267:
Heavy Chain (NMMHC-IIA). Second, in the late endosome, the low pH causes fusion activity in the membrane of the Gc protein. Uukuniemi virus (UUKV) penetration is promoted by the expression of vesicle-associated membrane protein 3 (VAMP3). Additionally, the fusion of Rift Valley fever virus (RVFV) in late endosomes is inhibited by the interferon-induced transmembrane proteins 2 and 3 (IFITIM2 and IFITIM3). Third, the viral and endosomal membranes are fused to allow for the release of the viral ribonucleoprotein complexes into the cytoplasm (Also the site of viral transcription and replication). Fourth, the precursor protein, Gn/Gc, is translated at the rough endoplasmic reticulum (ER). This precursor protein is cleaved by signal peptidase. Synthesis of the viral nucleoprotein and viral polymerase in the cytoplasm combines with the newly formed genomic RNA (gRNA) ribonucleic protein complexes (RNP). Fifth, two ER chaperones, binding immunoglobulin protein (BiP) and calnexin, are required to ensure proper folding of GN/Gc. Gn/Gc are similarly catalyzed by protein-disulfide-isomerase through the formation of disulfide bonds. At the same time, calreticulin prevents any misfolded Gn/Gc from being exported to the Golgi. Sixth, The correctly folded Gn/Gc heterodimers are transported to the Golgi apparatus. The cytoplasmic tails of Gn in the budding process associate with RNPs during this time. Seventh, once the budding of the new virus particles is completed, vesicles that contain the virus are transported to the plasma membrane to be released by exocytosis.
280:
an icosahedral lattice with a triangulation number of 12. Also included in the lattice composition are 110 hexametric and 12 pentameric capsomeres. For RVFV in particular, 720 Gn/Gc heterodimers are included in the capsomeres. In these cases, Gn forms the spikes of the capsomere while Gc is closer to the lipid membrane, thus placing it underneath. The pH surrounding the capsomere ultimately determines its shape. This is largely due to protonation triggering conformational changes in Gc, commonly included with membrane fusion. An assembly model for the RVFV envelope has been proposed which consists of Gc dimers positioned horizontally with respect to the viral membrane. This is known because the RVFV Gc ectodomain is crystallized as a dimer. This is opposed to the virion interior of bunyaviruses which has no matrix protein, and thus, has no defining organization. This means that on the virion surface, the Gc and Gn proteins must be present in a highly ordered placement.
291:(GAG), which is an unbranched polysaccharide made of disaccharide repeats, that results in the creation of a proteoglycan. Cell lines with defined glycosylation defects were analyzed and showed that HS is necessary for the entry of RVFV. This was confirmed by the removal of HS using an enzyme. HS is charge-dependent in their interactions with virus particles, and studies showed that there are basic amino acid clusters on the P78 protein that interact with negative sulfate clusters on HS. In comparison, there were no identified HS binding sites on Gn and Gc. The P78 protein is plentiful in RVFV-infected cells in insects, while infected cells in mammals produce significantly less P78 proteins. The P78 protein is much more efficiently produced in RVFV in mosquito cells as it is required for viral spread in mosquitos. Overall, the cell line is heavily dependent on HS in RVFV entry.
405:
95% or greater identity in the amino acid sequences of their respective RNA-dependent RNA plymerase (RdRp). Currently, the genus consists of 67 species. Some of the viruses have other hematophagous arthropods as their main vectors. Examples of this include mosquitos for the Rift Valley fever virus, and the Mukawa virus, which has been isolated from ticks, but remains in
Phlebovirus despite the common shift of tick-borne viruses from Phlebovirus to Uukuvirus. In addition to ticks and mosquitos, some Phleboviruses have been isolated from vertebrates like rodents in America and opossums or sloths in Africa. This wide variety of sources shows the possible presence of diversified epidemiological cycles.
295:
class II proteins. An internal fusion loop was discovered, which is a critical aspect of all class II proteins. The location of a histidine in Gc resembled a pH sensing feature, which matches class II characteristics. Although there were many similarities within the structure of the RVFV Gc and class II proteins, the interface between domains I and II in RVFV Gc is more rigid and bigger than other class II proteins. Additionally, RVFV Gc has more disulfide bridges centralized in different locations than the other compared proteins. However, its overall structure and functionality is most closely resembling a class II membrane fusion protein compared to any other class.
316:
and a high case-fatality rate. After this discovery, SFTSV was reported in Japan and Korea as well. North
America had a similar case, which was found to be a result of the Heartland virus, which is transmitted by ticks. These two discoveries changed the perception of the effect of tick-borne phleboviruses with regards to public health. These discoveries caused the re-classification of the Bhanja virus (BHAV) into the tick-borne phlebovirus group. Novel associations of phlebovirus diseases have been reported in the Mediterranean area. Examples include the sandfly fever Turkey virus, Adria virus, Granada virus, Adana virus, and Medjerda virus, among others.
303:
has finished. The Gn and Gc phlebovirus proteins are encoded on the M-segment and undergo synthesis. The precursor Gn/Gc protein cannot be detected in a cell already infected with phlebovirus. It only is visible after the expression of the M-segment. If microsomal membranes are present, the precursor is cleaved, indicating cleaving by a host factor during the synthesis of the viral protein. The signal peptidase complex in the ER membrane is responsible for the cleaving of the precursor. This precursor is then translocated into the ER lumen, in which two hydrophobic domains are inserted with a third, cleaved hydrophobic domain in between.
320:
genus of flying, blood-sucking dipteran in sandy areas. For example, "sandfly" in the United States refers to horse flies or members of the family
Ceratopogonidae. In other parts of the world, "sandfly" refers to members of the subfamily Phlebotominae within Psychodidae. Two of the three main genera were found in the Old World, Phlebotomus and Sergentomyia, and contain the prominent species that transmit the viral pathogens. The third genera was found in the New World and is called Lutzomyia. Other examples are biting midges, or Austrosimulium, a black fly found in New Zealand.
71:
246:
53:
44:
312:
Maintenance of the viruses is mainly completed through the vector species by means of vertical (transovarial) transmission. Concerns over the potential introduction of RVFV into susceptible areas has grown due to the increasing spread of vector species. The potential ramifications of this spread could cause massive economic loss through the harming of livestock.
1283:
Yu, X. J.; Liang, M. F.; Zhang, S. Y.; Liu, Y.; Li, J. D.; Sun, Y. L.; Zhang, L.; Zhang, Q. F.; Popov, V. L.; Li, C.; Qu, J.; Li, Q.; Zhang, Y. P.; Hai, R.; Wu, W.; Wang, Q.; Zhan, F. X.; Wang, X. J.; Kan, B.; Wang, S. W.; Wan, K. L.; Jing, H. Q.; Lu, J. X.; Yin, W. W.; Zhou, H.; Guan, X. H.; Liu, J.
319:
The
Toscana virus has a high rate of vertical transmission, as demonstrated in sandflies through experimental infection. This suggests that there is an amplified role for vertebrate hosts despite the maintenance in nature coming mainly from sandflies. A sandfly is a name for members of any species or
302:
After the viral and endosomal membranes have been fused together, the L,M, and S genomic segments (associated with viral polymerase) are released into the cytoplasm. This initiates the transcription to genomic RNA into mRNA. Viral proteins begin undergoing translation before the transcription of mRNA
283:
To begin entry, phlebovirus particles bind to various components of the plasma membrane. These components interact with the viral glycoproteins of phlebovirus and regulate entry efficiency. While these components are not crucial to the actual entry of the virus itself, receptors are components of the
315:
Two new members of phlebovirus as causative agents of traumatic human disease have been identified. In rural China, SFTSV, which is transmitted by ticks, was identified as the result of increased cases of a febrile illness combined with thrombocytopenia, leukenocytopenia, multiple organ dysfunction,
311:
Phleboviruses are arboviruses taxonomically split into tick- and dipteran-borne viruses. Phlebotomus sandflies are the primary sources for dipteran-borne phleboviruses, with Rift Valley fever virus being the exception (RVFV is associated with mosquitos and has a greater variety in its vector range).
279:
showed that bunyavirus particles are pleomorphic. This known fact cased some surprise when studies showed that UUKKV and RVFV particles are spherically shaped and highly ordered. The configuration of Gn and Gc proteins in the viral envelope imposes the order of the particle. The viral envelope forms
257:
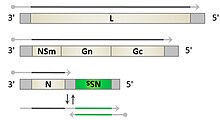
consisting of three segments. The small segment (S) codes for the viral N protein and a non structural protein, NSs via an ambisense coding strategy. The medium-sized segment (M) codes for a precursor of the viral glycoproteins and non-structural components. The product of the largest segment (L) is
404:
The name comes from the Greek root phlebos, which means "vein". Serological cross reactivity previously defined species in the genus. A change in classification rules was prompted due to the difficulty in detecting new phlebovirus in serological assays. As a result, viral species are now defined by
298:
The pH sensing feature in the Gc protein is of note. The membrane fusion activity of the phlebovirus Gc proteins is very dependent on pH, as a low pH triggers the transport of virions into endolysosomes. Elevating intravesicular pH inhibits phlebovirus entry. However, it is still unclear whether Gc
294:
A computational study provided evidence that phlebovirus Gc proteins might be class II membrane fusion proteins. Final proof for this theory was given by the elucidation of the ectodomain's structure of RVFV Gc in its pre-fusion status. Gc has three domains with characteristics that resembled other
266:
The
Phlebovirus replicates in a 7 step process. First, the cellular attachment is driven through the glycoprotein interactions with host cells. Examples of this are Dendritic Cell-Specific Intercellular Adhesion Molecule-3-Grabbing Non-Integrin (DC-SIGN), heparan sulfate (HS), or Non-Muscle Myosin
323:
The result of many of these viruses are some sort of fever. Pappataci fevers, or
Phlebotomus fevers, are mild 3-day fevers that are similar to influenza and have a rapid onset. They are most common in endemic areas during the summer months, particularly August, which is when sandflies are active.
1184:
Palacios, Gustavo; Tesh, Robert; Travassos da Rosa, Amelia; Savji, Nazir; Sze, Wilson; Jain, Komal; Serge, Robert; Guzman, Hilda; Guevara, Carolina; Nunes, Marcio; Nunes-Neto, Joaquim; Kochel, Tadeusz; Hutchinson, Stephen; Vasconcelos, Pedro; Lipkin, Ian (2011).
1174:, Beus I. Marton E. First natural clinical human Bhanja virus infection, p 297–301. 1980. In Vesenjak-Hirjan J, Porterfield JS, Arslanagí, c E (ed), Arboviruses in the Mediterranean countries: 6th FEMS Symposium. Fischer, Stuttgart, Germany.
1074:
Tchouassi, David P.; Marklewitz, Marco; Chepkorir, Edith; Zirkel, Florian; Agha, Sheila B.; Tigoi, Caroline C.; Koskei, Edith; Drosten, Christian; Borgemeister, Christian; Torto, Baldwyn; Junglen, Sandra; Sang, Rosemary (April 2019).
1443:"Comprehensive molecular detection of tick-borne phleboviruses leads to the retrospective identification of taxonomically unassigned bunyaviruses and the discovery of a novel member of the genus phlebovirus"
1394:"Comprehensive molecular detection of tick-borne phleboviruses leads to the retrospective identification of taxonomically unassigned bunyaviruses and the discovery of a novel member of the genus phlebovirus"
324:
Some more extreme cases are the
Toscana virus, which is associated with meningitis in humans, and the Rift Valley fever virus which has caused wide-spread epidemics in livestock in Africa.
284:
plasma membrane that bind to the glycoproteins and are critical for entry. In
Phleboviruses, it was determined that glycan-protein interactions play a crucial role in phlebovirus entry.
1001:
877:
As of 2015, within the phlebovirus there are four genetic groups of tick-borne phleboviruses: the SFTS group, the Bhanja group, the
Uukuniemi group, and the Kaisodi group.
1187:"Characterization of the Candiru antigenic complex (Bunyaviridae: Phlebovirus), a highly diverse and reassorting group of viruses affecting humans in tropical America"
1680:
1693:
1498:"Proteomics computational analyses suggest that the carboxyl terminal glycoproteins of Bunyaviruses are class II viral fusion protein (beta-penetrenes)"
1667:
985:
1392:
Matsuno, K; Weisend, C; Kajihara, M; Matysiak, C; Williamson, BN; Simuunza, M; Mweene, AS; Takada, A; Tesh, RB; Ebihara, H (January 2015).
1134:"Characterization of eight new phlebotomus fever serogroup arboviruses (Bunyaviridae: Phlebovirus) from the Amazon region of Brazil"
1441:
Matsuno, K; Weisend, C; Kajihara, M; Matysiak, C; Williamson, BN; Simuunza, M; Mweene, AS; Takada, A; Tesh, RB; Ebihara, H (2015).
1706:
349:
1540:
1548:"Phylogenetic relationships among members of the genus Phlebovirus (Bunyaviridae) based on partial M segment sequence analyses"
964:
A Greek-English
Lexicon. Revised and augmented throughout by Sir Henry Stuart Jones. With the assistance of Roderick McKenzie.
750:
1234:
Savage, HM; Godsey, MS; Lambert, A; Panella, NA; Burkhalter, KL; Harmon, JR; Lash, RR; Ashley, DC; Nicholson, WL (2013).
70:
1698:
886:
376:(the first tick-borne phlebovirus known to cause disease in the Western Hemisphere, discovered in 2009), and the
834:
764:
470:
715:
213:
757:
701:
659:
589:
389:
778:
708:
638:
617:
610:
603:
1734:
1594:
848:
743:
729:
722:
652:
645:
582:
575:
554:
547:
540:
456:
421:
377:
353:
869:
855:
771:
736:
673:
631:
596:
533:
505:
491:
463:
449:
1739:
841:
806:
799:
785:
694:
666:
568:
526:
519:
477:
428:
862:
820:
687:
561:
512:
484:
414:
624:
442:
1711:
1654:
813:
792:
435:
373:
361:
333:
65:
380:(discovered in China in 2011). They cause symptoms ranging from short self-limiting fevers, such as
827:
498:
365:
345:
341:
680:
385:
218:
1236:"First detection of heartland virus (Bunyaviridae: Phlebovirus) from field collected arthropods"
1685:
1641:
1569:
1529:
1474:
1423:
1374:
1356:
1315:
1265:
1216:
1153:
1114:
1096:
1056:
1038:
981:
357:
288:
1632:
951:
Phlebo: refers to phlebotomine vectors of sandfly fever group viruses; Greek phlebos, "vein".
1559:
1519:
1509:
1464:
1454:
1413:
1405:
1364:
1346:
1305:
1297:
1255:
1247:
1206:
1198:
1171:
1145:
1104:
1088:
1046:
1028:
287:
Heparan Sulfate (HS) is another crucial component aspect in Phlebovirus attachment. It is a
1077:"Sand Fly–Associated Phlebovirus with Evidence of Neutralizing Antibodies in Humans, Kenya"
400:
Phlebovirus is derived from Phlebotominae, which is the taxon of vectors of member species
1580:
381:
119:
52:
43:
1469:
1442:
1418:
1393:
1369:
1334:
1310:
1285:
1260:
1235:
1211:
1186:
1109:
1076:
1051:
1016:
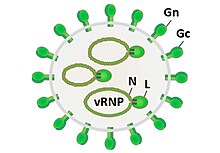
245:
223:
1524:
1497:
1132:
Travassos da Rosa AP, Tesh RB, Pinheiro FP, Travassos da Rosa JF, Peterson NE (1983).
931:
1728:
209:
198:
155:
107:
1546:
Liu DY, Tesh RB, Travassos Da Rosa AP, Peters CJ, Yang Z, Guzman H, Xiao SY (2003).
337:
204:
143:
131:
1646:
1626:
1617:
1149:
17:
1360:
1100:
1042:
1017:"The Role of Phlebovirus Glycoproteins in Viral Entry, Assembly and Release"
95:
1573:
1533:
1514:
1478:
1427:
1378:
1319:
1269:
1220:
1133:
1118:
1092:
1060:
1564:
1547:
1301:
1251:
1157:
1611:
1459:
1409:
1286:"Fever with Thrombocytopenia Associated with a Novel Bunyavirus in China"
1202:
233:, meaning "vein". The proper word for "vein" in ancient Greek is however
1672:
1351:
299:
proteins must bind to a receptor before being triggered by pH or not.
1335:"Taxonomy of Phleboviruses, Emphasizing Those That Are Sandfly-Borne"
1033:
254:
1588:
332:
The following twelve viruses have been linked to disease in humans:
912:. International Committee on Taxonomy of Viruses (ICTV). March 2021
244:
82:
976:
Modrow, Susanne; Falke, Dietrich; Truyen, Uwe; Schätzl, Hermann.
1592:
1015:
Spiegel, Martin; Plegge, Teresa; Pöhlmann, Stefan (July 2016).
909:
1659:
208:. The genus contains 66 species. It derives its name from
60:
Prototypic phlebovirus virion and genome organization.
940:
International Committee on Taxonomy of Viruses (ICTV)
1333:
Calisher, Charles H.; Calzolari, Mattia (May 2021).
307:
An emerging group of arthropod-transmitted pathogens
253:
Phleboviruses are viruses with a negative-sense RNA
1601:
226:
8:
1284:F.; Bi, Z. Q.; Liu, G. H.; Ren, J. (2011).
1589:
369:
222:, which is said to be ultimately from the
51:
42:
31:
1563:
1523:
1513:
1468:
1458:
1417:
1368:
1350:
1309:
1259:
1210:
1108:
1050:
1032:
258:the viral RNA-dependent RNA polymerase.
898:
962:Liddell, H.G. & Scott, R. (1940).
904:
902:
408:The following species are recognized:
271:Role of Gn and Gc in phlebovirus entry
196:is one of twenty genera of the family
7:
1002:"Replication cycle of phleboviruses"
1290:The New England Journal of Medicine
249:Replication cycle of phleboviruses.
25:
402:sandfly fever Naples phlebovirus.
350:Sandfly fever Naples phlebovirus
69:
910:"Virus Taxonomy: 2020 Release"
1:
751:Rift Valley fever phlebovirus
182:
1081:Emerging Infectious Diseases
1496:Garry CE, Garry RF (2004).
227:
1756:
1150:10.4269/ajtmh.1983.32.1164
835:Tres Almendras phlebovirus
980:. Springer. p. 460.
180:
175:
64:
59:
50:
41:
34:
966:Oxford: Clarendon Press.
932:"ICTV 9th Report (2011)
765:Saint Floris phlebovirus
471:Buenaventura phlebovirus
716:Pena Blanca phlebovirus
390:viral hemorrhagic fever
358:Rift Valley fever virus
1541:Course BS335: Virology
1515:10.1186/1742-4682-1-10
1093:10.3201/eid2504.180750
887:Saint Abb's Head virus
758:Rio Grande phlebovirus
744:Punta Toro phlebovirus
702:Odrenisrou phlebovirus
660:Mona Grita phlebovirus
590:Itaporanga phlebovirus
354:Punta Toro phlebovirus
275:A study of the family
250:
1565:10.1099/vir.0.18765-0
1302:10.1056/NEJMoa1010095
1252:10.4269/ajtmh.13-0209
1240:Am. J. Trop. Med. Hyg
1138:Am. J. Trop. Med. Hyg
779:Salehabad phlebovirus
709:Oriximina phlebovirus
639:Maldonado phlebovirus
618:La Gloria phlebovirus
611:Kiborgoch phlebovirus
604:Karimabad phlebovirus
328:Clinical significance
248:
1502:Theor Biol Med Model
1460:10.1128/JVI.02704-14
1410:10.1128/JVI.02704-14
1203:10.1128/JVI.02275-10
849:Uriurana phlebovirus
793:Sicilian phlebovirus
730:Perkerra phlebovirus
723:Penshurt phlebovirus
653:Medjerda phlebovirus
646:Massilia phlebovirus
583:Itaituba phlebovirus
576:Icoaraci phlebovirus
555:Frijoles phlebovirus
548:Embossos phlebovirus
541:Echarate phlebovirus
457:Arumowot phlebovirus
436:Alenquer phlebovirus
422:Aguacate phlebovirus
378:Sandfly Turkey virus
374:Heartland bandavirus
362:Sicilian phlebovirus
334:Alenquer phlebovirus
66:Virus classification
1191:Journal of Virology
1170:Vesenjak-Hirjan J,
870:Zerdali phlebovirus
856:Urucuri phlebovirus
828:Toscana phlebovirus
772:Salanga phlebovirus
737:Punique phlebovirus
674:Munguba phlebovirus
632:Leticia phlebovirus
597:Ixcanal phlebovirus
534:Durania phlebovirus
506:Chagres phlebovirus
499:Candiru phlebovirus
492:Campana phlebovirus
464:Bogoria phlebovirus
450:Anhanga phlebovirus
366:Toscana phlebovirus
342:Candiru phlebovirus
978:Molecular Virology
842:Turuna phlebovirus
807:Tehran phlebovirus
800:Tapara phlebovirus
786:Salobo phlabovirus
695:Ntepes phlebovirus
681:Naples phlebovirus
667:Mukawa phlebovirus
569:Gordil phlebovirus
527:Dashli phlebovirus
520:Corfou phlebovirus
478:Bujaru phlebovirus
429:Alcube phlebovirus
251:
219:Naples phlebovirus
216:of member species
1722:
1721:
1595:Taxon identifiers
1352:10.3390/v13050918
1296:(16): 1523–1532.
987:978-3-642-20718-1
863:Viola phlebovirus
821:Toros phlebovirus
688:Nique phlebovirus
562:Gabek phlebovirus
513:Cocle phlebovirus
485:Cacao phlebovirus
415:Adana phlebovirus
289:glycosaminoglycan
189:
188:
16:(Redirected from
1747:
1715:
1714:
1702:
1701:
1689:
1688:
1676:
1675:
1663:
1662:
1650:
1649:
1637:
1636:
1635:
1622:
1621:
1620:
1590:
1577:
1567:
1558:(Pt 2): 465–73.
1537:
1527:
1517:
1483:
1482:
1472:
1462:
1438:
1432:
1431:
1421:
1389:
1383:
1382:
1372:
1354:
1330:
1324:
1323:
1313:
1280:
1274:
1273:
1263:
1231:
1225:
1224:
1214:
1181:
1175:
1168:
1162:
1161:
1129:
1123:
1122:
1112:
1071:
1065:
1064:
1054:
1036:
1034:10.3390/v8070202
1012:
1006:
1005:
998:
992:
991:
973:
967:
960:
954:
953:
948:
946:
928:
922:
921:
919:
917:
906:
625:Lara phlebovirus
443:Ambe phlebovirus
232:
74:
73:
55:
46:
32:
27:Genus of viruses
21:
1755:
1754:
1750:
1749:
1748:
1746:
1745:
1744:
1725:
1724:
1723:
1718:
1710:
1705:
1697:
1692:
1684:
1679:
1671:
1666:
1658:
1653:
1645:
1640:
1631:
1630:
1625:
1616:
1615:
1610:
1597:
1545:
1495:
1492:
1487:
1486:
1440:
1439:
1435:
1391:
1390:
1386:
1332:
1331:
1327:
1282:
1281:
1277:
1233:
1232:
1228:
1183:
1182:
1178:
1169:
1165:
1131:
1130:
1126:
1073:
1072:
1068:
1014:
1013:
1009:
1000:
999:
995:
988:
975:
974:
970:
961:
957:
944:
942:
930:
929:
925:
915:
913:
908:
907:
900:
895:
883:
875:
814:Tico phebovirus
398:
382:pappataci fever
370:Uukuniemi virus
330:
309:
273:
264:
243:
121:Negarnaviricota
68:
28:
23:
22:
18:Uukuniemi virus
15:
12:
11:
5:
1753:
1751:
1743:
1742:
1737:
1727:
1726:
1720:
1719:
1717:
1716:
1703:
1690:
1677:
1664:
1651:
1638:
1623:
1607:
1605:
1599:
1598:
1593:
1587:
1586:
1578:
1543:
1538:
1491:
1490:External links
1488:
1485:
1484:
1453:(1): 594–604.
1433:
1404:(1): 594–604.
1384:
1325:
1275:
1226:
1197:(8): 3811–20.
1176:
1163:
1144:(5): 1164–71.
1124:
1087:(4): 681–690.
1066:
1007:
993:
986:
968:
955:
923:
897:
896:
894:
891:
890:
889:
882:
879:
874:
873:
866:
859:
852:
845:
838:
831:
824:
817:
810:
803:
796:
789:
782:
775:
768:
761:
754:
747:
740:
733:
726:
719:
712:
705:
698:
691:
684:
677:
670:
663:
656:
649:
642:
635:
628:
621:
614:
607:
600:
593:
586:
579:
572:
565:
558:
551:
544:
537:
530:
523:
516:
509:
502:
495:
488:
481:
474:
467:
460:
453:
446:
439:
432:
425:
418:
410:
397:
394:
329:
326:
308:
305:
272:
269:
263:
260:
242:
239:
187:
186:
178:
177:
173:
172:
165:
161:
160:
153:
149:
148:
141:
137:
136:
133:Ellioviricetes
129:
125:
124:
117:
113:
112:
105:
101:
100:
93:
86:
85:
80:
76:
75:
62:
61:
57:
56:
48:
47:
39:
38:
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
1752:
1741:
1738:
1736:
1735:Phleboviruses
1733:
1732:
1730:
1713:
1708:
1704:
1700:
1695:
1691:
1687:
1682:
1678:
1674:
1669:
1665:
1661:
1656:
1652:
1648:
1643:
1639:
1634:
1628:
1624:
1619:
1613:
1609:
1608:
1606:
1604:
1600:
1596:
1591:
1585:
1584:: Phlebovirus
1583:
1579:
1575:
1571:
1566:
1561:
1557:
1553:
1552:J. Gen. Virol
1549:
1544:
1542:
1539:
1535:
1531:
1526:
1521:
1516:
1511:
1507:
1503:
1499:
1494:
1493:
1489:
1480:
1476:
1471:
1466:
1461:
1456:
1452:
1448:
1444:
1437:
1434:
1429:
1425:
1420:
1415:
1411:
1407:
1403:
1399:
1395:
1388:
1385:
1380:
1376:
1371:
1366:
1362:
1358:
1353:
1348:
1344:
1340:
1336:
1329:
1326:
1321:
1317:
1312:
1307:
1303:
1299:
1295:
1291:
1287:
1279:
1276:
1271:
1267:
1262:
1257:
1253:
1249:
1246:(3): 445–52.
1245:
1241:
1237:
1230:
1227:
1222:
1218:
1213:
1208:
1204:
1200:
1196:
1192:
1188:
1180:
1177:
1173:
1167:
1164:
1159:
1155:
1151:
1147:
1143:
1139:
1135:
1128:
1125:
1120:
1116:
1111:
1106:
1102:
1098:
1094:
1090:
1086:
1082:
1078:
1070:
1067:
1062:
1058:
1053:
1048:
1044:
1040:
1035:
1030:
1026:
1022:
1018:
1011:
1008:
1003:
997:
994:
989:
983:
979:
972:
969:
965:
959:
956:
952:
941:
937:
935:
927:
924:
911:
905:
903:
899:
892:
888:
885:
884:
880:
878:
872:
871:
867:
865:
864:
860:
858:
857:
853:
851:
850:
846:
844:
843:
839:
837:
836:
832:
830:
829:
825:
823:
822:
818:
816:
815:
811:
809:
808:
804:
802:
801:
797:
795:
794:
790:
788:
787:
783:
781:
780:
776:
774:
773:
769:
767:
766:
762:
760:
759:
755:
753:
752:
748:
746:
745:
741:
739:
738:
734:
732:
731:
727:
725:
724:
720:
718:
717:
713:
711:
710:
706:
704:
703:
699:
697:
696:
692:
690:
689:
685:
683:
682:
678:
676:
675:
671:
669:
668:
664:
662:
661:
657:
655:
654:
650:
648:
647:
643:
641:
640:
636:
634:
633:
629:
627:
626:
622:
620:
619:
615:
613:
612:
608:
606:
605:
601:
599:
598:
594:
592:
591:
587:
585:
584:
580:
578:
577:
573:
571:
570:
566:
564:
563:
559:
557:
556:
552:
550:
549:
545:
543:
542:
538:
536:
535:
531:
529:
528:
524:
522:
521:
517:
515:
514:
510:
508:
507:
503:
501:
500:
496:
494:
493:
489:
487:
486:
482:
480:
479:
475:
473:
472:
468:
466:
465:
461:
459:
458:
454:
452:
451:
447:
445:
444:
440:
438:
437:
433:
431:
430:
426:
424:
423:
419:
417:
416:
412:
411:
409:
406:
403:
395:
393:
391:
387:
383:
379:
375:
371:
367:
363:
359:
355:
351:
347:
346:Chagres virus
343:
339:
335:
327:
325:
321:
317:
313:
306:
304:
300:
296:
292:
290:
285:
281:
278:
270:
268:
261:
259:
256:
247:
240:
238:
236:
231:
230:
225:
221:
220:
215:
211:
210:Phlebotominae
207:
206:
202:in the order
201:
200:
199:Phenuiviridae
195:
194:
185:
184:
179:
174:
171:
170:
166:
163:
162:
159:
158:
157:Phenuiviridae
154:
151:
150:
147:
146:
142:
139:
138:
135:
134:
130:
127:
126:
123:
122:
118:
115:
114:
111:
110:
109:Orthornavirae
106:
103:
102:
99:
98:
94:
91:
88:
87:
84:
81:
78:
77:
72:
67:
63:
58:
54:
49:
45:
40:
37:
33:
30:
19:
1740:Virus genera
1602:
1581:
1555:
1551:
1505:
1501:
1450:
1446:
1436:
1401:
1397:
1387:
1342:
1338:
1328:
1293:
1289:
1278:
1243:
1239:
1229:
1194:
1190:
1179:
1166:
1141:
1137:
1127:
1084:
1080:
1069:
1024:
1020:
1010:
996:
977:
971:
963:
958:
950:
943:. Retrieved
939:
934:Bunyaviridae
933:
926:
914:. Retrieved
876:
868:
861:
854:
847:
840:
833:
826:
819:
812:
805:
798:
791:
784:
777:
770:
763:
756:
749:
742:
735:
728:
721:
714:
707:
700:
693:
686:
679:
672:
665:
658:
651:
644:
637:
630:
623:
616:
609:
602:
595:
588:
581:
574:
567:
560:
553:
546:
539:
532:
525:
518:
511:
504:
497:
490:
483:
476:
469:
462:
455:
448:
441:
434:
427:
420:
413:
407:
401:
399:
386:encephalitis
338:Bhanja virus
331:
322:
318:
314:
310:
301:
297:
293:
286:
282:
277:Bunyaviridae
276:
274:
265:
252:
234:
228:
217:
205:Bunyavirales
203:
197:
192:
191:
190:
181:
168:
167:
156:
145:Bunyavirales
144:
132:
120:
108:
96:
89:
79:(unranked):
35:
29:
1633:Phlebovirus
1627:Wikispecies
1603:Phlebovirus
1172:Calisher CH
262:Replication
193:Phlebovirus
169:Phlebovirus
36:Phlebovirus
1729:Categories
1345:(5): 918.
1027:(7): 202.
945:31 January
893:References
388:and fatal
1582:Viralzone
1361:1999-4915
1101:1080-6040
1043:1999-4915
104:Kingdom:
97:Riboviria
1673:10123934
1618:Q3381330
1612:Wikidata
1574:12560581
1534:15544707
1479:25339769
1428:25339769
1379:34063467
1320:21410387
1270:23878186
1221:21289119
1119:30882303
1061:27455305
881:See also
396:Taxonomy
241:Virology
237:(φλέψ).
183:See text
176:Species
152:Family:
116:Phylum:
1686:1039879
1470:4301164
1447:J Virol
1419:4301164
1398:J Virol
1370:8156068
1339:Viruses
1311:3113718
1261:3771279
1212:3126144
1158:6312820
1110:6433041
1052:4974537
1021:Viruses
229:phlebos
214:vectors
164:Genus:
140:Order:
128:Class:
1712:600300
1572:
1532:
1525:535339
1522:
1508:: 10.
1477:
1467:
1426:
1416:
1377:
1367:
1359:
1318:
1308:
1268:
1258:
1219:
1209:
1156:
1117:
1107:
1099:
1059:
1049:
1041:
984:
916:19 May
255:genome
235:phleps
212:, the
1707:WoRMS
1699:11584
1681:IRMNG
1660:67734
384:, to
224:Greek
90:Realm
83:Virus
1694:NCBI
1668:GBIF
1647:6NP7
1570:PMID
1530:PMID
1475:PMID
1424:PMID
1375:PMID
1357:ISSN
1316:PMID
1266:PMID
1217:PMID
1154:PMID
1115:PMID
1097:ISSN
1057:PMID
1039:ISSN
982:ISBN
947:2019
918:2021
1655:EoL
1642:CoL
1560:doi
1520:PMC
1510:doi
1465:PMC
1455:doi
1414:PMC
1406:doi
1365:PMC
1347:doi
1306:PMC
1298:doi
1294:364
1256:PMC
1248:doi
1207:PMC
1199:doi
1146:doi
1105:PMC
1089:doi
1047:PMC
1029:doi
1731::
1709::
1696::
1683::
1670::
1657::
1644::
1629::
1614::
1568:.
1556:84
1554:.
1550:.
1528:.
1518:.
1504:.
1500:.
1473:.
1463:.
1451:89
1449:.
1445:.
1422:.
1412:.
1402:89
1400:.
1396:.
1373:.
1363:.
1355:.
1343:13
1341:.
1337:.
1314:.
1304:.
1292:.
1288:.
1264:.
1254:.
1244:89
1242:.
1238:.
1215:.
1205:.
1195:85
1193:.
1189:.
1152:.
1142:32
1140:.
1136:.
1113:.
1103:.
1095:.
1085:25
1083:.
1079:.
1055:.
1045:.
1037:.
1023:.
1019:.
949:.
938:.
901:^
392:.
372:,
368:,
364:,
360:,
356:,
352:,
348:,
344:,
340:,
336:,
92::
1576:.
1562::
1536:.
1512::
1506:1
1481:.
1457::
1430:.
1408::
1381:.
1349::
1322:.
1300::
1272:.
1250::
1223:.
1201::
1160:.
1148::
1121:.
1091::
1063:.
1031::
1025:8
1004:.
990:.
936:"
920:.
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.