucleophile will attack the double bond in the variable region, causing a chain reaction of electron pushing. Ultimately, one of the triple bonds of the enediyne is converted to a
100:
A nine- or ten-membered ring containing a double bond between two triple bonds is termed the warhead of the enediyne. In this state, the warhead is inactive. Enediynes are triggered into a chemically active state via
933:
Golik J, Clardy J, Dubay G, Groenewold G, Kawaguchi H, Konishi M, Krishnan B, Ohkuma H, Saitoh K (May 1987). "Esperamicins, a novel class of potent antitumor antibiotics. 2. Structure of esperamicin X".
357:
Golfomycin A is a synthetic enediyne molecule designed in an attempt to create a more easily manufactured antitumor antibiotic. DNA strand-scission induced by golfomycin A is pH dependent. Preliminary
190:, precursors that consist of seven or eight head-to-tail coupled acetate units. Enediyne assembly involves a highly conserved, iterative type I polyketide synthase (PKS) pathway Sequencing of enediyne
271:
of China. Unlike most enediynes, C-1027 does not undergo a triggering process to become an activated 1,4-benzenoid diradical. C-1027 has demonstrated potential efficacy against hypoxic tumors.
1069:
Unno R, Michishita H, Inagaki H, Suzuki Y, Baba Y, Jomori T, Nishikawa T, Isobe M (May 1997). "Synthesis and antitumor activity of water-soluble enediyne compounds related to dynemicin A".
299:
organisms. Calicheamicin γ1 exhibited significant antitumor activity against leukemia and melanoma cells in vivo. The calicheamicins are notably similar in structure to the esperamicins.
128:
Myers-Saito cyclization is another triggering mechanism by which an enediyne warhead becomes a diradical. This mechanism requires the alkene of the enediyne to be part of a
1109:
Nicolaou KC, Skokotas G, Furuya S, Suemune H, Nicolaou DC (September 1990). "Golfomycin A, a Novel
Designed Molecule with DNA-Cleaving Properties and Antitumor Activity".
206:
Notably, some microbes use CalC to sequester calicheamicin so that the reactive diradical abstracts hydrogens from a glycine inside of the protein instead of from DNA.
186:
Members of the enediyne family all share a unique enediyne core that is the cause of their potent cytotoxicity. The enediyne cores are derived from linear, probably
198:
maturation stage. These moieties can be either aromatic or sugars and define sequence specificity of DNA binding as well as the physical properties of the enediyne
849:
Xu YJ, Zhen YS, Goldberg IH (May 1994). "C1027 chromophore, a potent new enediyne antitumor antibiotic, induces sequence-specific double-strand DNA cleavage".
125:
group attached to their ring, making
Bergman cyclization unfavorable due to steric hindrance. For Bergman cyclization to occur, the epoxide must be removed.
313:, members of the esperamicin family include esperamicin A1, A1b, A2, A3, A4, B1, B2, and X. Esperamicin X is an inactive esperamicin naturally produced by
117:
Electron pushing that occurs during
Bergman cyclization in a generic enediyne molecule. A 1,4-benzenoid diradical fused to another ring is the result.
523:"Improvement of the enediyne antitumor antibiotic C-1027 production by manipulating its biosynthetic pathway regulation in Streptomyces globisporus"
349:
as a variable group attached to the enediyne core. Dynemycins have demonstrated strong antitumor activity against leukemia and melanoma cells.
309:
The esperamicins are a sub-family of enediynes that are considered among the most potent antitumor antibiotics discovered. First isolated in
229:
because they have an attached protein as a variable group. This protein is necessary for transport and stabilization of the enediyne group.
1162:
217:
Enediynes have been split into two sub-families: those with nine members in the core enediyne ring and those with ten-membered rings.
161:
Abstraction of hydrogen from either the C1, C4, or C5 position of the sugar phosphate backbone of DNA by a reactive 1,4-benzenoid
194:
has confirmed the polyketide origin of the enediyne core, and elucidated the biosynthetic pathways and mechanisms of enediynes.
214:
There are fourteen naturally occurring enediynes. The other existing classes of enediynes have been synthesized in the lab.
110:
resulting formation is a 1,4-benzenoid diradical fused to a ring composed of the leftover atoms from the original enediyne.
889:
Maiese WM, Lechevalier MP, Lechevalier HA, Korshalla J, Kuck N, Fantini A, Wildey MJ, Thomas J, Greenstein M (April 1989).
1028:
Konishi M, Ohkuma H, Matsumoto K, Tsuno T, Kamei H, Miyaki T, Oki T, Kawaguchi H, VanDuyne GD, Clardy J (September 1989).
718:"The biosynthetic genes encoding for the production of the dynemicin enediyne core in Micromonospora chersina ATCC53710"
113:
1277:
1272:
286:
92:
from healthy cells. Consequently, research is being conducted to increase the specificity of enediyne toxicity.
1262:
814:
Maeda H (March 2001). "SMANCS and polymer-conjugated macromolecular drugs: advantages in cancer chemotherapy".
263:
140:
1155:
459:
891:"Calicheamicins, a novel family of antitumor antibiotics: taxonomy, fermentation and biological properties"
296:
292:
971:"Nucleotide-specific cleavage and minor-groove interaction of DNA with esperamicin antitumor antibiotics"
1257:
982:
575:
Liang ZX (April 2010). "Complexity and simplicity in the biosynthesis of enediyne natural products".
413:
102:
1231:
1267:
1148:
268:
1126:
1086:
1051:
1010:
951:
912:
866:
831:
796:
747:
693:
644:
592:
552:
500:
441:
329:
The dynemicins are a sub-family of enediynes whose members are organic compounds generated in
137:. The cumulene and the remaining alkyne donate one electron each to form a new covalent bond.
1200:
1118:
1078:
1041:
1000:
990:
943:
902:
858:
823:
786:
778:
737:
729:
683:
675:
634:
626:
584:
542:
534:
492:
431:
421:
370:
237:
165:
The cyclization of the enediyne functional group creates a transient reactive 1,4-benzenoid
58:
613:
Shen B, Hindra, Yan X, Huang T, Ge H, Yang D, Teng Q, Rudolf JD, Lohman JR (January 2015).
1236:
986:
417:
742:
717:
688:
663:
639:
614:
547:
522:
1082:
1005:
970:
827:
791:
766:
1251:
1216:
733:
436:
401:
346:
291:. All calicheamicin family members demonstrate potent antimicrobial activity against
226:
149:
The diradicals generated by
Bergman and Myers-Saito cyclization are highly reactive.
44:
782:
615:"Enediynes: Exploration of microbial genomics to discover new anticancer drug leads"
361:
studies have demonstrated that golfomycin A can reduce carcinomas in bladder cells.
1195:
1030:"Dynemicin A, a novel antibiotic with the anthraquinone and 1,5-diyn-3-ene subunit"
191:
157:
85:
77:
1226:
1221:
1046:
1029:
969:
Sugiura Y, Uesawa Y, Takahashi Y, Kuwahara J, Golik J, Doyle TW (October 1989).
334:
199:
89:
66:
62:
975:
Proceedings of the
National Academy of Sciences of the United States of America
907:
890:
679:
630:
406:
Proceedings of the
National Academy of Sciences of the United States of America
337:
was the first member of this sub-family to be discovered. It was isolated from
1190:
261:
is one of the most potent antitumor enediynes. C-1027 was first isolated from
187:
171:
73:
1130:
955:
995:
426:
244:. It forms an apoprotein with a 113-amino acid polypeptide which can cleave
166:
81:
1122:
835:
800:
751:
697:
648:
596:
556:
38:
1090:
1055:
1014:
916:
870:
504:
445:
284:
The calicheamicins are a sub-family of enediynes that were isolated from
134:
947:
862:
24:
380:
342:
245:
122:
538:
496:
1185:
588:
258:
375:
318:
156:
139:
129:
112:
767:"Reinvestigation of the proteolytic activity of neocarzinostatin"
1144:
765:
Heyd B, Lerat G, Adjadj E, Minard P, Desmadril M (April 2000).
16:
Any organic compound containing one double and two triple bonds
483:
Smith AL, Nicolaou KC (May 1996). "The enediyne antibiotics".
345:
in India. Dynemicins are violet in color because they contain
72:
Enediynes are most notable for their limited use as antitumor
1140:
402:"Chemistry and biology of natural and designed enediynes"
178:, leading to double- or single-stranded DNA cleavage.
47:: An antitumor antibiotic featuring an enediyne unit
225:
The nine-membered enediynes are also referred to as
1209:
1178:
1111:Angewandte Chemie International Edition in English
170:DNA radicals that form can then cause interstrand
321:groups induce triggering among the esperamicins.
664:"Biosynthesis of enediyne antitumor antibiotics"
521:Chen Y, Yin M, Horsman GP, Shen B (March 2011).
1156:
8:
619:Bioorganic & Medicinal Chemistry Letters
608:
606:
516:
514:
400:Nicolaou KC, Smith AL, Yue EW (July 1993).
1163:
1149:
1141:
1045:
1004:
994:
906:
790:
741:
687:
638:
546:
435:
425:
341:in a soil sample taken from the state of
936:Journal of the American Chemical Society
392:
884:
882:
880:
1104:
1102:
1100:
928:
926:
711:
709:
707:
668:Current Topics in Medicinal Chemistry
7:
1071:Bioorganic & Medicinal Chemistry
570:
568:
566:
221:Nine-membered rings (chromoproteins)
80:). They are efficient at inducing
14:
734:10.1111/j.1574-6968.2008.01112.x
267:in a soil sample taken from the
37:
23:
783:10.1128/jb.182.7.1812-1818.2000
816:Advanced Drug Delivery Reviews
716:Gao Q, Thorson JS (May 2008).
485:Journal of Medicinal Chemistry
1:
1083:10.1016/s0968-0896(97)00037-0
828:10.1016/s0169-409x(00)00134-4
662:Van Lanen SG, Shen B (2008).
30:The enediyne functional group
242:Streptomyces carzinostaticus
1047:10.7164/antibiotics.42.1449
527:Journal of Natural Products
311:Actinomadura verrucosospora
88:, but cannot differentiate
1294:
1034:The Journal of Antibiotics
908:10.7164/antibiotics.42.558
895:The Journal of Antibiotics
680:10.2174/156802608783955656
631:10.1016/j.bmcl.2014.11.019
287:Micromonospora echinospora
722:FEMS Microbiology Letters
464:www.organic-chemistry.org
257:Also known as lidamycin,
264:Streptomyces globisporus
240:is a natural product of
96:Structure and reactivity
996:10.1073/pnas.86.20.7672
771:Journal of Bacteriology
577:Natural Product Reports
427:10.1073/pnas.90.13.5881
331:Micromonospora chersina
121:Some enediynes have an
1123:10.1002/anie.199010641
162:
146:
118:
78:anticancer antibiotics
460:"Bergman Cyclization"
160:
143:
116:
987:1989PNAS...86.7672S
948:10.1021/ja00245a048
863:10.1021/bi00185a036
418:1993PNAS...90.5881N
269:Qian-Jiang District
153:Mechanism of action
76:(known as enediyne
275:Ten-membered rings
163:
147:
119:
1278:Conjugated enynes
1273:Functional groups
1245:
1244:
1210:10 membered rings
942:(11): 3461–3462.
539:10.1021/np100825y
497:10.1021/jm9600398
317:. Compounds with
315:A. verrucosospora
59:organic compounds
1285:
1201:Neocarzinostatin
1179:9 membered rings
1165:
1158:
1151:
1142:
1135:
1134:
1117:(9): 1064–1067.
1106:
1095:
1094:
1066:
1060:
1059:
1049:
1025:
1019:
1018:
1008:
998:
966:
960:
959:
930:
921:
920:
910:
886:
875:
874:
846:
840:
839:
811:
805:
804:
794:
762:
756:
755:
745:
713:
702:
701:
691:
659:
653:
652:
642:
610:
601:
600:
589:10.1039/b908165h
572:
561:
560:
550:
518:
509:
508:
480:
474:
473:
471:
470:
456:
450:
449:
439:
429:
397:
371:Enyne metathesis
238:Neocarzinostatin
233:Neocarzinostatin
41:
27:
1293:
1292:
1288:
1287:
1286:
1284:
1283:
1282:
1263:Cancer research
1248:
1247:
1246:
1241:
1237:Shishijimicin A
1205:
1174:
1169:
1139:
1138:
1108:
1107:
1098:
1068:
1067:
1063:
1027:
1026:
1022:
968:
967:
963:
932:
931:
924:
888:
887:
878:
857:(19): 5947–54.
848:
847:
843:
822:(1–3): 169–85.
813:
812:
808:
764:
763:
759:
715:
714:
705:
661:
660:
656:
612:
611:
604:
574:
573:
564:
520:
519:
512:
491:(11): 2103–17.
482:
481:
477:
468:
466:
458:
457:
453:
399:
398:
394:
389:
367:
355:
327:
307:
282:
277:
255:
235:
223:
212:
184:
177:
174:or react with O
155:
98:
90:cancerous cells
61:containing two
52:
51:
50:
49:
48:
42:
33:
32:
31:
28:
17:
12:
11:
5:
1291:
1289:
1281:
1280:
1275:
1270:
1265:
1260:
1250:
1249:
1243:
1242:
1240:
1239:
1234:
1229:
1224:
1219:
1213:
1211:
1207:
1206:
1204:
1203:
1198:
1193:
1188:
1182:
1180:
1176:
1175:
1170:
1168:
1167:
1160:
1153:
1145:
1137:
1136:
1096:
1061:
1040:(9): 1449–52.
1020:
981:(20): 7672–6.
961:
922:
876:
841:
806:
757:
703:
654:
602:
583:(4): 499–528.
562:
510:
475:
451:
412:(13): 5881–8.
391:
390:
388:
385:
384:
383:
378:
373:
366:
363:
354:
351:
326:
323:
306:
303:
281:
280:Calicheamicins
278:
276:
273:
254:
251:
234:
231:
227:chromoproteins
222:
219:
211:
208:
183:
180:
175:
154:
151:
97:
94:
43:
36:
35:
34:
29:
22:
21:
20:
19:
18:
15:
13:
10:
9:
6:
4:
3:
2:
1290:
1279:
1276:
1274:
1271:
1269:
1266:
1264:
1261:
1259:
1256:
1255:
1253:
1238:
1235:
1233:
1230:
1228:
1225:
1223:
1220:
1218:
1217:Calicheamicin
1215:
1214:
1212:
1208:
1202:
1199:
1197:
1194:
1192:
1189:
1187:
1184:
1183:
1181:
1177:
1173:
1166:
1161:
1159:
1154:
1152:
1147:
1146:
1143:
1132:
1128:
1124:
1120:
1116:
1112:
1105:
1103:
1101:
1097:
1092:
1088:
1084:
1080:
1077:(5): 987–99.
1076:
1072:
1065:
1062:
1057:
1053:
1048:
1043:
1039:
1035:
1031:
1024:
1021:
1016:
1012:
1007:
1002:
997:
992:
988:
984:
980:
976:
972:
965:
962:
957:
953:
949:
945:
941:
937:
929:
927:
923:
918:
914:
909:
904:
901:(4): 558–63.
900:
896:
892:
885:
883:
881:
877:
872:
868:
864:
860:
856:
852:
845:
842:
837:
833:
829:
825:
821:
817:
810:
807:
802:
798:
793:
788:
784:
780:
777:(7): 1812–8.
776:
772:
768:
761:
758:
753:
749:
744:
739:
735:
731:
728:(1): 105–14.
727:
723:
719:
712:
710:
708:
704:
699:
695:
690:
685:
681:
677:
674:(6): 448–59.
673:
669:
665:
658:
655:
650:
646:
641:
636:
632:
628:
624:
620:
616:
609:
607:
603:
598:
594:
590:
586:
582:
578:
571:
569:
567:
563:
558:
554:
549:
544:
540:
536:
532:
528:
524:
517:
515:
511:
506:
502:
498:
494:
490:
486:
479:
476:
465:
461:
455:
452:
447:
443:
438:
433:
428:
423:
419:
415:
411:
407:
403:
396:
393:
386:
382:
379:
377:
374:
372:
369:
368:
364:
362:
360:
352:
350:
348:
347:anthraquinone
344:
340:
336:
332:
324:
322:
320:
316:
312:
304:
302:
300:
298:
297:Gram-negative
294:
293:Gram-positive
290:
288:
279:
274:
272:
270:
266:
265:
260:
252:
250:
247:
243:
239:
232:
230:
228:
220:
218:
215:
209:
207:
203:
201:
195:
193:
192:gene clusters
189:
181:
179:
173:
168:
159:
152:
150:
142:
138:
136:
131:
126:
124:
115:
111:
107:
104:
95:
93:
91:
87:
83:
79:
75:
70:
68:
64:
60:
56:
46:
45:Calicheamicin
40:
26:
1232:Golfomycin A
1196:Maduropeptin
1171:
1114:
1110:
1074:
1070:
1064:
1037:
1033:
1023:
978:
974:
964:
939:
935:
898:
894:
854:
851:Biochemistry
850:
844:
819:
815:
809:
774:
770:
760:
725:
721:
671:
667:
657:
622:
618:
580:
576:
533:(3): 420–4.
530:
526:
488:
484:
478:
467:. Retrieved
463:
454:
409:
405:
395:
358:
356:
353:Golfomycin A
338:
330:
328:
314:
310:
308:
305:Esperamicins
301:
285:
283:
262:
256:
241:
236:
224:
216:
213:
204:
200:chromophores
196:
185:
182:Biosynthesis
164:
148:
127:
120:
108:
99:
71:
63:triple bonds
54:
53:
1258:Antibiotics
1227:Esperamicin
1222:Dynemicin A
625:(1): 9–15.
339:M. chersina
335:Dynemicin A
289:calichensis
74:antibiotics
67:double bond
1252:Categories
1191:Kedarcidin
469:2018-05-05
387:References
325:Dynemicins
188:polyketide
172:crosslinks
1268:Enediynes
1172:Enediynes
1131:0570-0833
956:0002-7863
167:diradical
82:apoptosis
55:Enediynes
836:11259839
801:10714984
752:18328078
698:18397168
649:25434000
597:20336235
557:21250756
365:See also
359:in vitro
135:cumulene
65:and one
1091:9208107
1056:2793600
1015:2813351
983:Bibcode
917:2722671
871:8180224
743:5591436
689:3108100
640:4480864
548:3064734
505:8667354
446:8327459
414:Bibcode
381:Polyyne
343:Gujarat
246:histone
210:Classes
123:epoxide
103:Bergman
1186:C-1027
1129:
1089:
1054:
1013:
1006:298132
1003:
954:
915:
869:
834:
799:
792:101862
789:
750:
740:
696:
686:
647:
637:
595:
555:
545:
503:
444:
434:
259:C-1027
253:C-1027
145:rings.
437:46830
376:Enyne
319:thiol
130:diene
86:cells
1127:ISSN
1087:PMID
1052:PMID
1011:PMID
952:ISSN
913:PMID
867:PMID
832:PMID
797:PMID
748:PMID
694:PMID
645:PMID
593:PMID
553:PMID
501:PMID
442:PMID
295:and
57:are
1119:doi
1079:doi
1042:doi
1001:PMC
991:doi
944:doi
940:109
903:doi
859:doi
824:doi
787:PMC
779:doi
775:182
738:PMC
730:doi
726:282
684:PMC
676:doi
635:PMC
627:doi
585:doi
543:PMC
535:doi
493:doi
432:PMC
422:doi
84:in
1254::
1125:.
1115:29
1113:.
1099:^
1085:.
1073:.
1050:.
1038:42
1036:.
1032:.
1009:.
999:.
989:.
979:86
977:.
973:.
950:.
938:.
925:^
911:.
899:42
897:.
893:.
879:^
865:.
855:33
853:.
830:.
820:46
818:.
795:.
785:.
773:.
769:.
746:.
736:.
724:.
720:.
706:^
692:.
682:.
670:.
666:.
643:.
633:.
623:25
621:.
617:.
605:^
591:.
581:27
579:.
565:^
551:.
541:.
531:74
529:.
525:.
513:^
499:.
489:39
487:.
462:.
440:.
430:.
420:.
410:90
408:.
404:.
333:.
202:.
69:.
1164:e
1157:t
1150:v
1133:.
1121::
1093:.
1081::
1075:5
1058:.
1044::
1017:.
993::
985::
958:.
946::
919:.
905::
873:.
861::
838:.
826::
803:.
781::
754:.
732::
700:.
678::
672:8
651:.
629::
599:.
587::
559:.
537::
507:.
495::
472:.
448:.
424::
416::
176:2
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.