346:
reason vesicles have been used so frequently is that they are relatively easy to make. If a sample of dehydrated lipid is exposed to water it will spontaneously form vesicles. These initial vesicles are typically multilamellar (many-walled) and are of a wide range of sizes from tens of nanometers to several micrometres. Methods such as sonication or extrusion through a membrane are needed to break these initial vesicles into smaller, single-walled vesicles of uniform diameter known as small unilamellar vesicles (SUVs). SUVs typically have diameters between 50 and 200 nm. Alternatively, rather than synthesizing vesicles it is possible to simply isolate them from cell cultures or tissue samples. Vesicles are used to transport lipids, proteins and many other molecules within the cell as well as into or out of the cell. These naturally isolated vesicles are composed of a complex mixture of different lipids and proteins so, although they offer greater realism for studying specific biological phenomena, simple artificial vesicles are preferred for studies of fundamental lipid properties.
222:(iSCAT). When the bilayer is supported on top of a reflective surface, variations in intensity due to destructive interference from this interface can be used to calculate with angstrom accuracy the position of fluorophores within the bilayer. Both evanescent and interference techniques offer sub-wavelength resolution in only one dimension (z, or vertical). In many cases, this resolution is all that is needed. After all, bilayers are very small only in one dimension. Laterally, a bilayer can extend for many micrometres or even millimeters. But certain phenomena like dynamic phase rearrangement do occur in bilayers on a lateral sub-micrometre length scale. A promising approach to studying these structures is
175:
the study of lipid bilayers. One of the greatest advantages of the supported bilayer is its stability. SLBs will remain largely intact even when subject to high flow rates or vibration and, unlike black lipid membranes, the presence of holes will not destroy the entire bilayer. Because of this stability, experiments lasting weeks and even months are possible with supported bilayers while BLM experiments are usually limited to hours. Another advantage of the supported bilayer is that, because it is on a flat hard surface, it is amenable to a number of characterization tools which would be impossible or would offer lower resolution if performed on a freely floating sample.
260:, the most common experimental system. Because this layer is so thin there is extensive hydrodynamic coupling between the bilayer and the substrate, resulting in a lower diffusion coefficient in supported bilayers than for free bilayers of the same composition. A certain percentage of the supported bilayer will also be completely immobile, although the exact nature of and reason for these “pinned” sites is still uncertain. For high quality liquid phase supported bilayers the immobile fraction is typically around 1-5%. To quantify the diffusion coefficient and mobile fraction, researchers studying supported bilayers will often report
120:
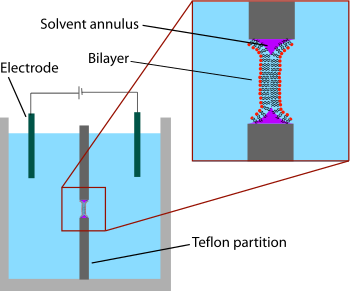
aqueous phases on either side of the lipid/solvent droplet. Because the walls of the aperture are hydrophobic the lipid/solvent solution wets this interface, thinning the droplet in the center. Once the two sides of the droplet come close enough together, the lipid monolayers fuse, rapidly excluding the small remaining volume of solution. At this point a bilayer is formed in the center of the aperture, but a significant annulus of solvent remains at the perimeter. This annulus is required to maintain stability by acting as a bridge between the ~5 nm bilayer and the tens of micrometers thick sheet in which the aperture is made.
374:
used to study membrane-remodeling and other protein-membrane interactions in vitro. A variety of methods exist to encapsulate proteins or other biological reactants within such vesicles, making GUVs an ideal system for the in vitro recreation (and investigation) of cell functions in cell-like model membrane environments. These methods include microfluidic methods, which allow for a high-yield production of vesicles with consistent sizes.
128:
allowing simple placement of large electrodes. For this reason, electrical characterization is one of the most important methods used in conjunction with painted lipid bilayers. Simple measurements indicate when a bilayer forms and when it breaks, as an intact bilayer has a large resistance (>GΩ) and a large capacitance (~2 μF/cm). More advanced electrical characterization has been particularly important in the study of
87:
218:(SPR) can offer extremely sensitive measurement of analyte binding and bilayer optical properties but can only function when the sample is supported on specialized optically functional materials. Another class of methods applicable only to supported bilayers is those based on optical interference such as fluorescence interference contrast microscopy (FLIC) and reflection interference contrast microscopy (RICM) or
167:
230:
338:
290:
383:
interfaces. DIBs can be formed to create tissue-like material with the ability to form asymmetric bilayers, reconstitute proteins and protein channels or made for use in studying electrophysiology. Extended DIB networks can be formed either by employing droplet microfluidic devices or using droplet printers.
409:
Bicelles are a related class of model membrane, typically made of two lipids, one of which forms a lipid bilayer while the other forms an amphipathic, micelle-like assembly shielding the bilayer center from surrounding solvent molecules. Bicelles can be thought of as a segment of bilayer encapsulated
243:
to demonstrate that stable composition gradients could be formed in bilayers, potentially allowing massively parallel studies of phase segregation, molecular binding and cellular response to artificial lipid membranes. Creative utilization of the corral concept has also allowed studies of the dynamic
140:
and added to the aqueous solution after the bilayer is formed. The detergent coating allows these proteins to spontaneously insert into the bilayer over a period of minutes. Additionally, initial experiments have been performed which combine electrophysiological and structural investigations of black
90:
Schematic of a painted bilayer experiment. A sheet of plastic with a small hole in the center separates the two sides of the chamber. The bilayer is formed across this hole, separating the two chambers. The electrical properties of the bilayer can be measured by putting an electrode into each side of
382:
Droplet
Interface Bilayers (DIBs) are phospholipid-encased droplets that form bilayers when they are put into contact. The droplets are surrounded by oil and phospholipids are dispersed in either the water or oil. As a result, the phospholipids spontaneously form a monolayer at each of the oil-water
368:
In spite of the fluorescent labeling, it is often difficult to perform detailed imaging on SUVs simply because they are so small. To combat this problem, researchers use giant unilamellar vesicles (GUVs). GUVs are large enough (1 - 200 μm) to be studied using traditional fluorescence microscopy
148:
The main problems associated with painted bilayers are residual solvent and limited lifetime. Some researchers believe that pockets of solvent trapped between the two bilayer leaflets can disrupt normal protein function. To overcome this limitation, Montal and
Mueller developed a modified deposition
190:
of single bilayers and to perform force spectroscopy on individual membrane proteins. These studies would be difficult or impossible without the use of supported bilayers since the surface of a cell or vesicle is relatively soft and would drift and fluctuate over time. Another example of a physical
174:
Unlike a vesicle or a cell membrane in which the lipid bilayer is rolled into an enclosed shell, a supported bilayer is a planar structure sitting on a solid support. Because of this, only the upper face of the bilayer is exposed to free solution. This layout has advantages and drawbacks related to
119:
After allowing the aperture to dry, salt solution (aqueous phase) is added to both sides of the chamber. The aperture is then "painted" with a lipid solution (generally the same solution that was used for pre-painting). A lipid monolayer spontaneously forms at the interface between the organic and
373:
in artificial lipid systems have been performed with GUVs for this reason. Compared to supported bilayers, GUVs present a more “natural” environment since there is no rigid surface that might induce defects, affect the properties of the membrane or denature proteins. Therefore, GUVs are frequently
345:
A vesicle is a lipid bilayer rolled up into a spherical shell, enclosing a small amount of water and separating it from the water outside the vesicle. Because of this fundamental similarity to the cell membrane, vesicles have been used extensively to study the properties of lipid bilayers. Another
251:
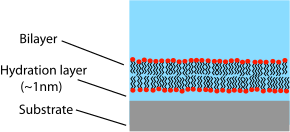
One of the primary limitations of supported bilayers is the possibility of unwanted interactions with the substrate. Although supported bilayers generally do not directly touch the substrate surface, they are separated by only a very thin water gap. The size and nature of this gap depends on the
153:
and waiting for the solvent to evaporate. The aperture is then lowered through the air-water interface and the two monolayers from the separate chambers are folded down against each other, forming a bilayer across the aperture. The stability issue has proven more difficult to solve. Typically, a
127:
with light reflecting off the front face. Indeed, this was one of the first clues that this technique produced a membrane of molecular-scale thickness. Black lipid membranes are also well suited to electrical characterization because the two chambers separated by the bilayer are both accessible,
69:
There are many different types of model bilayers, each having experimental advantages and disadvantages. The first system developed was the black lipid membrane or “painted” bilayer, which allows simple electrical characterization of bilayers but is short-lived and can be difficult to work with.
318:
The advantage of this approach is that because of the hydrophilic space of around 4 nm, the interaction with the substrate is minimal and the extra space allows the introduction of protein ion channels into the bilayer. Additionally the spacer layer creates an ionic reservoir that readily
238:
Another important capability of supported bilayers is the ability to pattern the surface to produce multiple isolated regions on the same substrate. This phenomenon was first demonstrated using scratches or metallic “corrals” to prevent mixing between adjacent regions while still allowing free
233:
Fluorescence micrograph of a supported bilayer on a substrate that has been patterned with a corral. This substrate was then sequentially exposed to two different populations of lipids (dyed red and green). Although the populations were kept largely separated there was some intermixing at the
420:
consist of a segment of bilayer encapsulated by an amphipathic protein coat, rather than a lipid or detergent layer. Nanodiscs are more stable than bicelles and micelles at low concentrations, and are very well-defined in size (depending on the type of protein coat, between 10 and 20
314:
The limitation of the intra-membrane mobility of supported lipid bilayers can be overcome by introducing half-membrane spanning tether lipids with benzyl disulphide (DPL) and synthetic archaea analogue full membrane spanning lipids with phytanoly chains to stabilize the structure and
149:
technique that eliminates the use of a heavy non-volatile solvent. In this method, the aperture starts out above the water surface, completely separating the two fluid chambers. On the surface of each chamber, a monolayer is formed by applying lipids in a volatile solvent such as
267:
Unwanted substrate interactions are a much greater problem when incorporating integral membrane proteins, particularly those with large domains sticking out beyond the core of the bilayer. Because the gap between bilayer and substrate is so thin these proteins will often become
158:. This lifetime can be extended by precisely structuring the supporting aperture, chemically crosslinking the lipids or gelling the surrounding solution to mechanically support the bilayer. Work is ongoing in this area and lifetimes of several hours will become feasible.
95:
The earliest model bilayer system developed was the “painted” bilayer, also known as a “black lipid membrane.” The term “painted” refers to the process by which these bilayers are made. First, a small aperture is created in a thin layer of a hydrophobic material such as
277:
which acts as a spacer and theoretically prevents denaturing substrate interactions. In practice, some percentage of the proteins will still lose mobility and functionality, probably due to interactions with the polymer/lipid anchors. Research in this area is ongoing.
141:
lipid membranes. In another variation of the BLM technique, termed the bilayer punch, a glass pipet (inner diameter ~10-40 μm) is used as the electrode on one side of the bilayer in order to isolate a small patch of membrane. This modification of the
226:(NSOM). Like AFM, NSOM relies on the scanning of a micromachined tip to give a highly localized signal. But unlike AFM, NSOM uses an optical rather than physical interaction with the sample, potentially perturbing delicate structures to a lesser extent.
298:
Gold can be used as a substrate because of its inert chemistry and thiolipids for covalent binding to the gold. Thiolipids are composed of lipid derivatives, extended at their polar head-groups by hydrophilic spacers which terminate in a
3898:
405:
molecules with their hydrophilic heads exposed to solvent and their hydrophobic tails in the center. Micelles can solubilize membrane proteins by partially encapsulating them and shielding their hydrophobic surfaces from solvent.
272:
on the substrate surface and therefore lose all functionality. One approach to circumvent this problem is the use of polymer tethered bilayers. In these systems the bilayer is supported on a loose network of hydrated polymers or
1215:
Mashaghi A, Swann M, Popplewell J, Textor M, Reimhult E (May 2008). "Optical anisotropy of supported lipid structures probed by waveguide spectroscopy and its application to study of supported lipid bilayer formation kinetics".
186:, formation of transmembrane nanopores followed by single protein molecule adsorption, and protein assembly with sub-nm accuracy without the need for a labeling dye. More recently, AFM has also been used to directly probe the
1600:
König BW, Krueger S, Orts WJ, Majkrzak CF, Berk NF, Silverton JV, Gawrisch K (1996). "Neutron reflectivity and atomic force microscopy studies of a lipid bilayer in water adsorbed to the surface of a silicon single crystal".
917:
Purrucker O, Hillebrandt H, Adlkofer K, Tanaka M (2001). "Deposition of highly resistive lipid bilayer on silicon-silicon dioxide electrode and incorporation of gramicidin studied by ac impedance spectroscopy".
100:. Typically the diameter of this hole is a few tens of micrometers up to hundreds of micrometers. To form a BLM, the area around the aperture is first "pre-painted" with a solution of lipids dissolved in a
3891:
357:
can measure unilamelar and multilamelar structures and insertion into and disruption of the vesicles in a label free assay format. Vesicles can also be labeled with fluorescent dyes to allow sensitive
123:
The term “black” bilayer refers to the fact that they are dark in reflected light because the thickness of the membrane is only a few nanometers, so light reflecting off the back face destructively
3068:
3008:
Roos C, Kai L, Proverbio D, Ghoshdastider U, Filipek S, Dötsch V, Bernhard F (February 2013). "Co-translational association of cell-free expressed membrane proteins with supplied lipid bilayers".
315:
polyethyleneglycol units as a hydrophilic spacer. Bilayer formation is achieved by exposure of the lipid coated gold substrate to outer layer lipids either in an ethanol solution or in liposomes.
136:
typically cannot be incorporated directly into the painted bilayer during formation because immersion in an organic solvent would denature the protein. Instead, the protein is solubilized with a
3884:
369:
and are within the same size range as most biological cells. Thus, they are used as mimicries of cell membranes for in vitro studies in molecular and cell biology. Many of the studies of
66:. The simplest model systems contain only a single pure synthetic lipid. More physiologically relevant model bilayers can be made with mixtures of several synthetic or natural lipids.
847:
Beerlink A, Wilbrandt PJ, Ziegler E, Carbone D, Metzger TH, Salditt T (May 2008). "X-ray structure analysis of free-standing lipid membranes facilitated by micromachined apertures".
2082:
F Szoka and D Papahadjopoulos."Comparative
Properties and Methods of Preparation of Lipid Vesicles (Liposomes)." Annual Review of Biophysics and Bioengineering. 9. (1980) 467-508.
1999:
Bangham AD, Horne RW (January 1964). "Negative staining of phospholipids and their structural modification by surface-active agents as observed in the electron microscope".
3061:
211:
2916:
Ritchie TK, Grinkova YV, Bayburt TH, Denisov IG, Zolnerciks JK, Atkins WM, Sligar SG (2009). "Reconstitution of
Membrane Proteins in Phospholipid Bilayer Nanodiscs".
286:
The use of a tethered bilayer lipid membrane (t-BLM) further increases the stability of supported membranes by chemically anchoring the lipids to the solid substrate.
2514:
Funakoshi K, Suzuki H, Takeuchi S (December 2006). "Lipid bilayer formation by contacting monolayers in a microfluidic device for membrane protein analysis".
3054:
2584:
Leptihn S, Thompson JR, Ellory JC, Tucker SJ, Wallace MI (June 2011). "In vitro reconstitution of eukaryotic ion channels using droplet interface bilayers".
2967:
Roos C, Kai L, Haberstock S, Proverbio D, Ghoshdastider U, Ma Y, et al. (2014). "High-Level Cell-Free
Production of Membrane Proteins with Nanodiscs".
490:
Mueller P, Rudin DO, Tien HT, Wescott WC (June 1962). "Reconstitution of cell membrane structure in vitro and its transformation into an excitable system".
54:. They are used to study the fundamental properties of biological membranes in a simplified and well-controlled environment, and increasingly in bottom-up
4454:
1349:
Crane JM, Kiessling V, Tamm LK (February 2005). "Measuring lipid asymmetry in planar supported bilayers by fluorescence interference contrast microscopy".
261:
178:
One of the clearest examples of this advantage is the use of mechanical probing techniques which require a direct physical interaction with the sample.
1859:
Naumann R, Jonczyk A, Kopp R, van Esch J, Ringsdorf H, Knoll W, Gräber P (1995). "Incorporation of
Membrane Proteins in Solid-Supported Lipid Layers".
1386:"Submicron structure in L-alpha-dipalmitoylphosphatidylcholine monolayers and bilayers probed with confocal, atomic force, and near-field microscopy"
1964:
Cornell BA, Krishna G, Osman PD, Pace RD, Wieczorek L (August 2001). "Tethered-bilayer lipid membranes as a support for membrane-active peptides".
2676:
Gross LC, Heron AJ, Baca SC, Wallace MI (December 2011). "Determining membrane capacitance by dynamic control of droplet interface bilayer area".
1484:
Groves JT, Ulman N, Cremer PS, Boxer SG (1998). "Substrate−Membrane
Interactions: Mechanisms for Imposing Patterns on a Fluid Bilayer Membrane".
1511:
Kam L, Boxer SG (2003). "Spatially
Selective Manipulation of Supported Lipid Bilayers by Laminar Flow: Steps Toward Biomembrane Microfluidics".
1188:
Ebara Y, Okahata (December 1994). "A kinetic study of concanavalin A binding to glycolipid monolayers by using a quartz-crystal microbalance".
223:
1886:
Cornell BA, Braach-Maksvytis VL, King LG, Osman PD, Raguse B, Wieczorek L, Pace RJ (June 1997). "A biosensor that uses ion-channel switches".
2984:
219:
2186:
Träuble H, Haynes DH (December 1971). "The volume change in lipid bilayer lamellae at the crystalline-liquid crystalline phase transition".
882:
Malmstadt N, Jeon TJ, Schmidt JJ (January 2008). "Long-Lived Planar Lipid
Bilayer Membranes Anchored to an In Situ Polymerized Hydrogel".
358:
2151:
Papahadjopoulos D, Miller N (September 1967). "Phospholipid model membranes. I. Structural characteristics of hydrated liquid crystals".
1145:
Oesterhelt F, Oesterhelt D, Pfeiffer M, Engel A, Gaub HE, Müller DJ (April 2000). "Unfolding pathways of individual bacteriorhodopsins".
4387:
1002:
Roiter Y, Ornatska M, Rammohan AR, Balakrishnan J, Heine DR, Minko S (March 2008). "Interaction of nanoparticles with lipid membrane".
4444:
2933:
4449:
354:
196:
104:
by applying this solution across the aperture with a brush, syringe, or glass applicator. The solvent used must have a very high
425:). Membrane proteins incorporated into and solubilized by Nanodiscs can be studied by a wide variety of biophysical techniques.
124:
145:
technique enables low noise recording, even at high potentials (up to 600 mV), at the expense of additional preparation time.
4372:
3722:
3317:
71:
4360:
3858:
3506:
183:
2873:, Wright PE (April 1999). "Improved low pH bicelle system for orienting macromolecules over a wide temperature range".
1538:
Parthasarathy R, Jackson BL, Lowery TJ, Wong AP (2004). "Nonequilibrium
Adhesion Patterns at Lipid Bilayer Junctions".
598:
Tien HT, Carbone S, Dawidowicz EA (1966). "Formation of "black" lipid membranes by oxidation products of cholesterol".
199:
is a high resolution optical tool for characterising the order and disruption in lipid bilayers during interactions or
74:
not possible in bulk solution. These advantages come at the cost of unwanted substrate interactions which can denature
3426:
3182:
1804:"Polymer-cushioned bilayers. II. An investigation of interaction forces and fusion using the surface forces apparatus"
1565:
Mager MD, Almquist B, Melosh NA (November 2008). "Formation and characterization of fluid lipid bilayers on alumina".
269:
192:
1937:
Lang H, Duschl C, Vogel H (1994). "A new class of thiolipids for the attachment of lipid bilayers on gold surfaces".
410:
and solubilized by a micelle. Bicelles are much smaller than liposomes, and so can be used in experiments such as
4514:
3727:
215:
129:
1253:"Direct observation and control of supported lipid bilayer formation with interferometric scattering microscopy"
4377:
3574:
3475:
3392:
1630:"Structure of an adsorbed dimyristoylphosphatidylcholine bilayer measured with specular reflection of neutrons"
1045:
Engel A, Müller DJ (September 2000). "Observing single biomolecules at work with the atomic force microscope".
308:
4466:
3601:
3377:
320:
187:
179:
3829:
3145:
349:
Since artificial SUVs can be made in large quantities they are suitable for bulk material studies such as
108:
and must be relatively viscous to prevent immediate rupture. The most common solvent used is a mixture of
1252:
947:"Lipid asymmetry in DLPC/DSPC-supported lipid bilayers: a combined AFM and fluorescence microscopy study"
4021:
3589:
3562:
105:
1690:"Lipid mono- and bilayer supported on polymer films: composite polymer-lipid films on solid substrates"
1441:
Groves JT, Ulman N, Boxer SG (January 1997). "Micropatterning fluid lipid bilayers on solid supports".
341:
Diagram of lipid vesicles showing a solution of molecules (green dots) trapped in the vesicle interior.
4476:
3876:
3533:
3528:
3404:
2784:
2727:
2632:
2549:
Hwang WL, Chen M, Cronin B, Holden MA, Bayley H (May 2008). "Asymmetric droplet interface bilayers".
2324:
2267:
2105:
1895:
1815:
1758:
1701:
1641:
1397:
1303:
1154:
1101:
1011:
958:
891:
801:
790:"Formation of bimolecular membranes from lipid monolayers and a study of their electrical properties"
695:
654:
607:
554:
499:
362:
332:
155:
4051:
3611:
3567:
3523:
3261:
2256:"Lipid bilayer vesicle fusion: intermediates captured by high-speed microfluorescence spectroscopy"
353:
to determine lattice spacing and differential scanning calorimetry to determine phase transitions.
70:
Supported bilayers are anchored to a solid substrate, increasing stability and allowing the use of
4345:
4267:
3819:
3033:
2898:
2393:
2368:
Litschel T, Schwille P (March 2021). "Protein Reconstitution Inside Giant Unilamellar Vesicles".
1919:
1466:
1070:
684:"Ion movement through gramicidin A channels. Single-channel measurements at very high potentials"
623:
523:
1628:
Johnson SJ, Bayerl TM, McDermott DC, Adam GW, Rennie AR, Thomas RK, Sackmann E (February 1991).
206:
Many modern fluorescence microscopy techniques also require a rigidly-supported planar surface.
2311:
Dietrich C, Bagatolli LA, Volovyk ZN, Thompson NL, Levi M, Jacobson K, Gratton E (March 2001).
4323:
4239:
4141:
3911:
3814:
3518:
3025:
2990:
2980:
2949:
2929:
2890:
2851:
2810:
2753:
2693:
2658:
2601:
2566:
2531:
2496:
2442:
2385:
2350:
2293:
2236:
2168:
2133:
2065:
2016:
1981:
1911:
1841:
1784:
1727:
1667:
1582:
1458:
1423:
1366:
1331:
1272:
1233:
1170:
1127:
1062:
1027:
984:
864:
829:
770:
721:
580:
515:
472:
398:
350:
75:
55:
3550:
3444:
3077:
3017:
2972:
2939:
2921:
2882:
2841:
2800:
2792:
2743:
2735:
2685:
2648:
2640:
2621:"Activation of bacterial channel MscL in mechanically stimulated droplet interface bilayers"
2593:
2558:
2523:
2486:
2478:
2432:
2424:
2377:
2340:
2332:
2283:
2275:
2226:
2195:
2160:
2123:
2113:
2055:
2047:
2008:
1973:
1946:
1903:
1868:
1831:
1823:
1774:
1766:
1717:
1709:
1657:
1649:
1610:
1574:
1547:
1520:
1493:
1450:
1413:
1405:
1358:
1321:
1311:
1264:
1225:
1197:
1162:
1117:
1109:
1054:
1019:
974:
966:
927:
899:
856:
819:
809:
760:
752:
711:
703:
662:
615:
570:
562:
507:
462:
454:
200:
4183:
3807:
3501:
3480:
3409:
3212:
2381:
304:
207:
59:
28:
2619:
Najem JS, Dunlap MD, Rowe ID, Freeman EC, Grant JW, Sukharev S, Leo DJ (September 2015).
1090:"Mechanical properties of pore-spanning lipid bilayers probed by atomic force microscopy"
2788:
2731:
2636:
2465:
Bayley H, Cronin B, Heron A, Holden MA, Hwang WL, Syeda R, et al. (December 2008).
2328:
2271:
2109:
1899:
1819:
1762:
1705:
1645:
1401:
1307:
1158:
1105:
1015:
962:
895:
805:
699:
658:
611:
558:
503:
4232:
4197:
4190:
4169:
4148:
4125:
4005:
3797:
3684:
3555:
3414:
3397:
3202:
3132:
2944:
2805:
2772:
2748:
2715:
2653:
2620:
2491:
2466:
2437:
2412:
2345:
2312:
2288:
2255:
2060:
2035:
1836:
1803:
1779:
1746:
1722:
1689:
1662:
1629:
1418:
1385:
1326:
1291:
1122:
1089:
979:
946:
765:
740:
716:
683:
575:
542:
467:
442:
401:, although they lack a lipid bilayer. In aqueous solutions, micelles are assemblies of
3046:
2925:
2336:
2279:
2128:
2093:
2012:
1827:
1713:
1653:
1409:
1088:
Steltenkamp S, Müller MM, Deserno M, Hennesthal C, Steinem C, Janshoff A (July 2006).
931:
824:
789:
707:
566:
4508:
4471:
4431:
4399:
4292:
4260:
4246:
4225:
4211:
4162:
4155:
4074:
3947:
3837:
3802:
3645:
3635:
3606:
3239:
3232:
3037:
2397:
2199:
2164:
240:
47:
17:
2902:
1470:
1074:
627:
86:
4488:
4421:
4285:
4253:
4218:
4204:
4176:
4102:
4081:
3824:
3667:
3627:
3594:
3366:
3352:
3250:
2771:
Restrepo Schild V, Booth MJ, Box SJ, Olof SN, Mahendran KR, Bayley H (April 2017).
1923:
527:
133:
51:
2918:
Chapter 11 - Reconstitution of membrane proteins in phospholipid bilayer nanodiscs
2413:"Stepwise synthesis of giant unilamellar vesicles on a microfluidic assembly line"
1770:
229:
3021:
2846:
2829:
2231:
2214:
1454:
1166:
239:
diffusion within any one region. Later work extended this concept by integrating
4350:
4095:
3848:
3714:
3662:
3657:
3170:
2976:
1113:
970:
402:
397:
are another class of model membranes that are commonly used to purify and study
370:
166:
142:
2098:
Proceedings of the National Academy of Sciences of the United States of America
1296:
Proceedings of the National Academy of Sciences of the United States of America
794:
Proceedings of the National Academy of Sciences of the United States of America
4481:
4428:
4015:
3928:
3756:
3751:
3704:
3697:
3692:
3677:
3672:
3513:
3448:
3197:
3139:
2886:
2870:
741:"Single molecule methods for monitoring changes in bilayer elastic properties"
253:
150:
101:
458:
4088:
4067:
3969:
3761:
3746:
3733:
3485:
3273:
3192:
3187:
3177:
3165:
3104:
2739:
2118:
1316:
814:
422:
391:
337:
137:
3029:
2994:
2953:
2894:
2855:
2814:
2757:
2697:
2662:
2605:
2570:
2535:
2500:
2446:
2389:
2354:
2297:
2240:
2020:
1985:
1872:
1845:
1788:
1747:"Solid supported lipid bilayers: From biophysical studies to sensor design"
1586:
1370:
1335:
1276:
1237:
1174:
1131:
1066:
1031:
988:
903:
868:
774:
519:
476:
443:"Artificial cell mimics as simplified models for the study of cell biology"
2213:
Popplewell JF, Swann MJ, Freeman NJ, McDonnell C, Ford RC (January 2007).
2172:
2137:
2069:
1915:
1731:
1671:
1462:
1427:
833:
725:
584:
252:
substrate material and lipid species but is generally about 1 nm for
4461:
4406:
3993:
3977:
3942:
3792:
3738:
3640:
3584:
3579:
3470:
3431:
3387:
3289:
3094:
417:
394:
274:
113:
43:
1950:
1201:
4493:
4411:
4392:
3952:
3937:
3774:
3652:
3436:
3421:
3124:
3099:
1977:
245:
39:
2796:
2689:
2644:
2597:
2562:
2527:
2428:
2051:
1614:
1578:
1551:
1524:
1497:
1362:
1268:
1229:
1023:
860:
667:
642:
4439:
4382:
4367:
4317:
4047:
3865:
3843:
3540:
3463:
3458:
3223:
3114:
3109:
2482:
619:
511:
257:
109:
97:
1251:
Andrecka J, Spillane KM, Ortega-Arroyo J, Kukura P (December 2013).
739:
Ingolfson H, Kapoor R, Collingwood SA, Andersen OS (November 2008).
154:
black lipid membrane will survive for less than an hour, precluding
4416:
4043:
3987:
3982:
3347:
3339:
3157:
3119:
1907:
1058:
543:"Analysis of the torus surrounding planar lipid bilayer membranes"
336:
300:
288:
228:
165:
85:
63:
2971:. Methods in Molecular Biology. Vol. 1118. pp. 109–30.
2830:"Membrane proteins, lipids and detergents: not just a soap opera"
2094:"VAMP-1: a synaptic vesicle-associated integral membrane protein"
756:
4355:
3959:
3907:
3853:
3545:
3086:
289:
3880:
3050:
2773:"Light-Patterned Current Generation in a Droplet Bilayer Array"
62:. A model bilayer can be made with either synthetic or natural
411:
1802:
Wong JY, Park CK, Seitz M, Israelachvili J (September 1999).
643:"Hard X-Ray Phase Contrast Imaging of Black lipid membranes"
641:
Beerlink A, Mell M, Tolkiehn M, Salditt T (November 2009).
1290:
de Wit G, Danial JS, Kukura P, Wallace MI (October 2015).
945:
Lin WC, Blanchette CD, Ratto TV, Longo ML (January 2006).
414:
spectroscopy where the larger vesicles are not an option.
2920:. Methods in Enzymology. Vol. 464. pp. 211–31.
195:(QCM) to study binding kinetics at the bilayer surface.
50:
or covering various sub-cellular structures like the
307:
group that forms a covalent bond with gold, forming
4336:
4310:
4277:
4133:
4119:
4112:
4059:
4042:
4035:
4003:
3968:
3926:
3919:
3785:
3713:
3626:
3494:
3376:
3365:
3338:
3310:
3260:
3249:
3222:
3211:
3156:
3085:
2834:Biochimica et Biophysica Acta (BBA) - Biomembranes
2219:Biochimica et Biophysica Acta (BBA) - Biomembranes
2215:"Quantifying the effects of melittin on liposomes"
2153:Biochimica et Biophysica Acta (BBA) - Biomembranes
1351:Langmuir: The ACS Journal of Surfaces and Colloids
849:Langmuir: The ACS Journal of Surfaces and Colloids
203:providing complementary data to QCM measurements.
1292:"Dynamic label-free imaging of lipid nanodomains"
212:total internal reflection fluorescence microscopy
2828:Seddon AM, Curnow P, Booth PJ (November 2004).
2092:Trimble WS, Cowan DM, Scheller RH (June 1988).
2709:
2707:
2313:"Lipid rafts reconstituted in model membranes"
441:Salehi-Reyhani A, Ces O, Elani Y (July 2017).
3892:
3062:
1683:
1681:
8:
2714:Villar G, Graham AD, Bayley H (April 2013).
4455:Reverse transcriptase-related cellular gene
1688:Kühner M, Tampé R, Sackmann E (July 1994).
244:reorganization of membrane proteins at the
4116:
4056:
4039:
3923:
3899:
3885:
3877:
3373:
3257:
3219:
3069:
3055:
3047:
1745:Castellana ET, Cremer PS (November 2006).
234:interface as seen from the color gradient.
2943:
2845:
2804:
2747:
2652:
2490:
2436:
2344:
2287:
2230:
2127:
2117:
2059:
1835:
1778:
1721:
1661:
1417:
1325:
1315:
1121:
978:
823:
813:
764:
715:
666:
574:
466:
27:For theoretical models of membranes, see
2586:Journal of the American Chemical Society
2551:Journal of the American Chemical Society
2417:Journal of the American Chemical Society
1190:Journal of the American Chemical Society
282:Tethered bilayer lipid membranes (t-BLM)
4436:Retroelements not elsewhere classified
433:
46:, as opposed to the bilayer of natural
2254:Lei G, MacDonald RC (September 2003).
224:near field scanning optical microscopy
2460:
2458:
2456:
2411:Matosevic S, Paegel BM (March 2011).
2382:10.1146/annurev-biophys-100620-114132
788:Montal M, Mueller P (December 1972).
220:interferometric scattering microscopy
7:
2036:"The mechanism of vesicle formation"
4388:Integrative and conjugative element
293:Diagram showing formation of t-BLM.
182:(AFM) has been used to image lipid
25:
4445:Diversity-generating retroelement
1384:Hollars CW, Dunn RC (July 1998).
745:Journal of Visualized Experiments
447:Experimental Biology and Medicine
4450:Telomerase reverse transcriptase
4022:Microbes with highly unusual DNA
2716:"A tissue-like printed material"
1966:Biochemical Society Transactions
387:Micelles, bicelles and nanodiscs
355:Dual polarisation interferometry
323:measurement across the bilayer.
197:Dual polarisation interferometry
2188:Chemistry and Physics of Lipids
1540:Journal of Physical Chemistry B
1449:(5300). New York, N.Y.: 651–3.
4373:Defective interfering particle
3723:Last universal common ancestor
3318:Defective interfering particle
170:Diagram of a supported bilayer
162:Supported lipid bilayers (SLB)
1:
4361:Clonally transmissible cancer
3859:Clonally transmissible cancer
3295:Satellite-like nucleic acids
2926:10.1016/s0076-6879(09)64011-8
2337:10.1016/S0006-3495(01)76114-0
2280:10.1016/S0006-3495(03)74590-1
2013:10.1016/S0022-2836(64)80115-7
1828:10.1016/s0006-3495(99)76993-6
1771:10.1016/j.surfrep.2006.06.001
1714:10.1016/s0006-3495(94)80472-2
1654:10.1016/S0006-3495(91)82222-6
1410:10.1016/s0006-3495(98)77518-6
932:10.1016/s0013-4686(01)00759-9
708:10.1016/S0006-3495(83)84414-2
682:Andersen OS (February 1983).
567:10.1016/s0006-3495(72)86095-8
3022:10.3109/09687688.2012.693212
2847:10.1016/j.bbamem.2004.04.011
2467:"Droplet interface bilayers"
2232:10.1016/j.bbamem.2006.05.016
2200:10.1016/0009-3084(71)90010-7
2165:10.1016/0005-2736(67)90094-6
2001:Journal of Molecular Biology
1455:10.1126/science.275.5300.651
1167:10.1126/science.288.5463.143
132:. Membrane proteins such as
3914:, and comparable structures
2977:10.1007/978-1-62703-782-2_7
2969:Cell-Free Protein Synthesis
2875:Journal of Biomolecular NMR
2370:Annual Review of Biophysics
1114:10.1529/biophysj.106.081398
971:10.1529/biophysj.105.067066
193:quartz crystal microbalance
82:Black lipid membranes (BLM)
4531:
3415:Class II or DNA transposon
3410:Class I or retrotransposon
3010:Molecular Membrane Biology
2034:Lasic DD (November 1988).
378:Droplet Interface Bilayers
330:
130:voltage gated ion channels
26:
3728:Earliest known life forms
3602:Repeated sequences in DNA
1047:Nature Structural Biology
309:self assembled monolayers
216:surface plasmon resonance
4378:Endogenous viral element
3575:Endogenous viral element
3393:Horizontal gene transfer
459:10.1177/1535370217711441
191:probe is the use of the
58:for the construction of
3272:dsDNA satellite virus (
2887:10.1023/a:1008360022444
2740:10.1126/science.1229495
2119:10.1073/pnas.85.12.4538
2040:The Biochemical Journal
1751:Surface Science Reports
1317:10.1073/pnas.1508483112
815:10.1073/pnas.69.12.3561
647:Applied Physics Letters
541:White SH (April 1972).
180:Atomic force microscopy
3830:Helper dependent virus
3146:Biological dark matter
1873:10.1002/anie.199520561
904:10.1002/adma.200700810
342:
294:
235:
171:
92:
72:characterization tools
3590:Endogenous retrovirus
3563:Origin of replication
3279:ssDNA satellite virus
3269:ssRNA satellite virus
340:
292:
232:
188:mechanical properties
169:
156:long-term experiments
106:partition coefficient
89:
18:Black lipid membranes
4477:Transposable element
4467:Spiegelman's Monster
3534:Secondary chromosome
3529:Extrachromosomal DNA
3405:Transposable element
2516:Analytical Chemistry
2471:Molecular BioSystems
1218:Analytical Chemistry
333:Unilamellar liposome
321:electrical impedance
256:lipids supported on
3770:Model lipid bilayer
3612:Interspersed repeat
2789:2017NatSR...746585R
2732:2013Sci...340...48V
2637:2015NatSR...513726N
2329:2001BpJ....80.1417D
2317:Biophysical Journal
2272:2003BpJ....85.1585L
2260:Biophysical Journal
2110:1988PNAS...85.4538T
1951:10.1021/la00013a029
1900:1997Natur.387..580C
1820:1999BpJ....77.1458W
1808:Biophysical Journal
1763:2006SurSR..61..429C
1706:1994BpJ....67..217K
1694:Biophysical Journal
1646:1991BpJ....59..289J
1634:Biophysical Journal
1402:1998BpJ....75..342H
1390:Biophysical Journal
1308:2015PNAS..11212299D
1202:10.1021/ja00104a001
1159:2000Sci...288..143O
1106:2006BpJ....91..217S
1094:Biophysical Journal
1016:2008NanoL...8..941R
963:2006BpJ....90..228L
951:Biophysical Journal
920:Electrochimica Acta
896:2008AdM....20...84M
806:1972PNAS...69.3561M
700:1983BpJ....41..119A
688:Biophysical Journal
659:2009ApPhL..95t3703B
612:1966Natur.212..718T
559:1972BpJ....12..432W
547:Biophysical Journal
504:1962Natur.194..979M
102:hydrophobic solvent
36:model lipid bilayer
4346:Bio-like structure
4268:Tolecusatellitidae
3080:organic structures
2777:Scientific Reports
2625:Scientific Reports
1978:10.1042/BST0290613
884:Advanced Materials
343:
295:
236:
172:
93:
4502:
4501:
4332:
4331:
4306:
4305:
4302:
4301:
4240:Portogloboviridae
4142:Alphasatellitidae
4036:Non-cellular life
4031:
4030:
3912:non-cellular life
3874:
3873:
3815:Non-cellular life
3622:
3621:
3361:
3360:
3334:
3333:
3288:ssRNA satellite (
2986:978-1-62703-781-5
2797:10.1038/srep46585
2690:10.1021/la203081v
2645:10.1038/srep13726
2598:10.1021/ja200128n
2563:10.1021/ja802089s
2528:10.1021/ac0613479
2429:10.1021/ja109137s
2052:10.1042/bj2560001
1867:(18): 2056–2058.
1615:10.1021/la950580r
1579:10.1021/la802726u
1552:10.1021/jp035543k
1525:10.1021/la0263413
1498:10.1021/la9711701
1363:10.1021/la047654w
1302:(40): 12299–303.
1269:10.1021/nn403367c
1230:10.1021/ac800027s
1024:10.1021/nl080080l
861:10.1021/la703704x
668:10.1063/1.3263946
606:(5063): 718–719.
453:(13): 1309–1317.
399:membrane proteins
351:x-ray diffraction
201:phase transitions
76:membrane proteins
56:synthetic biology
16:(Redirected from
4522:
4515:Membrane biology
4117:
4057:
4040:
3924:
3901:
3894:
3887:
3878:
3551:Gene duplication
3374:
3370:self-replication
3258:
3220:
3078:Self-replicating
3071:
3064:
3057:
3048:
3042:
3041:
3005:
2999:
2998:
2964:
2958:
2957:
2947:
2913:
2907:
2906:
2866:
2860:
2859:
2849:
2825:
2819:
2818:
2808:
2768:
2762:
2761:
2751:
2711:
2702:
2701:
2684:(23): 14335–42.
2673:
2667:
2666:
2656:
2616:
2610:
2609:
2581:
2575:
2574:
2546:
2540:
2539:
2511:
2505:
2504:
2494:
2483:10.1039/b808893d
2477:(12): 1191–208.
2462:
2451:
2450:
2440:
2408:
2402:
2401:
2365:
2359:
2358:
2348:
2308:
2302:
2301:
2291:
2251:
2245:
2244:
2234:
2210:
2204:
2203:
2183:
2177:
2176:
2148:
2142:
2141:
2131:
2121:
2089:
2083:
2080:
2074:
2073:
2063:
2031:
2025:
2024:
1996:
1990:
1989:
1961:
1955:
1954:
1934:
1928:
1927:
1883:
1877:
1876:
1856:
1850:
1849:
1839:
1799:
1793:
1792:
1782:
1742:
1736:
1735:
1725:
1685:
1676:
1675:
1665:
1625:
1619:
1618:
1609:(5): 1343–1350.
1597:
1591:
1590:
1562:
1556:
1555:
1535:
1529:
1528:
1519:(5): 1624–1631.
1508:
1502:
1501:
1481:
1475:
1474:
1438:
1432:
1431:
1421:
1381:
1375:
1374:
1346:
1340:
1339:
1329:
1319:
1287:
1281:
1280:
1263:(12): 10662–70.
1248:
1242:
1241:
1212:
1206:
1205:
1196:(25): 11209–12.
1185:
1179:
1178:
1142:
1136:
1135:
1125:
1085:
1079:
1078:
1042:
1036:
1035:
999:
993:
992:
982:
942:
936:
935:
914:
908:
907:
879:
873:
872:
844:
838:
837:
827:
817:
785:
779:
778:
768:
736:
730:
729:
719:
679:
673:
672:
670:
638:
632:
631:
620:10.1038/212718a0
595:
589:
588:
578:
538:
532:
531:
512:10.1038/194979a0
498:(4832): 979–80.
487:
481:
480:
470:
438:
210:methods such as
208:Evanescent field
184:phase separation
60:artificial cells
21:
4530:
4529:
4525:
4524:
4523:
4521:
4520:
4519:
4505:
4504:
4503:
4498:
4338:
4328:
4298:
4273:
4184:Finnlakeviridae
4129:
4108:
4050:
4046:
4027:
3999:
3964:
3915:
3905:
3875:
3870:
3820:Synthetic virus
3808:Artificial cell
3781:
3709:
3618:
3507:RNA replication
3502:DNA replication
3490:
3481:Group II intron
3379:
3369:
3357:
3348:Mammalian prion
3330:
3306:
3285:dsRNA satellite
3282:ssDNA satellite
3252:
3245:
3214:
3207:
3152:
3081:
3075:
3045:
3007:
3006:
3002:
2987:
2966:
2965:
2961:
2936:
2915:
2914:
2910:
2868:
2867:
2863:
2840:(1–2): 105–17.
2827:
2826:
2822:
2770:
2769:
2765:
2726:(6128): 48–52.
2713:
2712:
2705:
2675:
2674:
2670:
2618:
2617:
2613:
2583:
2582:
2578:
2548:
2547:
2543:
2522:(24): 8169–74.
2513:
2512:
2508:
2464:
2463:
2454:
2423:(9): 2798–800.
2410:
2409:
2405:
2367:
2366:
2362:
2310:
2309:
2305:
2253:
2252:
2248:
2212:
2211:
2207:
2185:
2184:
2180:
2150:
2149:
2145:
2104:(12): 4538–42.
2091:
2090:
2086:
2081:
2077:
2033:
2032:
2028:
1998:
1997:
1993:
1972:(Pt 4): 613–7.
1963:
1962:
1958:
1936:
1935:
1931:
1894:(6633): 580–3.
1885:
1884:
1880:
1858:
1857:
1853:
1801:
1800:
1796:
1757:(10): 429–444.
1744:
1743:
1739:
1687:
1686:
1679:
1627:
1626:
1622:
1599:
1598:
1594:
1573:(22): 12734–7.
1564:
1563:
1559:
1537:
1536:
1532:
1510:
1509:
1505:
1492:(12): 3347–50.
1483:
1482:
1478:
1440:
1439:
1435:
1383:
1382:
1378:
1348:
1347:
1343:
1289:
1288:
1284:
1250:
1249:
1245:
1224:(10): 3666–76.
1214:
1213:
1209:
1187:
1186:
1182:
1153:(5463): 143–6.
1144:
1143:
1139:
1087:
1086:
1082:
1044:
1043:
1039:
1001:
1000:
996:
944:
943:
939:
916:
915:
911:
881:
880:
876:
846:
845:
841:
787:
786:
782:
738:
737:
733:
681:
680:
676:
640:
639:
635:
597:
596:
592:
540:
539:
535:
489:
488:
484:
440:
439:
435:
431:
389:
380:
335:
329:
297:
284:
164:
84:
32:
29:Membrane models
23:
22:
15:
12:
11:
5:
4528:
4526:
4518:
4517:
4507:
4506:
4500:
4499:
4497:
4496:
4491:
4486:
4485:
4484:
4474:
4469:
4464:
4459:
4458:
4457:
4452:
4447:
4442:
4434:
4426:
4425:
4424:
4414:
4409:
4404:
4395:
4390:
4385:
4380:
4375:
4370:
4365:
4364:
4363:
4358:
4348:
4342:
4340:
4334:
4333:
4330:
4329:
4327:
4326:
4321:
4314:
4312:
4308:
4307:
4304:
4303:
4300:
4299:
4297:
4296:
4289:
4281:
4279:
4275:
4274:
4272:
4271:
4264:
4257:
4250:
4243:
4236:
4233:Polydnaviridae
4229:
4222:
4215:
4208:
4201:
4198:Globuloviridae
4194:
4191:Fuselloviridae
4187:
4180:
4173:
4170:Bicaudaviridae
4166:
4159:
4152:
4149:Ampullaviridae
4145:
4137:
4135:
4131:
4130:
4126:Naldaviricetes
4123:
4121:
4114:
4110:
4109:
4107:
4106:
4099:
4092:
4085:
4078:
4071:
4063:
4061:
4054:
4037:
4033:
4032:
4029:
4028:
4026:
4025:
4019:
4011:
4009:
4006:Incertae sedis
4001:
4000:
3998:
3997:
3990:
3985:
3980:
3974:
3972:
3966:
3965:
3963:
3962:
3957:
3956:
3955:
3950:
3940:
3934:
3932:
3921:
3917:
3916:
3906:
3904:
3903:
3896:
3889:
3881:
3872:
3871:
3869:
3868:
3863:
3862:
3861:
3856:
3846:
3840:
3834:
3833:
3832:
3827:
3817:
3812:
3811:
3810:
3805:
3795:
3789:
3787:
3783:
3782:
3780:
3779:
3778:
3777:
3772:
3764:
3759:
3754:
3749:
3743:
3742:
3741:
3730:
3725:
3719:
3717:
3711:
3710:
3708:
3707:
3702:
3701:
3700:
3695:
3687:
3685:Kappa organism
3682:
3681:
3680:
3675:
3670:
3665:
3660:
3650:
3649:
3648:
3643:
3632:
3630:
3624:
3623:
3620:
3619:
3617:
3616:
3615:
3614:
3609:
3599:
3598:
3597:
3592:
3587:
3582:
3572:
3571:
3570:
3560:
3559:
3558:
3556:Non-coding DNA
3553:
3548:
3538:
3537:
3536:
3531:
3526:
3521:
3511:
3510:
3509:
3498:
3496:
3492:
3491:
3489:
3488:
3483:
3478:
3476:Group I intron
3473:
3468:
3467:
3466:
3456:
3455:
3454:
3451:
3442:
3439:
3434:
3429:
3419:
3418:
3417:
3412:
3402:
3401:
3400:
3398:Genomic island
3395:
3384:
3382:
3378:Mobile genetic
3371:
3363:
3362:
3359:
3358:
3356:
3355:
3350:
3344:
3342:
3336:
3335:
3332:
3331:
3329:
3328:
3327:
3326:
3323:
3314:
3312:
3308:
3307:
3305:
3304:
3303:
3302:
3299:
3293:
3286:
3283:
3280:
3277:
3270:
3266:
3264:
3255:
3247:
3246:
3244:
3243:
3236:
3228:
3226:
3217:
3209:
3208:
3206:
3205:
3203:dsDNA-RT virus
3200:
3198:ssRNA-RT virus
3195:
3193:(−)ssRNA virus
3190:
3188:(+)ssRNA virus
3185:
3180:
3175:
3174:
3173:
3162:
3160:
3154:
3153:
3151:
3150:
3149:
3148:
3143:
3133:Incertae sedis
3129:
3128:
3127:
3122:
3117:
3112:
3102:
3097:
3091:
3089:
3083:
3082:
3076:
3074:
3073:
3066:
3059:
3051:
3044:
3043:
3000:
2985:
2959:
2934:
2908:
2861:
2820:
2763:
2703:
2668:
2611:
2592:(24): 9370–5.
2576:
2557:(18): 5878–9.
2541:
2506:
2452:
2403:
2360:
2323:(3): 1417–28.
2303:
2266:(3): 1585–99.
2246:
2205:
2178:
2143:
2084:
2075:
2026:
2007:(5): 660–668.
1991:
1956:
1929:
1878:
1851:
1814:(3): 1458–68.
1794:
1737:
1677:
1620:
1592:
1557:
1530:
1503:
1476:
1433:
1376:
1357:(4): 1377–88.
1341:
1282:
1243:
1207:
1180:
1137:
1080:
1037:
994:
937:
926:(5): 791–798.
909:
874:
839:
800:(12): 3561–6.
780:
731:
674:
653:(20): 203703.
633:
590:
533:
482:
432:
430:
427:
388:
385:
379:
376:
328:
325:
283:
280:
163:
160:
83:
80:
48:cell membranes
24:
14:
13:
10:
9:
6:
4:
3:
2:
4527:
4516:
4513:
4512:
4510:
4495:
4492:
4490:
4487:
4483:
4480:
4479:
4478:
4475:
4473:
4472:Tandem repeat
4470:
4468:
4465:
4463:
4460:
4456:
4453:
4451:
4448:
4446:
4443:
4441:
4438:
4437:
4435:
4433:
4430:
4427:
4423:
4420:
4419:
4418:
4415:
4413:
4410:
4408:
4405:
4402:
4401:
4400:Nanobacterium
4396:
4394:
4391:
4389:
4386:
4384:
4381:
4379:
4376:
4374:
4371:
4369:
4366:
4362:
4359:
4357:
4354:
4353:
4352:
4349:
4347:
4344:
4343:
4341:
4335:
4325:
4322:
4319:
4316:
4315:
4313:
4309:
4295:
4294:
4293:Rhizidiovirus
4290:
4288:
4287:
4283:
4282:
4280:
4276:
4270:
4269:
4265:
4263:
4262:
4261:Thaspiviridae
4258:
4256:
4255:
4251:
4249:
4248:
4247:Pospiviroidae
4244:
4242:
4241:
4237:
4235:
4234:
4230:
4228:
4227:
4226:Plasmaviridae
4223:
4221:
4220:
4216:
4214:
4213:
4212:Halspiviridae
4209:
4207:
4206:
4202:
4200:
4199:
4195:
4193:
4192:
4188:
4186:
4185:
4181:
4179:
4178:
4174:
4172:
4171:
4167:
4165:
4164:
4163:Avsunviroidae
4160:
4158:
4157:
4156:Anelloviridae
4153:
4151:
4150:
4146:
4144:
4143:
4139:
4138:
4136:
4132:
4128:
4127:
4122:
4118:
4115:
4111:
4105:
4104:
4100:
4098:
4097:
4093:
4091:
4090:
4086:
4084:
4083:
4079:
4077:
4076:
4075:Duplodnaviria
4072:
4070:
4069:
4065:
4064:
4062:
4058:
4055:
4053:
4049:
4045:
4041:
4038:
4034:
4023:
4020:
4018:
4017:
4013:
4012:
4010:
4008:
4007:
4002:
3995:
3991:
3989:
3986:
3984:
3981:
3979:
3976:
3975:
3973:
3971:
3967:
3961:
3958:
3954:
3951:
3949:
3948:Mitochondrion
3946:
3945:
3944:
3941:
3939:
3936:
3935:
3933:
3930:
3925:
3922:
3920:Cellular life
3918:
3913:
3909:
3902:
3897:
3895:
3890:
3888:
3883:
3882:
3879:
3867:
3864:
3860:
3857:
3855:
3852:
3851:
3850:
3847:
3845:
3841:
3839:
3838:Nanobacterium
3835:
3831:
3828:
3826:
3823:
3822:
3821:
3818:
3816:
3813:
3809:
3806:
3804:
3803:Cell division
3801:
3800:
3799:
3796:
3794:
3791:
3790:
3788:
3784:
3776:
3773:
3771:
3768:
3767:
3765:
3763:
3760:
3758:
3755:
3753:
3750:
3748:
3744:
3740:
3737:
3736:
3735:
3731:
3729:
3726:
3724:
3721:
3720:
3718:
3716:
3712:
3706:
3703:
3699:
3696:
3694:
3691:
3690:
3688:
3686:
3683:
3679:
3676:
3674:
3671:
3669:
3666:
3664:
3661:
3659:
3656:
3655:
3654:
3651:
3647:
3646:Hydrogenosome
3644:
3642:
3639:
3638:
3637:
3636:Mitochondrion
3634:
3633:
3631:
3629:
3628:Endosymbiosis
3625:
3613:
3610:
3608:
3607:Tandem repeat
3605:
3604:
3603:
3600:
3596:
3593:
3591:
3588:
3586:
3583:
3581:
3578:
3577:
3576:
3573:
3569:
3566:
3565:
3564:
3561:
3557:
3554:
3552:
3549:
3547:
3544:
3543:
3542:
3539:
3535:
3532:
3530:
3527:
3525:
3522:
3520:
3517:
3516:
3515:
3512:
3508:
3505:
3504:
3503:
3500:
3499:
3497:
3495:Other aspects
3493:
3487:
3484:
3482:
3479:
3477:
3474:
3472:
3469:
3465:
3462:
3461:
3460:
3457:
3452:
3450:
3446:
3443:
3440:
3438:
3435:
3433:
3430:
3428:
3425:
3424:
3423:
3420:
3416:
3413:
3411:
3408:
3407:
3406:
3403:
3399:
3396:
3394:
3391:
3390:
3389:
3386:
3385:
3383:
3381:
3375:
3372:
3368:
3364:
3354:
3351:
3349:
3346:
3345:
3343:
3341:
3337:
3324:
3321:
3320:
3319:
3316:
3315:
3313:
3309:
3300:
3297:
3296:
3294:
3291:
3287:
3284:
3281:
3278:
3275:
3271:
3268:
3267:
3265:
3263:
3259:
3256:
3254:
3248:
3242:
3241:
3240:Avsunviroidae
3237:
3235:
3234:
3233:Pospiviroidae
3230:
3229:
3227:
3225:
3221:
3218:
3216:
3210:
3204:
3201:
3199:
3196:
3194:
3191:
3189:
3186:
3184:
3181:
3179:
3176:
3172:
3169:
3168:
3167:
3164:
3163:
3161:
3159:
3155:
3147:
3144:
3142:
3141:
3137:
3136:
3135:
3134:
3130:
3126:
3123:
3121:
3118:
3116:
3113:
3111:
3108:
3107:
3106:
3103:
3101:
3098:
3096:
3093:
3092:
3090:
3088:
3087:Cellular life
3084:
3079:
3072:
3067:
3065:
3060:
3058:
3053:
3052:
3049:
3039:
3035:
3031:
3027:
3023:
3019:
3015:
3011:
3004:
3001:
2996:
2992:
2988:
2982:
2978:
2974:
2970:
2963:
2960:
2955:
2951:
2946:
2941:
2937:
2935:9780123749697
2931:
2927:
2923:
2919:
2912:
2909:
2904:
2900:
2896:
2892:
2888:
2884:
2881:(4): 387–91.
2880:
2876:
2872:
2869:Cavagnero S,
2865:
2862:
2857:
2853:
2848:
2843:
2839:
2835:
2831:
2824:
2821:
2816:
2812:
2807:
2802:
2798:
2794:
2790:
2786:
2782:
2778:
2774:
2767:
2764:
2759:
2755:
2750:
2745:
2741:
2737:
2733:
2729:
2725:
2721:
2717:
2710:
2708:
2704:
2699:
2695:
2691:
2687:
2683:
2679:
2672:
2669:
2664:
2660:
2655:
2650:
2646:
2642:
2638:
2634:
2630:
2626:
2622:
2615:
2612:
2607:
2603:
2599:
2595:
2591:
2587:
2580:
2577:
2572:
2568:
2564:
2560:
2556:
2552:
2545:
2542:
2537:
2533:
2529:
2525:
2521:
2517:
2510:
2507:
2502:
2498:
2493:
2488:
2484:
2480:
2476:
2472:
2468:
2461:
2459:
2457:
2453:
2448:
2444:
2439:
2434:
2430:
2426:
2422:
2418:
2414:
2407:
2404:
2399:
2395:
2391:
2387:
2383:
2379:
2375:
2371:
2364:
2361:
2356:
2352:
2347:
2342:
2338:
2334:
2330:
2326:
2322:
2318:
2314:
2307:
2304:
2299:
2295:
2290:
2285:
2281:
2277:
2273:
2269:
2265:
2261:
2257:
2250:
2247:
2242:
2238:
2233:
2228:
2224:
2220:
2216:
2209:
2206:
2201:
2197:
2194:(4): 324–35.
2193:
2189:
2182:
2179:
2174:
2170:
2166:
2162:
2159:(4): 624–38.
2158:
2154:
2147:
2144:
2139:
2135:
2130:
2125:
2120:
2115:
2111:
2107:
2103:
2099:
2095:
2088:
2085:
2079:
2076:
2071:
2067:
2062:
2057:
2053:
2049:
2045:
2041:
2037:
2030:
2027:
2022:
2018:
2014:
2010:
2006:
2002:
1995:
1992:
1987:
1983:
1979:
1975:
1971:
1967:
1960:
1957:
1952:
1948:
1944:
1940:
1933:
1930:
1925:
1921:
1917:
1913:
1909:
1908:10.1038/42432
1905:
1901:
1897:
1893:
1889:
1882:
1879:
1874:
1870:
1866:
1862:
1855:
1852:
1847:
1843:
1838:
1833:
1829:
1825:
1821:
1817:
1813:
1809:
1805:
1798:
1795:
1790:
1786:
1781:
1776:
1772:
1768:
1764:
1760:
1756:
1752:
1748:
1741:
1738:
1733:
1729:
1724:
1719:
1715:
1711:
1707:
1703:
1700:(1): 217–26.
1699:
1695:
1691:
1684:
1682:
1678:
1673:
1669:
1664:
1659:
1655:
1651:
1647:
1643:
1640:(2): 289–94.
1639:
1635:
1631:
1624:
1621:
1616:
1612:
1608:
1604:
1596:
1593:
1588:
1584:
1580:
1576:
1572:
1568:
1561:
1558:
1553:
1549:
1546:(2): 649–57.
1545:
1541:
1534:
1531:
1526:
1522:
1518:
1514:
1507:
1504:
1499:
1495:
1491:
1487:
1480:
1477:
1472:
1468:
1464:
1460:
1456:
1452:
1448:
1444:
1437:
1434:
1429:
1425:
1420:
1415:
1411:
1407:
1403:
1399:
1396:(1): 342–53.
1395:
1391:
1387:
1380:
1377:
1372:
1368:
1364:
1360:
1356:
1352:
1345:
1342:
1337:
1333:
1328:
1323:
1318:
1313:
1309:
1305:
1301:
1297:
1293:
1286:
1283:
1278:
1274:
1270:
1266:
1262:
1258:
1254:
1247:
1244:
1239:
1235:
1231:
1227:
1223:
1219:
1211:
1208:
1203:
1199:
1195:
1191:
1184:
1181:
1176:
1172:
1168:
1164:
1160:
1156:
1152:
1148:
1141:
1138:
1133:
1129:
1124:
1119:
1115:
1111:
1107:
1103:
1100:(1): 217–26.
1099:
1095:
1091:
1084:
1081:
1076:
1072:
1068:
1064:
1060:
1059:10.1038/78929
1056:
1052:
1048:
1041:
1038:
1033:
1029:
1025:
1021:
1017:
1013:
1009:
1005:
998:
995:
990:
986:
981:
976:
972:
968:
964:
960:
957:(1): 228–37.
956:
952:
948:
941:
938:
933:
929:
925:
921:
913:
910:
905:
901:
897:
893:
889:
885:
878:
875:
870:
866:
862:
858:
855:(9): 4952–8.
854:
850:
843:
840:
835:
831:
826:
821:
816:
811:
807:
803:
799:
795:
791:
784:
781:
776:
772:
767:
762:
758:
754:
751:(21): e1032.
750:
746:
742:
735:
732:
727:
723:
718:
713:
709:
705:
701:
697:
694:(2): 119–33.
693:
689:
685:
678:
675:
669:
664:
660:
656:
652:
648:
644:
637:
634:
629:
625:
621:
617:
613:
609:
605:
601:
594:
591:
586:
582:
577:
572:
568:
564:
560:
556:
553:(4): 432–45.
552:
548:
544:
537:
534:
529:
525:
521:
517:
513:
509:
505:
501:
497:
493:
486:
483:
478:
474:
469:
464:
460:
456:
452:
448:
444:
437:
434:
428:
426:
424:
419:
415:
413:
407:
404:
400:
396:
393:
386:
384:
377:
375:
372:
366:
364:
360:
356:
352:
347:
339:
334:
326:
324:
322:
316:
312:
310:
306:
302:
291:
287:
281:
279:
276:
271:
265:
263:
259:
255:
249:
247:
242:
241:microfluidics
231:
227:
225:
221:
217:
213:
209:
204:
202:
198:
194:
189:
185:
181:
176:
168:
161:
159:
157:
152:
146:
144:
139:
135:
131:
126:
121:
117:
115:
111:
107:
103:
99:
88:
81:
79:
77:
73:
67:
65:
61:
57:
53:
49:
45:
41:
37:
30:
19:
4489:Transpoviron
4422:Fungal prion
4398:
4291:
4286:Dinodnavirus
4284:
4266:
4259:
4254:Spiraviridae
4252:
4245:
4238:
4231:
4224:
4219:Ovaliviridae
4217:
4210:
4205:Guttaviridae
4203:
4196:
4189:
4182:
4177:Clavaviridae
4175:
4168:
4161:
4154:
4147:
4140:
4124:
4103:Varidnaviria
4101:
4094:
4087:
4082:Monodnaviria
4080:
4073:
4066:
4014:
4004:
3825:Viral vector
3769:
3668:Gerontoplast
3595:Transpoviron
3367:Nucleic acid
3353:Fungal prion
3251:Helper-virus
3238:
3231:
3138:
3131:
3016:(1): 75–89.
3013:
3009:
3003:
2968:
2962:
2917:
2911:
2878:
2874:
2864:
2837:
2833:
2823:
2780:
2776:
2766:
2723:
2719:
2681:
2677:
2671:
2628:
2624:
2614:
2589:
2585:
2579:
2554:
2550:
2544:
2519:
2515:
2509:
2474:
2470:
2420:
2416:
2406:
2373:
2369:
2363:
2320:
2316:
2306:
2263:
2259:
2249:
2225:(1): 13–20.
2222:
2218:
2208:
2191:
2187:
2181:
2156:
2152:
2146:
2101:
2097:
2087:
2078:
2043:
2039:
2029:
2004:
2000:
1994:
1969:
1965:
1959:
1942:
1938:
1932:
1891:
1887:
1881:
1864:
1860:
1854:
1811:
1807:
1797:
1754:
1750:
1740:
1697:
1693:
1637:
1633:
1623:
1606:
1602:
1595:
1570:
1566:
1560:
1543:
1539:
1533:
1516:
1512:
1506:
1489:
1485:
1479:
1446:
1442:
1436:
1393:
1389:
1379:
1354:
1350:
1344:
1299:
1295:
1285:
1260:
1256:
1246:
1221:
1217:
1210:
1193:
1189:
1183:
1150:
1146:
1140:
1097:
1093:
1083:
1053:(9): 715–8.
1050:
1046:
1040:
1010:(3): 941–4.
1007:
1004:Nano Letters
1003:
997:
954:
950:
940:
923:
919:
912:
887:
883:
877:
852:
848:
842:
797:
793:
783:
757:10.3791/1032
748:
744:
734:
691:
687:
677:
650:
646:
636:
603:
599:
593:
550:
546:
536:
495:
491:
485:
450:
446:
436:
416:
408:
390:
381:
367:
348:
344:
317:
313:
296:
285:
266:
254:zwitterionic
250:
237:
205:
177:
173:
147:
134:ion channels
122:
118:
94:
91:the chamber.
68:
35:
33:
4432:microsphere
4351:Cancer cell
4096:Ribozyviria
3849:Cancer cell
3715:Abiogenesis
3663:Chromoplast
3658:Chloroplast
3441:Degradative
3183:dsRNA virus
3178:ssDNA virus
3171:Giant virus
3166:dsDNA virus
2376:: 525–548.
2046:(1): 1–11.
1945:: 197–210.
1861:Angew. Chem
890:(1): 84–9.
403:amphipathic
371:lipid rafts
319:enables ac
248:interface.
214:(TIRF) and
143:patch clamp
4482:Retroposon
4429:Proteinoid
4339:structures
4337:Comparable
4113:Unassigned
4016:Parakaryon
3929:Prokaryota
3757:Proteinoid
3752:Coacervate
3705:Nitroplast
3698:Trophosome
3693:Bacteriome
3678:Apicoplast
3673:Leucoplast
3514:Chromosome
3432:Resistance
3140:Parakaryon
429:References
331:See also:
305:disulphide
151:chloroform
125:interferes
42:assembled
4089:Riboviria
4068:Adnaviria
4052:Satellite
3970:Eukaryota
3766:Research
3747:Protocell
3486:Retrozyme
3445:Virulence
3427:Fertility
3274:Virophage
3262:Satellite
3253:dependent
3105:Eukaryota
3038:207503256
2783:: 46585.
2631:: 13726.
2398:232131463
418:Nanodiscs
392:Detergent
270:denatured
138:detergent
4509:Category
4462:Ribozyme
4407:Phagemid
4134:Families
3994:Protista
3978:Animalia
3943:Bacteria
3793:Organism
3786:See also
3762:Sulphobe
3739:Ribozyme
3734:RNA life
3641:Mitosome
3585:Prophage
3580:Provirus
3568:Replicon
3524:Circular
3471:Phagemid
3388:Mobilome
3380:elements
3290:Virusoid
3213:Subviral
3125:Protista
3110:Animalia
3095:Bacteria
3030:22716775
2995:24395412
2954:19903557
2903:22774774
2895:10353198
2871:Dyson HJ
2856:15519311
2815:28417964
2758:23559243
2698:21978255
2678:Langmuir
2663:26348441
2606:21591742
2571:18407631
2536:17165804
2501:19396383
2447:21309555
2390:33667121
2355:11222302
2298:12944275
2241:17092481
2021:14187392
1986:11498038
1939:Langmuir
1846:10465756
1789:32287559
1603:Langmuir
1587:18942863
1567:Langmuir
1513:Langmuir
1486:Langmuir
1471:30939780
1371:15697284
1336:26401022
1277:24251388
1257:ACS Nano
1238:18422336
1175:10753119
1132:16617084
1075:20571172
1067:10966636
1032:18254602
989:16214871
869:18370435
775:19066527
628:34363724
520:14476933
477:28580796
395:micelles
365:assays.
327:Vesicles
275:hydrogel
246:synaptic
114:squalene
44:in vitro
4494:Xenobot
4412:Plasmid
4393:Jeewanu
4324:Obelisk
4120:Classes
3988:Plantae
3953:Plastid
3938:Archaea
3775:Jeewanu
3689:Organs
3653:Plastid
3453:Cryptic
3422:Plasmid
3120:Plantae
3100:Archaea
2945:4196316
2806:5394532
2785:Bibcode
2749:3750497
2728:Bibcode
2720:Science
2654:4562232
2633:Bibcode
2492:2763081
2438:3048828
2346:1301333
2325:Bibcode
2289:1303334
2268:Bibcode
2173:4167394
2138:3380805
2106:Bibcode
2070:3066342
2061:1135360
1924:4348659
1916:9177344
1896:Bibcode
1837:1300433
1816:Bibcode
1780:7114318
1759:Bibcode
1732:7918990
1723:1225352
1702:Bibcode
1672:2009353
1663:1281145
1642:Bibcode
1463:9005848
1443:Science
1428:9649391
1419:1299703
1398:Bibcode
1327:4603517
1304:Bibcode
1155:Bibcode
1147:Science
1123:1479081
1102:Bibcode
1012:Bibcode
980:1367021
959:Bibcode
892:Bibcode
834:4509315
802:Bibcode
766:2954507
726:6188500
717:1329161
696:Bibcode
655:Bibcode
608:Bibcode
585:5019479
576:1484121
555:Bibcode
528:2110051
500:Bibcode
468:5528198
361:-based
311:(SAM).
52:nucleus
40:bilayer
38:is any
4440:Retron
4383:Fosmid
4368:Cosmid
4318:Nanobe
4278:Genera
4060:Realms
4048:Viroid
3866:Virome
3844:Nanobe
3541:Genome
3519:Linear
3464:Fosmid
3459:Cosmid
3224:Viroid
3215:agents
3036:
3028:
2993:
2983:
2952:
2942:
2932:
2901:
2893:
2854:
2813:
2803:
2756:
2746:
2696:
2661:
2651:
2604:
2569:
2534:
2499:
2489:
2445:
2435:
2396:
2388:
2353:
2343:
2296:
2286:
2239:
2171:
2136:
2129:280466
2126:
2068:
2058:
2019:
1984:
1922:
1914:
1888:Nature
1844:
1834:
1787:
1777:
1730:
1720:
1670:
1660:
1585:
1469:
1461:
1426:
1416:
1369:
1334:
1324:
1275:
1236:
1173:
1130:
1120:
1073:
1065:
1030:
987:
977:
867:
832:
825:389821
822:
773:
763:
724:
714:
626:
600:Nature
583:
573:
526:
518:
492:Nature
475:
465:
363:fusion
264:data.
258:silica
110:decane
98:Teflon
64:lipids
4417:Prion
4311:Other
4044:Virus
3983:Fungi
3340:Prion
3311:Other
3158:Virus
3115:Fungi
3034:S2CID
2899:S2CID
2394:S2CID
1920:S2CID
1467:S2CID
1071:S2CID
624:S2CID
524:S2CID
301:thiol
4356:HeLa
3960:LUCA
3908:Life
3854:HeLa
3798:Cell
3546:Gene
3026:PMID
2991:PMID
2981:ISBN
2950:PMID
2930:ISBN
2891:PMID
2852:PMID
2838:1666
2811:PMID
2754:PMID
2694:PMID
2659:PMID
2602:PMID
2567:PMID
2532:PMID
2497:PMID
2443:PMID
2386:PMID
2351:PMID
2294:PMID
2237:PMID
2223:1768
2169:PMID
2134:PMID
2066:PMID
2017:PMID
1982:PMID
1912:PMID
1842:PMID
1785:PMID
1728:PMID
1668:PMID
1583:PMID
1459:PMID
1424:PMID
1367:PMID
1332:PMID
1273:PMID
1234:PMID
1171:PMID
1128:PMID
1063:PMID
1028:PMID
985:PMID
865:PMID
830:PMID
771:PMID
722:PMID
581:PMID
516:PMID
473:PMID
359:FRET
262:FRAP
112:and
4320:(?)
4024:(?)
3437:Col
3325:DNA
3322:RNA
3301:DNA
3298:RNA
3018:doi
2973:doi
2940:PMC
2922:doi
2883:doi
2842:doi
2801:PMC
2793:doi
2744:PMC
2736:doi
2724:340
2686:doi
2649:PMC
2641:doi
2594:doi
2590:133
2559:doi
2555:130
2524:doi
2487:PMC
2479:doi
2433:PMC
2425:doi
2421:133
2378:doi
2341:PMC
2333:doi
2284:PMC
2276:doi
2227:doi
2196:doi
2161:doi
2157:135
2124:PMC
2114:doi
2056:PMC
2048:doi
2044:256
2009:doi
1974:doi
1947:doi
1904:doi
1892:387
1869:doi
1832:PMC
1824:doi
1775:PMC
1767:doi
1718:PMC
1710:doi
1658:PMC
1650:doi
1611:doi
1575:doi
1548:doi
1544:108
1521:doi
1494:doi
1451:doi
1447:275
1414:PMC
1406:doi
1359:doi
1322:PMC
1312:doi
1300:112
1265:doi
1226:doi
1198:doi
1194:116
1163:doi
1151:288
1118:PMC
1110:doi
1055:doi
1020:doi
975:PMC
967:doi
928:doi
900:doi
857:doi
820:PMC
810:doi
761:PMC
753:doi
712:PMC
704:doi
663:doi
616:doi
604:212
571:PMC
563:doi
508:doi
496:194
463:PMC
455:doi
451:242
412:NMR
303:or
4511::
3910:,
3449:Ti
3032:.
3024:.
3014:30
3012:.
2989:.
2979:.
2948:.
2938:.
2928:.
2897:.
2889:.
2879:13
2877:.
2850:.
2836:.
2832:.
2809:.
2799:.
2791:.
2779:.
2775:.
2752:.
2742:.
2734:.
2722:.
2718:.
2706:^
2692:.
2682:27
2680:.
2657:.
2647:.
2639:.
2627:.
2623:.
2600:.
2588:.
2565:.
2553:.
2530:.
2520:78
2518:.
2495:.
2485:.
2473:.
2469:.
2455:^
2441:.
2431:.
2419:.
2415:.
2392:.
2384:.
2374:50
2372:.
2349:.
2339:.
2331:.
2321:80
2319:.
2315:.
2292:.
2282:.
2274:.
2264:85
2262:.
2258:.
2235:.
2221:.
2217:.
2190:.
2167:.
2155:.
2132:.
2122:.
2112:.
2102:85
2100:.
2096:.
2064:.
2054:.
2042:.
2038:.
2015:.
2003:.
1980:.
1970:29
1968:.
1943:10
1941:.
1918:.
1910:.
1902:.
1890:.
1865:34
1863:.
1840:.
1830:.
1822:.
1812:77
1810:.
1806:.
1783:.
1773:.
1765:.
1755:61
1753:.
1749:.
1726:.
1716:.
1708:.
1698:67
1696:.
1692:.
1680:^
1666:.
1656:.
1648:.
1638:59
1636:.
1632:.
1607:12
1605:.
1581:.
1571:24
1569:.
1542:.
1517:19
1515:.
1490:14
1488:.
1465:.
1457:.
1445:.
1422:.
1412:.
1404:.
1394:75
1392:.
1388:.
1365:.
1355:21
1353:.
1330:.
1320:.
1310:.
1298:.
1294:.
1271:.
1259:.
1255:.
1232:.
1222:80
1220:.
1192:.
1169:.
1161:.
1149:.
1126:.
1116:.
1108:.
1098:91
1096:.
1092:.
1069:.
1061:.
1049:.
1026:.
1018:.
1006:.
983:.
973:.
965:.
955:90
953:.
949:.
924:47
922:.
898:.
888:20
886:.
863:.
853:24
851:.
828:.
818:.
808:.
798:69
796:.
792:.
769:.
759:.
749:21
747:.
743:.
720:.
710:.
702:.
692:41
690:.
686:.
661:.
651:95
649:.
645:.
622:.
614:.
602:.
579:.
569:.
561:.
551:12
549:.
545:.
522:.
514:.
506:.
494:.
471:.
461:.
449:.
445:.
423:nm
116:.
78:.
34:A
4403:"
4397:"
3996:'
3992:'
3931:"
3927:"
3900:e
3893:t
3886:v
3842:?
3836:?
3745:†
3732:?
3447:/
3292:)
3276:)
3070:e
3063:t
3056:v
3040:.
3020::
2997:.
2975::
2956:.
2924::
2905:.
2885::
2858:.
2844::
2817:.
2795::
2787::
2781:7
2760:.
2738::
2730::
2700:.
2688::
2665:.
2643::
2635::
2629:5
2608:.
2596::
2573:.
2561::
2538:.
2526::
2503:.
2481::
2475:4
2449:.
2427::
2400:.
2380::
2357:.
2335::
2327::
2300:.
2278::
2270::
2243:.
2229::
2202:.
2198::
2192:7
2175:.
2163::
2140:.
2116::
2108::
2072:.
2050::
2023:.
2011::
2005:8
1988:.
1976::
1953:.
1949::
1926:.
1906::
1898::
1875:.
1871::
1848:.
1826::
1818::
1791:.
1769::
1761::
1734:.
1712::
1704::
1674:.
1652::
1644::
1617:.
1613::
1589:.
1577::
1554:.
1550::
1527:.
1523::
1500:.
1496::
1473:.
1453::
1430:.
1408::
1400::
1373:.
1361::
1338:.
1314::
1306::
1279:.
1267::
1261:7
1240:.
1228::
1204:.
1200::
1177:.
1165::
1157::
1134:.
1112::
1104::
1077:.
1057::
1051:7
1034:.
1022::
1014::
1008:8
991:.
969::
961::
934:.
930::
906:.
902::
894::
871:.
859::
836:.
812::
804::
777:.
755::
728:.
706::
698::
671:.
665::
657::
630:.
618::
610::
587:.
565::
557::
530:.
510::
502::
479:.
457::
31:.
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.