555:(insertion or deletion), or chromosomal rearrangement; any such errors may render the gene products coded at that location non-functional. Because this activity can vary depending on the species, cell type, target gene, and nuclease used, it should be monitored when designing new systems. A simple heteroduplex cleavage assay can be run which detects any difference between two alleles amplified by PCR. Cleavage products can be visualized on simple agarose gels or slab gel systems.
75:
473:
cleavage activity. The FokI domain functions as a dimer, requiring two constructs with unique DNA binding domains for sites in the target genome with proper orientation and spacing. Both the number of amino acid residues between the TALE DNA binding domain and the FokI cleavage domain and the number of bases between the two individual TALEN binding sites appear to be important parameters for achieving high levels of activity.
31:
506:
531:
and enter the nucleus to access the genome. Alternatively, TALEN constructs can be delivered to the cells as mRNAs, which removes the possibility of genomic integration of the TALEN-expressing protein. Using an mRNA vector can also dramatically increase the level of homology directed repair (HDR) and
653:
The off-target activity of an active nuclease may lead to unwanted double-strand breaks and may consequently yield chromosomal rearrangements and/or cell death. Studies have been carried out to compare the relative nuclease-associated toxicity of available technologies. Based on these studies and
472:
that are active in a yeast assay. These reagents are also active in plant cells and in animal cells. Initial TALEN studies used the wild-type FokI cleavage domain, but some subsequent TALEN studies also used FokI cleavage domain variants with mutations designed to improve cleavage specificity and
455:
recognition. This straightforward relationship between amino acid sequence and DNA recognition has allowed for the engineering of specific DNA-binding domains by selecting a combination of repeat segments containing the appropriate RVDs. Notably, slight changes in the RVD and the incorporation of
644:
relies on ribonucleotide complex formation instead of protein/DNA recognition. gRNAs have occasionally limitations regarding feasibility due to lack of PAM sites in the target sequence and even though they can be cheaply produced, the current development lead to a remarkable decrease of cost for
497:
oligonucleotide assembly followed by whole gene amplification. A number of modular assembly schemes for generating engineered TALE constructs have also been reported. Both methods offer a systematic approach to engineering DNA binding domains that is conceptually similar to the modular assembly
509:
Workflow of genome editing of Your
Favorite Gene (YFG) using TALEN. The target sequence is identified, a corresponding TALEN sequence is engineered and inserted into a plasmid. The plasmid is inserted into the target cell where it is translated to produce the functional TALEN, which enters the
633:. The DNA binding region of a TAL effector can be combined with the cleavage domain of a meganuclease to create a hybrid architecture combining the ease of engineering and highly specific DNA binding activity of a TAL effector with the low site frequency and specificity of a meganuclease.
599:. Moreover, the method can be used to generate knockin organisms. Wu et al.obtained a Sp110 knockin cattle using Talen nickases to induce increased resistance of tuberculosis. This approach has also been used to generate knockin rats by TALEN mRNA microinjection in one-cell embryos.
391:
which cuts DNA strands). Transcription activator-like effectors (TALEs) can be engineered to bind to practically any desired DNA sequence, so when combined with a nuclease, DNA can be cut at specific locations. The restriction enzymes can be introduced into cells, for use in
1157:
Miller JC, Tan S, Qiao G, Barlow KA, Wang J, Xia DF, Meng X, Paschon DE, Leung E, Hinkley SJ, Dulay GP, Hua KL, Ankoudinova I, Cost GJ, Urnov FD, Zhang HS, Holmes MC, Zhang L, Gregory PD, Rebar EJ (February 2011). "A TALE nuclease architecture for efficient genome editing".
640:, TALEN recognizes single nucleotides. It's far more straightforward to engineer interactions between TALEN DNA binding domains and their target nucleotides than it is to create interactions with ZFNs and their target nucleotide triplets. On the other hand,
2572:
Poirot L, Philip B, Schiffer-Mannioui C, Le Clerre D, Chion-Sotinel I, Derniame S, Potrel P, Bas C, Lemaire L, Galetto R, Lebuhotel C, Eyquem J, Cheung GW, Duclert A, Gouble A, Arnould S, Peggs K, Pule M, Scharenberg AM, Smith J (September 2015).
618:. Recently, it was shown that TALEN can be used as tools to harness the immune system to fight cancers; TALEN-mediated targeting can generate T cells that are resistant to chemotherapeutic drugs and show anti-tumor activity.
621:
In theory, the genome-wide specificity of engineered TALEN fusions allows for correction of errors at individual genetic loci via homology-directed repair from a correct exogenous template. In reality, however, the
1405:
Doyon Y, Vo TD, Mendel MC, Greenberg SG, Wang J, Xia DF, Miller JC, Urnov FD, Gregory PD, Holmes MC (January 2011). "Enhancing zinc-finger-nuclease activity with improved obligate heterodimeric architectures".
572:
TALEN has been used to efficiently modify plant genomes, creating economically important food crops with favorable nutritional qualities. They have also been harnessed to develop tools for the production of
510:
nucleus and binds and cleaves the target sequence. Depending on the application, this can be used to introduce an error (to knock out a target gene) or to introduce a new DNA sequence into the target gene.
1206:
Hockemeyer D, Wang H, Kiani S, Lai CS, Gao Q, Cassady JP, Cost GJ, Zhang L, Santiago Y, Miller JC, Zeitler B, Cherone JM, Meng X, Hinkley SJ, Rebar EJ, Gregory PD, Urnov FD, Jaenisch R (July 2011).
2474:
Osborn MJ, Starker CG, McElroy AN, Webber BR, Riddle MJ, Xia L, DeFeo AP, Gabriel R, Schmidt M, von Kalle C, Carlson DF, Maeder ML, Joung JK, Wagner JE, Voytas DF, Blazar BR, Tolar J (June 2013).
551:(NHEJ) directly ligates DNA from either side of a double-strand break where there is very little or no sequence overlap for annealing. This repair mechanism induces errors in the genome via
451:
sequence with divergent 12th and 13th amino acids. These two positions, referred to as the Repeat
Variable Diresidue (RVD), are highly variable and show a strong correlation with specific
778:
Boch J, Scholze H, Schornack S, Landgraf A, Hahn S, Kay S, Lahaye T, Nickstadt A, Bonas U (December 2009). "Breaking the code of DNA binding specificity of TAL-type III effectors".
654:
the maximal theoretical distance between DNA binding and nuclease activity, TALEN constructs are believed to have the greatest precision of the currently available technologies.
356:
2380:
Ramalingam S, Annaluru N, Kandavelou K, Chandrasegaran S (2014). "TALEN-mediated generation and genetic correction of disease-specific human induced pluripotent stem cells".
626:
application of TALEN is currently limited by the lack of an efficient delivery mechanism, unknown immunogenic factors, and uncertainty in the specificity of TALEN binding.
1999:
Daboussi F, Leduc S, Maréchal A, Dubois G, Guyot V, Perez-Michaut C, Amato A, Falciatore A, Juillerat A, Beurdeley M, Voytas DF, Cavarec L, Duchateau P (May 2014).
481:
The simple relationship between amino acid sequence and DNA recognition of the TALE binding domain allows for the efficient engineering of proteins. In this case,
1258:
Wood AJ, Lo TW, Zeitler B, Pickle CS, Ralston EJ, Lee AH, Amora R, Miller JC, Leung E, Meng X, Zhang L, Rebar EJ, Gregory PD, Urnov FD, Meyer BJ (July 2011).
1958:
Haun W, Coffman A, Clasen BM, Demorest ZL, Lowy A, Ray E, Retterath A, Stoddard T, Juillerat A, Cedrone F, Mathis L, Voytas DF, Zhang F (September 2014).
489:
of the repetitive sequence found in the TALE binding domain. One solution to this is to use a publicly available software program (DNAWorks) to calculate
349:
663:
407:
305:
1037:"De novo-engineered transcription activator-like effector (TALE) hybrid nuclease with novel DNA binding specificity creates double-strand breaks"
2051:
Wienert B, Funnell AP, Norton LJ, Pearson RC, Wilkinson-White LE, Lester K, Vadolas J, Porteus MH, Matthews JM, Quinlan KG, Crossley M (2015).
2614:
Boissel S, Jarjour J, Astrakhan A, Adey A, Gouble A, Duchateau P, Shendure J, Stoddard BL, Certo MT, Baker D, Scharenberg AM (February 2014).
1318:
Tesson L, Usal C, Ménoret S, Leung E, Niles BJ, Remy S, Santiago Y, Vincent AI, Meng X, Zhang L, Gregory PD, Anegon I, Cost GJ (August 2011).
1669:
342:
2663:
2415:
Dupuy A, Valton J, Leduc S, Armier J, Galetto R, Gouble A, Lebuhotel C, Stary A, Pâques F, Duchateau P, Sarasin A, Daboussi F (2013).
564:
can also introduce foreign DNA at the DSB as the transfected double-stranded sequences are used as templates for the repair enzymes.
2758:
2743:
177:
880:
Juillerat A, Pessereau C, Dubois G, Guyot V, Maréchal A, Valton J, Daboussi F, Poirot L, Duclert A, Duchateau P (January 2015).
2723:
1362:
Huang P, Xiao A, Zhou M, Zhu Z, Lin S, Zhang B (August 2011). "Heritable gene targeting in zebrafish using customized TALENs".
602:
TALEN has also been utilized experimentally to correct the genetic errors that underlie disease. For example, it has been used
187:
205:
170:
558:
Alternatively, DNA can be introduced into a genome through NHEJ in the presence of exogenous double-stranded DNA fragments.
582:
96:
87:
545:
TALEN can be used to edit genomes by inducing double-strand breaks (DSB), which cells respond to with repair mechanisms.
2733:
147:
56:
1099:
Cermak T, Doyle EL, Christian M, Wang L, Zhang Y, Schmidt C, Baller JA, Somia NV, Bogdanove AJ, Voytas DF (July 2011).
486:
440:
152:
123:
113:
108:
2682:
636:
In comparison to other genome editing techniques, TALEN falls in the middle in terms of difficulty and cost. Unlike
2748:
1744:"Modularly assembled designer TAL effector nucleases for targeted gene knockout and gene replacement in eukaryotes"
548:
272:
137:
132:
101:
2053:"Editing the genome to introduce a beneficial naturally occurring mutation associated with increased fetal globin"
2102:"Site specific mutation of the Zic2 locus by microinjection of TALEN mRNA in mouse CD1, C3H and C57BL/6J oocytes"
494:
482:
277:
227:
142:
118:
561:
17:
629:
Another emerging application of TALEN is its ability to combine with other genome engineering tools, such as
2753:
2738:
310:
237:
182:
2575:"Multiplex Genome-Edited T-cell Manufacturing Platform for "Off-the-Shelf" Adoptive T-cell Immunotherapies"
2523:
Valton J, Guyot V, Marechal A, Filhol JM, Juillerat A, Duclert A, Duchateau P, Poirot L (September 2015).
743:
Boch J, Bonas U (September 2010). "Xanthomonas AvrBs3 family-type III effectors: discovery and function".
587:
1101:"Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting"
615:
611:
411:
1501:"Directed evolution of an enhanced and highly efficient FokI cleavage domain for zinc finger nucleases"
937:
Christian M, Cermak T, Doyle EL, Schmidt C, Zhang F, Hummel A, Bogdanove AJ, Voytas DF (October 2010).
1553:"A novel TALE nuclease scaffold enables high genome editing activity in combination with low toxicity"
1451:"Structure-based redesign of the dimerization interface reduces the toxicity of zinc-finger nucleases"
1450:
2428:
2280:
2221:
2113:
2064:
2012:
1863:
1804:
1271:
1048:
893:
838:
829:
Moscou MJ, Bogdanove AJ (December 2009). "A simple cipher governs DNA recognition by TAL effectors".
787:
668:
637:
325:
607:
578:
65:
1605:"Efficient construction of sequence-specific TAL effectors for modulating mammalian transcription"
1960:"Improved soybean oil quality by targeted mutagenesis of the fatty acid desaturase 2 gene family"
1481:
1431:
1387:
1183:
862:
811:
725:
384:
377:
320:
988:"TAL nucleases (TALNs): hybrid proteins composed of TAL effectors and FokI DNA-cleavage domain"
2645:
2596:
2554:
2505:
2456:
2397:
2362:
2308:
2249:
2190:
2141:
2082:
2030:
1981:
1940:
1891:
1832:
1791:
Geissler R, Scholze H, Hahn S, Streubel J, Bonas U, Behrens SE, Boch J (2011). Shiu SH (ed.).
1773:
1724:
1675:
1665:
1652:
Hoover D (2012). "Using DNAWorks in
Designing Oligonucleotides for PCR-Based Gene Synthesis".
1634:
1582:
1530:
1473:
1423:
1379:
1341:
1297:
1237:
1175:
1130:
1076:
1017:
968:
919:
854:
803:
760:
717:
330:
210:
36:
2210:"TALE nickase-mediated SP110 knockin endows cattle with increased resistance to tuberculosis"
1909:
Zhang Y, Zhang F, Li X, Baller JA, Qi Y, Starker CG, Bogdanove AJ, Voytas DF (January 2013).
645:
TALENs, so that they are in a similar price and time range like CRISPR based genome editing.
2635:
2627:
2586:
2544:
2536:
2495:
2487:
2446:
2436:
2389:
2352:
2344:
2298:
2288:
2239:
2229:
2180:
2172:
2131:
2121:
2072:
2020:
1971:
1930:
1922:
1881:
1871:
1822:
1812:
1763:
1755:
1742:
Li T, Huang S, Zhao X, Wright DA, Carpenter S, Spalding MH, Weeks DP, Yang B (August 2011).
1714:
1706:
1657:
1624:
1616:
1572:
1564:
1520:
1512:
1465:
1415:
1371:
1331:
1287:
1279:
1227:
1219:
1167:
1120:
1112:
1066:
1056:
1007:
999:
958:
950:
909:
901:
846:
795:
752:
709:
490:
1911:"Transcription activator-like effector nucleases enable efficient plant genome engineering"
528:
444:
215:
74:
2432:
2284:
2225:
2117:
2068:
2016:
1867:
1808:
1275:
1052:
897:
842:
791:
756:
2640:
2615:
2549:
2524:
2500:
2475:
2451:
2416:
2357:
2332:
2303:
2268:
2244:
2209:
2185:
2160:
2136:
2101:
1935:
1910:
1886:
1851:
1827:
1792:
1768:
1743:
1719:
1694:
1629:
1604:
1577:
1552:
1525:
1500:
1292:
1259:
1232:
1207:
1125:
1100:
1071:
1036:
1012:
987:
963:
938:
914:
881:
419:
393:
315:
244:
30:
2525:"A Multidrug-resistant Engineered CAR T Cell for Allogeneic Combination Immunotherapy"
2393:
1551:
Mussolino C, Morbitzer R, Lütge F, Dannemann N, Lahaye T, Cathomen T (November 2011).
34:
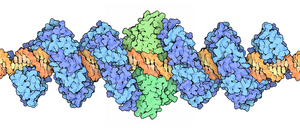
Spacefill drawing of dimeric TALE-FokI fusion (blue: TALE; green: FokI) bound to DNA (
2717:
815:
289:
222:
2616:"megaTALs: a rare-cleaving nuclease architecture for therapeutic genome engineering"
2417:"Targeted gene therapy of xeroderma pigmentosum cells using meganuclease and TALEN™"
2159:
Sander JD, Cade L, Khayter C, Reyon D, Peterson RT, Joung JK, Yeh JR (August 2011).
2001:"Genome engineering empowers the diatom Phaeodactylum tricornutum for biotechnology"
1850:
Weber E, Gruetzner R, Werner S, Engler C, Marillonnet S (2011). Bendahmane M (ed.).
1485:
1435:
1391:
1187:
2728:
866:
673:
630:
592:
524:
430:
381:
380:
that can be engineered to cut specific sequences of DNA. They are made by fusing a
284:
232:
2591:
2574:
729:
2441:
2293:
2126:
1876:
1817:
1661:
986:
Li T, Huang S, Jiang WZ, Wright D, Spalding MH, Weeks DP, Yang B (January 2011).
2707:
954:
499:
435:
2214:
Proceedings of the
National Academy of Sciences of the United States of America
1449:
Szczepek M, Brondani V, Büchel J, Serrano L, Segal DJ, Cathomen T (July 2007).
1041:
Proceedings of the
National Academy of Sciences of the United States of America
2269:"Generation of TALEN-mediated GRdim knock-in rats by homologous recombination"
1516:
452:
448:
2161:"Targeted gene disruption in somatic zebrafish cells using engineered TALENs"
1793:"Transcriptional activators of human genes with programmable DNA-specificity"
1035:
Mahfouz MM, Li L, Shamimuzzaman M, Wibowo A, Fang X, Zhu JK (February 2011).
2234:
2100:
Davies B, Davies G, Preece C, Puliyadi R, Szumska D, Bhattacharya S (2013).
1283:
1061:
850:
799:
596:
2649:
2600:
2558:
2509:
2460:
2401:
2366:
2312:
2253:
2194:
2145:
2086:
2034:
1985:
1944:
1895:
1836:
1777:
1728:
1679:
1638:
1603:
Zhang F, Cong L, Lodato S, Kosuri S, Church GM, Arlotta P (February 2011).
1586:
1534:
1477:
1427:
1383:
1345:
1301:
1241:
1179:
1134:
1080:
1021:
972:
923:
858:
807:
764:
721:
2631:
1926:
1759:
1710:
1568:
1116:
1003:
469:
388:
2540:
2348:
2491:
2077:
2052:
2025:
2000:
1419:
574:
520:
505:
402:
1976:
1959:
905:
519:
Once the TALEN constructs have been assembled, they are inserted into
44:
40:
2176:
1695:"Assembly of custom TALE-type DNA binding domains by modular cloning"
1620:
1375:
1336:
1319:
1223:
1208:"Genetic engineering of human pluripotent cells using TALE nucleases"
1171:
713:
678:
641:
415:
397:
254:
2267:
Ponce de León V, Mérillat AM, Tesson L, Anegón I, Hummler E (2014).
1469:
585:(IPSCs) clones and human erythroid cell lines, to generate knockout
447:. The DNA binding domain contains a repeated highly conserved 33–34
882:"Optimized tuning of TALEN specificity using non-conventional RVDs"
577:. In addition, it has been used to engineer stably modified human
456:"nonconventional" RVD sequences can improve targeting specificity.
2208:
Wu H, Wang Y, Zhang Y, Yang M, Lv J, Liu J, Zhang Y (March 2015).
552:
249:
29:
1656:. Methods in Molecular Biology. Vol. 852. pp. 215–23.
939:"Targeting DNA double-strand breaks with TAL effector nucleases"
465:
2710:
An entry in the
Protein Database's monthly structural highlight
1260:"Targeted genome editing across species using ZFNs and TALENs"
2683:"Boston Consulting Group - Report on Gene Editing Precision"
1320:"Knockout rats generated by embryo microinjection of TALENs"
606:
to correct the genetic defects that cause disorders such as
1852:"Assembly of designer TAL effectors by Golden Gate cloning"
1693:
Morbitzer R, Elsaesser J, Hausner J, Lahaye T (July 2011).
2701:
464:
The non-specific DNA cleavage domain from the end of the
2476:"TALEN-based gene correction for epidermolysis bullosa"
2331:
Carlson DF, Fahrenkrug SC, Hackett PB (January 2012).
700:
Boch J (February 2011). "TALEs of genome targeting".
532:the success of introgression during gene editing.
1201:
1199:
1197:
1546:
1544:
2664:"Pros and Cons Of ZFNS, TALENS, AND CRISPR/CAS"
2046:
2044:
370:Transcription activator-like effector nucleases
18:Transcription Activator-Like Effector Nuclease
1313:
1311:
1152:
1150:
1148:
1146:
1144:
527:with the plasmids, and the gene products are
468:endonuclease can be used to construct hybrid
350:
8:
1598:
1596:
1357:
1355:
1094:
1092:
1090:
418:, TALEN is a prominent tool in the field of
1253:
1251:
2326:
2324:
2322:
695:
693:
357:
343:
51:
2639:
2590:
2548:
2499:
2450:
2440:
2356:
2302:
2292:
2243:
2233:
2184:
2135:
2125:
2076:
2024:
1975:
1934:
1885:
1875:
1826:
1816:
1767:
1718:
1628:
1576:
1524:
1335:
1291:
1231:
1124:
1070:
1060:
1011:
962:
913:
664:Genome editing with engineered nucleases
504:
408:genome editing with engineered nucleases
27:Enzymes that cleave DNA in specific ways
2333:"Targeting DNA With Fingers and TALENs"
689:
306:Genetically modified food controversies
297:
264:
197:
162:
86:
63:
2704:A comprehensive tool for TALEN design
1499:Guo J, Gaj T, Barbas CF (July 2010).
7:
493:suitable for assembly in a two step
757:10.1146/annurev-phyto-080508-081936
485:is problematic because of improper
433:are proteins that are secreted by
25:
2681:Boglioli, Elsy; Richard, Magali.
2394:10.2174/1566523214666140918101725
2337:Molecular Therapy: Nucleic Acids
73:
745:Annual Review of Phytopathology
649:TAL effector nuclease precision
188:Cartagena Protocol on Biosafety
595:, knockout mice, and knockout
88:Genetically modified organisms
1:
2592:10.1158/0008-5472.CAN-14-3321
583:induced pluripotent stem cell
2442:10.1371/journal.pone.0078678
2294:10.1371/journal.pone.0088146
2127:10.1371/journal.pone.0060216
1877:10.1371/journal.pone.0019722
1818:10.1371/journal.pone.0019509
1662:10.1007/978-1-61779-564-0_16
1505:Journal of Molecular Biology
523:; the target cells are then
477:Engineering TALEN constructs
387:to a DNA cleavage domain (a
1964:Plant Biotechnology Journal
955:10.1534/genetics.110.120717
48:), by David Goodsell
2775:
549:Non-homologous end joining
273:Genetically modified crops
2708:PDB Molecule of the Month
1517:10.1016/j.jmb.2010.04.060
502:DNA recognition domains.
483:artificial gene synthesis
441:type III secretion system
2759:Repetitive DNA sequences
2744:History of biotechnology
562:Homology directed repair
2235:10.1073/pnas.1421587112
1284:10.1126/science.1207773
1062:10.1073/pnas.1019533108
851:10.1126/science.1178817
800:10.1126/science.1178811
426:TALE DNA-binding domain
406:, a technique known as
311:GMO conspiracy theories
183:Substantial equivalence
2724:Biological engineering
2668:The Jackson Laboratory
2620:Nucleic Acids Research
1748:Nucleic Acids Research
1699:Nucleic Acids Research
1557:Nucleic Acids Research
1105:Nucleic Acids Research
992:Nucleic Acids Research
511:
498:method for generating
163:History and regulation
49:
2057:Nature Communications
2005:Nature Communications
1927:10.1104/pp.112.205179
616:epidermolysis bullosa
612:xeroderma pigmentosum
508:
412:zinc finger nucleases
33:
2382:Current Gene Therapy
2165:Nature Biotechnology
1609:Nature Biotechnology
1458:Nature Biotechnology
1364:Nature Biotechnology
1324:Nature Biotechnology
1212:Nature Biotechnology
1160:Nature Biotechnology
702:Nature Biotechnology
669:Zinc finger nuclease
326:StarLink corn recall
2734:Genetic engineering
2632:10.1093/nar/gkt1224
2541:10.1038/mt.2015.104
2433:2013PLoSO...878678D
2349:10.1038/mtna.2011.5
2285:2014PLoSO...988146P
2226:2015PNAS..112E1530W
2118:2013PLoSO...860216D
2069:2015NatCo...6.7085W
2017:2014NatCo...5.3831D
1868:2011PLoSO...619722W
1809:2011PLoSO...619509G
1276:2011Sci...333..307W
1053:2011PNAS..108.2623M
898:2015NatSR...5E8150J
843:2009Sci...326.1501M
792:2009Sci...326.1509B
608:sickle cell disease
579:embryonic stem cell
460:DNA cleavage domain
439:bacteria via their
378:restriction enzymes
66:Genetic engineering
2492:10.1038/mt.2013.56
2078:10.1038/ncomms8085
2026:10.1038/ncomms4831
1760:10.1093/nar/gkr188
1711:10.1093/nar/gkr151
1569:10.1093/nar/gkr597
1420:10.1038/nmeth.1539
1117:10.1093/nar/gkr218
1004:10.1093/nar/gkq704
886:Scientific Reports
512:
385:DNA-binding domain
50:
2749:Molecular biology
2529:Molecular Therapy
2480:Molecular Therapy
1977:10.1111/pbi.12201
1671:978-1-61779-563-3
906:10.1038/srep08150
786:(5959): 1509–12.
367:
366:
331:He Jiankui affair
211:Molecular cloning
16:(Redirected from
2766:
2690:
2689:
2687:
2678:
2672:
2671:
2660:
2654:
2653:
2643:
2611:
2605:
2604:
2594:
2569:
2563:
2562:
2552:
2520:
2514:
2513:
2503:
2471:
2465:
2464:
2454:
2444:
2412:
2406:
2405:
2377:
2371:
2370:
2360:
2328:
2317:
2316:
2306:
2296:
2264:
2258:
2257:
2247:
2237:
2205:
2199:
2198:
2188:
2177:10.1038/nbt.1934
2156:
2150:
2149:
2139:
2129:
2097:
2091:
2090:
2080:
2048:
2039:
2038:
2028:
1996:
1990:
1989:
1979:
1955:
1949:
1948:
1938:
1915:Plant Physiology
1906:
1900:
1899:
1889:
1879:
1847:
1841:
1840:
1830:
1820:
1788:
1782:
1781:
1771:
1739:
1733:
1732:
1722:
1690:
1684:
1683:
1649:
1643:
1642:
1632:
1621:10.1038/nbt.1775
1600:
1591:
1590:
1580:
1548:
1539:
1538:
1528:
1496:
1490:
1489:
1455:
1446:
1440:
1439:
1402:
1396:
1395:
1376:10.1038/nbt.1939
1359:
1350:
1349:
1339:
1337:10.1038/nbt.1940
1315:
1306:
1305:
1295:
1255:
1246:
1245:
1235:
1224:10.1038/nbt.1927
1203:
1192:
1191:
1172:10.1038/nbt.1755
1154:
1139:
1138:
1128:
1096:
1085:
1084:
1074:
1064:
1032:
1026:
1025:
1015:
983:
977:
976:
966:
934:
928:
927:
917:
877:
871:
870:
826:
820:
819:
775:
769:
768:
740:
734:
733:
714:10.1038/nbt.1767
697:
491:oligonucleotides
359:
352:
345:
77:
52:
47:
21:
2774:
2773:
2769:
2768:
2767:
2765:
2764:
2763:
2714:
2713:
2698:
2693:
2685:
2680:
2679:
2675:
2662:
2661:
2657:
2626:(4): 2591–601.
2613:
2612:
2608:
2585:(18): 3853–64.
2579:Cancer Research
2571:
2570:
2566:
2522:
2521:
2517:
2473:
2472:
2468:
2414:
2413:
2409:
2379:
2378:
2374:
2330:
2329:
2320:
2266:
2265:
2261:
2220:(13): E1530-9.
2207:
2206:
2202:
2158:
2157:
2153:
2099:
2098:
2094:
2050:
2049:
2042:
1998:
1997:
1993:
1957:
1956:
1952:
1908:
1907:
1903:
1849:
1848:
1844:
1790:
1789:
1785:
1754:(14): 6315–25.
1741:
1740:
1736:
1692:
1691:
1687:
1672:
1651:
1650:
1646:
1602:
1601:
1594:
1563:(21): 9283–93.
1550:
1549:
1542:
1498:
1497:
1493:
1470:10.1038/nbt1317
1453:
1448:
1447:
1443:
1404:
1403:
1399:
1361:
1360:
1353:
1317:
1316:
1309:
1257:
1256:
1249:
1205:
1204:
1195:
1156:
1155:
1142:
1098:
1097:
1088:
1034:
1033:
1029:
985:
984:
980:
936:
935:
931:
879:
878:
874:
828:
827:
823:
777:
776:
772:
742:
741:
737:
699:
698:
691:
687:
660:
651:
570:
543:
538:
517:
479:
462:
428:
363:
321:Séralini affair
216:Recombinant DNA
35:
28:
23:
22:
15:
12:
11:
5:
2772:
2770:
2762:
2761:
2756:
2754:Non-coding RNA
2751:
2746:
2741:
2739:Genome editing
2736:
2731:
2726:
2716:
2715:
2712:
2711:
2705:
2697:
2696:External links
2694:
2692:
2691:
2673:
2655:
2606:
2564:
2535:(9): 1507–18.
2515:
2466:
2427:(11): e78678.
2407:
2372:
2318:
2259:
2200:
2151:
2092:
2040:
1991:
1950:
1901:
1842:
1783:
1734:
1705:(13): 5790–9.
1685:
1670:
1654:Gene Synthesis
1644:
1592:
1540:
1491:
1441:
1408:Nature Methods
1397:
1370:(8): 699–700.
1351:
1307:
1247:
1193:
1140:
1086:
1027:
978:
929:
872:
837:(5959): 1501.
821:
770:
735:
688:
686:
683:
682:
681:
676:
671:
666:
659:
656:
650:
647:
569:
566:
542:
539:
537:
536:Genome editing
534:
516:
513:
478:
475:
461:
458:
427:
424:
420:genome editing
365:
364:
362:
361:
354:
347:
339:
336:
335:
334:
333:
328:
323:
318:
316:Pusztai affair
313:
308:
300:
299:
295:
294:
293:
292:
287:
282:
281:
280:
267:
266:
262:
261:
260:
259:
258:
257:
252:
245:Genome editing
242:
241:
240:
235:
230:
228:Transformation
220:
219:
218:
208:
200:
199:
195:
194:
193:
192:
191:
190:
185:
174:
173:
165:
164:
160:
159:
158:
157:
156:
155:
150:
145:
140:
129:
128:
127:
126:
121:
116:
105:
104:
99:
91:
90:
84:
83:
79:
78:
70:
69:
61:
60:
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
2771:
2760:
2757:
2755:
2752:
2750:
2747:
2745:
2742:
2740:
2737:
2735:
2732:
2730:
2727:
2725:
2722:
2721:
2719:
2709:
2706:
2703:
2700:
2699:
2695:
2684:
2677:
2674:
2670:. March 2014.
2669:
2665:
2659:
2656:
2651:
2647:
2642:
2637:
2633:
2629:
2625:
2621:
2617:
2610:
2607:
2602:
2598:
2593:
2588:
2584:
2580:
2576:
2568:
2565:
2560:
2556:
2551:
2546:
2542:
2538:
2534:
2530:
2526:
2519:
2516:
2511:
2507:
2502:
2497:
2493:
2489:
2486:(6): 1151–9.
2485:
2481:
2477:
2470:
2467:
2462:
2458:
2453:
2448:
2443:
2438:
2434:
2430:
2426:
2422:
2418:
2411:
2408:
2403:
2399:
2395:
2391:
2388:(6): 461–72.
2387:
2383:
2376:
2373:
2368:
2364:
2359:
2354:
2350:
2346:
2342:
2338:
2334:
2327:
2325:
2323:
2319:
2314:
2310:
2305:
2300:
2295:
2290:
2286:
2282:
2279:(2): e88146.
2278:
2274:
2270:
2263:
2260:
2255:
2251:
2246:
2241:
2236:
2231:
2227:
2223:
2219:
2215:
2211:
2204:
2201:
2196:
2192:
2187:
2182:
2178:
2174:
2170:
2166:
2162:
2155:
2152:
2147:
2143:
2138:
2133:
2128:
2123:
2119:
2115:
2112:(3): e60216.
2111:
2107:
2103:
2096:
2093:
2088:
2084:
2079:
2074:
2070:
2066:
2062:
2058:
2054:
2047:
2045:
2041:
2036:
2032:
2027:
2022:
2018:
2014:
2010:
2006:
2002:
1995:
1992:
1987:
1983:
1978:
1973:
1970:(7): 934–40.
1969:
1965:
1961:
1954:
1951:
1946:
1942:
1937:
1932:
1928:
1924:
1920:
1916:
1912:
1905:
1902:
1897:
1893:
1888:
1883:
1878:
1873:
1869:
1865:
1862:(5): e19722.
1861:
1857:
1853:
1846:
1843:
1838:
1834:
1829:
1824:
1819:
1814:
1810:
1806:
1803:(5): e19509.
1802:
1798:
1794:
1787:
1784:
1779:
1775:
1770:
1765:
1761:
1757:
1753:
1749:
1745:
1738:
1735:
1730:
1726:
1721:
1716:
1712:
1708:
1704:
1700:
1696:
1689:
1686:
1681:
1677:
1673:
1667:
1663:
1659:
1655:
1648:
1645:
1640:
1636:
1631:
1626:
1622:
1618:
1615:(2): 149–53.
1614:
1610:
1606:
1599:
1597:
1593:
1588:
1584:
1579:
1574:
1570:
1566:
1562:
1558:
1554:
1547:
1545:
1541:
1536:
1532:
1527:
1522:
1518:
1514:
1511:(1): 96–107.
1510:
1506:
1502:
1495:
1492:
1487:
1483:
1479:
1475:
1471:
1467:
1464:(7): 786–93.
1463:
1459:
1452:
1445:
1442:
1437:
1433:
1429:
1425:
1421:
1417:
1413:
1409:
1401:
1398:
1393:
1389:
1385:
1381:
1377:
1373:
1369:
1365:
1358:
1356:
1352:
1347:
1343:
1338:
1333:
1329:
1325:
1321:
1314:
1312:
1308:
1303:
1299:
1294:
1289:
1285:
1281:
1277:
1273:
1270:(6040): 307.
1269:
1265:
1261:
1254:
1252:
1248:
1243:
1239:
1234:
1229:
1225:
1221:
1217:
1213:
1209:
1202:
1200:
1198:
1194:
1189:
1185:
1181:
1177:
1173:
1169:
1165:
1161:
1153:
1151:
1149:
1147:
1145:
1141:
1136:
1132:
1127:
1122:
1118:
1114:
1110:
1106:
1102:
1095:
1093:
1091:
1087:
1082:
1078:
1073:
1068:
1063:
1058:
1054:
1050:
1047:(6): 2623–8.
1046:
1042:
1038:
1031:
1028:
1023:
1019:
1014:
1009:
1005:
1001:
998:(1): 359–72.
997:
993:
989:
982:
979:
974:
970:
965:
960:
956:
952:
949:(2): 757–61.
948:
944:
940:
933:
930:
925:
921:
916:
911:
907:
903:
899:
895:
891:
887:
883:
876:
873:
868:
864:
860:
856:
852:
848:
844:
840:
836:
832:
825:
822:
817:
813:
809:
805:
801:
797:
793:
789:
785:
781:
774:
771:
766:
762:
758:
754:
750:
746:
739:
736:
731:
727:
723:
719:
715:
711:
707:
703:
696:
694:
690:
684:
680:
677:
675:
672:
670:
667:
665:
662:
661:
657:
655:
648:
646:
643:
639:
634:
632:
631:meganucleases
627:
625:
619:
617:
613:
609:
605:
600:
598:
594:
593:knockout rats
590:
589:
584:
580:
576:
567:
565:
563:
559:
556:
554:
550:
546:
540:
535:
533:
530:
526:
522:
514:
507:
503:
501:
496:
492:
488:
484:
476:
474:
471:
467:
459:
457:
454:
450:
446:
445:infect plants
442:
438:
437:
432:
431:TAL effectors
425:
423:
421:
417:
413:
409:
405:
404:
399:
395:
390:
386:
383:
379:
375:
371:
360:
355:
353:
348:
346:
341:
340:
338:
337:
332:
329:
327:
324:
322:
319:
317:
314:
312:
309:
307:
304:
303:
302:
301:
298:Controversies
296:
291:
290:Designer baby
288:
286:
283:
279:
276:
275:
274:
271:
270:
269:
268:
263:
256:
253:
251:
248:
247:
246:
243:
239:
236:
234:
231:
229:
226:
225:
224:
223:Gene delivery
221:
217:
214:
213:
212:
209:
207:
204:
203:
202:
201:
196:
189:
186:
184:
181:
180:
179:
176:
175:
172:
169:
168:
167:
166:
161:
154:
151:
149:
146:
144:
141:
139:
136:
135:
134:
131:
130:
125:
122:
120:
117:
115:
112:
111:
110:
107:
106:
103:
100:
98:
95:
94:
93:
92:
89:
85:
81:
80:
76:
72:
71:
68:
67:
62:
58:
54:
53:
46:
42:
38:
32:
19:
2676:
2667:
2658:
2623:
2619:
2609:
2582:
2578:
2567:
2532:
2528:
2518:
2483:
2479:
2469:
2424:
2420:
2410:
2385:
2381:
2375:
2340:
2336:
2276:
2272:
2262:
2217:
2213:
2203:
2171:(8): 697–8.
2168:
2164:
2154:
2109:
2105:
2095:
2060:
2056:
2008:
2004:
1994:
1967:
1963:
1953:
1918:
1914:
1904:
1859:
1855:
1845:
1800:
1796:
1786:
1751:
1747:
1737:
1702:
1698:
1688:
1653:
1647:
1612:
1608:
1560:
1556:
1508:
1504:
1494:
1461:
1457:
1444:
1411:
1407:
1400:
1367:
1363:
1330:(8): 695–6.
1327:
1323:
1267:
1263:
1218:(8): 731–4.
1215:
1211:
1166:(2): 143–8.
1163:
1159:
1108:
1104:
1044:
1040:
1030:
995:
991:
981:
946:
942:
932:
889:
885:
875:
834:
830:
824:
783:
779:
773:
748:
744:
738:
708:(2): 135–6.
705:
701:
674:Meganuclease
652:
635:
628:
623:
620:
603:
601:
586:
571:
568:Applications
560:
557:
547:
544:
518:
515:Transfection
480:
463:
434:
429:
410:. Alongside
401:
394:gene editing
382:TAL effector
373:
369:
368:
285:Gene therapy
265:Applications
238:Transduction
233:Transfection
64:
2702:E-TALEN.org
1921:(1): 20–7.
1414:(1): 74–9.
1111:(12): e82.
525:transfected
500:zinc finger
436:Xanthomonas
416:CRISPR/Cas9
2718:Categories
751:: 419–36.
685:References
588:C. elegans
541:Mechanisms
453:nucleotide
449:amino acid
443:when they
206:Techniques
178:Regulation
138:Maize/corn
2343:(3): e3.
816:206522347
597:zebrafish
529:expressed
487:annealing
470:nucleases
2650:24285304
2601:26183927
2559:26061646
2510:23546300
2461:24236034
2421:PLOS ONE
2402:25245091
2367:23344620
2313:24523878
2273:PLOS ONE
2254:25733846
2195:21822241
2146:23555929
2106:PLOS ONE
2087:25971621
2063:: 7085.
2035:24871200
2011:: 3831.
1986:24851712
1945:23124327
1896:21625552
1856:PLOS ONE
1837:21625585
1797:PLOS ONE
1778:21459844
1729:21421566
1680:22328436
1639:21248753
1587:21813459
1535:20447404
1486:22079561
1478:17603476
1436:14334237
1428:21131970
1392:28802632
1384:21822242
1346:21822240
1302:21700836
1242:21738127
1188:53549397
1180:21179091
1135:21493687
1081:21262818
1022:20699274
973:20660643
943:Genetics
924:25632877
892:: 8150.
859:19933106
808:19933107
765:19400638
722:21301438
658:See also
604:in vitro
575:biofuels
521:plasmids
400:editing
389:nuclease
97:Bacteria
57:a series
55:Part of
2641:3936731
2550:4817890
2501:3677309
2452:3827243
2429:Bibcode
2358:3381595
2304:3921256
2281:Bibcode
2245:4386332
2222:Bibcode
2186:3154023
2137:3610929
2114:Bibcode
2065:Bibcode
2013:Bibcode
1936:3532252
1887:3098256
1864:Bibcode
1828:3098229
1805:Bibcode
1769:3152341
1720:3141260
1630:3084533
1578:3241638
1526:2885538
1293:3489282
1272:Bibcode
1264:Science
1233:3152587
1126:3130291
1072:3038751
1049:Bibcode
1013:3017587
964:2942870
915:4311247
894:Bibcode
867:6648530
839:Bibcode
831:Science
788:Bibcode
780:Science
624:in situ
403:in situ
396:or for
198:Process
171:History
148:Soybean
124:Insects
114:Mammals
109:Animals
102:Viruses
2648:
2638:
2599:
2557:
2547:
2508:
2498:
2459:
2449:
2400:
2365:
2355:
2311:
2301:
2252:
2242:
2193:
2183:
2144:
2134:
2085:
2033:
1984:
1943:
1933:
1894:
1884:
1835:
1825:
1776:
1766:
1727:
1717:
1678:
1668:
1637:
1627:
1585:
1575:
1533:
1523:
1484:
1476:
1434:
1426:
1390:
1382:
1344:
1300:
1290:
1240:
1230:
1186:
1178:
1133:
1123:
1079:
1069:
1020:
1010:
971:
961:
922:
912:
865:
857:
814:
806:
763:
730:304571
728:
720:
679:CRISPR
642:CRISPR
614:, and
553:indels
398:genome
376:) are
255:CRISPR
153:Potato
133:Plants
82:
2686:(PDF)
1482:S2CID
1454:(PDF)
1432:S2CID
1388:S2CID
1184:S2CID
863:S2CID
812:S2CID
726:S2CID
374:TALEN
250:TALEN
2646:PMID
2597:PMID
2555:PMID
2506:PMID
2457:PMID
2398:PMID
2363:PMID
2309:PMID
2250:PMID
2191:PMID
2142:PMID
2083:PMID
2031:PMID
1982:PMID
1941:PMID
1892:PMID
1833:PMID
1774:PMID
1725:PMID
1676:PMID
1666:ISBN
1635:PMID
1583:PMID
1531:PMID
1474:PMID
1424:PMID
1380:PMID
1342:PMID
1298:PMID
1238:PMID
1176:PMID
1131:PMID
1077:PMID
1018:PMID
969:PMID
920:PMID
855:PMID
804:PMID
761:PMID
718:PMID
638:ZFNs
581:and
466:FokI
414:and
278:food
143:Rice
119:Fish
45:3UGM
41:1FOK
2729:DNA
2636:PMC
2628:doi
2587:doi
2545:PMC
2537:doi
2496:PMC
2488:doi
2447:PMC
2437:doi
2390:doi
2353:PMC
2345:doi
2299:PMC
2289:doi
2240:PMC
2230:doi
2218:112
2181:PMC
2173:doi
2132:PMC
2122:doi
2073:doi
2021:doi
1972:doi
1931:PMC
1923:doi
1919:161
1882:PMC
1872:doi
1823:PMC
1813:doi
1764:PMC
1756:doi
1715:PMC
1707:doi
1658:doi
1625:PMC
1617:doi
1573:PMC
1565:doi
1521:PMC
1513:doi
1509:400
1466:doi
1416:doi
1372:doi
1332:doi
1288:PMC
1280:doi
1268:333
1228:PMC
1220:doi
1168:doi
1121:PMC
1113:doi
1067:PMC
1057:doi
1045:108
1008:PMC
1000:doi
959:PMC
951:doi
947:186
910:PMC
902:doi
847:doi
835:326
796:doi
784:326
753:doi
710:doi
495:PCR
37:PDB
2720::
2666:.
2644:.
2634:.
2624:42
2622:.
2618:.
2595:.
2583:75
2581:.
2577:.
2553:.
2543:.
2533:23
2531:.
2527:.
2504:.
2494:.
2484:21
2482:.
2478:.
2455:.
2445:.
2435:.
2423:.
2419:.
2396:.
2386:14
2384:.
2361:.
2351:.
2339:.
2335:.
2321:^
2307:.
2297:.
2287:.
2275:.
2271:.
2248:.
2238:.
2228:.
2216:.
2212:.
2189:.
2179:.
2169:29
2167:.
2163:.
2140:.
2130:.
2120:.
2108:.
2104:.
2081:.
2071:.
2059:.
2055:.
2043:^
2029:.
2019:.
2007:.
2003:.
1980:.
1968:12
1966:.
1962:.
1939:.
1929:.
1917:.
1913:.
1890:.
1880:.
1870:.
1858:.
1854:.
1831:.
1821:.
1811:.
1799:.
1795:.
1772:.
1762:.
1752:39
1750:.
1746:.
1723:.
1713:.
1703:39
1701:.
1697:.
1674:.
1664:.
1633:.
1623:.
1613:29
1611:.
1607:.
1595:^
1581:.
1571:.
1561:39
1559:.
1555:.
1543:^
1529:.
1519:.
1507:.
1503:.
1480:.
1472:.
1462:25
1460:.
1456:.
1430:.
1422:.
1410:.
1386:.
1378:.
1368:29
1366:.
1354:^
1340:.
1328:29
1326:.
1322:.
1310:^
1296:.
1286:.
1278:.
1266:.
1262:.
1250:^
1236:.
1226:.
1216:29
1214:.
1210:.
1196:^
1182:.
1174:.
1164:29
1162:.
1143:^
1129:.
1119:.
1109:39
1107:.
1103:.
1089:^
1075:.
1065:.
1055:.
1043:.
1039:.
1016:.
1006:.
996:39
994:.
990:.
967:.
957:.
945:.
941:.
918:.
908:.
900:.
888:.
884:.
861:.
853:.
845:.
833:.
810:.
802:.
794:.
782:.
759:.
749:48
747:.
724:.
716:.
706:29
704:.
692:^
610:,
591:,
422:.
59:on
43:,
39::
2688:.
2652:.
2630::
2603:.
2589::
2561:.
2539::
2512:.
2490::
2463:.
2439::
2431::
2425:8
2404:.
2392::
2369:.
2347::
2341:1
2315:.
2291::
2283::
2277:9
2256:.
2232::
2224::
2197:.
2175::
2148:.
2124::
2116::
2110:8
2089:.
2075::
2067::
2061:6
2037:.
2023::
2015::
2009:5
1988:.
1974::
1947:.
1925::
1898:.
1874::
1866::
1860:6
1839:.
1815::
1807::
1801:6
1780:.
1758::
1731:.
1709::
1682:.
1660::
1641:.
1619::
1589:.
1567::
1537:.
1515::
1488:.
1468::
1438:.
1418::
1412:8
1394:.
1374::
1348:.
1334::
1304:.
1282::
1274::
1244:.
1222::
1190:.
1170::
1137:.
1115::
1083:.
1059::
1051::
1024:.
1002::
975:.
953::
926:.
904::
896::
890:5
869:.
849::
841::
818:.
798::
790::
767:.
755::
732:.
712::
372:(
358:e
351:t
344:v
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.