32:
20:
120:
It is possible to identify a limited class of pseudoknots using dynamic programming, but these methods are not exhaustive and scale worse as a function of sequence length than non-pseudoknotted algorithms. The general problem of predicting lowest free energy structures with pseudoknots has been shown
153:
Many types of pseudoknots exist, differing by how they cross and how many times they cross. To reflect this difference, pseudoknots are classed into H-, K-, L-, M-types, with each successive type adding a layer of step intercalation. The simple telomerase P2b-P3 example in the article, for example,
161:
indicating basepairs in a stem and dots representing loops. The interrupted stems of pseudoknots mean that such notation must be extended with extra brackets, or even letters, so that different sets of stems can be represented. One such extension uses, in nesting order,
173:.]]]]]]. drawing 1 CGCGCGCUGUUUUUCUCGCUGACUUUCAGCGGGCGA---AAAAAAUGUCAGCU 50 ALIGN |.||||||||||||||||||||||||| .|.| |||||| ||||||. 1ymo 1 ---GGGCUGUUUUUCUCGCUGACUUUCAGC--CCCAAACAAAAAA-GUCAGCA 47 ((((((........
97:
in pseudoknots is not well nested; that is, base pairs occur that "overlap" one another in sequence position. This makes the presence of pseudoknots in RNA sequences more difficult to
588:
336:
Dirks, R.M. Pierce N.A. (2004) An algorithm for computing nucleic acid base-pairing probabilities including pseudoknots. "J Computation
Chemistry". 25:1295-1304, 2004.
93:
The structural configuration of pseudoknots does not lend itself well to bio-computational detection due to its context-sensitivity or "overlapping" nature. The
145:
contains a pseudoknot that is critical for activity, and several viruses use a pseudoknot structure to form a tRNA-like motif to infiltrate the host cell.
180:
Note that U bulge at the end is normally present in telomerase RNA. It was removed in the 1ymo solution model for enhanced stability of the pseudoknot.
105:, which use a recursive scoring system to identify paired stems and consequently, most cannot detect non-nested base pairs. The newer method of
686:
581:
110:
117:
will not predict pseudoknot structures present in a query sequence; they will only identify the more stable of the two pseudoknot stems.
73:
structures in which half of one stem is intercalated between the two halves of another stem. The pseudoknot was first recognized in the
691:
681:
98:
671:
66:
35:
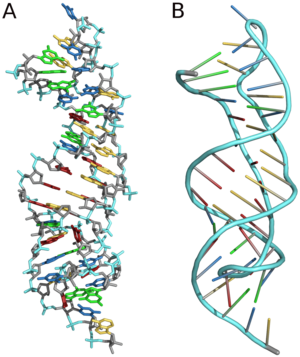
Three dimensional structure of almost the same pseudoknot from telomerase RNA. (A) sticks (B) backbone. The pdb-file is based on
676:
574:
106:
638:
628:
133:
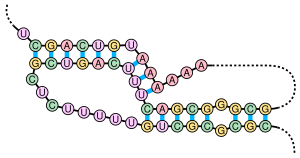
Several important biological processes rely on RNA molecules that form pseudoknots, which are often RNAs with extensive
701:
618:
114:
623:
74:
757:
613:
520:"Structure of the human telomerase RNA pseudoknot reveals conserved tertiary interactions essential for function"
320:
Rivas E, Eddy S. (1999). "A dynamic programming algorithm for RNA structure prediction including pseudoknots".
142:
658:
648:
597:
189:
134:
361:
Lyngsø, R. B. (2004). Complexity of pseudoknot prediction in simple models. Paper presented at the ICALP.
666:
157:
RNA secondary structure is usually represented by the dot-bracket notation, with pairing round brackets
224:
174:
166:
for closing. The structure for the two (slightly varying) telomerase examples, in this notation, is:
170:
727:
696:
102:
78:
633:
605:
541:
500:
451:
402:
303:
252:
37:
752:
531:
490:
482:
441:
433:
392:
384:
293:
283:
242:
232:
82:
77:
in 1982. Pseudoknots fold into knot-shaped three-dimensional conformations but are not true
471:"New algorithms to represent complex pseudoknotted RNA structures in dot-bracket notation"
109:
suffers from the same problem. Thus, popular secondary structure prediction methods like
469:
Antczak, M; Popenda, M; Zok, T; Zurkowski, M; Adamiak, RW; Szachniuk, M (15 April 2018).
228:
23:
This example of a naturally occurring pseudoknot is found in the RNA component of human
722:
643:
495:
470:
446:
421:
298:
271:
247:
212:
397:
372:
774:
486:
437:
345:
Lyngsø RB, Pedersen CN. (2000). "RNA pseudoknot prediction in energy-based models".
780:
737:
536:
519:
288:
122:
217:
Proceedings of the
National Academy of Sciences of the United States of America
732:
31:
24:
388:
237:
94:
70:
545:
504:
455:
307:
256:
85:
framework, which, in contrast to knot theory, is a contact-based approach.
406:
566:
213:"Functional analysis of the pseudoknot structure in human telomerase RNA"
717:
138:
19:
41:
81:. These structures are categorized as cross (X) topology within the
420:
Kucharík, M; Hofacker, IL; Stadler, PF; Qin, J (15 January 2016).
561:
570:
141:
is one of the most conserved elements in all of evolution. The
747:
742:
373:"A new principle of RNA folding based on pseudoknotting"
272:"Pseudoknots: RNA structures with diverse functions"
710:
657:
604:
518:Theimer, CA; Blois, CA; Feigon, J (4 March 2005).
582:
8:
589:
575:
567:
535:
494:
445:
396:
297:
287:
246:
236:
137:. For example, the pseudoknot region of
30:
27:. Sequence from Chen and Greider (2005).
18:
422:"Pseudoknots in RNA folding landscapes"
200:
371:Pleij CW, Rietveld K, Bosch L (1985).
206:
204:
211:Chen, JL; Greider, CW (7 June 2005).
7:
270:Staple DW, Butcher SE (June 2005).
14:
562:Rfam entry for PK-HAV pseudoknot
107:stochastic context-free grammars
67:nucleic acid secondary structure
16:Nucleic acid secondary structure
1:
487:10.1093/bioinformatics/btx783
438:10.1093/bioinformatics/btv572
89:Prediction and identification
537:10.1016/j.molcel.2005.01.017
289:10.1371/journal.pbio.0030213
797:
101:by the standard method of
75:turnip yellow mosaic virus
758:Nucleic acid double helix
154:is an H-type pseudoknot.
149:Representing pseudoknots
143:telomerase RNA component
69:containing at least two
238:10.1073/pnas.0502259102
129:Biological significance
659:Nucleic acid structure
598:Biomolecular structure
190:Long range pseudoknots
58:
28:
389:10.1093/nar/13.5.1717
34:
22:
728:Protein engineering
229:2005PNAS..102.8080C
103:dynamic programming
135:tertiary structure
59:
29:
766:
765:
606:Protein structure
377:Nucleic Acids Res
169:(((.(((((........
79:topological knots
45:. Colors:
788:
753:Structural motif
591:
584:
577:
568:
550:
549:
539:
515:
509:
508:
498:
481:(8): 1304–1312.
466:
460:
459:
449:
417:
411:
410:
400:
368:
362:
359:
353:
343:
337:
334:
328:
318:
312:
311:
301:
291:
267:
261:
260:
250:
240:
208:
175:)))) ))........
171:))))).))). ...
165:
160:
83:circuit topology
57:
54:
51:
48:
44:
796:
795:
791:
790:
789:
787:
786:
785:
771:
770:
767:
762:
706:
653:
600:
595:
558:
553:
517:
516:
512:
468:
467:
463:
419:
418:
414:
370:
369:
365:
360:
356:
352:(3–4): 409–427.
344:
340:
335:
331:
327:(5): 2053–2068.
319:
315:
269:
268:
264:
210:
209:
202:
198:
186:
178:
163:
158:
151:
131:
91:
55:
52:
49:
46:
36:
17:
12:
11:
5:
794:
792:
784:
783:
773:
772:
764:
763:
761:
760:
755:
750:
745:
740:
735:
730:
725:
723:Protein domain
720:
714:
712:
708:
707:
705:
704:
702:Thermodynamics
699:
694:
689:
684:
679:
674:
669:
663:
661:
655:
654:
652:
651:
649:Thermodynamics
646:
641:
636:
631:
626:
621:
616:
610:
608:
602:
601:
596:
594:
593:
586:
579:
571:
565:
564:
557:
556:External links
554:
552:
551:
524:Molecular Cell
510:
475:Bioinformatics
461:
426:Bioinformatics
412:
383:(5): 1717–31.
363:
354:
338:
329:
313:
262:
223:(23): 8077–9.
199:
197:
194:
193:
192:
185:
182:
168:
150:
147:
130:
127:
90:
87:
15:
13:
10:
9:
6:
4:
3:
2:
793:
782:
779:
778:
776:
769:
759:
756:
754:
751:
749:
746:
744:
741:
739:
736:
734:
731:
729:
726:
724:
721:
719:
716:
715:
713:
709:
703:
700:
698:
695:
693:
690:
688:
687:Determination
685:
683:
680:
678:
675:
673:
670:
668:
665:
664:
662:
660:
656:
650:
647:
645:
642:
640:
637:
635:
634:Determination
632:
630:
627:
625:
622:
620:
617:
615:
612:
611:
609:
607:
603:
599:
592:
587:
585:
580:
578:
573:
572:
569:
563:
560:
559:
555:
547:
543:
538:
533:
530:(5): 671–82.
529:
525:
521:
514:
511:
506:
502:
497:
492:
488:
484:
480:
476:
472:
465:
462:
457:
453:
448:
443:
439:
435:
432:(2): 187–94.
431:
427:
423:
416:
413:
408:
404:
399:
394:
390:
386:
382:
378:
374:
367:
364:
358:
355:
351:
348:
347:J Comput Biol
342:
339:
333:
330:
326:
323:
317:
314:
309:
305:
300:
295:
290:
285:
281:
277:
273:
266:
263:
258:
254:
249:
244:
239:
234:
230:
226:
222:
218:
214:
207:
205:
201:
195:
191:
188:
187:
183:
181:
176:
172:
167:
155:
148:
146:
144:
140:
136:
128:
126:
124:
118:
116:
112:
108:
104:
100:
96:
88:
86:
84:
80:
76:
72:
68:
64:
43:
39:
33:
26:
21:
768:
738:Nucleic acid
527:
523:
513:
478:
474:
464:
429:
425:
415:
380:
376:
366:
357:
349:
346:
341:
332:
324:
321:
316:
279:
275:
265:
220:
216:
179:
156:
152:
132:
119:
95:base pairing
92:
62:
60:
282:(6): e213.
123:NP-complete
733:Proteasome
692:Prediction
682:Quaternary
639:Prediction
629:Quaternary
322:J Mol Biol
196:References
63:pseudoknot
25:telomerase
672:Secondary
619:Secondary
276:PLOS Biol
177:.]]]]]].
71:stem-loop
775:Category
711:See also
677:Tertiary
624:Tertiary
546:15749017
505:29236971
456:26428288
308:15941360
257:15849264
184:See also
718:Protein
667:Primary
614:Primary
496:5905660
447:4708108
407:4000943
299:1149493
248:1149427
225:Bibcode
139:RNase P
99:predict
697:Design
644:Design
544:
503:
493:
454:
444:
405:
398:341107
395:
306:
296:
255:
245:
121:to be
115:Pfold
111:Mfold
65:is a
542:PMID
501:PMID
452:PMID
403:PMID
304:PMID
253:PMID
113:and
42:1YMO
781:RNA
748:RNA
743:DNA
532:doi
491:PMC
483:doi
442:PMC
434:doi
393:PMC
385:doi
325:285
294:PMC
284:doi
243:PMC
233:doi
221:102
38:PDB
777::
540:.
528:17
526:.
522:.
499:.
489:.
479:34
477:.
473:.
450:.
440:.
430:32
428:.
424:.
401:.
391:.
381:13
379:.
375:.
302:.
292:.
278:.
274:.
251:.
241:.
231:.
219:.
215:.
203:^
164:()
159:()
125:.
61:A
40::
590:e
583:t
576:v
548:.
534::
507:.
485::
458:.
436::
409:.
387::
350:7
310:.
286::
280:3
259:.
235::
227::
56:G
53:C
50:U
47:A
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.