58:
128:. Molecular screens and fingerprints can contain both 2D- and 3D-information. However, the 2D-fingerprints, which are a kind of binary fragment descriptors, dominate in this area. Fragment-based structural keys, like MDL keys, are sufficiently good for handling small and medium-sized chemical databases, whereas processing of large databases is performed with fingerprints having much higher information density. Fragment-based Daylight, BCI, and UNITY 2D (Tripos) fingerprints are the best known examples. The most popular
120:(a kind of ligand-based virtual screening) assumes that all compounds in a database that are similar to a query compound have similar biological activity. Although this hypothesis is not always valid, quite often the set of retrieved compounds is considerably enriched with actives. To achieve high efficacy of similarity-based screening of databases containing millions of compounds, molecular structures are usually represented by
84:. It plays an important role in modern approaches to predicting the properties of chemical compounds, designing chemicals with a predefined set of properties and, especially, in conducting drug design studies by screening large databases containing structures of available (or potentially available) chemicals. These studies are based on the similar property principle of Johnson and Maggiora, which states:
626:— a Java-based software library for calculating Maximum Common Subgraph (MCS) between small molecules. This enables us to find similarity/distance between molecules. MCS is also used for screening drug like compounds by hitting molecules, which share common subgraph (substructure).
171:. Recently, 3D chemical similarity networks based on 3D ligand conformation have also been developed, which can be used to identify scaffold hopping ligands.
227:
46:
partners in inorganic or biological settings. Biological effects and thus also similarity of effects are usually quantified using the
245:
389:
Martin, Y. C.; Kofron, J. L.; Traphagen, L. M. (2002). "Do structurally similar molecules have similar biological activity?".
425:
Durant, J. L.; Leland, B. A.; Henry, D. R.; Nourse, J. G. (2002). "Reoptimization of MDL Keys for Use in Drug
Discovery".
57:
272:
Ralaivola, Liva; Swamidass, Sanjay J.; Hiroto, Saigo; Baldi, Pierre (2005). "Graph kernels for chemical informatics".
427:
315:
274:
634:
650:
655:
579:
Bender, Andreas; Glen, Robert C. (2004). "Molecular similarity: a key technique in molecular informatics".
143:> 0.85 (for Daylight fingerprints). However, it is a common misunderstanding that a similarity of
486:
190:
47:
660:
612:
129:
39:
508:
465:
604:
596:
551:
443:
407:
344:
291:
223:
117:
51:
43:
35:
588:
543:
435:
399:
371:
334:
324:
283:
254:
195:
65:
27:
180:
105:
97:
81:
243:
N. Nikolova; J. Jaworska (2003). "Approaches to
Measure Chemical Similarity - a Review".
339:
310:
164:
156:
644:
535:
391:
133:
616:
623:
309:
Rahman, S. A.; Bashton, M.; Holliday, G. L.; Schrader, R.; Thornton, J. M. (2009).
160:
155:
The concept of chemical similarity can be expanded to consider chemical similarity
530:
287:
362:
Kubinyi, H. (1998). "Similarity and
Dissimilarity: A Medicinal Chemist's View".
185:
168:
61:
375:
132:
for comparing chemical structures represented by means of fingerprints is the
629:
600:
608:
555:
447:
411:
348:
329:
295:
258:
42:
or functional qualities, i.e. the effect that the chemical compound has on
637:— a similarity analysis tool based on molecular interaction fields.
101:
31:
147:> 0.85 reflects similar bioactivities in general ("the 0.85 myth").
547:
439:
403:
592:
630:
Kernel-based
Similarity for Clustering, regression and QSAR Modeling
56:
108:, that measure the structural similarity of chemical compounds.
104:
in descriptor space. Examples for inverse distance measures are
50:
of a compound. In general terms, function can be related to the
529:
Maggiora, G.; Vogt, M.; Stumpfe, D.; Bajorath, J. (2014).
461:
490:
504:
124:(structural keys) or by fixed-size or variable-size
139:. Two structures are usually considered similar if
587:(22). Royal Society of Chemistry (RSC): 3204–18.
311:"Small Molecule Subgraph Detector (SMSD) toolkit"
220:Concepts and Applications of Molecular Similarity
213:
211:
531:"Molecular Similarity in Medicinal Chemistry"
96:Chemical similarity is often described as an
8:
462:"Daylight Chemical Information Systems Inc"
159:, where descriptive network properties and
80:) is one of the most important concepts in
167:, estimate chemical diversity and predict
364:Perspectives in Drug Discovery and Design
338:
328:
86:similar compounds have similar properties
218:Johnson, A. M.; Maggiora, G. M. (1990).
624:Small Molecule Subgraph Detector (SMSD)
207:
112:Similarity search and virtual screening
7:
581:Organic & Biomolecular Chemistry
222:. New York: John Wiley & Sons.
487:"Barnard Chemical Information Ltd"
14:
134:Tanimoto (or Jaccard) coefficient
246:QSAR & Combinatorial Science
163:can be applied to analyze large
511:from the original on 2012-04-19
468:from the original on 2012-12-05
26:) refers to the similarity of
1:
54:of compounds (among others).
288:10.1016/j.neunet.2005.07.009
151:Chemical similarity network
677:
428:J. Chem. Inf. Comput. Sci.
316:Journal of Cheminformatics
376:10.1023/A:1027221424359
38:with respect to either
330:10.1186/1758-2946-1-12
259:10.1002/qsar.200330831
126:molecular fingerprints
69:
116:The similarity-based
60:
78:molecular similarity
24:molecular similarity
253:(9–10): 1006–1026.
191:Substructure search
102:measure of distance
92:Similarity measures
74:chemical similarity
48:biological activity
20:Chemical similarity
130:similarity measure
70:
36:chemical compounds
548:10.1021/jm401411z
440:10.1021/ci010132r
404:10.1021/jm020155c
398:(19): 4350–4358.
229:978-0-471-62175-1
122:molecular screens
118:virtual screening
52:chemical activity
28:chemical elements
668:
620:
593:10.1039/b409813g
566:
565:
563:
562:
542:(8): 3186–3204.
526:
520:
519:
517:
516:
501:
495:
494:
489:. Archived from
483:
477:
476:
474:
473:
458:
452:
451:
434:(6): 1273–1280.
422:
416:
415:
386:
380:
379:
359:
353:
352:
342:
332:
306:
300:
299:
282:(8): 1093–1110.
269:
263:
262:
240:
234:
233:
215:
196:Ternary compound
106:molecule kernels
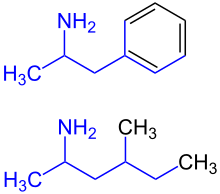
66:Methylhexanamine
676:
675:
671:
670:
669:
667:
666:
665:
651:Cheminformatics
641:
640:
578:
575:
570:
569:
560:
558:
528:
527:
523:
514:
512:
503:
502:
498:
485:
484:
480:
471:
469:
460:
459:
455:
424:
423:
419:
388:
387:
383:
361:
360:
356:
308:
307:
303:
275:Neural Networks
271:
270:
266:
242:
241:
237:
230:
217:
216:
209:
204:
181:Me-too compound
177:
153:
114:
94:
82:cheminformatics
17:
12:
11:
5:
674:
672:
664:
663:
658:
656:Drug discovery
653:
643:
642:
639:
638:
632:
627:
621:
574:
573:External links
571:
568:
567:
521:
496:
493:on 2008-10-11.
478:
453:
417:
381:
354:
301:
264:
235:
228:
206:
205:
203:
200:
199:
198:
193:
188:
183:
176:
173:
165:chemical space
157:network theory
152:
149:
113:
110:
93:
90:
72:The notion of
15:
13:
10:
9:
6:
4:
3:
2:
673:
662:
659:
657:
654:
652:
649:
648:
646:
636:
633:
631:
628:
625:
622:
618:
614:
610:
606:
602:
598:
594:
590:
586:
582:
577:
576:
572:
557:
553:
549:
545:
541:
538:
537:
536:J. Med. Chem.
532:
525:
522:
510:
506:
500:
497:
492:
488:
482:
479:
467:
463:
457:
454:
449:
445:
441:
437:
433:
430:
429:
421:
418:
413:
409:
405:
401:
397:
394:
393:
392:J. Med. Chem.
385:
382:
377:
373:
369:
365:
358:
355:
350:
346:
341:
336:
331:
326:
322:
318:
317:
312:
305:
302:
297:
293:
289:
285:
281:
277:
276:
268:
265:
260:
256:
252:
248:
247:
239:
236:
231:
225:
221:
214:
212:
208:
201:
197:
194:
192:
189:
187:
184:
182:
179:
178:
174:
172:
170:
166:
162:
158:
150:
148:
146:
142:
138:
135:
131:
127:
123:
119:
111:
109:
107:
103:
99:
91:
89:
87:
83:
79:
75:
67:
63:
59:
55:
53:
49:
45:
41:
37:
33:
29:
25:
21:
16:Chemical term
584:
580:
559:. Retrieved
539:
534:
524:
513:. Retrieved
505:"Tripos Inc"
499:
491:the original
481:
470:. Retrieved
456:
431:
426:
420:
395:
390:
384:
367:
363:
357:
320:
314:
304:
279:
273:
267:
250:
244:
238:
219:
161:graph theory
154:
144:
140:
136:
125:
121:
115:
95:
85:
77:
73:
71:
23:
19:
18:
370:: 225–252.
186:Drug design
169:drug target
62:Amphetamine
645:Categories
561:2023-11-13
515:2022-07-19
472:2022-07-19
323:(12): 12.
202:References
68:similarity
40:structural
661:Chemistry
601:1477-0520
32:molecules
617:16399588
609:15534697
556:24151987
509:Archived
466:Archived
448:12444722
412:12213076
349:20298518
296:16157471
175:See also
44:reaction
340:2820491
98:inverse
635:Brutus
615:
607:
599:
554:
446:
410:
347:
337:
294:
226:
613:S2CID
100:of a
605:PMID
597:ISSN
552:PMID
444:PMID
408:PMID
368:9–11
345:PMID
292:PMID
224:ISBN
76:(or
64:and
22:(or
589:doi
544:doi
436:doi
400:doi
372:doi
335:PMC
325:doi
284:doi
255:doi
34:or
647::
611:.
603:.
595:.
583:.
550:.
540:57
533:.
507:.
464:.
442:.
432:42
406:.
396:45
366:.
343:.
333:.
319:.
313:.
290:.
280:18
278:.
251:22
249:.
210:^
88:.
30:,
619:.
591::
585:2
564:.
546::
518:.
475:.
450:.
438::
414:.
402::
378:.
374::
351:.
327::
321:1
298:.
286::
261:.
257::
232:.
145:T
141:T
137:T
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.