199:
used to calculate the genetic distance between the markers if they are close on the same chromosome. Tetrad analyses have also contributed to detection and study of the phenomena of gene conversion and post-meiotic segregation. These studies have proven central to understanding the mechanism of meiotic recombination, which in turn is a key to understanding the adaptive function of sexual reproduction. The use of tetrads in fine-structure genetic analysis is described in the articles
27:
61:
221:, then the resulting diploids are transferred to sporulation media to form a tetrad containing four haploid spores. Tetrads can then be prepared with Zymolyase, or another enzyme, to digest the wall of the ascus. The spores are then separated with a micromanipulator needle and deposited in separate positions on a
233:
Traditionally, tetrad dissection has a reputation as "black art". However, instruments have since been developed specifically for tetrad dissection; the most advanced allow easy and semi-automated separation of tetrads. Most micromanipulators use a glass fiber needle to which the spores adhere due
198:
Tetrad analysis can be used to confirm whether a phenotype is caused by a specific mutation, construction of strains, and for investigating gene interaction. Since the frequency of tetrad segregation types is influenced by the recombination frequency for the two markers, the segregation data can be
46:
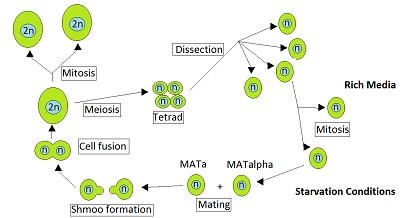
or other alga, or a plant. After parent haploids mate, they produce diploids. Under appropriate environmental conditions, diploids sporulate and undergo meiosis. The meiotic products, spores, remain packaged in the parental cell body to produce the tetrad.
154:
Tetratype is a tetrad containing four different genotypes, two parental and two recombinant. A spore arrangement in ascomycetes that consists of two parental and two recombinant spores indicates a single crossover between two linked loci.
178:
has become a powerful tool of yeast geneticists, and is used in conjunction with the many established procedures utilizing the versatility of yeasts as
344:
190:
techniques allows the four haploid spores of a yeast tetrad to be separated and germinated individually to form isolated spore colonies.
320:
115:
336:
101:
297:
Perkins, D.D. (1962) Crossing-over and interference in a multiply marked chromosome arm of
Neurosopora. Genetics 47, 1253-1274.
20:
125:
If the two parents have a mutation in two different genes, the tetrad can segregate these genes as the parental ditype (
79:
151:(assuming two segregating loci). A tetrad type containing two different genotypes, both of which are recombinant.
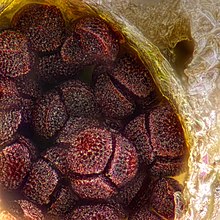
374:
268:
257:
379:
251:
331:
Yeast
Protocols: Methods in Cell and Molecular Biology, Ivor Howell Evan, Published by Humana Press, 1996,
71:
273:
83:
163:
The ratio between the different segregation types arising after the sporulation is a measure of the
340:
332:
316:
300:
235:
201:
187:
19:
For the microscopic chromosomal structures formed during the initial phases of meiosis, see
218:
206:
164:
26:
179:
368:
359:
298:
147:
Non-parental ditype (NPD) is a spore that contains only the two recombinant-type
137:
222:
183:
148:
141:
278:
133:
304:
114:
42:
is the four spores produced after meiosis of a yeast or other
Ascomycota,
263:
25:
239:
315:
54:
129:), the non-parental ditype (NPD) or as the tetratype (TT).
132:
Parental ditype is a tetrad type containing two different
136:, both of which are parental. A spore arrangement in
217:
Crosses are performed between haploid MATa and MATα
140:that contains only the two non-recombinant-type
16:Product of meiosis in spore-producing organisms
34:still associated in tetrads following meiosis.
8:
82:. There might be a discussion about this on
102:Learn how and when to remove this message
113:
290:
7:
126:
14:
360:"Crosses, diploids, and tetrads"
59:
21:tetrad (chromosomal formation)
1:
234:to the formation of a water
396:
18:
269:Homologous recombination
258:Saccharomyces cerevisiae
119:Saccharomyces cerevisiae
252:Synthetic genetic array
167:between the two genes.
122:
35:
274:Genetic recombination
117:
29:
72:confusing or unclear
51:Genetic typification
80:clarify the section
123:
36:
345:978-0-89603-319-1
213:General procedure
202:Neurospora crassa
188:micromanipulation
176:Tetrad dissection
171:Tetrad dissection
112:
111:
104:
387:
347:
329:
323:
313:
307:
295:
242:and the needle.
182:. Use of modern
159:Linkage analysis
107:
100:
96:
93:
87:
63:
62:
55:
32:Riccia sorocarpa
395:
394:
390:
389:
388:
386:
385:
384:
375:Fungus genetics
365:
364:
356:
351:
350:
330:
326:
314:
310:
296:
292:
287:
248:
231:
215:
207:Gene conversion
196:
180:model organisms
173:
161:
108:
97:
91:
88:
77:
64:
60:
53:
24:
17:
12:
11:
5:
393:
391:
383:
382:
380:Plant genetics
377:
367:
366:
363:
362:
355:
354:External links
352:
349:
348:
324:
321:978-0471102052
308:
289:
288:
286:
283:
282:
281:
276:
271:
266:
261:
254:
247:
244:
230:
227:
219:mating strains
214:
211:
195:
192:
172:
169:
160:
157:
110:
109:
67:
65:
58:
52:
49:
15:
13:
10:
9:
6:
4:
3:
2:
392:
381:
378:
376:
373:
372:
370:
361:
358:
357:
353:
346:
342:
338:
337:0-89603-319-8
334:
328:
325:
322:
318:
312:
309:
306:
302:
299:
294:
291:
284:
280:
277:
275:
272:
270:
267:
265:
262:
260:
259:
255:
253:
250:
249:
245:
243:
241:
237:
228:
226:
224:
220:
212:
210:
208:
204:
203:
193:
191:
189:
185:
181:
177:
170:
168:
166:
158:
156:
152:
150:
145:
143:
139:
135:
130:
128:
120:
116:
106:
103:
95:
92:November 2017
85:
84:the talk page
81:
75:
73:
68:This section
66:
57:
56:
50:
48:
45:
44:Chlamydomonas
41:
33:
28:
22:
327:
311:
293:
256:
238:between the
232:
216:
200:
197:
175:
174:
162:
153:
146:
131:
124:
118:
98:
89:
78:Please help
69:
43:
39:
37:
31:
138:ascomycetes
369:Categories
285:References
223:petri dish
184:microscopy
149:ascospores
142:ascospores
74:to readers
30:Spores of
279:Ascospore
134:genotypes
305:13942437
246:See also
236:meniscus
165:linkage
70:may be
343:
335:
319:
303:
121:tetrad
40:tetrad
264:Ascus
229:Tools
341:ISBN
333:ISBN
317:ISBN
301:PMID
240:agar
205:and
194:Uses
186:and
38:The
371::
339:,
225:.
209:.
144:.
127:PD
105:)
99:(
94:)
90:(
86:.
76:.
23:.
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.