142:
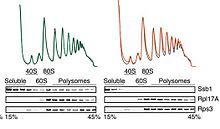
can first be detected in the 40S fraction, then nearly disappears from the 60S fraction (the separations on these gradients are not absolute), then reappears in the 80S and polysome fractions. This indicates that there is at most very little of the protein found in the cell that is not part of the small subunit. In contrast, in the upper row of the immunoblot figure, a soluble protein appears in the soluble fractions and associated with ribosomes and polysomes. The particular protein is a
17:
122:
132:
After centrifugation, the contents of the tube are collected as fractions from the top (smaller, slower traveling) to bottom (bigger, faster traveling) and the optical density of the fractions is determined. The first fractions removed have a large amount of relatively small molecules, such as tRNAs,
163:
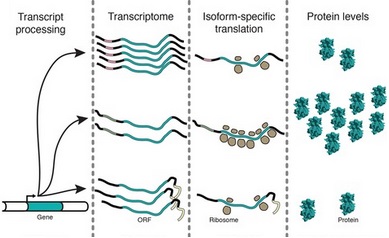
The technique can also be used to study the degree of translation of a particular mRNA In these experiments, 5' and 3' sequences of an mRNA were investigated for their effects on amount of mRNA produced and how well the mRNAs were translated. As shown, not all mRNA isoforms are translated with the
141:
It is possible to use this technique to study the overall degree of translation in cells (for examples), but it can be used much more specifically to study individual proteins and their mRNAs. As an example shown in the lower portion of the figure, a protein that composes part of the small subunit
113:
and thus propels them through the gradient based upon how "big" the individual components are. The small (40S) subunits travel less far into the gradient than the large (60S) subunits. The 80S ribosomes on an mRNA travel further (note that the contribution of the size of the mRNA to the distance
102:) of the lysate is then layered gently on top of the gradient in the tube. The lysate, even though it contains a large amount of soluble material, is much less dense than 15% sucrose, and so it can be kept as a separate layer at the top of the tube if this is done gently.
293:
90:
tube. At the concentrations used (15-45% in the example), sucrose does not disrupt the association of ribosomes and mRNA. The 15% portion of the gradient is at the top of the tube, while the 45% portion is at the bottom because of their different
47:, but the data they generate are at very different levels of specificity. When employed by experts, the technique is remarkably reproducible: the 3 profiles in the first image are from 3 different experiments.
114:
traveled is not significant). Polysomes composed of 2 ribosomes travel further, polysomes with 3 ribosomes travel further still, and on and on. The "size" of the components is designated by S, the
154:
294:"The antidepressant sertraline inhibits translation initiation by curtailing mammalian target of rapamycin signaling"
490:
237:"Multivalent contacts of the Hsp70 Ssb contribute to its architecture on ribosomes and nascent chain interaction"
105:
In order to separate the components of the lysate, the preparation is subjected to centrifugation. This
248:
118:
unit. Note that one S = 10 seconds, and that the concept of "big" is actually an oversimplification.
143:
214:
40:
383:"The rate of metabolism as a factor determining longevity of the Saccharomyces cerevisiae yeast"
466:
412:
363:
318:
274:
206:
28:
456:
446:
402:
394:
353:
345:
308:
264:
256:
198:
186:
110:
99:
252:
461:
434:
407:
382:
358:
337:
269:
236:
160:
in the paper showed, there is a direct association of the chaperone with the ribosome.
484:
75:) and large (60S in eukaryotes) ribosomal subunits, "free" mRNA and a host of other
338:"Analysis of translation initiation during stress conditions by polysome profiling"
218:
106:
313:
56:
44:
398:
126:
87:
72:
187:"Translational control of immune responses: from transcripts to translatomes"
60:
470:
416:
367:
322:
278:
210:
153:
121:
43:. Both techniques have been reviewed and both are used in analysis of the
115:
68:
36:
451:
260:
147:
92:
83:
76:
16:
202:
349:
120:
32:
15:
435:"Tunable protein synthesis by transcript isoforms in human cells"
64:
39:. It is important to note that this technique is different from
152:
150:
as it is being extruded from the ribosome. As other work
63:, monosomes (composed of one ribosome residing on an
109:the components of the lysate with many times the
86:gradient of continuously variable density in a
82:The procedure continues by making a continuous
59:of the cells of interest. This lysate contains
230:
228:
146:, which (in brief) helps to fold the nascent
8:
428:
426:
460:
450:
406:
357:
312:
268:
31:that is used to study the association of
177:
168:their coding sequences are the same.
7:
185:Piccirillo, CA; et al. (2014).
235:Hanebuth, MA; et al. (2016).
98:A specific amount (as measured by
14:
342:Journal of Visualized Experiments
55:The procedure begins by making a
336:Coudert, L; et al. (2014).
433:Floor, SN; Doudna, JA (2016).
381:Molon, M; et al. (2016).
1:
314:10.1158/0008-5472.CAN-09-4072
292:Lin, CJ; et al. (2010).
387:Age (Dordrecht, Netherlands)
507:
133:individual proteins, etc.
399:10.1007/s11357-015-9868-8
157:
129:
21:
241:Nature Communications
156:
125:sucrose gradient and
124:
79:cellular components.
19:
452:10.7554/eLife.10921
261:10.1038/ncomms13695
253:2016NatCo...713695H
158:
130:
41:ribosome profiling
27:is a technique in
25:Polysome profiling
22:
491:Molecular biology
191:Nature Immunology
144:chaperone protein
29:molecular biology
498:
475:
474:
464:
454:
430:
421:
420:
410:
378:
372:
371:
361:
333:
327:
326:
316:
307:(8): 3199–3208.
298:
289:
283:
282:
272:
232:
223:
222:
182:
164:same efficiency
111:force of gravity
67:), the small (40
506:
505:
501:
500:
499:
497:
496:
495:
481:
480:
479:
478:
432:
431:
424:
380:
379:
375:
335:
334:
330:
301:Cancer Research
296:
291:
290:
286:
234:
233:
226:
203:10.1038/ni.2891
184:
183:
179:
174:
139:
100:optical density
53:
12:
11:
5:
504:
502:
494:
493:
483:
482:
477:
476:
422:
373:
328:
284:
224:
197:(6): 503–511.
176:
175:
173:
170:
138:
135:
52:
49:
13:
10:
9:
6:
4:
3:
2:
503:
492:
489:
488:
486:
472:
468:
463:
458:
453:
448:
444:
440:
436:
429:
427:
423:
418:
414:
409:
404:
400:
396:
392:
388:
384:
377:
374:
369:
365:
360:
355:
351:
350:10.3791/51164
347:
343:
339:
332:
329:
324:
320:
315:
310:
306:
302:
295:
288:
285:
280:
276:
271:
266:
262:
258:
254:
250:
246:
242:
238:
231:
229:
225:
220:
216:
212:
208:
204:
200:
196:
192:
188:
181:
178:
171:
169:
167:
161:
155:
151:
149:
145:
136:
134:
128:
123:
119:
117:
112:
108:
103:
101:
96:
94:
89:
85:
80:
78:
74:
70:
66:
62:
58:
51:The procedure
50:
48:
46:
42:
38:
34:
30:
26:
18:
442:
438:
390:
386:
376:
341:
331:
304:
300:
287:
244:
240:
194:
190:
180:
165:
162:
159:
140:
137:Applications
131:
104:
97:
81:
54:
24:
23:
166:even though
107:accelerates
57:cell lysate
45:translatome
172:References
127:immunoblot
88:centrifuge
73:eukaryotes
393:(1): 11.
247:: 13695.
61:polysomes
37:ribosomes
20:Polysomes
485:Category
471:26735365
417:26783001
368:24893838
323:20354178
279:27917864
211:24840981
116:svedberg
462:4764583
408:5005888
359:4193336
270:5150220
249:Bibcode
219:6269940
148:peptide
93:density
84:sucrose
77:soluble
469:
459:
415:
405:
366:
356:
344:(87).
321:
277:
267:
217:
209:
439:eLife
297:(PDF)
215:S2CID
35:with
33:mRNAs
467:PMID
413:PMID
364:PMID
319:PMID
275:PMID
207:PMID
65:mRNA
457:PMC
447:doi
403:PMC
395:doi
354:PMC
346:doi
309:doi
265:PMC
257:doi
199:doi
71:in
487::
465:.
455:.
445:.
441:.
437:.
425:^
411:.
401:.
391:38
389:.
385:.
362:.
352:.
340:.
317:.
305:70
303:.
299:.
273:.
263:.
255:.
243:.
239:.
227:^
213:.
205:.
195:15
193:.
189:.
95:.
473:.
449::
443:5
419:.
397::
370:.
348::
325:.
311::
281:.
259::
251::
245:7
221:.
201::
69:S
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.