441:
76:
Degeneracy results because there are more codons than encodable amino acids. For example, if there were two bases per codon, then only 16 amino acids could be coded for (4²=16). Because at least 21 codes are required (20 amino acids plus stop) and the next largest number of bases is three, then 4³
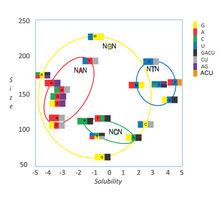
464:). For example, a codon of NUN (where N = any nucleotide) tends to code for hydrophobic amino acids, NCN yields amino acid residues that are small in size and moderate in hydropathy, and NAN encodes average size hydrophilic residues. These tendencies may result from the shared ancestry of the
118:
A position is said to be non-degenerate if any mutation at this position changes the amino acid. For example, all three positions of methionine's AUG are non-degenerate, because the only codon coding for methionine is AUG. The same goes for tryptophan's UGG.
57:
Degeneracy of the genetic code was identified by
Lagerkvist. For instance, codons GAA and GAG both specify glutamic acid and exhibit redundancy; but, neither specifies any other amino acid and thus are not ambiguous or demonstrate no ambiguity.
61:
The codons encoding one amino acid may differ in any of their three positions; however, more often than not, this difference is in the second or third position. For instance, the amino acid
429:
restricts this in practice in many organisms; twofold degenerate codons can withstand silence mutation rather than
Missense or Nonsense point mutations at the third position. Since
45:, exhibited as the multiplicity of three-base pair codon combinations that specify an amino acid. The degeneracy of the genetic code is what accounts for the existence of
452:
A practical consequence of redundancy is that some errors in the genetic code cause only a synonymous mutation, or an error that would not affect the protein because the
437:(purine to pyrimidine or vice versa) mutations, the equivalence of purines or that of pyrimidines at twofold degenerate sites adds a further fault-tolerance.
96:
of four possible nucleotides (A, C, G, T) at this position specify the same amino acid. A nucleotide substitution at a 4-fold degenerate site is always a
759:
927:
558:
492:
512:
811:
791:
899:
649:
666:
694:"The G x U wobble base pair. A fundamental building block of RNA structure crucial to RNA function in diverse biological systems"
1053:
997:
992:
1144:
1080:
1011:
752:
816:
1134:
425:. For example, in theory, fourfold degenerate codons can tolerate any point mutation at the third position, although
69:
is specified by UUA, UUG, CUU, CUC, CUA, CUG codons (difference in the first or third position); and the amino acid
1031:
465:
1139:
1075:
863:
776:
745:
1036:
853:
838:
107:
on some of the substitutions. An example (and the only) 3-fold degenerate site is the third position of an
1041:
966:
858:
104:
956:
941:
821:
461:
430:
147:
593:"Degeneracy of the genetic code and stability of the base pair at the second position of the anticodon"
1063:
961:
879:
471:
These variable codes for amino acids are allowed because of modified bases in the first base of the
889:
97:
73:
is specified by UCA, UCG, UCC, UCU, AGU, AGC (difference in the first, second, or third position).
46:
1024:
907:
513:"The Information in DNA Determines Cellular Function via Translation | Learn Science at Scitable"
1108:
723:
645:
622:
554:
440:
713:
705:
612:
604:
476:
445:
426:
134:. Only two amino acids are specified by a single codon each. One of these is the amino-acid
1085:
922:
768:
138:, specified by the codon AUG, which also specifies the start of translation; the other is
433:
mutations (purine to purine or pyrimidine to pyrimidine mutations) are more likely than
912:
884:
718:
693:
678:
617:
592:
422:
65:
is specified by GAA and GAG codons (difference in the third position); the amino acid
1128:
1113:
800:
62:
987:
917:
786:
434:
42:
709:
1090:
1019:
457:
453:
17:
115:. In computation, this position is often treated as a twofold degenerate site.
1048:
667:"Genetic Algorithms and Recursive Ensemble Mutagenesis in Protein Engineering"
574:
139:
135:
112:
108:
472:
727:
626:
1068:
1058:
982:
131:
608:
804:
480:
127:
66:
421:
These properties of the genetic code make it more fault-tolerant for
123:
70:
77:
gives 64 possible codons, meaning that some degeneracy must exist.
826:
737:
439:
38:
642:
Reaction centers of photosynthetic bacteria: Feldafing-II-Meeting
549:
Watson JD, Baker TA, Bell SP, Gann A, Levine M, Oosick R (2008).
741:
111:
codon. AUU, AUC, or AUA all encode isoleucine, but AUG encodes
796:
146:
Inverse table for the standard genetic code (compressed using
122:
There are three amino acids encoded by six different codons:
444:
Grouping of codons by amino acid residue molar volume and
644:. Vol. 6. Berlin: Springer-Verlag. pp. 209–18.
575:
Two out of three: An alternative method for codon reading
460:
is maintained by equivalent substitution of amino acids (
640:
Yang; et al. (1990). Michel-Beyerle, M. E. (ed.).
1101:
1010:
975:
949:
940:
898:
872:
846:
837:
775:
475:of the tRNA, and the base-pair formed is called a
753:
8:
553:. San Francisco: Pearson/Benjamin Cummings.
85:The examples all refer to the standard code.
946:
908:Precursor mRNA (pre-mRNA / hnRNA)
843:
760:
746:
738:
717:
616:
144:
27:Redundancy of codons in the genetic code
504:
483:and the Non-Watson-Crick U-G basepair.
103:A less degenerate site would produce a
591:Lehmann, J; Libchaber, A (July 2008).
88:A position of a codon is said to be a
928:Histone acetylation and deacetylation
544:
542:
540:
538:
536:
534:
532:
493:Neutral theory of molecular evolution
7:
993:Ribosome-nascent chain complex (RNC)
692:Varani G, McClain WH (July 2000).
100:with no change on the amino acid.
25:
998:Post-translational modification
142:, specified by the codon UGG.
92:-fold degenerate site if only
1:
551:Molecular Biology of the Gene
479:. The modified bases include
310:UCU, UCC, UCA, UCG; AGU, AGC
211:CUU, CUC, CUA, CUG; UUA, UUG
200:CGU, CGC, CGA, CGG; AGA, AGG
665:Füllen G, Youvan DC (1994).
710:10.1093/embo-reports/kvd001
1161:
466:aminoacyl tRNA synthetases
163:
84:
468:related to these codons.
349:
252:
41:is the redundancy of the
1059:sequestration (P-bodies)
671:Complexity International
573:Lagerkvist, U. (1978.) "
1037:Gene regulatory network
1042:cis-regulatory element
449:
105:nonsynonymous mutation
462:conservative mutation
443:
1145:Protein biosynthesis
1064:alternative splicing
1054:Post-transcriptional
880:Transcription factor
47:synonymous mutations
988:Transfer RNA (tRNA)
609:10.1261/rna.1029808
387:GUU, GUC, GUA, GUG
358:GGU, GGC, GGA, GGG
341:CAA, CAG; GAA, GAG
330:ACU, ACC, ACA, ACG
290:CCU, CCC, CCA, CCG
261:AAU, AAC; GAU, GAC
180:GCU, GCC, GCA, GCG
151:
98:synonymous mutation
1135:Molecular genetics
1102:Influential people
1081:Post-translational
900:Post-transcription
450:
145:
1122:
1121:
1006:
1005:
936:
935:
812:Special transfers
560:978-0-8053-9592-1
414:
413:
16:(Redirected from
1152:
947:
844:
762:
755:
748:
739:
732:
731:
721:
689:
683:
682:
677:. Archived from
662:
656:
655:
637:
631:
630:
620:
588:
582:
571:
565:
564:
546:
527:
526:
524:
523:
509:
477:wobble base pair
427:codon usage bias
152:
21:
18:Codon redundancy
1160:
1159:
1155:
1154:
1153:
1151:
1150:
1149:
1140:Gene expression
1125:
1124:
1123:
1118:
1097:
1032:Transcriptional
1002:
971:
932:
923:Polyadenylation
894:
868:
833:
827:Protein→Protein
778:
771:
769:Gene expression
766:
736:
735:
691:
690:
686:
664:
663:
659:
652:
639:
638:
634:
590:
589:
585:
572:
568:
561:
548:
547:
530:
521:
519:
511:
510:
506:
501:
489:
423:point mutations
419:
215:
204:
86:
83:
55:
28:
23:
22:
15:
12:
11:
5:
1158:
1156:
1148:
1147:
1142:
1137:
1127:
1126:
1120:
1119:
1117:
1116:
1111:
1109:François Jacob
1105:
1103:
1099:
1098:
1096:
1095:
1094:
1093:
1088:
1078:
1073:
1072:
1071:
1066:
1061:
1051:
1046:
1045:
1044:
1039:
1029:
1028:
1027:
1016:
1014:
1008:
1007:
1004:
1003:
1001:
1000:
995:
990:
985:
979:
977:
973:
972:
970:
969:
964:
959:
953:
951:
944:
938:
937:
934:
933:
931:
930:
925:
920:
915:
910:
904:
902:
896:
895:
893:
892:
887:
885:RNA polymerase
882:
876:
874:
870:
869:
867:
866:
861:
856:
850:
848:
841:
835:
834:
832:
831:
830:
829:
824:
819:
809:
808:
807:
789:
783:
781:
773:
772:
767:
765:
764:
757:
750:
742:
734:
733:
684:
681:on 2011-03-15.
657:
650:
632:
583:
566:
559:
528:
517:www.nature.com
503:
502:
500:
497:
496:
495:
488:
485:
458:hydrophobicity
454:hydrophilicity
418:
415:
412:
411:
408:
407:UAA, UGA, UAG
405:
402:
399:
398:AUG, CUG, UUG
396:
392:
391:
388:
385:
382:
379:
376:
372:
371:
368:
365:
362:
359:
356:
352:
351:
348:
345:
342:
339:
338:Gln or Glu, Z
335:
334:
331:
328:
325:
322:
319:
315:
314:
311:
308:
305:
302:
299:
295:
294:
291:
288:
285:
282:
279:
275:
274:
271:
268:
265:
262:
259:
258:Asn or Asp, B
255:
254:
251:
248:
245:
242:
238:
237:
234:
231:
228:
225:
222:
218:
217:
212:
209:
206:
201:
198:
194:
193:
190:
189:AUU, AUC, AUA
187:
184:
181:
178:
174:
173:
170:
167:
164:
162:
159:
156:
148:IUPAC notation
82:
79:
54:
51:
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
1157:
1146:
1143:
1141:
1138:
1136:
1133:
1132:
1130:
1115:
1114:Jacques Monod
1112:
1110:
1107:
1106:
1104:
1100:
1092:
1089:
1087:
1084:
1083:
1082:
1079:
1077:
1076:Translational
1074:
1070:
1067:
1065:
1062:
1060:
1057:
1056:
1055:
1052:
1050:
1047:
1043:
1040:
1038:
1035:
1034:
1033:
1030:
1026:
1023:
1022:
1021:
1018:
1017:
1015:
1013:
1009:
999:
996:
994:
991:
989:
986:
984:
981:
980:
978:
974:
968:
965:
963:
960:
958:
955:
954:
952:
948:
945:
943:
939:
929:
926:
924:
921:
919:
916:
914:
911:
909:
906:
905:
903:
901:
897:
891:
888:
886:
883:
881:
878:
877:
875:
871:
865:
862:
860:
857:
855:
852:
851:
849:
845:
842:
840:
839:Transcription
836:
828:
825:
823:
820:
818:
815:
814:
813:
810:
806:
802:
798:
795:
794:
793:
792:Central dogma
790:
788:
785:
784:
782:
780:
774:
770:
763:
758:
756:
751:
749:
744:
743:
740:
729:
725:
720:
715:
711:
707:
703:
699:
695:
688:
685:
680:
676:
672:
668:
661:
658:
653:
651:3-540-53420-2
647:
643:
636:
633:
628:
624:
619:
614:
610:
606:
603:(7): 1264–9.
602:
598:
594:
587:
584:
581:, 75:1759-62.
580:
576:
570:
567:
562:
556:
552:
545:
543:
541:
539:
537:
535:
533:
529:
518:
514:
508:
505:
498:
494:
491:
490:
486:
484:
482:
478:
474:
469:
467:
463:
459:
455:
447:
442:
438:
436:
432:
428:
424:
416:
409:
406:
403:
400:
397:
394:
393:
389:
386:
383:
380:
377:
374:
373:
369:
366:
363:
360:
357:
354:
353:
346:
343:
340:
337:
336:
332:
329:
326:
323:
320:
317:
316:
312:
309:
306:
303:
300:
297:
296:
292:
289:
286:
283:
280:
277:
276:
272:
269:
266:
263:
260:
257:
256:
249:
246:
243:
240:
239:
235:
232:
229:
226:
223:
220:
219:
214:CUN, UUR; or
213:
210:
207:
202:
199:
196:
195:
191:
188:
185:
182:
179:
176:
175:
171:
168:
165:
160:
157:
154:
153:
149:
143:
141:
137:
133:
129:
125:
120:
116:
114:
110:
106:
101:
99:
95:
91:
80:
78:
74:
72:
68:
64:
63:glutamic acid
59:
52:
50:
48:
44:
40:
36:
32:
19:
1091:irreversible
976:Key elements
873:Key elements
787:Genetic code
777:Introduction
704:(1): 18–23.
701:
697:
687:
679:the original
674:
670:
660:
641:
635:
600:
596:
586:
578:
569:
550:
520:. Retrieved
516:
507:
470:
451:
435:transversion
420:
417:Implications
203:CGN, AGR; or
121:
117:
102:
93:
89:
87:
75:
60:
56:
43:genetic code
34:
30:
29:
942:Translation
779:to genetics
172:Compressed
161:Compressed
81:Terminology
1129:Categories
1086:reversible
1049:lac operon
1025:imprinting
1020:Epigenetic
1012:Regulation
967:Eukaryotic
913:5' capping
864:Eukaryotic
522:2021-07-14
499:References
446:hydropathy
431:transition
169:DNA codons
166:Amino acid
158:DNA codons
155:Amino acid
140:tryptophan
136:methionine
113:methionine
109:isoleucine
53:Background
35:redundancy
31:Degeneracy
957:Bacterial
854:Bacterial
473:anticodon
410:URA, UAR
378:CAU, CAC
367:UAU, UAC
321:GAA, GAG
313:UCN, AGY
301:CAA, CAG
281:UGU, UGC
270:UUU, UUC
244:GAU, GAC
233:AAA, AAG
224:AAU, AAC
216:CUY, YUR
205:CGY, MGR
1069:microRNA
983:Ribosome
962:Archaeal
918:Splicing
890:Promoter
859:Archaeal
803: →
799: →
728:11256617
698:EMBO Rep
627:18495942
487:See also
132:arginine
822:RNA→DNA
817:RNA→RNA
805:Protein
719:1083677
618:2441979
481:inosine
384:Val, V
375:His, H
364:Tyr, Y
355:Gly, G
347:Trp, W
327:Thr, T
318:Glu, E
307:Ser, S
298:Gln, Q
287:Pro, P
278:Cys, C
267:Phe, F
250:Met, M
241:Asp, D
230:Lys, K
221:Asn, N
208:Leu, L
197:Arg, R
186:Ile, I
177:Ala, A
128:leucine
67:leucine
726:
716:
648:
625:
615:
557:
395:START
130:, and
124:serine
71:serine
39:codons
950:Types
847:Types
404:STOP
724:PMID
646:ISBN
623:PMID
579:PNAS
555:ISBN
401:HUG
390:GUN
381:CAY
370:UAY
361:GGN
350:UGG
344:SAR
333:ACN
324:GAR
304:CAR
293:CCN
284:UGY
273:UUY
264:RAY
253:AUG
247:GAY
236:AAR
227:AAY
192:AUH
183:GCN
801:RNA
797:DNA
714:PMC
706:doi
613:PMC
605:doi
597:RNA
577:",
456:or
37:of
33:or
1131::
722:.
712:.
700:.
696:.
673:.
669:.
621:.
611:.
601:14
599:.
595:.
531:^
515:.
150:)
126:,
49:.
761:e
754:t
747:v
730:.
708::
702:1
675:1
654:.
629:.
607::
563:.
525:.
448:.
94:n
90:n
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.