425:. In development, neutral networks are clusters of GRNs that differ in only one interaction between two nodes (e.g. replacing transcription with suppression) and yet produce the same phenotypic outcome. In this sense, an individual phenotype within a population could be mapped to several equivalent GRNs, that together constitute a neutral network. Conversely, a GRN that differs in one interaction and causes a different phenotype is considered non-neutral. Given this architecture, the probability of mutating from one phenotype to another will depend on the number of neutral-neighbors relative to non-neutral neighbors for a particular GRN, and thus, phenotypic change will be influenced by the position of a GRN within the network and will be biased towards changes that require few mutations to reach a neighboring non-neutral GRN.
164:. One way to study the distribution of phenotypic variation is through depicting the volume of the morphospace occupied by a set of organisms or species. Theoretically, there can exist a natural process that generates an almost-evenly (quasi stochastic) distributed pattern of phenotypes in the morphospace, regarding that new species necessary tend to occupy a point in the morphospace that is close to those of its phylogenetic relatives. However, it is now widely acknowledged that organisms are not evenly distributed along the morphospace, i.e. isotropic variation, but instead are nonrandomly distributed, i.e. anisotropic variation. In other words, there exists a discordance between the apparent (or theoretical) possible phenotypes and their actual accessibility.
168:
219:
44:
303:, the mutational matrix (M-matrix), also known as the distribution of mutational effects, has been shown to be of equivalent importance. The M-matrix describes the potential effects of new mutations on the existing genetic variances and covariances, and these effects will depend on the epistatic and pleiotropic interactions of the underlying genes. In other words, the M-matrix determines the G-matrix, and thus, the response to selection of a population. Similarly to the P-matrix, the G-matrix describes the main axis of variation.
101:
312:
401:
under selection but few effects on other traits, are expected to accumulate a higher proportion of mutations that cause evolutionary change. These strategically-positioned genes have the potential to filter random genetic variation and translate it to nonrandom functionally integrated phenotypes, making adaptive variants effectively accessible to selection, and, thus, many of the mutations contributing to phenotypic evolution may be concentrated in these genes.
202:
363:
231:
three-jointed thumb due to the extension of the first toe. However, 20 toes were found much more frequently and then 22, 24 or 26 toes with decreasing frequency. Odd total numbers of toes on the feet were less common. There is another bias between the number of toes on the front and rear feet, and a left-right asymmetry in the number of toes. Random bistability during the development process could explain the observed bias.
238:) have been used to understand the logic behind the mechanisms that produce variation. For example, in a wide range of animals, from fish to humans, two-headed organisms are much more common than three-headed organisms; similarly, Siamese twins theoretically could ‘fuse’ through any region in the body but the fusion occurs more frequently in the abdominal region. This trend was referred to as
143:
93:. Some kinds of reprogramming are more likely to occur than others given the nature of the genotype–phenotype map, which determines the propensity of a system to vary in a particular direction, thus, creating a bias. In other words, the underlying architecture of the developmental systems influences the kinds of possible phenotypic outcomes.
256:
321:
however, if the main axis of variation is orthogonal to the direction of selection, covariation will constraint the rate of adaptive evolution. In general, for a population under the influence of a single fitness optimum, the rate of morphological divergence (from an ancestral to a new phenotype or between pairs of
388:
provides multiple inputs to other genes, creating a complex array of interactions, and information regarding the timing, place and amount of gene expression generally flows from few high-level control genes through multiple intermediate genes to peripheral gene batteries that ultimately determine the
209:
In a classical natural example of bias it was shown that only a small proportion of all possible snail shell shapes was realized in nature and actual species were confined to discrete regions of the shell-morphospace rather than being continuously distributed. In another natural example, it was shown
151:
The morphospace is a quantitative representation of phenotypes in a multidimensional space, where each dimension corresponds to a trait. The phenotype of each organism or species is then represented as a point in that space that summarizes the combination of values or states at each particular trait.
400:
effects are more likely to be targets of evolution, thus, the hierarchical architecture of developmental pathways may bias the genetic basis of evolutionary change. For instance, genes within GRNs with "optimally pleiotropic" effects, that is, genes that have the most widespread effect on the trait
315:
Morphospace and fitness landscape with a single fitness optimum. For a population undergoing directional selection, the main axis of variation (largest axis of the white ellipse) will bias the main direction of the trajectory toward the fitness optimum (arrow). The rate of morphological change will
268:
changes are widespread in nature and can account for a wide variety of realized morphologies and subsequent ecological and physiological changes. Under this approach, phenotype is seen as an integrated system where each trait develops and evolves in concert with the other traits, and thus, a change
259:
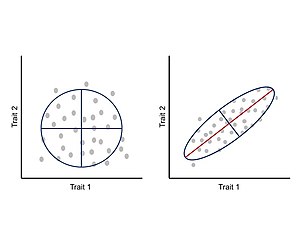
Representation of the relationship between two traits. Left: No trait covariation. Each trait changes independently of the other. Right: Trait covariation causes a positive correlation between traits where increase in one trait is correlated with an increase in the other trait (covariation can also
214:
have an enormous variation in the number of pairs of legs, the lowest being 27 and the highest 191 pairs; however, there are no species with an even number of leg pairs, which suggests that either these phenotypes are somehow restricted during development or that there is a developmental drive into
320:
A general consequence of the P-matrices and G-matrices is that evolution will tend to follow the ‘path of least resistance’. In other words, if the main axis of variation is aligned with the direction of selection, covariation (genetic or phenotypic) will facilitate the rate of adaptive evolution;
128:
Developmental drive is the inherent natural tendency of organisms and their ontogenetic trajectories to change in a particular direction (i.e. a bias towards a certain ontogenetic trajectory). This type of bias is thought to facilitate adaptive evolution by aligning phenotypic variability with the
298:
and covariance between traits. Thus, a population’s immediate ability to respond to selection is determined by the G-matrix, in which the variance is a function of standing genetic variation, and the covariance arises from pleiotropy and linkage disequilibrium. Although the G-matrix is one of the
96:
However, developmental bias can evolve through natural selection, and both processes simultaneously influence phenotypic evolution. For example, developmental bias can affect the rate or path to an adaptive peak (high-fitness phenotype), and conversely, strong directional selection can modify the
88:
and mutation does not in itself produce phenotypic variation, thus, there is a conceptual gap regarding the connection between a mutation and the potential change in phenotype. For a mutation to readily alter a phenotype, and hence be visible to natural selection, it has to modify the ontogenetic
188:
An important distinction between structuralism and functionalism regards primarily with the interpretation of the causes of the empty regions in the morphospace (that is, the inexistent phenotypes): Under the functionalist view, empty spaces correspond to phenotypes that are both ontogenetically
175:
Thus, some phenotypes are inaccessible (or impossible) due to the underlying architecture of the developmental trajectory, while others are accessible (or possible). However, of the possible phenotypes, some are ‘easier’ or more probable to occur than others. For example, a phenotype such as the
333:: ability of a developmental system to change in the direction of natural selection. In the latter, the main axis of phenotypic variation is aligned with the direction of selection. Similarly, from the G-matrix, the most important parameter that describes the propensity of variation is the lead
184:
characters (e.g. birds and bats), and, thus, are mutually exclusive. On the other hand, if two phenotypes are possible (and equally fit), but one form of reprogramming requires only one mutation while the other requires two or more, the former will be more likely to occur (assuming that genetic
285:
is a statistical framework mainly concerned with modeling the evolution of continuous characters. Under this framework, correlation between traits could be the result of two processes: 1) natural selection acting simultaneously on several traits ensuring that they are inherited together (i.e.
104:
Developmental bias for continuous characters. If the main axis of variation (red arrows) is orthogonal to the direction of selection (dashed line), trait covariation will constraint adaptive evolution. Conversely, if the main axis of variation is aligned with the direction of selection, trait
230:
showed that the number of additional toes was variable (plastic) and contained a bias. The Maine Coon cat (as the basic model of the
Hemingway mutants) has 18 toes in the wild. Polydactyly occurred in some cases with an unchanged number of toes (18 toes), whereby the deviation consisted of a
119:
Developmental constraints are limitations on phenotypic variability (or absence of variation) caused by the inherent structure and dynamics of the developmental system. Constraints are a bias against a certain ontogenetic trajectory, and consequently are thought to limit adaptive evolution.
193:. In contrast, under the structuralist view, empty spaces correspond to ontogenetically impossible or improbable phenotypes, thus, implying a bias in the types of phenotypes that can be produced assuming equal amounts of variation (genetic mutations) in both models.
273:
effects of underlying genes. This correlated change between traits can be measured and analyzed through a phenotypic variance-covariance matrix (P-matrix) which summarizes the dimensions of phenotypic variability and the main axis of variation.
146:
Multidimensional representation of species in the morphospace. Each axis corresponds to a trait, and dots correspond to organisms with particular trait values combinations. In this case, the axes represent the form of the fish
290:), or 2) natural selection acting on one trait causing correlated change in other traits due to pleiotropic effects of genes. For a set of traits, the equation that describe the variance among traits is the multivariate
73:(also “adaptationist”, “pan-selectionist” or “externalist”) view in which phenotypic evolution results only from the interaction between the deterministic action of natural selection and variation caused by mutation.
328:
From the P-matrix for a set of characters, two broadly important measures of the propensity of variation can be extracted: 1) Respondability: ability of a developmental system to change in any direction, and 2)
409:
The genotype–phenotype map perspective establishes that the way in which genotypic variation can be mapped to phenotypic variation is critical for the ability of a system to evolve. The prevalence of
35:. Historically, the term was synonymous with developmental constraint, however, the latter has been more recently interpreted as referring solely to the negative role of development in evolution.
264:
Integration or covariation among traits during development has been suggested to constrain phenotypic evolution to certain regions of the morphospace and limit adaptive evolution. These
269:
in one trait affects the interacting parts in a correlated manner. The correlation between traits is a consequence of the architecture of the genotype–phenotype map, particularly the
1210:
Chartier, Marion; Jabbour, Florian; Gerber, Sylvain; Mitteroecker, Philipp; Sauquet, Hervé; von
Balthazar, Maria; Staedler, Yannick; Crane, Peter R.; Schönenberger, Jürg (2014).
218:
1544:
Lange, Axel; Nemeschkal, Hans L.; Müller, Gerd B. (2014). "Biased
Polyphenism in Polydactylous Cats Carrying a Single Point Mutation: The Hemingway Model for Digit Novelty".
76:
The rationale behind the role of the organism, or more specifically the embryo, as a causal force in evolution and for the existence of bias is as follows: The traditional,
325:) is inversely proportional to the angle formed by the main axis of variation and the direction of selection, causing a curved trajectory through the morphospace.
167:
2346:
Schuster, Peter; Fontana, Walter; Stadler, Peter F.; Hofacker, Ivo L. (1994). "From sequences to shapes and back: a case study in RNA secondary structures".
2506:
393:
affecting multiple downstream genes, whereas intermediate and peripheral genes tend to have moderate to low pleiotropic effects, respectively.
294:Δz = β x G, where Δz is the vector of differences in trait means, β is a vector of selection coefficients, and G is a matrix of the
1591:
1191:
1154:
28:
which ultimately influence the direction and outcome of evolutionary change by affecting the rates, magnitudes, directions and limits of
910:
Arthur, Wallace (2000). "The concept of developmental reprogramming and the quest for an inclusive theory of evolutionary mechanisms".
2226:
753:
152:
This approach is used to study the evolution of realized phenotypes compared to those that are theoretically possible but inexistent.
840:
316:
be inversely proportional to the angle (beta) formed between the direction of selection (dashed line) and the main axis of variation.
1340:
886:
858:
80:, approach to explain the process behind evolutionary change is natural selection acting upon heritable variation caused by genetic
260:
produce negative correlation). The red line within the ellipse represents the main eigenvector of the variance-covariance matrix.
242:, suggesting the existence of profound historical rules governing the expression of abnormal forms in distantly related species.
568:; Lewontin, R. C. (1979). "The spandrels of San Marco and the Panglossian paradigm: a critique of the adaptationist programme".
630:(1989). "A Developmental Constraint in Cerion, with Comments of the Definition and Interpretation of Constraint in Evolution".
176:
classical figure of a dragon (i.e. a giant reptile-like creature with two pairs of limbs and an anterior pair of wings) may be
2250:
Kopp, A. (2009). "Metamodels and phylogenetic replication: A systematic approach to the evolution of developmental pathways".
1137:
Altenberg, L. (1995). "Genome Growth and the
Evolution of the Genotype-Phenotype Map". In Banzhaf, W.; Eeckman, F. H. (eds.).
2501:
1913:
Steppan, Scott J.; Patrick C. Phillips; David Houle (2002). "Comparative quantitative genetics: evolution of the G matrix".
160:
Describing and understanding the drivers of the distribution of phenotypic variation in nature is one of the main goals in
334:
43:
1957:
Jones, Adam G.; Arnold, Stevan J.; Bürger, Reinhard (2007). "The
Mutation Matrix and the Evolution of Evolvability".
1272:
Gerber, Sylvain (2014). "Not all roads can be taken: development induces anisotropic accessibility in morphospace".
2409:
2286:
1497:"The interaction between developmental bias and natural selection: from centipede segments to a general hypothesis"
678:
Arthur, Wallace (2001). "Developmental drive: an important determinant of the direction of phenotypic evolution".
1170:
Altenberg, L. (2005). "Modularity in
Evolution: Some Low-Level Questions". In Callebaut, W.; Rasskin-Gutman, D.;
342:
295:
47:
Haeckel's drawings of "lower" (fish, salamander) and "higher" (tortoise, chick) vertebrates at comparable stages
61:
force of evolutionary change. In the
Structuralist view, phenotypic evolution is the result of the action of
2000:
Cheverud, James M. (1984). "Quantitative genetics and developmental constraints on evolution by selection".
381:
371:
77:
53:
51:
In modern evolutionary biology, the idea of developmental bias is embedded into a current of thought called
291:
421:, and a consequence of this "many-to-few" relationship between genotype and phenotype is the existence of
287:
282:
396:
In general, it is expected that newly arisen mutations with higher dominance and fewer pleiotropic and
345:
for a set of continuous traits within populations. For a population undergoing directional selection, g
100:
2355:
2009:
1733:
796:
577:
385:
161:
17:
1824:
Lande, Russell; Arnold, Stevan J. (1983). "The
Measurement of Selection on Correlated Characters".
1183:
1039:
Uller, Tobias; Moczek, Armin P.; Watson, Richard A.; Brakefield, Paul M.; Laland, Kevin N. (2018).
968:
389:
fate of each cell. This type of architecture implies that high-level control genes tend to be more
311:
1724:
Emlen, Douglas J. (2001-02-23). "Costs and the
Diversification of Exaggerated Animal Structures".
1332:
513:"The effect of development on the direction of evolution: toward a twenty-first century consensus"
2387:
2196:
2076:
1892:
1841:
1765:
1706:
1477:
1305:
943:
711:
647:
609:
488:
464:
181:
787:(1989). "The logic of monsters: Evidence for internal constraint in development and evolution".
1146:
2440:
2432:
2379:
2371:
2328:
2310:
2267:
2232:
2222:
2188:
2180:
2133:
2125:
2084:
2033:
2025:
1982:
1974:
1930:
1884:
1849:
1806:
1757:
1749:
1698:
1663:
1645:
1597:
1587:
1561:
1526:
1518:
1450:
1442:
1402:
1384:
1336:
1297:
1289:
1249:
1231:
1187:
1150:
1119:
1078:
1060:
1002:
994:
935:
927:
892:
882:
854:
812:
759:
749:
703:
695:
655:
627:
601:
593:
565:
542:
534:
190:
62:
2157:
1626:"The macroevolutionary consequences of phenotypic integration: from development to deep time"
384:
are modular, multilayered, and semi-hierarchically systems of genes and their products: each
2424:
2363:
2318:
2302:
2259:
2172:
2115:
2068:
2017:
1966:
1922:
1876:
1833:
1796:
1741:
1690:
1653:
1637:
1553:
1508:
1434:
1392:
1376:
1328:
1281:
1239:
1223:
1175:
1171:
1138:
1109:
1068:
1052:
984:
919:
846:
804:
687:
639:
585:
524:
480:
410:
201:
29:
362:
1176:
691:
467:; Burian, R.; Kauffman, S.; Alberch, P.; Campbell, J.; Goodwin, B.; Lande, R.; Raup, D.;
2359:
2013:
1737:
800:
581:
97:
developmental bias to increase the phenotypic variation in the direction of selection.
2323:
2290:
2056:
1837:
1694:
1658:
1625:
1397:
1364:
1244:
1211:
1073:
989:
972:
227:
211:
189:
possible and equally probable but are eliminated by natural selection due to their low
2021:
1926:
1325:
The Origin of Higher Taxa: Palaeobiological, developmental and ecological perspectives
1212:"The floral morphospace - a modern comparative approach to study angiosperm evolution"
808:
2495:
2306:
2263:
1970:
1801:
1139:
923:
529:
512:
468:
2391:
1769:
1710:
1309:
947:
715:
492:
2200:
1896:
784:
613:
434:
330:
300:
1785:"Phenotypic integration: studying the ecology and evolution of complex phenotypes"
1178:
Modularity: Understanding the
Development and Evolution of Natural Complex Systems
1468:
Raup, David M. (1966). "Geometric Analysis of Shell Coiling: General Problems".
1056:
2428:
1438:
1601:
1557:
1380:
1114:
1097:
896:
763:
439:
390:
270:
235:
2436:
2375:
2314:
2236:
2184:
2129:
2104:"Genetics, development and evolution of adaptive pigmentation in vertebrates"
2029:
1978:
1934:
1810:
1753:
1649:
1565:
1522:
1446:
1388:
1293:
1235:
1064:
998:
931:
850:
816:
699:
597:
538:
1745:
1040:
418:
397:
265:
85:
32:
2444:
2367:
2332:
2271:
2192:
2137:
2120:
2103:
2088:
1986:
1888:
1853:
1761:
1667:
1641:
1530:
1513:
1496:
1454:
1406:
1365:"Approaches to Macroevolution: 1. General Concepts and Origin of Variation"
1301:
1253:
1123:
1082:
1006:
939:
707:
659:
589:
546:
142:
2383:
2037:
1702:
414:
367:
180:
because in vertebrates the fore-limbs and the anterior pair of wings are
81:
66:
25:
1096:
Drost, Hajk-Georg; Janitza, Philipp; Grosse, Ivo; Quint, Marcel (2017).
605:
2080:
1845:
1584:
Freaks of Nature What Anomalies Tell Us About Development and Evolution
1481:
651:
322:
2059:(1996). "Adaptive Radiation Along Genetic Lines of Least Resistance".
1425:
Olson, M.E. (2012). "The developmental renaissance in adaptationism".
1285:
1227:
226:
A study of the polydactyl toe counts of 375 Hemingway mutants of the
2176:
2072:
1784:
1681:
Gould, S.J. (1966). "Allometry and Size in Ontogeny and Phylogeny".
1041:"Developmental Bias and Evolution: A Regulatory Network Perspective"
973:"Perspective: Complex Adaptations and the Evolution of Evolvability"
643:
255:
1880:
1624:
Goswami, A.; Smaers, J. B.; Soligo, C.; Polly, P. D. (2014-08-19).
484:
166:
42:
2221:. Greenwood Village, Colorado: Roberts and Company Publishers.
1141:
Evolution and Biocomputation: Computational Models of Evolution
2291:"The Loci of Evolution: How Predictable is Genetic Evolution?"
222:
Biased number of polydactylous toes in a Main Coon population
1867:
Arnold, S.J. (1992). "Constraints on phenotypic evolution".
353:
Biased phenotypes II: Properties of gene regulatory networks
217:
156:
Nonrandom (anisotropic) distribution of phenotypic variation
2410:"Genotype networks shed light on evolutionary constraints"
1098:"Cross-kingdom comparison of the developmental hourglass"
2158:"The evolution of hierarchical gene regulatory networks"
65:
on previously ‘filtered’ variation during the course of
2219:
Evolution, development, & the predictable genome
877:
Zimmer, Carl.; Emlen D.; Perkins, Alison EH (2013).
413:
in nature implies that biological systems have more
24:
refers to the production against or towards certain
471:(1985). "Developmental constraints and evolution".
2486:Evolution, development, and the predictable genome
57:, which emphasizes the role of the organism as a
349:will bias the main direction of the trajectory.
105:covariation will facilitate adaptive evolution.
2156:Erwin, Douglas H.; Davidson, Eric H. (2009).
1102:Current Opinion in Genetics & Development
341:), which describes the direction of greatest
8:
2480:Homology, Genes, and Evolutionary Innovation
2348:Proceedings of the Royal Society of London B
746:Homology, Genes, and Evolutionary Innovation
570:Proceedings of the Royal Society of London B
234:Conversely, developmental abnormalities (or
197:Classical examples of anisotropic variation
366:Different vertebrate species have evolved
251:Developmental integration and the P-matrix
2322:
2119:
1800:
1657:
1512:
1396:
1333:10.1093/acprof:oso/9780199691883.001.0001
1243:
1113:
1072:
988:
845:. Cambridge: Cambridge University Press.
528:
246:Biased phenotypes I: Continuous variation
361:
310:
254:
200:
141:
99:
451:
2403:
2401:
2212:
2210:
2151:
2149:
2147:
2051:
2049:
2047:
1952:
1950:
1948:
1946:
1944:
1908:
1906:
1619:
1617:
1615:
1613:
1611:
1577:
1575:
1420:
1418:
1416:
1358:
1356:
1354:
1352:
963:
961:
959:
957:
278:Quantitative genetics and the G-matrix
1267:
1265:
1263:
1205:
1203:
1182:. Cambridge, MA: MIT Press. pp.
1034:
1032:
1030:
1028:
1026:
1024:
1022:
1020:
1018:
1016:
872:
870:
370:forms from parallel mutations at the
89:trajectory, a process referred to as
84:. However, natural selection acts on
7:
834:
832:
830:
828:
826:
779:
777:
775:
773:
739:
737:
735:
733:
731:
729:
727:
725:
692:10.1046/j.1525-142x.2001.003004271.x
673:
671:
669:
560:
558:
556:
506:
504:
502:
459:
457:
455:
133:Distribution of phenotypic variation
2474:Evolution: A developmental approach
171:Ontogenetically impossible creature
1838:10.1111/j.1558-5646.1983.tb00236.x
1695:10.1111/j.1469-185X.1966.tb01624.x
990:10.1111/j.1558-5646.1996.tb02339.x
881:. Greenwood Village, CO: Roberts.
299:most relevant parameters to study
14:
2417:Trends in Ecology & Evolution
1915:Trends in Ecology & Evolution
1427:Trends in Ecology & Evolution
2307:10.1111/j.1558-5646.2008.00450.x
2264:10.1111/j.1558-5646.2009.00761.x
1971:10.1111/j.1558-5646.2007.00071.x
1802:10.1046/j.1461-0248.2003.00428.x
924:10.1046/j.1525-142x.2000.00028.x
530:10.1111/j.1525-142x.2004.04033.x
358:Hierarchy and optimal pleiotropy
2507:Extended evolutionary synthesis
879:Evolution: Making sense of life
473:The Quarterly Review of Biology
2102:Hoekstra, H. E. (2006-07-05).
2002:Journal of Theoretical Biology
748:. Princeton University Press.
1:
2022:10.1016/s0022-5193(84)80050-8
1927:10.1016/S0169-5347(02)02505-3
1145:. Berlin: Springer. pp.
809:10.1016/s0016-6995(89)80006-3
2468:Biased Embryos and Evolution
842:Biased Embryos and Evolution
1586:. Oxford University Press.
1327:. Oxford University Press.
1274:Evolution & Development
1057:10.1534/genetics.118.300995
185:mutations occur randomly).
91:developmental reprogramming
2523:
2429:10.1016/j.tree.2011.07.001
1439:10.1016/j.tree.2011.12.005
744:Wagner, Gunter P. (2014).
1558:10.1007/s11692-013-9267-y
1381:10.1007/s11692-017-9420-0
1115:10.1016/j.gde.2017.03.003
971:; Altenberg, Lee (1996).
912:Evolution and Development
680:Evolution and Development
517:Evolution and Development
343:additive genetic variance
307:Paths of least resistance
296:additive genetic variance
205:Shell variation in nature
115:Developmental constraints
2408:Wagner, Andreas (2011).
1495:Arthur, Wallace (2002).
851:10.1017/cbo9780511606830
839:Arthur, Wallace (2004).
511:Arthur, Wallace (2004).
240:transpecific parallelism
212:soil-dwelling centipedes
129:direction of selection.
69:. It contrasts with the
26:ontogenetic trajectories
2165:Nature Reviews Genetics
1869:The American Naturalist
1746:10.1126/science.1056607
1582:Blumberg, M.S. (2009).
1470:Journal of Paleontology
2462:Ontogeny and Phylogeny
2368:10.1098/rspb.1994.0040
2121:10.1038/sj.hdy.6800861
1642:10.1098/rstb.2013.0254
1630:Phil. Trans. R. Soc. B
1514:10.1038/sj.hdy.6800139
1363:Jablonski, D. (2017).
590:10.1098/rspb.1979.0086
378:
317:
288:linkage disequilibrium
261:
223:
206:
172:
148:
106:
48:
39:The role of the embryo
2502:Developmental biology
1783:Pigliucci, M (2003).
365:
314:
283:Quantitative genetics
258:
221:
204:
170:
145:
103:
46:
2217:Stern, D.L. (2011).
1546:Evolutionary Biology
1369:Evolutionary Biology
386:transcription factor
162:evolutionary biology
18:evolutionary biology
2360:1994RSPSB.255..279S
2014:1984JThBi.110..155C
1738:2001Sci...291.1534E
1732:(5508): 1534–1536.
1323:Kemp, T.S. (2016).
801:1989Geobi..22...21A
582:1979RSPSB.205..581G
465:Maynard Smith, John
124:Developmental drive
1636:(1649): 20130254.
628:Gould, Stephen Jay
379:
318:
292:breeder’s equation
262:
224:
207:
173:
149:
107:
49:
22:developmental bias
2354:(1344): 279–284.
2258:(11): 2771–2789.
1593:978-0-1997-5064-1
1286:10.1111/ede.12098
1228:10.1111/nph.12969
1193:978-0-262-03326-8
1172:Simon, Herbert A.
1156:978-3-540-49176-7
969:Wagner, Günter P.
576:(1161): 581–598.
411:neutral mutations
63:natural selection
2514:
2449:
2448:
2414:
2405:
2396:
2395:
2343:
2337:
2336:
2326:
2301:(9): 2155–2177.
2282:
2276:
2275:
2247:
2241:
2240:
2214:
2205:
2204:
2162:
2153:
2142:
2141:
2123:
2099:
2093:
2092:
2067:(5): 1766–1774.
2053:
2042:
2041:
1997:
1991:
1990:
1954:
1939:
1938:
1910:
1901:
1900:
1864:
1858:
1857:
1832:(6): 1210–1226.
1821:
1815:
1814:
1804:
1780:
1774:
1773:
1721:
1715:
1714:
1678:
1672:
1671:
1661:
1621:
1606:
1605:
1579:
1570:
1569:
1541:
1535:
1534:
1516:
1492:
1486:
1485:
1476:(5): 1178–1190.
1465:
1459:
1458:
1422:
1411:
1410:
1400:
1360:
1347:
1346:
1320:
1314:
1313:
1269:
1258:
1257:
1247:
1207:
1198:
1197:
1181:
1167:
1161:
1160:
1144:
1134:
1128:
1127:
1117:
1093:
1087:
1086:
1076:
1036:
1011:
1010:
992:
965:
952:
951:
907:
901:
900:
874:
865:
864:
836:
821:
820:
781:
768:
767:
741:
720:
719:
675:
664:
663:
624:
618:
617:
562:
551:
550:
532:
508:
497:
496:
461:
423:neutral networks
405:Neutral networks
2522:
2521:
2517:
2516:
2515:
2513:
2512:
2511:
2492:
2491:
2458:
2456:Further reading
2453:
2452:
2423:(11): 577–584.
2412:
2407:
2406:
2399:
2345:
2344:
2340:
2284:
2283:
2279:
2249:
2248:
2244:
2229:
2216:
2215:
2208:
2177:10.1038/nrg2499
2160:
2155:
2154:
2145:
2101:
2100:
2096:
2073:10.2307/2410734
2057:Schluter, Dolph
2055:
2054:
2045:
1999:
1998:
1994:
1956:
1955:
1942:
1912:
1911:
1904:
1866:
1865:
1861:
1823:
1822:
1818:
1789:Ecology Letters
1782:
1781:
1777:
1723:
1722:
1718:
1680:
1679:
1675:
1623:
1622:
1609:
1594:
1581:
1580:
1573:
1543:
1542:
1538:
1494:
1493:
1489:
1467:
1466:
1462:
1424:
1423:
1414:
1362:
1361:
1350:
1343:
1322:
1321:
1317:
1271:
1270:
1261:
1216:New Phytologist
1209:
1208:
1201:
1194:
1169:
1168:
1164:
1157:
1136:
1135:
1131:
1095:
1094:
1090:
1038:
1037:
1014:
967:
966:
955:
909:
908:
904:
889:
876:
875:
868:
861:
838:
837:
824:
783:
782:
771:
756:
743:
742:
723:
677:
676:
667:
644:10.2307/2409056
626:
625:
621:
564:
563:
554:
510:
509:
500:
463:
462:
453:
448:
431:
407:
360:
355:
348:
340:
309:
280:
253:
248:
199:
158:
140:
135:
126:
117:
112:
41:
12:
11:
5:
2520:
2518:
2510:
2509:
2504:
2494:
2493:
2490:
2489:
2483:
2482:(Wagner, 2014)
2477:
2476:(Arthur, 2010)
2471:
2470:(Arthur, 2004)
2465:
2457:
2454:
2451:
2450:
2397:
2338:
2277:
2242:
2228:978-1936221011
2227:
2206:
2171:(2): 141–148.
2143:
2114:(3): 222–234.
2094:
2043:
2008:(2): 155–171.
1992:
1965:(4): 727–745.
1940:
1921:(7): 320–327.
1902:
1881:10.1086/285398
1859:
1816:
1795:(3): 265–272.
1775:
1716:
1689:(4): 587–640.
1673:
1607:
1592:
1571:
1552:(2): 262–275.
1536:
1507:(4): 239–246.
1487:
1460:
1433:(5): 278–287.
1412:
1375:(4): 427–450.
1348:
1341:
1315:
1280:(6): 373–381.
1259:
1222:(4): 841–853.
1199:
1192:
1162:
1155:
1129:
1088:
1051:(4): 949–966.
1012:
983:(3): 967–976.
953:
902:
887:
866:
859:
822:
769:
755:978-0691180670
754:
721:
686:(4): 271–278.
665:
638:(3): 516–539.
619:
552:
523:(4): 282–288.
498:
485:10.1086/414425
479:(3): 265–287.
450:
449:
447:
444:
443:
442:
437:
430:
427:
406:
403:
359:
356:
354:
351:
346:
338:
308:
305:
279:
276:
252:
249:
247:
244:
228:Maine Coon cat
215:odd numbers.
198:
195:
157:
154:
139:
136:
134:
131:
125:
122:
116:
113:
111:
108:
40:
37:
13:
10:
9:
6:
4:
3:
2:
2519:
2508:
2505:
2503:
2500:
2499:
2497:
2488:(Stern, 2011)
2487:
2484:
2481:
2478:
2475:
2472:
2469:
2466:
2464:(Gould, 1977)
2463:
2460:
2459:
2455:
2446:
2442:
2438:
2434:
2430:
2426:
2422:
2418:
2411:
2404:
2402:
2398:
2393:
2389:
2385:
2381:
2377:
2373:
2369:
2365:
2361:
2357:
2353:
2349:
2342:
2339:
2334:
2330:
2325:
2320:
2316:
2312:
2308:
2304:
2300:
2296:
2292:
2288:
2285:Stern, D.L.;
2281:
2278:
2273:
2269:
2265:
2261:
2257:
2253:
2246:
2243:
2238:
2234:
2230:
2224:
2220:
2213:
2211:
2207:
2202:
2198:
2194:
2190:
2186:
2182:
2178:
2174:
2170:
2166:
2159:
2152:
2150:
2148:
2144:
2139:
2135:
2131:
2127:
2122:
2117:
2113:
2109:
2105:
2098:
2095:
2090:
2086:
2082:
2078:
2074:
2070:
2066:
2062:
2058:
2052:
2050:
2048:
2044:
2039:
2035:
2031:
2027:
2023:
2019:
2015:
2011:
2007:
2003:
1996:
1993:
1988:
1984:
1980:
1976:
1972:
1968:
1964:
1960:
1953:
1951:
1949:
1947:
1945:
1941:
1936:
1932:
1928:
1924:
1920:
1916:
1909:
1907:
1903:
1898:
1894:
1890:
1886:
1882:
1878:
1874:
1870:
1863:
1860:
1855:
1851:
1847:
1843:
1839:
1835:
1831:
1827:
1820:
1817:
1812:
1808:
1803:
1798:
1794:
1790:
1786:
1779:
1776:
1771:
1767:
1763:
1759:
1755:
1751:
1747:
1743:
1739:
1735:
1731:
1727:
1720:
1717:
1712:
1708:
1704:
1700:
1696:
1692:
1688:
1684:
1677:
1674:
1669:
1665:
1660:
1655:
1651:
1647:
1643:
1639:
1635:
1631:
1627:
1620:
1618:
1616:
1614:
1612:
1608:
1603:
1599:
1595:
1589:
1585:
1578:
1576:
1572:
1567:
1563:
1559:
1555:
1551:
1547:
1540:
1537:
1532:
1528:
1524:
1520:
1515:
1510:
1506:
1502:
1498:
1491:
1488:
1483:
1479:
1475:
1471:
1464:
1461:
1456:
1452:
1448:
1444:
1440:
1436:
1432:
1428:
1421:
1419:
1417:
1413:
1408:
1404:
1399:
1394:
1390:
1386:
1382:
1378:
1374:
1370:
1366:
1359:
1357:
1355:
1353:
1349:
1344:
1342:9780199691883
1338:
1334:
1330:
1326:
1319:
1316:
1311:
1307:
1303:
1299:
1295:
1291:
1287:
1283:
1279:
1275:
1268:
1266:
1264:
1260:
1255:
1251:
1246:
1241:
1237:
1233:
1229:
1225:
1221:
1217:
1213:
1206:
1204:
1200:
1195:
1189:
1185:
1180:
1179:
1173:
1166:
1163:
1158:
1152:
1148:
1143:
1142:
1133:
1130:
1125:
1121:
1116:
1111:
1107:
1103:
1099:
1092:
1089:
1084:
1080:
1075:
1070:
1066:
1062:
1058:
1054:
1050:
1046:
1042:
1035:
1033:
1031:
1029:
1027:
1025:
1023:
1021:
1019:
1017:
1013:
1008:
1004:
1000:
996:
991:
986:
982:
978:
974:
970:
964:
962:
960:
958:
954:
949:
945:
941:
937:
933:
929:
925:
921:
917:
913:
906:
903:
898:
894:
890:
888:9781319202590
884:
880:
873:
871:
867:
862:
860:9780511606830
856:
852:
848:
844:
843:
835:
833:
831:
829:
827:
823:
818:
814:
810:
806:
802:
798:
794:
790:
786:
785:Alberch, Pere
780:
778:
776:
774:
770:
765:
761:
757:
751:
747:
740:
738:
736:
734:
732:
730:
728:
726:
722:
717:
713:
709:
705:
701:
697:
693:
689:
685:
681:
674:
672:
670:
666:
661:
657:
653:
649:
645:
641:
637:
633:
629:
623:
620:
615:
611:
607:
603:
599:
595:
591:
587:
583:
579:
575:
571:
567:
561:
559:
557:
553:
548:
544:
540:
536:
531:
526:
522:
518:
514:
507:
505:
503:
499:
494:
490:
486:
482:
478:
474:
470:
466:
460:
458:
456:
452:
445:
441:
438:
436:
433:
432:
428:
426:
424:
420:
416:
412:
404:
402:
399:
394:
392:
387:
383:
376:
374:
369:
364:
357:
352:
350:
344:
336:
332:
326:
324:
313:
306:
304:
302:
297:
293:
289:
284:
277:
275:
272:
267:
257:
250:
245:
243:
241:
237:
232:
229:
220:
216:
213:
203:
196:
194:
192:
186:
183:
179:
169:
165:
163:
155:
153:
144:
137:
132:
130:
123:
121:
114:
110:Types of bias
109:
102:
98:
94:
92:
87:
83:
79:
78:neo-Darwinian
74:
72:
71:Functionalist
68:
64:
60:
56:
55:
54:Structuralism
45:
38:
36:
34:
31:
27:
23:
19:
2485:
2479:
2473:
2467:
2461:
2420:
2416:
2351:
2347:
2341:
2298:
2294:
2287:Orgogozo, V.
2280:
2255:
2251:
2245:
2218:
2168:
2164:
2111:
2107:
2097:
2064:
2060:
2005:
2001:
1995:
1962:
1958:
1918:
1914:
1875:: S85–S107.
1872:
1868:
1862:
1829:
1825:
1819:
1792:
1788:
1778:
1729:
1725:
1719:
1686:
1682:
1676:
1633:
1629:
1583:
1549:
1545:
1539:
1504:
1500:
1490:
1473:
1469:
1463:
1430:
1426:
1372:
1368:
1324:
1318:
1277:
1273:
1219:
1215:
1177:
1165:
1140:
1132:
1105:
1101:
1091:
1048:
1044:
980:
976:
918:(1): 49–57.
915:
911:
905:
878:
841:
792:
788:
745:
683:
679:
635:
631:
622:
573:
569:
566:Gould, S. J.
520:
516:
476:
472:
435:Evolvability
422:
408:
395:
380:
372:
331:Evolvability
327:
319:
301:evolvability
281:
263:
239:
236:teratologies
233:
225:
208:
187:
177:
174:
159:
150:
127:
118:
95:
90:
75:
70:
58:
52:
50:
21:
15:
469:Wolpert, L.
391:pleiotropic
335:eigenvector
271:pleiotropic
138:Morphospace
2496:Categories
1602:1058406207
897:1051973071
764:1005108561
446:References
440:Speciation
419:phenotypes
266:allometric
182:homologous
178:impossible
86:phenotypes
2437:0169-5347
2376:0962-8452
2315:0014-3820
2295:Evolution
2252:Evolution
2237:762460688
2185:1471-0056
2130:0018-067X
2061:Evolution
2030:0022-5193
1979:0014-3820
1959:Evolution
1935:0169-5347
1826:Evolution
1811:1461-023X
1754:0036-8075
1683:Biol. Rev
1650:0962-8436
1566:0071-3260
1523:0018-067X
1447:0169-5347
1389:0071-3260
1294:1520-541X
1236:0028-646X
1108:: 69–75.
1065:0016-6731
999:0014-3820
977:Evolution
932:1520-541X
817:0016-6995
795:: 21–57.
700:1520-541X
632:Evolution
598:0080-4649
539:1520-541X
415:genotypes
398:epistatic
82:mutations
33:evolution
2445:21840080
2392:12021473
2333:18616572
2289:(2008).
2272:19545263
2193:19139764
2138:16823403
2108:Heredity
2089:28565589
1987:17439608
1889:19426028
1854:28556011
1770:24821274
1762:11222856
1711:28606846
1668:25002699
1531:12242638
1501:Heredity
1455:22326724
1407:29142333
1310:21562182
1302:25212955
1254:25539005
1174:(eds.).
1124:28347942
1083:30049818
1045:Genetics
1007:28565291
948:11972625
940:11256417
716:41698287
708:11478524
660:28568388
547:15230968
493:85201850
429:See also
147:species.
67:ontogeny
2384:7517565
2356:Bibcode
2324:2613234
2201:7613857
2081:2410734
2038:6492829
2010:Bibcode
1897:5965825
1846:2408842
1734:Bibcode
1726:Science
1703:5342162
1659:4084539
1482:1301992
1398:5661017
1245:5526441
1074:6063245
797:Bibcode
789:Geobios
652:2409056
614:2129408
578:Bibcode
368:melanic
337:of G (g
323:species
191:fitness
2443:
2435:
2390:
2382:
2374:
2331:
2321:
2313:
2270:
2235:
2225:
2199:
2191:
2183:
2136:
2128:
2087:
2079:
2036:
2028:
1985:
1977:
1933:
1895:
1887:
1852:
1844:
1809:
1768:
1760:
1752:
1709:
1701:
1666:
1656:
1648:
1600:
1590:
1564:
1529:
1521:
1480:
1453:
1445:
1405:
1395:
1387:
1339:
1308:
1300:
1292:
1252:
1242:
1234:
1190:
1186:–128.
1153:
1149:–259.
1122:
1081:
1071:
1063:
1005:
997:
946:
938:
930:
895:
885:
857:
815:
762:
752:
714:
706:
698:
658:
650:
612:
604:
596:
545:
537:
491:
59:causal
2413:(PDF)
2388:S2CID
2197:S2CID
2161:(PDF)
2077:JSTOR
1893:S2CID
1842:JSTOR
1766:S2CID
1707:S2CID
1478:JSTOR
1306:S2CID
944:S2CID
712:S2CID
648:JSTOR
610:S2CID
606:42062
489:S2CID
417:than
210:that
30:trait
2441:PMID
2433:ISSN
2380:PMID
2372:ISSN
2329:PMID
2311:ISSN
2268:PMID
2233:OCLC
2223:ISBN
2189:PMID
2181:ISSN
2134:PMID
2126:ISSN
2085:PMID
2034:PMID
2026:ISSN
1983:PMID
1975:ISSN
1931:ISSN
1885:PMID
1850:PMID
1807:ISSN
1758:PMID
1750:ISSN
1699:PMID
1664:PMID
1646:ISSN
1598:OCLC
1588:ISBN
1562:ISSN
1527:PMID
1519:ISSN
1451:PMID
1443:ISSN
1403:PMID
1385:ISSN
1337:ISBN
1298:PMID
1290:ISSN
1250:PMID
1232:ISSN
1188:ISBN
1151:ISBN
1120:PMID
1079:PMID
1061:ISSN
1003:PMID
995:ISSN
936:PMID
928:ISSN
893:OCLC
883:ISBN
855:ISBN
813:ISSN
760:OCLC
750:ISBN
704:PMID
696:ISSN
656:PMID
602:PMID
594:ISSN
543:PMID
535:ISSN
382:GRNs
375:gene
373:mc1r
2425:doi
2364:doi
2352:255
2319:PMC
2303:doi
2260:doi
2173:doi
2116:doi
2069:doi
2018:doi
2006:110
1967:doi
1923:doi
1877:doi
1873:140
1834:doi
1797:doi
1742:doi
1730:291
1691:doi
1654:PMC
1638:doi
1634:369
1554:doi
1509:doi
1435:doi
1393:PMC
1377:doi
1329:doi
1282:doi
1240:PMC
1224:doi
1220:204
1147:205
1110:doi
1069:PMC
1053:doi
1049:209
985:doi
920:doi
847:doi
805:doi
688:doi
640:doi
586:doi
574:205
525:doi
481:doi
347:max
339:max
16:In
2498::
2439:.
2431:.
2421:26
2419:.
2415:.
2400:^
2386:.
2378:.
2370:.
2362:.
2350:.
2327:.
2317:.
2309:.
2299:62
2297:.
2293:.
2266:.
2256:63
2254:.
2231:.
2209:^
2195:.
2187:.
2179:.
2169:10
2167:.
2163:.
2146:^
2132:.
2124:.
2112:97
2110:.
2106:.
2083:.
2075:.
2065:50
2063:.
2046:^
2032:.
2024:.
2016:.
2004:.
1981:.
1973:.
1963:61
1961:.
1943:^
1929:.
1919:17
1917:.
1905:^
1891:.
1883:.
1871:.
1848:.
1840:.
1830:37
1828:.
1805:.
1791:.
1787:.
1764:.
1756:.
1748:.
1740:.
1728:.
1705:.
1697:.
1687:41
1685:.
1662:.
1652:.
1644:.
1632:.
1628:.
1610:^
1596:.
1574:^
1560:.
1550:41
1548:.
1525:.
1517:.
1505:89
1503:.
1499:.
1474:40
1472:.
1449:.
1441:.
1431:27
1429:.
1415:^
1401:.
1391:.
1383:.
1373:44
1371:.
1367:.
1351:^
1335:.
1304:.
1296:.
1288:.
1278:16
1276:.
1262:^
1248:.
1238:.
1230:.
1218:.
1214:.
1202:^
1184:99
1118:.
1106:45
1104:.
1100:.
1077:.
1067:.
1059:.
1047:.
1043:.
1015:^
1001:.
993:.
981:50
979:.
975:.
956:^
942:.
934:.
926:.
914:.
891:.
869:^
853:.
825:^
811:.
803:.
793:22
791:.
772:^
758:.
724:^
710:.
702:.
694:.
682:.
668:^
654:.
646:.
636:43
634:.
608:.
600:.
592:.
584:.
572:.
555:^
541:.
533:.
519:.
515:.
501:^
487:.
477:60
475:.
454:^
377:.
20:,
2447:.
2427::
2394:.
2366::
2358::
2335:.
2305::
2274:.
2262::
2239:.
2203:.
2175::
2140:.
2118::
2091:.
2071::
2040:.
2020::
2012::
1989:.
1969::
1937:.
1925::
1899:.
1879::
1856:.
1836::
1813:.
1799::
1793:6
1772:.
1744::
1736::
1713:.
1693::
1670:.
1640::
1604:.
1568:.
1556::
1533:.
1511::
1484:.
1457:.
1437::
1409:.
1379::
1345:.
1331::
1312:.
1284::
1256:.
1226::
1196:.
1159:.
1126:.
1112::
1085:.
1055::
1009:.
987::
950:.
922::
916:2
899:.
863:.
849::
819:.
807::
799::
766:.
718:.
690::
684:3
662:.
642::
616:.
588::
580::
549:.
527::
521:6
495:.
483::
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.