26:
125:
267:"The Schizosaccharomyces pombe spindle checkpoint protein mad2p blocks anaphase and genetically interacts with the anaphase-promoting complex"
324:
Aravind L, Koonin EV (1998). "The HORMA domain: a common structural denominator in mitotic checkpoints, chromosome synapsis and DNA repair".
73:
362:
145:
395:
133:
129:
278:
86:
190:
341:
306:
120:
333:
296:
286:
112:
218:
194:
282:
30:
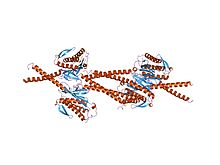
crystal structure of mad1-mad2 reveals a conserved mad2 binding motif in mad1 and cdc20.
178:
337:
389:
301:
266:
78:
54:
108:
381:
66:
366:
210:
186:
214:
206:
213:, and MAD2 is a spindle checkpoint protein which prevents progression of the
291:
182:
345:
310:
82:
25:
377:
61:
237:
233:
202:
198:
174:
245:
241:
140:
373:
102:
49:
372:
This article incorporates text from the public domain
139:
119:
101:
96:
72:
60:
48:
40:
35:
18:
229:Humans proteins containing this domain include:
8:
197:and act as an adaptor that recruits other
93:
24:
300:
290:
205:-specific protein, Rev7 is required for
257:
15:
181:that has been suggested to recognise
7:
265:He X, Patterson TE, Sazer S (1997).
14:
363:Eukaryotic Linear Motif resource
193:breaks or non-attachment to the
217:upon detection of a defect in
1:
338:10.1016/S0968-0004(98)01257-2
97:Available protein structures:
412:
371:
157:In molecular biology, the
92:
23:
271:Proc Natl Acad Sci U S A
292:10.1073/pnas.94.15.7965
185:states resulting from
283:1997PNAS...94.7965H
326:Trends Biochem Sci
161:(named after the
155:
154:
151:
150:
146:structure summary
403:
350:
349:
321:
315:
314:
304:
294:
262:
94:
28:
16:
411:
410:
406:
405:
404:
402:
401:
400:
396:Protein domains
386:
385:
384:
359:
354:
353:
323:
322:
318:
277:(15): 7965–70.
264:
263:
259:
254:
227:
219:mitotic spindle
191:double stranded
31:
12:
11:
5:
409:
407:
399:
398:
388:
387:
370:
369:
358:
357:External links
355:
352:
351:
316:
256:
255:
253:
250:
249:
248:
226:
223:
179:protein domain
153:
152:
149:
148:
143:
137:
136:
123:
117:
116:
106:
99:
98:
90:
89:
76:
70:
69:
64:
58:
57:
52:
46:
45:
42:
38:
37:
33:
32:
29:
21:
20:
13:
10:
9:
6:
4:
3:
2:
408:
397:
394:
393:
391:
383:
379:
375:
368:
364:
361:
360:
356:
347:
343:
339:
335:
331:
327:
320:
317:
312:
308:
303:
298:
293:
288:
284:
280:
276:
272:
268:
261:
258:
251:
247:
243:
239:
235:
232:
231:
230:
224:
222:
220:
216:
212:
208:
204:
200:
196:
192:
188:
184:
180:
176:
172:
168:
164:
160:
147:
144:
142:
138:
135:
131:
127:
124:
122:
118:
114:
110:
107:
104:
100:
95:
91:
88:
84:
80:
77:
75:
71:
68:
65:
63:
59:
56:
53:
51:
47:
43:
39:
34:
27:
22:
17:
365:motif class
332:(8): 284–6.
329:
325:
319:
274:
270:
260:
228:
201:. Hop1 is a
170:
166:
162:
159:HORMA domain
158:
156:
19:HORMA domain
221:integrity.
211:mutagenesis
187:DNA adducts
36:Identifiers
252:References
215:cell cycle
207:DNA damage
109:structures
382:IPR003511
183:chromatin
169:ev7p and
67:IPR003511
390:Category
378:InterPro
367:LIG_MAD2
225:Examples
209:induced
199:proteins
175:proteins
126:RCSB PDB
62:InterPro
346:9757827
311:9223296
279:Bibcode
238:HORMAD2
234:HORMAD1
203:meiosis
195:spindle
177:) is a
55:PF02301
344:
309:
299:
246:MAD2L2
242:MAD2L1
141:PDBsum
115:
105:
87:SUPFAM
41:Symbol
302:21538
165:p1p,
83:SCOPe
74:SCOP2
44:HORMA
376:and
374:Pfam
342:PMID
307:PMID
134:PDBj
130:PDBe
113:ECOD
103:Pfam
79:1duj
50:Pfam
334:doi
297:PMC
287:doi
173:D2
121:PDB
392::
380::
340:.
330:23
328:.
305:.
295:.
285:.
275:94
273:.
269:.
244:,
240:,
236:,
189:,
171:MA
163:Ho
132:;
128:;
111:/
85:/
81:/
348:.
336::
313:.
289::
281::
167:R
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.