1121:
field. Different parametrization procedures have been developed for the parametrization of different substances, e.g. metals, ions, and molecules. For different material types, usually different parametrization strategies are used. In general, two main types can be distinguished for the parametrization, either using data/ information from the atomistic level, e.g. from quantum mechanical calculations or spectroscopic data, or using data from macroscopic properties, e.g. the hardness or compressibility of a given material. Often a combination of these routes is used. Hence, one way or the other, the force field parameters are always determined in an empirical way. Nevertheless, the term 'empirical' is often used in the context of force field parameters when macroscopic material property data was used for the fitting. Experimental data (microscopic and macroscopic) included for the fit, for example, the
1137:, and various spectroscopic properties such as vibrational frequencies. Often, for molecular systems, quantum mechanical calculations in the gas phase are used for parametrizing intramolecular interactions and parametrizing intermolecular dispersive interactions by using macroscopic properties such as liquid densities. The assignment of atomic charges often follows quantum mechanical protocols with some heuristics, which can lead to significant deviation in representing specific properties.
7631:
1369:
program DYANA (calculations from NMR data), or with programs for crystallographic refinement that use no energy functions at all. These shortcomings are related to interatomic potentials and to the inability to sample the conformation space of large molecules effectively. Thereby also the development of parameters to tackle such large-scale problems requires new approaches. A specific problem area is
1532:(GROningen MOlecular Simulation) – a force field that comes as part of the GROMOS software, a general-purpose molecular dynamics computer simulation package for the study of biomolecular systems. GROMOS force field A-version has been developed for application to aqueous or apolar solutions of proteins, nucleotides, and sugars. A B-version to simulate gas phase isolated molecules is also available.
1283:(DFT) calculations, with access to million times larger systems and time scales, to random guesses depending on the force field. The use of accurate representations of chemical bonding, combined with reproducible experimental data and validation, can lead to lasting interatomic potentials of high quality with much fewer parameters and assumptions in comparison to DFT-level quantum methods.
33:
250:
1505:, Lifson, and coworkers as a general method for unifying studies of energies, structures, and vibration of general molecules and molecular crystals. The CFF program, developed by Levitt and Warshel, is based on the Cartesian representation of all the atoms, and it served as the basis for many subsequent simulation programs.
1589:, where a particle's effective charge can be influenced by electrostatic interactions with its neighbors. Core-shell models are common, which consist of a positively charged core particle, representing the polarizable atom, and a negatively charged particle attached to the core atom through a spring-like
1827:– An AMBER-based forcefield and webtool for modeling common post-translational modifications of amino acids in proteins developed by Chris Floudas and coworkers. It uses the ff03 charge model and has several side-chain torsion corrections parameterized to match the quantum chemical rotational surface.
1665:
XED (eXtended
Electron Distribution) - a polarizable force-field created as a modification of an atom-centered charge model, developed by Andy Vinter. Partially charged monopoles are placed surrounding atoms to simulate more geometrically accurate electrostatic potentials at a fraction of the expense
1210:
Heuristic force field parametrization procedures have been very successfully for many year, but recently criticized. since they are usually not fully automated and therefore subject to some subjectivity of the developers, which also brings problems regarding the reproducibility of the parametrization
857:
Though the formula of Hooke's law provides a reasonable level of accuracy at bond lengths near the equilibrium distance, it is less accurate as one moves away. In order to model the Morse curve better one could employ cubic and higher powers. However, for most practical applications these differences
1331:
state that the interaction energy of two dissimilar atoms (e.g., C...N) is an average of the interaction energies of corresponding identical atom pairs (i.e., C...C and N...N). According to McLachlan's theory, the interactions of particles in media can even be fully repulsive, as observed for liquid
1255:
and transferability. When functional forms of the potential terms vary or are mixed, the parameters from one interatomic potential function can typically not be used together with another interatomic potential function. In some cases, modifications can be made with minor effort, for example, between
1618:
COSMOS-NMR (Computer
Simulation of Molecular Structure) – developed by Ulrich Sternberg and coworkers. Hybrid QM/MM force field enables explicit quantum-mechanical calculation of electrostatic properties using localized bond orbitals with fast BPT formalism. Atomic charge fluctuation is possible in
1535:
IFF (Interface Force Field) – covers metals, minerals, 2D materials, and polymers. It uses 12-6 LJ and 9-6 LJ interactions. IFF was developed as for compounds across the periodic table. It assigs consistent charges, utilizes standard conditions as a reference state, reproduces structures, energies,
1368:
applications, especially using program XPLOR. However, the refinement is driven mainly by a set of experimental constraints and the interatomic potentials serve mainly to remove interatomic hindrances. The results of calculations were practically the same with rigid sphere potentials implemented in
1120:
In addition to the functional form of the potentials, a force fields consists of the parameters of these functions. Together, they specify the interactions on the atomistic level. The parametrization, i.e. determining of the parameter values, is crucial for the accuracy and reliability of the force
1661:
WASABe v1.0 PFF (for Water, orgAnic
Solvents, And Battery electrolytes) An isotropic atomic dipole polarizable force field for accurate description of battery electrolytes in terms of thermodynamic and dynamic properties for high lithium salt concentrations in sulfonate solvent by Oleg Starovoytov
1318:
theory of these forces applies only in a vacuum. A more general theory of van der Waals forces in condensed media was developed by A. D. McLachlan in 1963 and included the original London's approach as a special case. The McLachlan theory predicts that van der Waals attractions in media are weaker
1140:
A large number of workflows and parametrization procedures have been employed in the past decades using different data and optimization strategies for determining the force field parameters. They differ significantly, which is also due to different focuses of different developments. The parameters
1742:
VAMM (Virtual atom molecular mechanics) – a coarse-grained force field developed by Korkut and
Hendrickson for molecular mechanics calculations such as large scale conformational transitions based on the virtual interactions of C-alpha atoms. It is a knowledge based force field and formulated to
1694:
and coworkers. It is slower than classical MD (50x), needs parameter sets with specific validation, and has no validation for surface and interfacial energies. Parameters are non-interpretable. It can be used atomistic-scale dynamical simulations of chemical reactions. Parallelized ReaxFF allows
1348:". Such effects are represented in molecular dynamics through pairwise interactions that are spatially more dense in the condensed phase relative to the gas phase and reproduced once the parameters for all phases are validated to reproduce chemical bonding, density, and cohesive/surface energy.
1230:
A large number of force fields has been published in the past decades - mostly in scientific publications. In recent years, some databases have attempted to collect, categorize and make force fields digitally available. Therein, different databases, focus on different types of force fields. For
1099:
Atomistic interactions in crystal systems significantly deviate from those in molecular systems, e.g. of organic molecules. For crystal systems, in particular multi-body interactions are important and cannot be neglected if a high accuracy of the force field is the aim. For crystal systems with
1514:
COSMOS-NMR – hybrid QM/MM force field adapted to various inorganic compounds, organic compounds, and biological macromolecules, including semi-empirical calculation of atomic charges NMR properties. COSMOS-NMR is optimized for NMR-based structure elucidation and implemented in COSMOS molecular
1090:
Atomic charges can make dominant contributions to the potential energy, especially for polar molecules and ionic compounds, and are critical to simulate the geometry, interaction energy, and the reactivity. The assignment of charges usually uses some heuristic approach, with different possible
1657:
GFN-FF (Geometry, Frequency, and
Noncovalent Interaction Force-Field) – a completely automated partially polarizable generic force-field for the accurate description of structures and dynamics of large molecules across the periodic table developed by Stefan Grimme and Sebastian Spicher at the
1575:
UFF (Universal Force Field) – A general force field with parameters for the full periodic table up to and including the actinoids, developed at
Colorado State University. The reliability is known to be poor due to lack of validation and interpretation of the parameters for nearly all claimed
1521:
ECEPP – first force field for polypeptide molecules - developed by F.A. Momany, H.A. Scheraga and colleagues. ECEPP was developed specifically for the modeling of peptides and proteins. It uses fixed geometries of amino acid residues to simplify the potential energy surface. Thus, the energy
196:
There are various criteria that can be used for categorizing force field parametrization strategies. An important differentiation is 'component-specific' and 'transferable'. For a component-specific parametrization, the considered force field is developed solely for describing a single given
1290:
charge distributions. The remedy is that point charges have a clear interpretation and virtual electrons can be added to capture essential features of the electronic structure, such additional polarizability in metallic systems to describe the image potential, internal multipole moments in
4891:
Momany FA, McGuire RF, Burgess AW, Scheraga HA (October 1975). "Energy parameters in polypeptides. VII. Geometric parameters, partial atomic charges, nonbonded interactions, hydrogen bond interactions, and intrinsic torsional potentials for the naturally occurring amino acids".
1222:. The use of semi-automation or full automation, without input from chemical knowledge, is likely to increase inconsistencies at the level of atomic charges, for the assignment of remaining parameters, and likely to dilute the interpretability and performance of parameters.
1558:
and other small organic molecules. It is designed to reproduce the equilibrium covalent geometry of molecules as precisely as possible. It implements a large set of parameters that is continuously refined and updated for many different classes of organic compounds (MM3 and
1778:Δ-ML not a force field method but a model that adds learnt correctional energy terms to approximate and relatively computationally cheap quantum chemical methods in order to provide an accuracy level of a higher order, more computationally expensive quantum chemical model.
1833:- An AMBER-based forcefield and webtool for modeling common non-natural amino acids in proteins in condensed-phase simulations using the ff03 charge model. The charges have been reported to be correlated with hydration free energies of corresponding side-chain analogs.
99:
1728:, initially developed for molecular dynamics simulations of lipids, later extended to various other molecules. The force field applies a mapping of four heavy atoms to one CG interaction site and is parameterized with the aim of reproducing thermodynamic properties.
1351:
Limitations have been strongly felt in protein structure refinement. The major underlying challenge is the huge conformation space of polymeric molecules, which grows beyond current computational feasibility when containing more than ~20 monomers. Participants in
486:. A large number of force fields based on this or similar energy expressions have been proposed in the past decades for modeling different types of materials such as molecular substances, metals, glasses etc. - see below for a comprehensive list of force fields.
1738:
SIRAH – a coarse-grained force field developed by
Pantano and coworkers of the Biomolecular Simulations Group, Institut Pasteur of Montevideo, Uruguay; developed for molecular dynamics of water, DNA, and proteins. Free available for AMBER and GROMACS
1600:
AMOEBA (Atomic
Multipole Optimized Energetics for Biomolecular Applications) – force field developed by Pengyu Ren (University of Texas at Austin) and Jay W. Ponder (Washington University). AMOEBA force field is gradually moving to more physics-rich
1540:(IFF) assumes one single energy expression for all compounds across the periodic (with 9-6 and 12-6 LJ options). The IFF is in most parts non-polarizable, but also comprises polarizable parts, e.g. for some metals (Au, W) and pi-conjugated molecules
1709:) – This is a method commonly applied in chemical engineering. It is typically used for studying the hydrodynamics of various simple and complex fluids which require consideration of time and length scales larger than those accessible to classical
3978:
Pramanik C, Jamil T, Gissinger JR, Guittet D, Arias-Monje PJ, Kumar S, Heinz H (2019-10-03). "Polyacrylonitrile
Interactions with Carbon Nanotubes in Solution: Conformations and Binding as a Function of Solvent, Temperature, and Concentration".
1628:
PIPF – The polarizable intermolecular potential for fluids is an induced point-dipole force field for organic liquids and biopolymers. The molecular polarization is based on Thole's interacting dipole (TID) model and was developed by Jiali Gao
1011:
1522:
minimization is conducted in the space of protein torsion angles. Both MM2 and ECEPP include potentials for H-bonds and torsion potentials for describing rotations around single bonds. ECEPP/3 was implemented (with some modifications) in
5313:
Warshel A, Levitt M (1974). QCFF/PI: A Program for the
Consistent Force Field Evaluation of Equilibrium Geometries and Vibrational Frequencies of Molecules (Report). Indiana University: Quantum Chemistry Program Exchange. p. QCPE
1717:
workshop in 2008. Recently, work has been undertaken to capture some of the chemical subtitles relevant to solutions. This has led to work considering automated parameterisation of the DPD interaction potentials against experimental
1753:
MACE (Multi Atomic Cluster Expansion) is a highly accurate machine learning force field architecture that combines the rigorous many-body expansion of the total potential energy with rotationally equivariant representations of the
446:
394:
1843:
LFMM (Ligand Field Molecular Mechanics) - functions for the coordination sphere around transition metals based on the angular overlap model (AOM). Implemented in the Molecular Operating Environment (MOE) as DommiMOE and in
1428:
is significantly smaller than enthalpy of sublimation. Hence, the potentials describing protein folding or ligand binding need more consistent parameterization protocols, e.g., as described for IFF. Indeed, the energies of
2845:
Dauber-Osguthorpe P, Roberts VA, Osguthorpe DJ, Wolff J, Genest M, Hagler AT (1988). "Structure and energetics of ligand binding to proteins: Escherichia coli dihydrofolate reductase-trimethoprim, a drug-receptor system".
2454:
Mishra RK, Fernández-Carrasco L, Flatt RJ, Heinz H (July 2014). "A force field for tricalcium aluminate to characterize surface properties, initial hydration, and organically modified interfaces in atomic resolution".
7363:"Forcefield_NCAA: ab initio charge parameters to aid in the discovery and design of therapeutic proteins and peptides with unnatural amino acids and their application to complement inhibitors of the compstatin family"
597:
328:
3081:
Dharmawardhana CC, Kanhaiya K, Lin TJ, Garley A, Knecht MR, Zhou J, Miao J, Heinz H (2017-06-19). "Reliable computational design of biological-inorganic materials to the large nanometer scale using Interface-FF".
1680:) – reactive force field introduced by Warshel and coworkers for use in modeling chemical reactions in different environments. The EVB facilitates calculating activation free energies in condensed phases and in
450:
The bond and angle terms are usually modeled by quadratic energy functions that do not allow bond breaking. A more realistic description of a covalent bond at higher stretching is provided by the more expensive
166:, with the main difference being that the force field parameters in chemistry describe the energy landscape on the atomistic level. From a force field, the acting forces on every particle are derived as a
3448:
Ruiz VG, Liu W, Tkatchenko A (2016-01-15). "Density-functional theory with screened van der Waals interactions applied to atomic and molecular adsorbates on close-packed and non-close-packed surfaces".
5967:
Engkvist O, Astrand PO, Karlström G (November 2000). "Accurate Intermolecular Potentials Obtained from Molecular Wave Functions: Bridging the Gap between Quantum Chemistry and Molecular Simulations".
5778:
Anisimov VM, Lamoureux G, Vorobyov IV, Huang N, Roux B, MacKerell AD (January 2005). "Determination of Electrostatic Parameters for a Polarizable Force Field Based on the Classical Drude Oscillator".
5286:
Warshel A (1973). "Quantum mechanical consistent force field (QCFF/PI) method: Calculations of energies, conformations and vibronic interactions of ground and excited states of conjugated molecules".
862:
can be employed instead to enable bond breaking and higher accuracy, even though it is less efficient to compute. For reactive force fields, bond breaking and bond orders are additionally considered.
1648:
Gaussian Electrostatic Model (GEM) – a polarizable force field based on Density Fitting developed by Thomas A. Darden and G. Andrés Cisneros at NIEHS; and Jean-Philip Piquemal at Paris VI University.
1651:
Atomistic Polarizable Potential for Liquids, Electrolytes, and Polymers(APPLE&P), developed by Oleg Borogin, Dmitry Bedrov and coworkers, which is distributed by Wasatch Molecular Incorporated.
1286:
Possible limitations include atomic charges, also called point charges. Most force fields rely on point charges to reproduce the electrostatic potential around molecules, which works less well for
5010:
Heinz H, Lin TJ, Mishra RK, Emami FS (February 2013). "Thermodynamically consistent force fields for the assembly of inorganic, organic, and biological nanostructures: the INTERFACE force field".
6078:
Maple JR, Cao Y, Damm W, Halgren TA, Kaminski GA, Zhang LY, Friesner RA (July 2005). "A Polarizable Force Field and Continuum Solvation Methodology for Modeling of Protein-Ligand Interactions".
6450:
Starovoytov ON (September 2021). "Development of Polarizable Force Field for Molecular Dynamics Simulation of Lithium-Ion Battery Electrolytes: Sulfonate Based Solvents and Lithium Salts".
5259:
Lii JH, Allinger NL (November 1989). "Molecular mechanics. The MM3 force field for hydrocarbons. 3. The van der Waals' potentials and crystal data for aliphatic and aromatic hydrocarbons".
4811:
Gavezzotti A, Filippini G (May 1994). "Geometry of the intermolecular XH. cntdot.. cntdot.. cntdot. Y (X, Y= N, O) hydrogen bond and the calibration of empirical hydrogen-bond potentials".
1303:. However, application of one value of dielectric constant is a coarse approximation in the highly heterogeneous environments of proteins, biological membranes, minerals, or electrolytes.
3371:
Heinz H, Ramezani-Dakhel H (January 2016). "Simulations of inorganic-bioorganic interfaces to discover new materials: insights, comparisons to experiment, challenges, and opportunities".
1336:, however, the lack of vaporization and presence of a freezing point contradicts a theory of purely repulsive interactions. Measurements of attractive forces between different materials (
1388:
It was also argued that some protein force fields operate with energies that are irrelevant to protein folding or ligand binding. The parameters of proteins force fields reproduce the
919:
1901:
1481:
7592:
858:
are negligible, and inaccuracies in predictions of bond lengths are on the order of the thousandth of an angstrom, which is also the limit of reliability for common force fields. A
4633:
Shakhnovich EI, Finkelstein AV (October 1989). "Theory of cooperative transitions in protein molecules. I. Why denaturation of globular protein is a first-order phase transition".
2805:
Jorgensen WL, Maxwell DS, Tirado-Rives J (January 1996). "Development and Testing of the OPLS All-Atom Force Field on Conformational Energetics and Properties of Organic Liquids".
1604:
CHARMM – polarizable force field developed by S. Patel (University of Delaware) and C. L. Brooks III (University of Michigan). Based on the classical Drude oscillator developed by
3177:
McDonagh JL, Shkurti A, Bray DJ, Anderson RL, Pyzer-Knapp EO (October 2019). "Utilizing Machine Learning for Efficient Parameterization of Coarse Grained Molecular Force Fields".
197:
substance (e.g. water). For a transferable force field, all or some parameters are designed as building blocks and become transferable/ applicable for different substances (e.g.
5862:
Warshel A, Levitt M (May 1976). "Theoretical studies of enzymic reactions: dielectric, electrostatic and steric stabilization of the carbonium ion in the reaction of lysozyme".
5325:
Rappé AK, Casewit CJ, Colwell KS, Goddard III WA, Skiff WM (December 1992). "UFF, a full periodic table force field for molecular mechanics and molecular dynamics simulations".
193:). Depending on the material, different functional forms are usually chosen for the force fields since different types of atomistic interactions dominate the material behavior.
7410:
Khoury GA, Bhatia N, Floudas CA (2014). "Hydration free energies calculated using the AMBER ff03 charge model for natural and unnatural amino acids and multiple water models".
5360:
Heinz H, Koerner H, Anderson KL, Vaia RA, Farmer BL (November 2005). "Force Field for Mica-Type Silicates and Dynamics of Octadecylammonium Chains Grafted to Montmorillonite".
4954:
Schaumann T, Braun W, Wüthrich K (March 1990). "The program FANTOM for energy refinement of polypeptides and proteins using a Newton–Raphson minimizer in torsion angle space".
3642:
Xu R, Chen CC, Wu L, Scott MC, Theis W, Ophus C, et al. (November 2015). "Three-dimensional coordinates of individual atoms in materials revealed by electrontomography".
1501:
CFF (Consistent Force Field) – a family of force fields adapted to a broad variety of organic compounds, includes force fields for polymers, metals, etc. CFF was developed by
1757:
ANI (Artificial Narrow Intelligence) is a transferable neural network potential, built from atomic environment vectors, and able to provide DFT accuracy in terms of energies.
1353:
130:
model that is used to describe the forces between atoms (or collections of atoms) within molecules or between molecules as well as in crystals. Force fields are a variety of
6113:
Chelli R, Procacci P (November 2002). "A transferable polarizable electrostatic force field for molecular mechanics based on the chemical potential equalization principle".
1358:
a central embarrassment of molecular mechanics, namely that energy minimization or molecular dynamics generally leads to a model that is less like the experimental structure
398:
333:
1713:. The potential was originally proposed by Hoogerbrugge and Koelman with later modifications by Español and Warren The current state of the art was well documented in a
1412:
of mobile molecules in condensed media. Thus, free energy changes during protein folding or ligand binding are expected to represent a combination of an energy similar to
470:
The nonbonded terms are computationally most intensive. A popular choice is to limit interactions to pairwise energies. The van der Waals term is usually computed with a
2924:
Kirschner KN, Lins RD, Maass A, Soares TA (November 2012). "A Glycam-Based Force Field for Simulations of Lipopolysaccharide Membranes: Parametrization and Validation".
769:
693:
455:. The functional form for dihedral energy is variable from one force field to another. Additional, "improper torsional" terms may be added to enforce the planarity of
3596:
Mahoney MW, Jorgensen WL (2000-05-22). "A five-site model for liquid water and the reproduction of the density anomaly by rigid, nonpolarizable potential functions".
2889:
Aduri R, Psciuk BT, Saro P, Taniga H, Schlegel HB, SantaLucia J (July 2007). "AMBER Force Field Parameters for the Naturally Occurring Modified Nucleosides in RNA".
1041:
844:
802:
657:
627:
898:
4676:
Graziano G, Catanzano F, Del Vecchio P, Giancola C, Barone G (1996). "Thermodynamic stability of globular proteins: a reliable model from small molecule studies".
3706:
Pramanik C, Gissinger JR, Kumar S, Heinz H (December 2017). "Carbon Nanotube Dispersion in Solvents and Polymer Solutions: Mechanisms, Assembly, and Preferences".
463:, and "cross-terms" that describe the coupling of different internal variables, such as angles and bond lengths. Some force fields also include explicit terms for
6309:"Generalization of the Gaussian electrostatic model: extension to arbitrary angular momentum, distributed multipoles, and speedup with reciprocal space methods"
5535:
Patel S, Brooks CL (January 2004). "CHARMM fluctuating charge force field for proteins: I parameterization and application to bulk organic liquid simulations".
1811:(flexible SPC), ST2, and mW. Other solvents and methods of solvent representation are also applied within computational chemistry and physics; these are termed
1593:
potential. Recent examples include polarizable models with virtual electrons that reproduce image charges in metals and polarizable biomolecular force fields.
1081:
1061:
733:
713:
5731:"CHARMM fluctuating charge force field for proteins: II protein/solvent properties from molecular dynamics simulations using a nonadditive electrostatic model"
4241:
Gohlke H, Klebe G (August 2002). "Approaches to the description and prediction of the binding affinity of small-molecule ligands to macromolecular receptors".
501:
5897:
Sternberg U, Koch FT, Möllhoff M (May 1994). "New approach to the semiempirical calculation of atomic charges for polypeptides and large molecular systems".
280:
7647:
6945:
Hughes ZE, Thacker JC, Wilson AL, Popelier PL (January 2019). "Description of Potential Energy Surfaces of Molecules Using FFLUX Machine Learning Models".
277:. The specific decomposition of the terms depends on the force field, but a general form for the total energy in an additive force field can be written as
2959:
Gross KC, Seybold PG, Hadad CM (2002). "Comparison of different atomic charge schemes for predicting pKa variations in substituted anilines and phenols".
7652:
7585:
6203:"Anisotropic, Polarizable Molecular Mechanics Studies of Inter- and Intramolecular Interactions and Ligand-Macromolecule Complexes. A Bottom-Up Strategy"
6864:
4838:
Möllhoff M, Sternberg U (May 2001). "Molecular mechanics with fluctuating atomic charges–a new force field with a semi-empirical charge calculation".
1876:
1473:
57:
7080:
Ramakrishnan R, Dral PO, Rupp M, von Lilienfeld OA (May 2015). "Big Data Meets Quantum Chemistry Approximations: The Δ-Machine Learning Approach".
1456:
were -4 to -6 kcal/mol, which is related to re-forming existing hydrogen bonds and not forming hydrogen bonds from scratch. The depths of modified
5232:
Lii JH, Allinger NL (November 1989). "Molecular mechanics. The MM3 force field for hydrocarbons. 2. Vibrational frequencies and thermodynamics".
6148:
Cioce CR, McLaughlin K, Belof JL, Space B (December 2013). "A Polarizable and Transferable PHAST N2 Potential for Use in Materials Simulation".
5578:
Yang L, Tan CH, Hsieh MJ, Wang J, Duan Y, Cieplak P, Caldwell J, Kollman PA, Luo R (July 2006). "New-generation amber united-atom force field".
5066:
Mishra RK, Mohamed AK, Geissbühler D, Manzano H, Jamil T, Shahsavari R, Kalinichev AG, Galmarini S, Tao L, Heinz H, Pellenq R (December 2017).
7578:
7560:
7541:
7522:
3898:
1950:
1164:
Atom types are defined for different elements as well as for the same elements in sufficiently different chemical environments. For example,
4391:"Improving physical realism, stereochemistry, and side-chain accuracy in homology modeling: Four approaches that performed well in CASP8"
6559:
6002:
Gao J, Habibollazadeh D, Shao L (November 1995). "A polarizable intermolecular potential function for simulation of liquid alcohols".
6551:
1565:(Optimized Potential for Liquid Simulations) (variants include OPLS-AA, OPLS-UA, OPLS-2001, OPLS-2005, OPLS3e, OPLS4) – developed by
1511:(Chemistry at HARvard Molecular Mechanics) – originally developed at Harvard, widely used for both small molecules and macromolecules
1310:
are also strongly environment-dependent because these forces originate from interactions of induced and "instantaneous" dipoles (see
7942:
4525:
3833:
Schutz CN, Warshel A (September 2001). "What are the dielectric "constants" of proteins and how to validate electrostatic models?".
2080:
1615:
CFF/ind and ENZYMIX – The first polarizable force field which has subsequently been used in many applications to biological systems.
1263:
In many cases, force fields can be straight forwardly combined. Yet, often, additional specifications and assumptions are required.
1256:
9-6 Lennard-Jones potentials to 12-6 Lennard-Jones potentials. Transfers from Buckingham potentials to harmonic potentials, or from
75:
7206:
O. T. Unke and M. Meuwly (2019). "PhysNet: A Neural Network for Predicting Energies, Forces, Dipole Moments, and Partial Charges".
4919:
Arnautova YA, Jagielska A, Scheraga HA (March 2006). "A new force field (ECEPP-05) for peptides, proteins, and organic molecules".
4254:
154:
simulations. The parameters for a chosen energy function may be derived from classical laboratory experiment data, calculations in
1251:
Functional forms and parameter sets have been defined by the developers of interatomic potentials and feature variable degrees of
5160:
Allinger NL (December 1977). "Conformational analysis. 130. MM2. A hydrocarbon force field utilizing V1 and V2 torsional terms".
2698:"An Empirical Polarizable Force Field Based on the Classical Drude Oscillator Model: Development History and Recent Applications"
2996:"Modeling the Partial Atomic Charges in Inorganometallic Molecules and Solids and Charge Redistribution in Lithium-Ion Cathodes"
3316:
Schmitt, Sebastian; Kanagalingam, Gajanan; Fleckenstein, Florian; Froescher, Daniel; Hasse, Hans; Stephan, Simon (2023-11-27).
7947:
5187:
201:
in alkane transferable force fields). A different important differentiation addresses the physical structure of the models: A
7846:
1323:
rule, which means that different types of atoms interact more weakly than identical types of atoms. This is in contrast to
1706:
1625:
NEMO (Non-Empirical Molecular Orbital) – procedure developed by Gunnar Karlström and coworkers at Lund University (Sweden)
1523:
4754:"Calorimetric determination of the enthalpy change for the alpha-helix to coil transition of an alanine peptide in water"
3504:"Density-functional theory with screened van der Waals interactions for the modeling of hybrid inorganic-organic systems"
494:
As it is rare for bonds to deviate significantly from their equilibrium values, the most simplistic approaches utilize a
7957:
7615:
6618:
Hoogerbrugge PJ, Koelman JM (1992). "Simulating Microscopic Hydrodynamic Phenomena with Dissipative Particle Dynamics".
92:
850:
simulations. The stronger the bond is between atoms, the higher is the value of the force constant, and the higher the
48:
6773:
1605:
1544:
261:
for modeling molecular systems includes intramolecular interaction terms for interactions of atoms that are linked by
7437:
Deeth RJ (2001). "The ligand field molecular mechanics model and the stereoelectronic effects of d and s electrons".
6364:
Borodin O (August 2009). "Polarizable force field development and molecular dynamics simulations of ionic liquids".
1666:
of using quantum mechanical methods. Primarily used by software packages supplied by Cresset Biomolecular Discovery.
1654:
Polarizable procedure based on the Kim-Gordon approach developed by Jürg Hutter and coworkers (University of Zürich)
7952:
7723:
7639:
6547:
1691:
1203:. The bonded terms refer to pairs, triplets, and quadruplets of bonded atoms, and include values for the effective
2318:
Sun H, Mumby SJ, Maple JR, Hagler AT (April 1994). "An ab Initio CFF93 All-Atom Force Field for Polycarbonates".
2031:
Siu SW, Pluhackova K, Böckmann RA (April 2012). "Optimization of the OPLS-AA Force Field for Long Hydrocarbons".
1891:
1830:
1477:
1280:
1824:
1788:
PhysNet is a Neural Network-based energy function to predict energies, forces and (fluctuating) partial charges.
7899:
7882:
1803:
The set of parameters used to model water or aqueous solutions (basically a force field for water) is called a
1785:
utilising continuous-filter convolutional layers, to predict chemical properties and potential energy surfaces.
1179:
are classified as different force field types. Typical molecular force field parameter sets include values for
1122:
5105:"Insight into induced charges at metal surfaces and biointerfaces using a polarizable Lennard-Jones potential"
3269:
Eggimann, Becky L.; Sunnarborg, Amara J.; Stern, Hudson D.; Bliss, Andrew P.; Siepmann, J. Ilja (2014-01-02).
1460:
derived from protein engineering data were also smaller than in typical potential parameters and followed the
5205:
Allinger NL, Yuh YH, Lii JH (November 1989). "Molecular mechanics. The MM3 force field for hydrocarbons. 1".
3408:"Force Field and a Surface Model Database for Silica to Simulate Interfacial Properties in Atomic Resolution"
1977:
Abascal JL, Vega C (December 2005). "A general purpose model for the condensed phases of water: TIP4P/2005".
7746:
7662:
7314:"Ab Initio Charge and AMBER Forcefield Parameters for Frequently Occurring Post-Translational Modifications"
1732:
1725:
1457:
1292:
1260:
to harmonic potentials, on the contrary, would require many additional assumptions and may not be possible.
1188:
1134:
1130:
471:
127:
1735:
fitted to liquid phase densities and vapor pressures of pure compounds by using the SAFT equation of state.
7793:
7259:
Molinero V, Moore EB (April 2009). "Water modeled as an intermediate element between carbon and silicon".
7142:
7000:"Multipolar Electrostatic Energy Prediction for all 20 Natural Amino Acids Using Kriging Machine Learning"
6576:
5067:
1743:
capture features dependent on secondary structure and on residue-specific contact information in proteins.
1677:
1417:
330:
where the components of the covalent and noncovalent contributions are given by the following summations:
135:
7464:
Foscato M, Deeth RJ, Jensen VR (June 2015). "Integration of Ligand Field Molecular Mechanics in Tinker".
6669:
Koelman JM, Hoogerbrugge PJ (1993). "Dynamic Simulations of Hard-Sphere Suspensions Under Steady Shear".
5068:"A force field database for cementitious materials including validations, applications and opportunities"
4598:
Murphy KP, Gill SJ (December 1991). "Solid model compounds and the thermodynamics of protein unfolding".
7678:
7601:
6495:"Extended electron distributions applied to the molecular mechanics of some intermolecular interactions"
1911:
1906:
1446:
1393:
1361:
1272:
805:
159:
131:
3798:
Warshel A, Sharma PK, Kato M, Parson WW (November 2006). "Modeling electrostatic effects in proteins".
2610:"Embedded-atom method: Derivation and application to impurities, surfaces, and other defects in metals"
1235:
focuses on interatomic functions describing the individual interactions between specific elements. The
3561:
Kramer C, Spinn A, Liedl KR (October 2014). "Charge Anisotropy: Where Atomic Multipoles Matter Most".
7821:
7164:
7099:
6817:
6729:
6678:
6627:
6568:
6320:
6263:
6122:
6031:"Development of a polarizable intermolecular potential function (PIPF) for liquid amides and alkanes"
5498:
5447:
5404:
5395:
Dick BG, Overhauser AW (1958-10-01). "Theory of the Dielectric Constants of Alkali Halide Crystals".
5116:
4765:
4708:
4355:
3661:
3605:
3515:
3458:
3233:
3047:
2754:
2621:
2566:
2517:
2279:
2102:
1986:
1916:
1858:
1768:
or Quantum chemical topology energy terms including electrostatic, exchange and electron correlation.
1566:
1311:
1101:
6581:
4311:
Lazaridis T, Karplus M (April 2000). "Effective energy functions for protein structure prediction".
3270:
43:
7887:
7866:
6898:"ANI-1: an extensible neural network potential with DFT accuracy at force field computational cost"
6865:"MACE: Higher order equivariant message passing neural networks for fast and accurate force fields"
5815:"Simulating Monovalent and Divalent Ions in Aqueous Solution Using a Drude Polarizable Force Field"
3748:"Interatomic potentials and solvation parameters from protein engineering data for buried residues"
1921:
1896:
1886:
1862:
1854:
1775:
provides a short-range potential, whilst more traditional potentials add screened long-range terms.
1590:
1434:
1341:
1307:
1300:
1257:
1105:
274:
119:
1006:{\displaystyle E_{\text{Coulomb}}={\frac {1}{4\pi \varepsilon _{0}}}{\frac {q_{i}q_{j}}{r_{ij}}},}
7914:
7826:
7803:
7784:
7764:
7728:
7294:
7268:
7241:
7215:
7188:
7154:
7123:
7089:
6980:
6876:
6753:
6702:
6651:
6475:
5949:
5914:
5813:
Yu H, Whitfield TW, Harder E, Lamoureux G, Vorobyov I, Anisimov VM, Mackerell AD, Roux B (2010).
5760:
5560:
5471:
4971:
4855:
4658:
4371:
4223:
4174:
4125:
4047:
4004:
3960:
3908:
3858:
3685:
3651:
3353:
3298:
3202:
3107:
2871:
2433:
2248:
2193:
2162:
2140:
2114:
2010:
1881:
1765:
1710:
1639:
SP-basis Chemical Potential Equalization (CPE) – approach developed by R. Chelli and P. Procacci.
1239:
focuses on transferable force fields of organic molecules (developed by the Siepmann group). The
1196:
1083:. The total Coulomb energy is a sum over all pairwise combinations of atoms and usually excludes
847:
809:
151:
147:
115:
6532:
3038:
Heinz H, Suter UW (November 2004). "Atomic Charges for Classical Simulations of Polar Systems".
5489:
Yu H, van Gunsteren WF (November 2005). "Accounting for polarization in molecular simulation".
4276:
Edgcomb SP, Murphy KP (February 2000). "Structural energetics of protein folding and binding".
1731:
SAFT – A top-down coarse-grained model developed in the Molecular Systems Engineering group at
7919:
7816:
7789:
7700:
7690:
7683:
7657:
7556:
7537:
7518:
7491:
7392:
7343:
7286:
7233:
7180:
7115:
7062:
7021:
6972:
6927:
6845:
6745:
6694:
6643:
6514:
6467:
6432:
6381:
6346:
6289:
6232:
6165:
6095:
6060:
5984:
5879:
5844:
5795:
5752:
5711:
5662:
5595:
5552:
5514:
5463:
5420:
5377:
5342:
5142:
5027:
4936:
4793:
4734:
4650:
4615:
4580:
4531:
4521:
4508:
Rohl CA, Strauss CE, Misura KM, Baker D (2004). "Protein Structure Prediction Using Rosetta".
4490:
4455:
4420:
4328:
4293:
4258:
4215:
4166:
4117:
4082:
4065:
Brunger AT, Adams PD (June 2002). "Molecular dynamics applied to X-ray structure refinement".
4039:
3996:
3952:
3894:
3850:
3815:
3777:
3723:
3677:
3621:
3578:
3543:
3484:
3430:
3388:
3345:
3337:
3290:
3251:
3194:
3156:
3099:
3063:
3017:
2976:
2941:
2906:
2863:
2822:
2727:
2678:
2637:
2590:
2582:
2535:
2482:
2425:
2384:
2335:
2297:
2240:
2232:
2185:
2132:
2076:
2048:
2002:
1956:
1946:
1845:
1370:
905:
813:
738:
662:
460:
163:
155:
111:
1360:". Force fields have been applied successfully for protein structure refinement in different
241:, and multi-component complexes, sacrifice chemical details for higher computing efficiency.
7861:
7811:
7481:
7473:
7446:
7419:
7382:
7374:
7333:
7325:
7278:
7225:
7172:
7107:
7052:
7011:
6962:
6954:
6917:
6909:
6835:
6825:
6737:
6686:
6635:
6586:
6506:
6459:
6422:
6412:
6373:
6336:
6328:
6279:
6271:
6222:
6214:
6157:
6130:
6087:
6050:
6042:
6011:
5976:
5941:
5906:
5871:
5834:
5826:
5787:
5742:
5701:
5693:
5652:
5644:
5587:
5544:
5506:
5455:
5412:
5369:
5334:
5295:
5268:
5241:
5214:
5169:
5132:
5124:
5082:
5019:
4963:
4928:
4901:
4847:
4820:
4783:
4773:
4724:
4716:
4642:
4607:
4570:
4562:
4513:
4482:
4447:
4410:
4402:
4363:
4320:
4285:
4250:
4205:
4156:
4109:
4074:
4031:
3988:
3944:
3886:
3842:
3807:
3767:
3759:
3715:
3669:
3613:
3570:
3533:
3523:
3474:
3466:
3422:
3380:
3329:
3282:
3241:
3186:
3146:
3138:
3091:
3055:
3007:
2968:
2933:
2898:
2855:
2814:
2785:
2717:
2709:
2668:
2629:
2574:
2525:
2472:
2464:
2415:
2374:
2366:
2355:"CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data"
2327:
2287:
2224:
2177:
2124:
2040:
1994:
1365:
1337:
1176:
1158:
1112:
potentials have been developed, which describe a form of attachment of electrons to nuclei.
1016:
913:
819:
777:
632:
602:
479:
258:
222:
174:
143:
4752:
Scholtz JM, Marqusee S, Baldwin RL, York EJ, Stewart JM, Santoro M, Bolen DW (April 1991).
4145:"163 Enhanced sampling of peptides and proteins with a new biasing replica exchange method"
876:
7856:
7851:
7779:
7774:
7741:
7668:
7536:. Interdisciplinary Applied Mathematics: Mathematical Biology. New York: Springer-Verlag.
6777:
6594:
4873:
4100:
Güntert P (May 1998). "Structure calculation of biological macromolecules from NMR data".
3921:
3538:
3479:
2213:"A child of prediction. On the History, Ontology, and Computation of the Lennard-Jonesium"
1551:
1405:
1397:
1378:
1252:
1204:
859:
452:
5633:"Implementation of Geometry-Dependent Charge Flux into the Polarizable AMOEBA+ Potential"
4022:
Koehl P, Levitt M (February 1999). "A brighter future for protein structure prediction".
482:. However, both can be buffered or scaled by a constant factor to account for electronic
7168:
7103:
6821:
6733:
6682:
6631:
6572:
6324:
6267:
6126:
5502:
5451:
5408:
5120:
4769:
4712:
4359:
3665:
3609:
3519:
3462:
3407:
3406:
Emami FS, Puddu V, Berry RJ, Varshney V, Patwardhan SV, Perry CC, Heinz H (2014-04-22).
3237:
3220:
Tadmor, E. B.; Elliott, R. S.; Sethna, J. P.; Miller, R. E.; Becker, C. A. (July 2011).
3051:
2758:
2625:
2570:
2521:
2283:
1990:
1373:
of proteins. Meanwhile, alternative empirical scoring functions have been developed for
1243:
focuses on molecular and ionic force fields (both component-specific and transferable).
7904:
7620:
7387:
7362:
7361:
Khoury GA, Smadbeck J, Tamamis P, Vandris AC, Kieslich CA, Floudas CA (December 2014).
7338:
7313:
6922:
6897:
6840:
6805:
6494:
6427:
6400:
6341:
6308:
6284:
6251:
6227:
6202:
6055:
6030:
5839:
5814:
5706:
5681:
5657:
5632:
5137:
5104:
4729:
4696:
4575:
4550:
4415:
4390:
3890:
3772:
3747:
3151:
3126:
2722:
2697:
2379:
2354:
1782:
1772:
1609:
1586:
1425:
1413:
1401:
1382:
1374:
1327:
or Slater-Kirkwood equation applied for development of the classical force fields. The
1200:
1184:
1066:
1046:
871:
718:
698:
495:
483:
98:
7450:
7423:
6720:
Español P, Warren P (1995). "Statistical Mechanics of Dissipative Particle Dynamics".
5438:
Mitchell PJ, Fincham D (1993-02-22). "Shell model simulations by adiabatic dynamics".
4720:
4517:
4486:
4451:
4438:
Gordon DB, Marshall SA, Mayo SL (August 1999). "Energy functions for protein design".
4324:
4289:
3885:
Israelachvili JN (2011). "Intermolecular and Surface Forces". Elsevier. pp. iii.
2103:"MolMod – an open access database of force fields for molecular simulations of fluids"
1645:
ORIENT – procedure developed by Anthony J. Stone (Cambridge University) and coworkers.
441:{\displaystyle E_{\text{nonbonded}}=E_{\text{electrostatic}}+E_{\text{van der Waals}}}
389:{\displaystyle E_{\text{bonded}}=E_{\text{bond}}+E_{\text{angle}}+E_{\text{dihedral}}}
7936:
7892:
6771:
Dissipative Particle Dynamics: Addressing deficiencies and establishing new frontiers
6741:
6706:
6690:
6655:
6639:
6479:
5875:
5475:
4788:
4753:
4611:
4473:
Mendes J, Guerois R, Serrano L (August 2002). "Energy estimation in protein design".
4008:
3357:
3206:
2673:
2656:
2252:
2144:
1812:
1502:
1142:
909:
901:
475:
464:
270:
262:
230:
17:
7298:
7245:
7127:
6984:
6757:
5953:
5764:
5564:
5459:
5086:
4975:
4859:
4662:
4375:
4227:
4178:
4129:
3317:
3302:
3222:"The potential of atomistic simulations and the knowledgebase of interatomic models"
3111:
2437:
2197:
1346:
the interaction between hydrocarbons across water is about 10% of that across vacuum
7909:
7192:
6790:
5918:
4051:
3964:
3862:
3689:
3528:
3503:
2875:
2014:
1536:
and energy derivatives, and quantifies limitations for all included compounds. The
1498:(Assisted Model Building and Energy Refinement) – widely used for proteins and DNA.
1315:
238:
218:
198:
7145:(June 2018). "SchNet - A deep learning architecture for molecules and materials".
7041:"Machine Learning of Dynamic Electron Correlation Energies from Topological Atoms"
5191:
3095:
2128:
6401:"Robust Atomistic Modeling of Materials, Organometallic, and Biochemical Systems"
4161:
4144:
3811:
3286:
2212:
7718:
7713:
5648:
2713:
2292:
2267:
2228:
1808:
1804:
1798:
1555:
1442:
1438:
1192:
1180:
1109:
771:
is at times differently defined or taken at different thermodynamic conditions.
6810:
Proceedings of the National Academy of Sciences of the United States of America
5128:
4758:
Proceedings of the National Academy of Sciences of the United States of America
3470:
1636:
Polarizable Force Field (PFF) – developed by Richard A. Friesner and coworkers.
1232:
1161:, which are more accessible for experimental studies and quantum calculations.
870:
Electrostatic interactions are represented by a Coulomb energy, which utilizes
7708:
6183:
5945:
5510:
5045:
4113:
3948:
3246:
3221:
2609:
2554:
1537:
1287:
851:
205:
force fields provide parameters for every type of atom in a system, including
7477:
7312:
Khoury GA, Thompson JP, Smadbeck J, Kieslich CA, Floudas CA (December 2013).
7229:
7111:
7057:
7040:
7016:
6999:
6958:
6749:
6698:
6647:
6518:
6463:
5697:
5518:
5467:
5424:
5381:
5346:
4170:
4000:
3625:
3488:
3434:
3341:
3333:
3294:
3255:
3190:
3103:
3067:
2980:
2826:
2790:
2682:
2641:
2633:
2586:
2578:
2539:
2339:
2301:
2236:
2136:
2101:
Stephan, Simon; Horsch, Martin T.; Vrabec, Jadran; Hasse, Hans (2019-07-03).
1960:
1807:. Many water models have been proposed; some examples are TIP3P, TIP4P, SPC,
7769:
6830:
3935:
Leckband D, Israelachvili J (May 2001). "Intermolecular forces in biology".
3719:
3127:"Building Force Fields: An Automatic, Systematic, and Reproducible Approach"
2163:"The MARTINI force field: coarse grained model for biomolecular simulations"
1764:
models which operate together to provide a molecular force field trained on
1421:
139:
107:
7495:
7396:
7347:
7290:
7237:
7184:
7119:
7066:
7025:
6976:
6931:
6849:
6770:
6471:
6436:
6417:
6385:
6350:
6293:
6236:
6169:
6099:
6064:
5988:
5848:
5799:
5756:
5715:
5666:
5599:
5556:
5416:
5299:
5146:
5031:
4940:
4778:
4584:
4535:
4494:
4459:
4424:
4332:
4297:
4262:
4219:
4086:
4043:
3992:
3956:
3854:
3819:
3781:
3727:
3681:
3582:
3547:
3392:
3349:
3198:
3160:
3021:
2945:
2910:
2859:
2731:
2486:
2429:
2388:
2244:
2189:
2161:
Marrink SJ, Risselada HJ, Yefimov S, Tieleman DP, de Vries AH (July 2007).
2052:
2006:
1690:– reactive force field (interatomic potential) developed by Adri van Duin,
5910:
5103:
Geada IL, Ramezani-Dakhel H, Jamil T, Sulpizi M, Heinz H (February 2018).
4967:
4851:
4797:
4738:
4654:
4646:
4619:
4255:
10.1002/1521-3773(20020802)41:15<2644::AID-ANIE2644>3.0.CO;2-O
4121:
2867:
2594:
1275:
are based on approximations and experimental data, therefore often termed
7673:
5883:
4566:
4389:
Krieger E, Joo K, Lee J, Lee J, Raman S, Thompson J, et al. (2009).
4367:
2555:"New empirical approach for the structure and energy of covalent systems"
1943:
Understanding molecular simulation : from algorithms to applications
1449:
1389:
1173:
1126:
592:{\displaystyle E_{\text{bond}}={\frac {k_{ij}}{2}}(l_{ij}-l_{0,ij})^{2},}
456:
206:
167:
7570:
6250:
Piquemal JP, Cisneros GA, Reinhardt P, Gresh N, Darden TA (March 2006).
6015:
5932:
Swart M, van Duijnen PT (May 2006). "DRF90: a polarizable force field".
5338:
5272:
5245:
5218:
5173:
4905:
4824:
2331:
1865:. It can be incorporated into other force fields such as CHARMM and UFF.
1695:
reactive simulations on >>1,000,000 atoms on large supercomputers.
1597:
AMBER – polarizable force field developed by Jim Caldwell and coworkers.
7838:
6967:
6913:
6510:
4406:
4210:
4193:
3763:
3384:
2468:
1850:
1761:
1724:– a coarse-grained potential developed by Marrink and coworkers at the
1721:
1453:
1146:
323:{\displaystyle E_{\text{total}}=E_{\text{bonded}}+E_{\text{nonbonded}}}
234:
186:
182:
7486:
7378:
7329:
7282:
7176:
6590:
6377:
6332:
6275:
6218:
6161:
6134:
6091:
6046:
5980:
5830:
5791:
5747:
5730:
5591:
5548:
5373:
5023:
4989:
4932:
4078:
3846:
3574:
3426:
3271:"An online parameter and property database for the TraPPE force field"
3142:
3059:
3012:
2995:
2972:
2937:
2902:
2818:
2530:
2501:
2477:
2420:
2403:
2370:
2181:
2044:
1998:
1104:
are usually used, e.g. Tersoff potentials. For metal systems, usually
3673:
3617:
2745:
Lorentz HA (1905). "The Motion of Electrons in Metallic Bodies, I.".
2655:
Daw, Murray S.; Foiles, Stephen M.; Baskes, Michael I. (March 1993).
1687:
1681:
1642:
PHAST – polarizable potential developed by Chris Cioce and coworkers.
1529:
1508:
1430:
1333:
1165:
253:
Molecular mechanics potential energy function with continuum solvent.
214:
146:
of a system on the atomistic level. Force fields are usually used in
2773:
1240:
173:
A large number of different force field types exist today (e.g. for
7220:
7159:
7094:
6881:
5613:
3656:
2119:
7273:
1714:
1495:
1169:
1085:
1, 2 bonded atoms, 1, 3 bonded atoms, as well as 1, 4 bonded atoms
249:
248:
190:
170:
of the potential energy with respect to the particle coordinates.
97:
5682:"AMOEBA+ Classical Potential for Modeling Molecular Interactions"
4035:
3502:
Ruiz VG, Liu W, Zojer E, Scheffler M, Tkatchenko A (April 2012).
7553:
Computer Modeling of Chemical Reactions in Enzymes and Solutions
6806:"A force field for virtual atom molecular mechanics of proteins"
4551:"Positioning of proteins in membranes: a computational approach"
3172:
3170:
2211:
Lenhard, Johannes; Stephan, Simon; Hasse, Hans (February 2024).
1562:
1518:
CVFF – also used broadly for small molecules and macromolecules.
7574:
7039:
McDonagh JL, Silva AF, Vincent MA, Popelier PL (January 2018).
4346:
Javidpour L (2012). "Computer Simulations of Protein Folding".
3318:"Extension of the MolMod Database to Transferable Force Fields"
2657:"The embedded-atom method: a review of theory and applications"
2500:
Ashcroft, N. W.; Mermin, N. D.; Smoluchowski, R. (1977-01-01).
2402:
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (July 2004).
735:
when all other terms in the force field are set to 0. The term
6201:
Gresh N, Cisneros GA, Darden TA, Piquemal JP (November 2007).
1396:, i.e., energy of evaporation of molecular crystals. However,
1154:
1150:
178:
26:
7534:
Molecular Modeling and Simulation: An Interdisciplinary Guide
3800:
Biochimica et Biophysica Acta (BBA) - Proteins and Proteomics
1236:
229:
potentials, which are often used in long-time simulations of
4549:
Lomize AL, Pogozheva ID, Lomize MA, Mosberg HI (June 2006).
1486:
Different force fields are designed for different purposes:
1157:
were often derived/ transferred from observations for small
2268:"On the history of key empirical intermolecular potentials"
1408:, or liquid-solid transitions as these processes represent
6780:, CECAM workshop, July 16–18, 2008, Lausanne, Switzerland.
1572:
QCFF/PI – A general force fields for conjugated molecules.
1416:(energy absorbed during melting of molecular crystals), a
1291:π-conjugated systems, and lone pairs in water. Electronic
102:
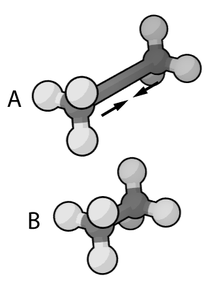
Part of force field of ethane for the C-C stretching bond.
6863:
Batatia I, Kovacs DP, Simm G, Ortner C, Csanyi G (2022).
2840:
2838:
2836:
1214:
Efforts to provide open source codes and methods include
4512:. Methods in Enzymology. Vol. 383. pp. 66–93.
2404:"Development and testing of a general amber force field"
1902:
Comparison of software for molecular mechanics modeling
1482:
Comparison of software for molecular mechanics modeling
1630:
1622:
DRF90 – developed by P. Th. van Duijnen and coworkers.
1547:) – developed at Merck for a broad range of molecules.
1445:
transition data, but the same energies estimated from
1191:
for every atom type, as well as equilibrium values of
6307:
Cisneros GA, Piquemal JP, Darden TA (November 2006).
3880:
3878:
3876:
3874:
3872:
2696:
Lemkul JA, Huang J, Roux B, MacKerell AD (May 2016).
1576:
compounds, especially metals and inorganic compounds.
1069:
1049:
1019:
922:
879:
822:
780:
741:
721:
701:
665:
635:
605:
504:
401:
336:
283:
7141:
Schütt KT, Sauceda HE, Kindermans PJ, Tkatchenko A,
3746:
Lomize AL, Reibarkh MY, Pogozheva ID (August 2002).
1433:
in proteins are ~ -1.5 kcal/mol when estimated from
1356:(CASP) did not try to refine their models to avoid "
1219:
1215:
158:, or both. Force fields utilize the same concept as
7875:
7837:
7802:
7757:
7699:
7638:
7608:
5729:Patel S, Mackerell AD, Brooks CL (September 2004).
4194:"Assessment of homology-based predictions in CASP5"
1354:
Critical Assessment of protein Structure Prediction
1295:of the environment may be better included by using
1279:. The performance varies from higher accuracy than
2266:Fischer, Johann; Wendland, Martin (October 2023).
1075:
1055:
1035:
1005:
892:
838:
796:
763:
727:
707:
687:
651:
621:
591:
440:
388:
322:
6869:Advances in Neural Information Processing Systems
6552:"ReaxFF: A Reactive Force Field for Hydrocarbons"
2096:
2094:
2092:
6252:"Towards a force field based on density fitting"
3701:
3699:
2073:Molecular Modelling: Principles and Applications
1853:- a function for angle bending that is based on
134:. More precisely, the force field refers to the
4697:"Hydrogen bonding stabilizes globular proteins"
1569:at the Yale University Department of Chemistry.
695:is the value for the bond length between atoms
265:and intermolecular (i.e. nonbonded also termed
6896:Smith JS, Isayev O, Roitberg AE (April 2017).
5061:
5059:
4149:Journal of Biomolecular Structure and Dynamics
3741:
3739:
3737:
2449:
2447:
2156:
2154:
213:interatomic potentials treat the hydrogen and
7586:
5098:
5096:
5005:
5003:
3033:
3031:
8:
7466:Journal of Chemical Information and Modeling
3322:Journal of Chemical Information and Modeling
3179:Journal of Chemical Information and Modeling
3125:Wang LP, Martinez TJ, Pande VS (June 2014).
2608:Daw, Murray S.; Baskes, M. I. (1984-06-15).
2217:Studies in History and Philosophy of Science
2066:
2064:
2062:
6804:Korkut A, Hendrickson WA (September 2009).
2994:Wang B, Li SL, Truhlar DG (December 2014).
2774:"Zur Elekronentheorie der Metalle. I. Teil"
1760:FFLUX (originally QCTFF) A set of trained
1381:, homology model refinement, computational
900:to represent chemical bonding ranging from
7593:
7579:
7571:
7318:Journal of Chemical Theory and Computation
7082:Journal of Chemical Theory and Computation
7045:Journal of Chemical Theory and Computation
7004:Journal of Chemical Theory and Computation
6947:Journal of Chemical Theory and Computation
6499:Journal of Computer-Aided Molecular Design
6207:Journal of Chemical Theory and Computation
6150:Journal of Chemical Theory and Computation
6080:Journal of Chemical Theory and Computation
6035:Journal of Chemical Theory and Computation
5819:Journal of Chemical Theory and Computation
5780:Journal of Chemical Theory and Computation
5686:Journal of Chemical Theory and Computation
5631:Liu C, Piquemal JP, Ren P (January 2020).
3793:
3791:
3637:
3635:
3563:Journal of Chemical Theory and Computation
3000:Journal of Chemical Theory and Computation
2961:International Journal of Quantum Chemistry
2926:Journal of Chemical Theory and Computation
2891:Journal of Chemical Theory and Computation
2313:
2311:
2033:Journal of Chemical Theory and Computation
7485:
7386:
7337:
7272:
7219:
7158:
7093:
7056:
7015:
6966:
6921:
6880:
6839:
6829:
6580:
6426:
6416:
6340:
6283:
6226:
6054:
6029:Xie W, Pu J, Mackerell AD, Gao J (2007).
5838:
5746:
5705:
5656:
5637:The Journal of Physical Chemistry Letters
5136:
4787:
4777:
4728:
4574:
4414:
4209:
4160:
4143:Ostermeir K, Zacharias M (January 2013).
3771:
3655:
3537:
3527:
3478:
3245:
3150:
3131:The Journal of Physical Chemistry Letters
3011:
2789:
2721:
2672:
2529:
2476:
2419:
2378:
2291:
2118:
1877:Comparison of force field implementations
1857:and works for large angular distortions,
1474:Comparison of force field implementations
1385:, and modeling of proteins in membranes.
1247:Transferability and mixing function types
1068:
1048:
1024:
1018:
989:
978:
968:
961:
952:
936:
927:
921:
884:
878:
827:
821:
785:
779:
746:
740:
720:
700:
670:
664:
640:
634:
610:
604:
580:
561:
545:
524:
518:
509:
503:
432:
419:
406:
400:
380:
367:
354:
341:
335:
314:
301:
288:
282:
76:Learn how and when to remove this message
5530:
5528:
5327:Journal of the American Chemical Society
5261:Journal of the American Chemical Society
5234:Journal of the American Chemical Society
5207:Journal of the American Chemical Society
5162:Journal of the American Chemical Society
2807:Journal of the American Chemical Society
2353:Huang J, MacKerell AD (September 2013).
2320:Journal of the American Chemical Society
1809:flexible simple point charge water model
1608:(University of Maryland, Baltimore) and
1585:Several force fields explicitly capture
1464:rule, as predicted by McLachlan theory.
1141:for molecular simulations of biological
804:can be determined from the experimental
6405:Angewandte Chemie International Edition
5680:Liu C, Piquemal JP, Ren P (July 2019).
2075:(2nd ed.). Harlow: Prentice Hall.
1933:
6998:Fletcher TL, Popelier PL (June 2016).
6399:Spicher S, Grimme S (September 2020).
4348:Computing in Science & Engineering
3917:
3906:
1554:mainly for conformational analysis of
846:determines vibrational frequencies in
4475:Current Opinion in Structural Biology
4440:Current Opinion in Structural Biology
4313:Current Opinion in Structural Biology
1108:are used. For metals, also so-called
269:) terms that describe the long-range
7:
7412:Computers & Chemical Engineering
5440:Journal of Physics: Condensed Matter
2026:
2024:
1972:
1970:
1468:Force fields available in literature
7555:. New York: John Wiley & Sons.
7261:The Journal of Physical Chemistry B
6560:The Journal of Physical Chemistry A
6546:van Duin AC, Dasgupta S, Lorant F,
6452:The Journal of Physical Chemistry B
6366:The Journal of Physical Chemistry B
5614:"Tinker Molecular Modeling Package"
5580:The Journal of Physical Chemistry B
4921:The Journal of Physical Chemistry B
3040:The Journal of Physical Chemistry B
2170:The Journal of Physical Chemistry B
854:(energy) in the IR/Raman spectrum.
6184:"Anthony Stone: Computer programs"
5899:Journal of Computational Chemistry
5735:Journal of Computational Chemistry
5537:Journal of Computational Chemistry
4695:Myers JK, Pace CN (October 1996).
4510:Numerical Computer Methods, Part D
3891:10.1016/b978-0-12-391927-4.10024-6
2408:Journal of Computational Chemistry
2359:Journal of Computational Chemistry
1043:is the distance between two atoms
245:Force fields for molecular systems
25:
7515:Intermolecular and surface forces
7424:10.1016/j.compchemeng.2014.07.017
6004:The Journal of Physical Chemistry
4894:The Journal of Physical Chemistry
4813:The Journal of Physical Chemistry
7629:
4278:Current Opinion in Biotechnology
1404:are thermodynamically closer to
1095:Force fields for crystal systems
478:and the electrostatic term with
31:
7147:The Journal of Chemical Physics
6313:The Journal of Chemical Physics
6256:The Journal of Chemical Physics
6115:The Journal of Chemical Physics
5491:Computer Physics Communications
5087:10.1016/j.cemconres.2017.09.003
4102:Quarterly Reviews of Biophysics
3937:Quarterly Reviews of Biophysics
3598:The Journal of Chemical Physics
1979:The Journal of Chemical Physics
1633:at the University of Minnesota.
7439:Coordination Chemistry Reviews
4192:Tramontano A, Morea V (2003).
3539:11858/00-001M-0000-000F-C6EA-3
3529:10.1103/physrevlett.108.146103
3480:11858/00-001M-0000-0029-3035-8
1319:than in vacuum and follow the
577:
538:
1:
7517:. San Diego: Academic Press.
7451:10.1016/S0010-8545(00)00354-4
5046:"Interface Force Field (IFF)"
4721:10.1016/S0006-3495(96)79401-8
4518:10.1016/S0076-6879(04)83004-0
4487:10.1016/s0959-440x(02)00345-7
4452:10.1016/S0959-440X(99)80072-4
4325:10.1016/s0959-440x(00)00063-4
4290:10.1016/s0958-1669(99)00055-5
4067:Accounts of Chemical Research
3981:Advanced Functional Materials
3096:10.1080/08927022.2017.1332414
2129:10.1080/08927022.2019.1601191
1707:Dissipative particle dynamics
1619:each molecular dynamics step.
1524:Internal Coordinate Mechanics
912:. The typical formula is the
774:The bond stretching constant
257:The basic functional form of
88:Concept on molecular modeling
7616:Principle of maximum entropy
6493:Vinter, J. G. (1994-12-01).
5876:10.1016/0022-2836(76)90311-9
5864:Journal of Molecular Biology
5075:Cement and Concrete Research
4612:10.1016/0022-2836(91)90506-2
4600:Journal of Molecular Biology
4162:10.1080/07391102.2013.786405
3812:10.1016/j.bbapap.2006.08.007
3287:10.1080/08927022.2013.842994
2674:10.1016/0920-2307(93)90001-u
1771:TensorMol, a mixed model, a
5649:10.1021/acs.jpclett.9b03489
5288:Israel Journal of Chemistry
2714:10.1021/acs.chemrev.5b00505
2293:10.1016/j.fluid.2023.113876
2229:10.1016/j.shpsa.2023.11.007
1545:Merck Molecular Force Field
816:calculations. The constant
225:as one interaction center.
142:sets used to calculate the
51:. The specific problem is:
7974:
7640:Statistical thermodynamics
6742:10.1209/0295-5075/30/4/001
6691:10.1209/0295-5075/21/3/018
6640:10.1209/0295-5075/19/3/001
5129:10.1038/s41467-018-03137-8
3471:10.1103/physrevb.93.035118
2553:Tersoff, J. (1988-04-15).
1863:transition metal complexes
1796:
1471:
866:Electrostatic interactions
90:
47:to meet Knowledge (XXG)'s
7627:
7513:Israelachvili JN (1992).
6722:Europhysics Letters (EPL)
6671:Europhysics Letters (EPL)
6620:Europhysics Letters (EPL)
5946:10.1080/08927020600631270
5511:10.1016/j.cpc.2005.01.022
5460:10.1088/0953-8984/5/8/006
4840:Molecular Modeling Annual
4114:10.1017/S0033583598003436
4024:Nature Structural Biology
3949:10.1017/S0033583501003687
3247:10.1007/s11837-011-0102-6
2661:Materials Science Reports
1892:Molecular design software
1478:Molecular design software
1340:) have been explained by
1281:density functional theory
1172:and an oxygen atoms in a
7943:Force fields (chemistry)
7900:Condensed matter physics
7883:Statistical field theory
7478:10.1021/acs.jcim.5b00098
7230:10.1021/acs.jctc.9b00181
7112:10.1021/acs.jctc.5b00099
7058:10.1021/acs.jctc.7b01157
7017:10.1021/acs.jctc.6b00457
6959:10.1021/acs.jctc.8b00806
6464:10.1021/acs.jpcb.1c05744
5698:10.1021/acs.jctc.9b00261
3373:Chemical Society Reviews
3334:10.1021/acs.jcim.3c01484
3191:10.1021/acs.jcim.9b00646
2791:10.1002/andp.19003060312
2634:10.1103/physrevb.29.6443
2579:10.1103/physrevb.37.6991
1612:(University of Chicago).
1458:Lennard-Jones potentials
1297:polarizable force fields
1189:Lennard-Jones parameters
1123:enthalpy of vaporization
1106:embedded atom potentials
812:spectrum, or high-level
764:{\displaystyle l_{0,ij}}
688:{\displaystyle l_{0,ij}}
659:is the bond length, and
7758:Mathematical approaches
7747:Lennard-Jones potential
7663:thermodynamic potential
6831:10.1073/pnas.0907674106
5188:"MM2 and MM3 home page"
3720:10.1021/acsnano.7b07684
3508:Physical Review Letters
2747:Proc. K. Ned. Akad. Wet
1733:Imperial College London
1726:University of Groningen
1299:or using a macroscopic
629:is the force constant,
472:Lennard-Jones potential
7794:conformal field theory
6418:10.1002/anie.202004239
6188:www-stone.ch.cam.ac.uk
5417:10.1103/physrev.112.90
5362:Chemistry of Materials
5300:10.1002/ijch.197300067
4779:10.1073/pnas.88.7.2854
4678:Gazetta Chim. Italiana
3993:10.1002/adfm.201905247
3415:Chemistry of Materials
2860:10.1002/prot.340040106
2272:Fluid Phase Equilibria
2071:Leach A (2001-01-30).
1678:Empirical valence bond
1418:conformational entropy
1273:interatomic potentials
1077:
1057:
1037:
1036:{\displaystyle r_{ij}}
1007:
894:
840:
839:{\displaystyle k_{ij}}
798:
797:{\displaystyle k_{ij}}
765:
729:
709:
689:
653:
652:{\displaystyle l_{ij}}
623:
622:{\displaystyle k_{ij}}
593:
442:
390:
324:
254:
132:interatomic potentials
103:
7948:Intermolecular forces
7709:Ferromagnetism models
7602:Statistical mechanics
7367:ACS Synthetic Biology
5911:10.1002/jcc.540150505
5109:Nature Communications
4968:10.1002/bip.360290403
4852:10.1007/s008940100008
4647:10.1002/bip.360281003
1912:Interatomic potential
1907:Statistical potential
1859:hypervalent molecules
1550:MM2 was developed by
1538:Interface force field
1472:Further information:
1362:X-ray crystallography
1226:Force field databases
1102:bond order potentials
1078:
1058:
1038:
1008:
895:
893:{\displaystyle q_{i}}
841:
799:
766:
730:
710:
690:
654:
624:
594:
443:
391:
325:
252:
101:
18:Universal force field
5934:Molecular Simulation
4567:10.1110/ps.062126106
4368:10.1109/MCSE.2012.21
3275:Molecular Simulation
3090:(13–16): 1394–1405.
3084:Molecular Simulation
2107:Molecular Simulation
1917:Bond order potential
1819:Modified amino acids
1631:Gao Research Group |
1567:William L. Jorgensen
1312:Intermolecular force
1308:van der Waals forces
1258:Embedded Atom Models
1207:for each potential.
1067:
1047:
1017:
920:
877:
820:
778:
739:
719:
699:
663:
633:
603:
502:
399:
334:
281:
275:van der Waals forces
91:For other uses, see
58:improve this article
7958:Molecular modelling
7888:elementary particle
7653:partition functions
7208:J. Chem. Theo. Chem
7169:2018JChPh.148x1722S
7104:2015arXiv150304987R
6822:2009PNAS..10615667K
6734:1995EL.....30..191E
6683:1993EL.....21..363K
6632:1992EL.....19..155H
6573:2001JPCA..105.9396V
6458:(40): 11242–11255.
6411:(36): 15665–15673.
6325:2006JChPh.125r4101C
6268:2006JChPh.124j4101P
6127:2002JChPh.117.9175C
6016:10.1021/j100044a039
5503:2005CoPhC.172...69Y
5452:1993JPCM....5.1031M
5409:1958PhRv..112...90D
5339:10.1021/ja00051a040
5333:(25): 10024–10035.
5273:10.1021/ja00205a003
5246:10.1021/ja00205a002
5219:10.1021/ja00205a001
5174:10.1021/ja00467a001
5121:2018NatCo...9..716G
4906:10.1021/j100589a006
4825:10.1021/j100069a010
4770:1991PNAS...88.2854S
4713:1996BpJ....71.2033M
4701:Biophysical Journal
4401:(Suppl 9): 114–22.
4360:2012CSE....14b..97J
4204:(Suppl 6): 352–68.
3714:(12): 12805–12816.
3666:2015NatMa..14.1099X
3610:2000JChPh.112.8910M
3520:2012PhRvL.108n6103R
3463:2016PhRvB..93c5118R
3238:2011JOM....63g..17T
3052:2004APS..MAR.Y8006H
3046:(47): 18341–18352.
2813:(45): 11225–11236.
2759:1904KNAB....7..438L
2626:1984PhRvB..29.6443D
2571:1988PhRvB..37.6991T
2522:1977PhT....30R..61A
2504:Solid State Physics
2457:Dalton Transactions
2332:10.1021/ja00086a030
2284:2023FlPEq.57313876F
1991:2005JChPh.123w4505A
1922:Embedded atom model
1897:Molecular modelling
1887:Molecular mechanics
1855:valence bond theory
1658:University of Bonn.
1606:Alexander MacKerell
1591:harmonic oscillator
1462:like dissolves like
1435:protein engineering
1342:Jacob Israelachvili
1329:combinatorial rules
1325:combinatorial rules
1321:like dissolves like
1301:dielectric constant
120:molecular modelling
7915:information theory
7822:correlation length
7817:Critical exponents
7804:Critical phenomena
7785:stochastic process
7765:Boltzmann equation
7658:equations of state
7551:Warshel A (1991).
7532:Schlick T (2002).
6914:10.1039/C6SC05720A
6776:2010-07-15 at the
6511:10.1007/BF00124013
5052:. 2 February 2016.
4878:biohpc.cornell.edu
4407:10.1002/prot.22570
4211:10.1002/prot.10543
3764:10.1110/ps.0307002
3385:10.1039/c5cs00890e
2778:Annalen der Physik
2469:10.1039/c4dt00438h
1945:. Academic Press.
1941:Frenkel D (2007).
1882:Molecular dynamics
1766:Atoms in molecules
1711:Molecular dynamics
1515:modelling package.
1420:contribution, and
1100:covalent bonding,
1073:
1053:
1033:
1003:
890:
848:molecular dynamics
836:
814:quantum-mechanical
794:
761:
725:
705:
685:
649:
619:
589:
461:conjugated systems
438:
386:
320:
255:
148:molecular dynamics
116:physical chemistry
106:In the context of
104:
7953:Molecular physics
7930:
7929:
7920:Boltzmann machine
7790:mean-field theory
7691:Maxwell relations
7562:978-0-471-53395-5
7543:978-0-387-95404-2
7524:978-0-12-375181-2
7379:10.1021/sb400168u
7330:10.1021/ct400556v
7324:(12): 5653–5674.
7283:10.1021/jp805227c
7177:10.1063/1.5019779
6591:10.1021/jp004368u
6567:(41): 9396–9409.
6533:"Cresset Science"
6378:10.1021/jp905220k
6333:10.1063/1.2363374
6276:10.1063/1.2173256
6219:10.1021/ct700134r
6162:10.1021/ct400526a
6135:10.1063/1.1515773
6092:10.1021/ct049855i
6047:10.1021/ct700146x
5981:10.1021/cr9900477
5831:10.1021/ct900576a
5792:10.1021/ct049930p
5748:10.1002/jcc.20077
5592:10.1021/jp060163v
5549:10.1002/jcc.10355
5374:10.1021/cm0509328
5368:(23): 5658–5669.
5024:10.1021/la3038846
4933:10.1021/jp054994x
4243:Angewandte Chemie
4079:10.1021/ar010034r
3916:Missing or empty
3900:978-0-12-391927-4
3847:10.1002/prot.1106
3604:(20): 8910–8922.
3575:10.1021/ct5005565
3451:Physical Review B
3427:10.1021/cm500365c
3328:(22): 7148–7158.
3185:(10): 4278–4288.
3143:10.1021/jz500737m
3060:10.1021/jp048142t
3013:10.1021/ct500790p
2973:10.1002/qua.10108
2938:10.1021/ct300534j
2903:10.1021/ct600329w
2819:10.1021/ja9621760
2620:(12): 6443–6453.
2614:Physical Review B
2565:(12): 6991–7000.
2559:Physical Review B
2531:10.1063/1.3037370
2421:10.1002/jcc.20035
2371:10.1002/jcc.23354
2182:10.1021/jp071097f
2045:10.1021/ct200908r
1999:10.1063/1.2121687
1952:978-0-12-267351-1
1424:free energy. The
1371:homology modeling
1159:organic molecules
1076:{\displaystyle j}
1056:{\displaystyle i}
998:
959:
930:
728:{\displaystyle j}
708:{\displaystyle i}
536:
512:
435:
422:
409:
383:
370:
357:
344:
317:
304:
291:
223:methylene bridges
175:organic molecules
164:classical physics
156:quantum mechanics
112:molecular physics
86:
85:
78:
49:quality standards
40:This article may
16:(Redirected from
7965:
7812:Phase transition
7633:
7632:
7595:
7588:
7581:
7572:
7566:
7547:
7528:
7500:
7499:
7489:
7461:
7455:
7454:
7434:
7428:
7427:
7407:
7401:
7400:
7390:
7358:
7352:
7351:
7341:
7309:
7303:
7302:
7276:
7256:
7250:
7249:
7223:
7214:(6): 3678–3693.
7203:
7197:
7196:
7162:
7138:
7132:
7131:
7097:
7077:
7071:
7070:
7060:
7036:
7030:
7029:
7019:
6995:
6989:
6988:
6970:
6942:
6936:
6935:
6925:
6908:(4): 3192–3203.
6902:Chemical Science
6893:
6887:
6886:
6884:
6860:
6854:
6853:
6843:
6833:
6816:(37): 15667–72.
6801:
6795:
6794:
6787:
6781:
6768:
6762:
6761:
6717:
6711:
6710:
6666:
6660:
6659:
6615:
6609:
6608:
6606:
6605:
6599:
6593:. Archived from
6584:
6556:
6543:
6537:
6536:
6529:
6523:
6522:
6490:
6484:
6483:
6447:
6441:
6440:
6430:
6420:
6396:
6390:
6389:
6372:(33): 11463–78.
6361:
6355:
6354:
6344:
6304:
6298:
6297:
6287:
6247:
6241:
6240:
6230:
6213:(6): 1960–1986.
6198:
6192:
6191:
6180:
6174:
6173:
6145:
6139:
6138:
6110:
6104:
6103:
6075:
6069:
6068:
6058:
6041:(6): 1878–1889.
6026:
6020:
6019:
5999:
5993:
5992:
5975:(11): 4087–108.
5969:Chemical Reviews
5964:
5958:
5957:
5929:
5923:
5922:
5894:
5888:
5887:
5859:
5853:
5852:
5842:
5810:
5804:
5803:
5775:
5769:
5768:
5750:
5726:
5720:
5719:
5709:
5692:(7): 4122–4139.
5677:
5671:
5670:
5660:
5628:
5622:
5621:
5618:dasher.wustl.edu
5610:
5604:
5603:
5586:(26): 13166–76.
5575:
5569:
5568:
5532:
5523:
5522:
5486:
5480:
5479:
5446:(8): 1031–1038.
5435:
5429:
5428:
5392:
5386:
5385:
5357:
5351:
5350:
5322:
5316:
5315:
5310:
5304:
5303:
5283:
5277:
5276:
5256:
5250:
5249:
5229:
5223:
5222:
5202:
5196:
5195:
5190:. Archived from
5184:
5178:
5177:
5157:
5151:
5150:
5140:
5100:
5091:
5090:
5072:
5063:
5054:
5053:
5050:Heinz Laboratory
5042:
5036:
5035:
5007:
4998:
4997:
4986:
4980:
4979:
4951:
4945:
4944:
4916:
4910:
4909:
4888:
4882:
4881:
4870:
4864:
4863:
4835:
4829:
4828:
4808:
4802:
4801:
4791:
4781:
4749:
4743:
4742:
4732:
4692:
4686:
4685:
4673:
4667:
4666:
4630:
4624:
4623:
4595:
4589:
4588:
4578:
4546:
4540:
4539:
4505:
4499:
4498:
4470:
4464:
4463:
4435:
4429:
4428:
4418:
4386:
4380:
4379:
4343:
4337:
4336:
4308:
4302:
4301:
4273:
4267:
4266:
4238:
4232:
4231:
4213:
4189:
4183:
4182:
4164:
4140:
4134:
4133:
4097:
4091:
4090:
4062:
4056:
4055:
4019:
4013:
4012:
3975:
3969:
3968:
3932:
3926:
3925:
3919:
3914:
3912:
3904:
3882:
3867:
3866:
3830:
3824:
3823:
3795:
3786:
3785:
3775:
3758:(8): 1984–2000.
3743:
3732:
3731:
3703:
3694:
3693:
3674:10.1038/nmat4426
3659:
3650:(11): 1099–103.
3644:Nature Materials
3639:
3630:
3629:
3618:10.1063/1.481505
3593:
3587:
3586:
3558:
3552:
3551:
3541:
3531:
3499:
3493:
3492:
3482:
3445:
3439:
3438:
3421:(8): 2647–2658.
3412:
3403:
3397:
3396:
3368:
3362:
3361:
3313:
3307:
3306:
3281:(1–3): 101–105.
3266:
3260:
3259:
3249:
3217:
3211:
3210:
3174:
3165:
3164:
3154:
3122:
3116:
3115:
3078:
3072:
3071:
3035:
3026:
3025:
3015:
2991:
2985:
2984:
2956:
2950:
2949:
2921:
2915:
2914:
2886:
2880:
2879:
2842:
2831:
2830:
2802:
2796:
2795:
2793:
2772:Drude P (1900).
2769:
2763:
2762:
2742:
2736:
2735:
2725:
2708:(9): 4983–5013.
2702:Chemical Reviews
2693:
2687:
2686:
2676:
2667:(7–8): 251–310.
2652:
2646:
2645:
2605:
2599:
2598:
2550:
2544:
2543:
2533:
2497:
2491:
2490:
2480:
2463:(27): 10602–16.
2451:
2442:
2441:
2423:
2399:
2393:
2392:
2382:
2350:
2344:
2343:
2326:(7): 2978–2987.
2315:
2306:
2305:
2295:
2263:
2257:
2256:
2208:
2202:
2201:
2167:
2158:
2149:
2148:
2122:
2098:
2087:
2086:
2068:
2057:
2056:
2028:
2019:
2018:
1974:
1965:
1964:
1938:
1748:Machine learning
1366:NMR spectroscopy
1344:. For example, "
1338:Hamaker constant
1314:). The original
1253:self-consistency
1233:openKim database
1177:functional group
1116:Parameterization
1086:
1082:
1080:
1079:
1074:
1062:
1060:
1059:
1054:
1042:
1040:
1039:
1034:
1032:
1031:
1012:
1010:
1009:
1004:
999:
997:
996:
984:
983:
982:
973:
972:
962:
960:
958:
957:
956:
937:
932:
931:
928:
899:
897:
896:
891:
889:
888:
845:
843:
842:
837:
835:
834:
803:
801:
800:
795:
793:
792:
770:
768:
767:
762:
760:
759:
734:
732:
731:
726:
714:
712:
711:
706:
694:
692:
691:
686:
684:
683:
658:
656:
655:
650:
648:
647:
628:
626:
625:
620:
618:
617:
598:
596:
595:
590:
585:
584:
575:
574:
553:
552:
537:
532:
531:
519:
514:
513:
510:
459:rings and other
447:
445:
444:
439:
437:
436:
433:
424:
423:
420:
411:
410:
407:
395:
393:
392:
387:
385:
384:
381:
372:
371:
368:
359:
358:
355:
346:
345:
342:
329:
327:
326:
321:
319:
318:
315:
306:
305:
302:
293:
292:
289:
259:potential energy
144:potential energy
81:
74:
70:
67:
61:
35:
34:
27:
21:
7973:
7972:
7968:
7967:
7966:
7964:
7963:
7962:
7933:
7932:
7931:
7926:
7871:
7833:
7798:
7780:BBGKY hierarchy
7775:Vlasov equation
7753:
7742:depletion force
7735:Particles with
7695:
7634:
7630:
7625:
7604:
7599:
7569:
7563:
7550:
7544:
7531:
7525:
7512:
7508:
7506:Further reading
7503:
7463:
7462:
7458:
7436:
7435:
7431:
7409:
7408:
7404:
7360:
7359:
7355:
7311:
7310:
7306:
7267:(13): 4008–16.
7258:
7257:
7253:
7205:
7204:
7200:
7140:
7139:
7135:
7079:
7078:
7074:
7038:
7037:
7033:
6997:
6996:
6992:
6944:
6943:
6939:
6895:
6894:
6890:
6875:: 11423–11436.
6862:
6861:
6857:
6803:
6802:
6798:
6789:
6788:
6784:
6778:Wayback Machine
6769:
6765:
6719:
6718:
6714:
6668:
6667:
6663:
6617:
6616:
6612:
6603:
6601:
6597:
6582:10.1.1.507.6992
6554:
6545:
6544:
6540:
6531:
6530:
6526:
6492:
6491:
6487:
6449:
6448:
6444:
6398:
6397:
6393:
6363:
6362:
6358:
6306:
6305:
6301:
6249:
6248:
6244:
6200:
6199:
6195:
6182:
6181:
6177:
6147:
6146:
6142:
6121:(20): 9175–89.
6112:
6111:
6107:
6077:
6076:
6072:
6028:
6027:
6023:
6010:(44): 16460–7.
6001:
6000:
5996:
5966:
5965:
5961:
5931:
5930:
5926:
5896:
5895:
5891:
5861:
5860:
5856:
5812:
5811:
5807:
5777:
5776:
5772:
5741:(12): 1504–14.
5728:
5727:
5723:
5679:
5678:
5674:
5630:
5629:
5625:
5612:
5611:
5607:
5577:
5576:
5572:
5534:
5533:
5526:
5488:
5487:
5483:
5437:
5436:
5432:
5397:Physical Review
5394:
5393:
5389:
5359:
5358:
5354:
5324:
5323:
5319:
5312:
5311:
5307:
5285:
5284:
5280:
5267:(23): 8576–82.
5258:
5257:
5253:
5240:(23): 8566–75.
5231:
5230:
5226:
5213:(23): 8551–66.
5204:
5203:
5199:
5186:
5185:
5181:
5168:(25): 8127–34.
5159:
5158:
5154:
5102:
5101:
5094:
5070:
5065:
5064:
5057:
5044:
5043:
5039:
5009:
5008:
5001:
4994:www.igc.ethz.ch
4988:
4987:
4983:
4962:(4–5): 679–94.
4953:
4952:
4948:
4927:(10): 5025–44.
4918:
4917:
4913:
4900:(22): 2361–81.
4890:
4889:
4885:
4872:
4871:
4867:
4837:
4836:
4832:
4810:
4809:
4805:
4751:
4750:
4746:
4694:
4693:
4689:
4675:
4674:
4670:
4641:(10): 1667–80.
4632:
4631:
4627:
4597:
4596:
4592:
4555:Protein Science
4548:
4547:
4543:
4528:
4507:
4506:
4502:
4472:
4471:
4467:
4437:
4436:
4432:
4388:
4387:
4383:
4345:
4344:
4340:
4310:
4309:
4305:
4275:
4274:
4270:
4249:(15): 2644–76.
4240:
4239:
4235:
4191:
4190:
4186:
4142:
4141:
4137:
4099:
4098:
4094:
4064:
4063:
4059:
4021:
4020:
4016:
3987:(50): 1905247.
3977:
3976:
3972:
3934:
3933:
3929:
3915:
3905:
3901:
3884:
3883:
3870:
3832:
3831:
3827:
3806:(11): 1647–76.
3797:
3796:
3789:
3752:Protein Science
3745:
3744:
3735:
3705:
3704:
3697:
3641:
3640:
3633:
3595:
3594:
3590:
3569:(10): 4488–96.
3560:
3559:
3555:
3501:
3500:
3496:
3447:
3446:
3442:
3410:
3405:
3404:
3400:
3370:
3369:
3365:
3315:
3314:
3310:
3268:
3267:
3263:
3219:
3218:
3214:
3176:
3175:
3168:
3137:(11): 1885–91.
3124:
3123:
3119:
3080:
3079:
3075:
3037:
3036:
3029:
3006:(12): 5640–50.
2993:
2992:
2988:
2958:
2957:
2953:
2932:(11): 4719–31.
2923:
2922:
2918:
2888:
2887:
2883:
2844:
2843:
2834:
2804:
2803:
2799:
2771:
2770:
2766:
2744:
2743:
2739:
2695:
2694:
2690:
2654:
2653:
2649:
2607:
2606:
2602:
2552:
2551:
2547:
2499:
2498:
2494:
2453:
2452:
2445:
2401:
2400:
2396:
2365:(25): 2135–45.
2352:
2351:
2347:
2317:
2316:
2309:
2265:
2264:
2260:
2210:
2209:
2205:
2176:(27): 7812–24.
2165:
2160:
2159:
2152:
2113:(10): 806–814.
2100:
2099:
2090:
2083:
2070:
2069:
2060:
2030:
2029:
2022:
1976:
1975:
1968:
1953:
1940:
1939:
1935:
1931:
1926:
1872:
1840:
1831:Forcefield_NCAA
1821:
1801:
1795:
1750:
1702:
1692:William Goddard
1673:
1583:
1552:Norman Allinger
1492:
1484:
1470:
1406:crystallization
1398:protein folding
1379:protein folding
1269:
1249:
1241:MolMod database
1237:TraPPE database
1228:
1205:spring constant
1201:dihedral angles
1118:
1097:
1084:
1065:
1064:
1045:
1044:
1020:
1015:
1014:
985:
974:
964:
963:
948:
941:
923:
918:
917:
880:
875:
874:
868:
860:Morse potential
823:
818:
817:
781:
776:
775:
742:
737:
736:
717:
716:
697:
696:
666:
661:
660:
636:
631:
630:
606:
601:
600:
576:
557:
541:
520:
505:
500:
499:
492:
490:Bond stretching
453:Morse potential
428:
415:
402:
397:
396:
376:
363:
350:
337:
332:
331:
310:
297:
284:
279:
278:
247:
136:functional form
96:
89:
82:
71:
65:
62:
55:
53:Grammar issues.
36:
32:
23:
22:
15:
12:
11:
5:
7971:
7969:
7961:
7960:
7955:
7950:
7945:
7935:
7934:
7928:
7927:
7925:
7924:
7923:
7922:
7917:
7912:
7905:Complex system
7902:
7897:
7896:
7895:
7890:
7879:
7877:
7873:
7872:
7870:
7869:
7864:
7859:
7854:
7849:
7843:
7841:
7835:
7834:
7832:
7831:
7830:
7829:
7824:
7814:
7808:
7806:
7800:
7799:
7797:
7796:
7787:
7782:
7777:
7772:
7767:
7761:
7759:
7755:
7754:
7752:
7751:
7750:
7749:
7744:
7733:
7732:
7731:
7726:
7721:
7716:
7705:
7703:
7697:
7696:
7694:
7693:
7688:
7687:
7686:
7681:
7676:
7671:
7660:
7655:
7650:
7644:
7642:
7636:
7635:
7628:
7626:
7624:
7623:
7621:ergodic theory
7618:
7612:
7610:
7606:
7605:
7600:
7598:
7597:
7590:
7583:
7575:
7568:
7567:
7561:
7548:
7542:
7529:
7523:
7509:
7507:
7504:
7502:
7501:
7472:(6): 1282–90.
7456:
7445:(212): 11–34.
7429:
7402:
7373:(12): 855–69.
7353:
7304:
7251:
7198:
7153:(24): 241722.
7133:
7088:(5): 2087–96.
7072:
7051:(1): 216–224.
7031:
7010:(6): 2742–51.
6990:
6953:(1): 116–126.
6937:
6888:
6855:
6796:
6782:
6763:
6728:(4): 191–196.
6712:
6677:(3): 363–368.
6661:
6626:(3): 155–160.
6610:
6538:
6524:
6505:(6): 653–668.
6485:
6442:
6391:
6356:
6319:(18): 184101.
6299:
6262:(10): 104101.
6242:
6193:
6175:
6156:(12): 5550–7.
6140:
6105:
6086:(4): 694–715.
6070:
6021:
5994:
5959:
5924:
5889:
5854:
5825:(3): 774–786.
5805:
5770:
5721:
5672:
5643:(2): 419–426.
5623:
5605:
5570:
5524:
5481:
5430:
5387:
5352:
5317:
5305:
5278:
5251:
5224:
5197:
5194:on 2009-01-23.
5179:
5152:
5092:
5055:
5037:
5018:(6): 1754–65.
4999:
4981:
4946:
4911:
4883:
4865:
4830:
4819:(18): 4831–7.
4803:
4744:
4687:
4668:
4625:
4606:(3): 699–709.
4590:
4561:(6): 1318–33.
4541:
4526:
4500:
4465:
4430:
4381:
4338:
4303:
4268:
4233:
4184:
4135:
4108:(2): 145–237.
4092:
4057:
4014:
3970:
3943:(2): 105–267.
3927:
3899:
3868:
3825:
3787:
3733:
3695:
3631:
3588:
3553:
3514:(14): 146103.
3494:
3440:
3398:
3363:
3308:
3261:
3212:
3166:
3117:
3073:
3027:
2986:
2967:(1): 445–458.
2951:
2916:
2897:(4): 1464–75.
2881:
2832:
2797:
2784:(3): 566–613.
2764:
2737:
2688:
2647:
2600:
2545:
2492:
2443:
2414:(9): 1157–74.
2394:
2345:
2307:
2258:
2203:
2150:
2088:
2081:
2058:
2039:(4): 1459–70.
2020:
1985:(23): 234505.
1966:
1951:
1932:
1930:
1927:
1925:
1924:
1919:
1914:
1909:
1904:
1899:
1894:
1889:
1884:
1879:
1873:
1871:
1868:
1867:
1866:
1848:
1839:
1836:
1835:
1834:
1828:
1825:Forcefield_PTM
1820:
1817:
1813:solvent models
1797:Main article:
1794:
1791:
1790:
1789:
1786:
1783:Neural network
1779:
1776:
1773:neural network
1769:
1758:
1755:
1749:
1746:
1745:
1744:
1740:
1736:
1729:
1719:
1701:
1700:Coarse-grained
1698:
1697:
1696:
1685:
1672:
1669:
1668:
1667:
1663:
1659:
1655:
1652:
1649:
1646:
1643:
1640:
1637:
1634:
1626:
1623:
1620:
1616:
1613:
1602:
1598:
1587:polarizability
1582:
1579:
1578:
1577:
1573:
1570:
1560:
1548:
1541:
1533:
1527:
1519:
1516:
1512:
1506:
1499:
1491:
1488:
1469:
1466:
1426:heat of fusion
1414:heat of fusion
1402:ligand binding
1383:protein design
1375:ligand docking
1268:
1265:
1248:
1245:
1227:
1224:
1143:macromolecules
1135:dipole moments
1117:
1114:
1096:
1093:
1072:
1052:
1030:
1027:
1023:
1002:
995:
992:
988:
981:
977:
971:
967:
955:
951:
947:
944:
940:
935:
926:
906:polar covalent
887:
883:
872:atomic charges
867:
864:
833:
830:
826:
791:
788:
784:
758:
755:
752:
749:
745:
724:
704:
682:
679:
676:
673:
669:
646:
643:
639:
616:
613:
609:
588:
583:
579:
573:
570:
567:
564:
560:
556:
551:
548:
544:
540:
535:
530:
527:
523:
517:
508:
491:
488:
484:polarizability
465:hydrogen bonds
431:
427:
418:
414:
405:
379:
375:
366:
362:
353:
349:
340:
313:
309:
300:
296:
287:
263:covalent bonds
246:
243:
231:macromolecules
227:Coarse-grained
87:
84:
83:
39:
37:
30:
24:
14:
13:
10:
9:
6:
4:
3:
2:
7970:
7959:
7956:
7954:
7951:
7949:
7946:
7944:
7941:
7940:
7938:
7921:
7918:
7916:
7913:
7911:
7908:
7907:
7906:
7903:
7901:
7898:
7894:
7893:superfluidity
7891:
7889:
7886:
7885:
7884:
7881:
7880:
7878:
7874:
7868:
7865:
7863:
7860:
7858:
7855:
7853:
7850:
7848:
7845:
7844:
7842:
7840:
7836:
7828:
7825:
7823:
7820:
7819:
7818:
7815:
7813:
7810:
7809:
7807:
7805:
7801:
7795:
7791:
7788:
7786:
7783:
7781:
7778:
7776:
7773:
7771:
7768:
7766:
7763:
7762:
7760:
7756:
7748:
7745:
7743:
7740:
7739:
7738:
7734:
7730:
7727:
7725:
7722:
7720:
7717:
7715:
7712:
7711:
7710:
7707:
7706:
7704:
7702:
7698:
7692:
7689:
7685:
7682:
7680:
7677:
7675:
7672:
7670:
7667:
7666:
7664:
7661:
7659:
7656:
7654:
7651:
7649:
7646:
7645:
7643:
7641:
7637:
7622:
7619:
7617:
7614:
7613:
7611:
7607:
7603:
7596:
7591:
7589:
7584:
7582:
7577:
7576:
7573:
7564:
7558:
7554:
7549:
7545:
7539:
7535:
7530:
7526:
7520:
7516:
7511:
7510:
7505:
7497:
7493:
7488:
7483:
7479:
7475:
7471:
7467:
7460:
7457:
7452:
7448:
7444:
7440:
7433:
7430:
7425:
7421:
7417:
7413:
7406:
7403:
7398:
7394:
7389:
7384:
7380:
7376:
7372:
7368:
7364:
7357:
7354:
7349:
7345:
7340:
7335:
7331:
7327:
7323:
7319:
7315:
7308:
7305:
7300:
7296:
7292:
7288:
7284:
7280:
7275:
7270:
7266:
7262:
7255:
7252:
7247:
7243:
7239:
7235:
7231:
7227:
7222:
7217:
7213:
7209:
7202:
7199:
7194:
7190:
7186:
7182:
7178:
7174:
7170:
7166:
7161:
7156:
7152:
7148:
7144:
7137:
7134:
7129:
7125:
7121:
7117:
7113:
7109:
7105:
7101:
7096:
7091:
7087:
7083:
7076:
7073:
7068:
7064:
7059:
7054:
7050:
7046:
7042:
7035:
7032:
7027:
7023:
7018:
7013:
7009:
7005:
7001:
6994:
6991:
6986:
6982:
6978:
6974:
6969:
6964:
6960:
6956:
6952:
6948:
6941:
6938:
6933:
6929:
6924:
6919:
6915:
6911:
6907:
6903:
6899:
6892:
6889:
6883:
6878:
6874:
6870:
6866:
6859:
6856:
6851:
6847:
6842:
6837:
6832:
6827:
6823:
6819:
6815:
6811:
6807:
6800:
6797:
6792:
6786:
6783:
6779:
6775:
6772:
6767:
6764:
6759:
6755:
6751:
6747:
6743:
6739:
6735:
6731:
6727:
6723:
6716:
6713:
6708:
6704:
6700:
6696:
6692:
6688:
6684:
6680:
6676:
6672:
6665:
6662:
6657:
6653:
6649:
6645:
6641:
6637:
6633:
6629:
6625:
6621:
6614:
6611:
6600:on 2018-03-21
6596:
6592:
6588:
6583:
6578:
6574:
6570:
6566:
6562:
6561:
6553:
6549:
6542:
6539:
6534:
6528:
6525:
6520:
6516:
6512:
6508:
6504:
6500:
6496:
6489:
6486:
6481:
6477:
6473:
6469:
6465:
6461:
6457:
6453:
6446:
6443:
6438:
6434:
6429:
6424:
6419:
6414:
6410:
6406:
6402:
6395:
6392:
6387:
6383:
6379:
6375:
6371:
6367:
6360:
6357:
6352:
6348:
6343:
6338:
6334:
6330:
6326:
6322:
6318:
6314:
6310:
6303:
6300:
6295:
6291:
6286:
6281:
6277:
6273:
6269:
6265:
6261:
6257:
6253:
6246:
6243:
6238:
6234:
6229:
6224:
6220:
6216:
6212:
6208:
6204:
6197:
6194:
6189:
6185:
6179:
6176:
6171:
6167:
6163:
6159:
6155:
6151:
6144:
6141:
6136:
6132:
6128:
6124:
6120:
6116:
6109:
6106:
6101:
6097:
6093:
6089:
6085:
6081:
6074:
6071:
6066:
6062:
6057:
6052:
6048:
6044:
6040:
6036:
6032:
6025:
6022:
6017:
6013:
6009:
6005:
5998:
5995:
5990:
5986:
5982:
5978:
5974:
5970:
5963:
5960:
5955:
5951:
5947:
5943:
5940:(6): 471–84.
5939:
5935:
5928:
5925:
5920:
5916:
5912:
5908:
5905:(5): 524–31.
5904:
5900:
5893:
5890:
5885:
5881:
5877:
5873:
5870:(2): 227–49.
5869:
5865:
5858:
5855:
5850:
5846:
5841:
5836:
5832:
5828:
5824:
5820:
5816:
5809:
5806:
5801:
5797:
5793:
5789:
5786:(1): 153–68.
5785:
5781:
5774:
5771:
5766:
5762:
5758:
5754:
5749:
5744:
5740:
5736:
5732:
5725:
5722:
5717:
5713:
5708:
5703:
5699:
5695:
5691:
5687:
5683:
5676:
5673:
5668:
5664:
5659:
5654:
5650:
5646:
5642:
5638:
5634:
5627:
5624:
5619:
5615:
5609:
5606:
5601:
5597:
5593:
5589:
5585:
5581:
5574:
5571:
5566:
5562:
5558:
5554:
5550:
5546:
5542:
5538:
5531:
5529:
5525:
5520:
5516:
5512:
5508:
5504:
5500:
5496:
5492:
5485:
5482:
5477:
5473:
5469:
5465:
5461:
5457:
5453:
5449:
5445:
5441:
5434:
5431:
5426:
5422:
5418:
5414:
5410:
5406:
5403:(1): 90–103.
5402:
5398:
5391:
5388:
5383:
5379:
5375:
5371:
5367:
5363:
5356:
5353:
5348:
5344:
5340:
5336:
5332:
5328:
5321:
5318:
5309:
5306:
5301:
5297:
5294:(5): 709–17.
5293:
5289:
5282:
5279:
5274:
5270:
5266:
5262:
5255:
5252:
5247:
5243:
5239:
5235:
5228:
5225:
5220:
5216:
5212:
5208:
5201:
5198:
5193:
5189:
5183:
5180:
5175:
5171:
5167:
5163:
5156:
5153:
5148:
5144:
5139:
5134:
5130:
5126:
5122:
5118:
5114:
5110:
5106:
5099:
5097:
5093:
5088:
5084:
5080:
5076:
5069:
5062:
5060:
5056:
5051:
5047:
5041:
5038:
5033:
5029:
5025:
5021:
5017:
5013:
5006:
5004:
5000:
4995:
4991:
4985:
4982:
4977:
4973:
4969:
4965:
4961:
4957:
4950:
4947:
4942:
4938:
4934:
4930:
4926:
4922:
4915:
4912:
4907:
4903:
4899:
4895:
4887:
4884:
4879:
4875:
4869:
4866:
4861:
4857:
4853:
4849:
4846:(4): 90–102.
4845:
4841:
4834:
4831:
4826:
4822:
4818:
4814:
4807:
4804:
4799:
4795:
4790:
4785:
4780:
4775:
4771:
4767:
4764:(7): 2854–8.
4763:
4759:
4755:
4748:
4745:
4740:
4736:
4731:
4726:
4722:
4718:
4714:
4710:
4707:(4): 2033–9.
4706:
4702:
4698:
4691:
4688:
4683:
4679:
4672:
4669:
4664:
4660:
4656:
4652:
4648:
4644:
4640:
4636:
4629:
4626:
4621:
4617:
4613:
4609:
4605:
4601:
4594:
4591:
4586:
4582:
4577:
4572:
4568:
4564:
4560:
4556:
4552:
4545:
4542:
4537:
4533:
4529:
4527:9780121827885
4523:
4519:
4515:
4511:
4504:
4501:
4496:
4492:
4488:
4484:
4480:
4476:
4469:
4466:
4461:
4457:
4453:
4449:
4446:(4): 509–13.
4445:
4441:
4434:
4431:
4426:
4422:
4417:
4412:
4408:
4404:
4400:
4396:
4392:
4385:
4382:
4377:
4373:
4369:
4365:
4361:
4357:
4354:(2): 97–103.
4353:
4349:
4342:
4339:
4334:
4330:
4326:
4322:
4319:(2): 139–45.
4318:
4314:
4307:
4304:
4299:
4295:
4291:
4287:
4283:
4279:
4272:
4269:
4264:
4260:
4256:
4252:
4248:
4244:
4237:
4234:
4229:
4225:
4221:
4217:
4212:
4207:
4203:
4199:
4195:
4188:
4185:
4180:
4176:
4172:
4168:
4163:
4158:
4155:(sup1): 106.
4154:
4150:
4146:
4139:
4136:
4131:
4127:
4123:
4119:
4115:
4111:
4107:
4103:
4096:
4093:
4088:
4084:
4080:
4076:
4073:(6): 404–12.
4072:
4068:
4061:
4058:
4053:
4049:
4045:
4041:
4037:
4033:
4030:(2): 108–11.
4029:
4025:
4018:
4015:
4010:
4006:
4002:
3998:
3994:
3990:
3986:
3982:
3974:
3971:
3966:
3962:
3958:
3954:
3950:
3946:
3942:
3938:
3931:
3928:
3923:
3910:
3902:
3896:
3892:
3888:
3881:
3879:
3877:
3875:
3873:
3869:
3864:
3860:
3856:
3852:
3848:
3844:
3841:(4): 400–17.
3840:
3836:
3829:
3826:
3821:
3817:
3813:
3809:
3805:
3801:
3794:
3792:
3788:
3783:
3779:
3774:
3769:
3765:
3761:
3757:
3753:
3749:
3742:
3740:
3738:
3734:
3729:
3725:
3721:
3717:
3713:
3709:
3702:
3700:
3696:
3691:
3687:
3683:
3679:
3675:
3671:
3667:
3663:
3658:
3653:
3649:
3645:
3638:
3636:
3632:
3627:
3623:
3619:
3615:
3611:
3607:
3603:
3599:
3592:
3589:
3584:
3580:
3576:
3572:
3568:
3564:
3557:
3554:
3549:
3545:
3540:
3535:
3530:
3525:
3521:
3517:
3513:
3509:
3505:
3498:
3495:
3490:
3486:
3481:
3476:
3472:
3468:
3464:
3460:
3457:(3): 035118.
3456:
3452:
3444:
3441:
3436:
3432:
3428:
3424:
3420:
3416:
3409:
3402:
3399:
3394:
3390:
3386:
3382:
3379:(2): 412–48.
3378:
3374:
3367:
3364:
3359:
3355:
3351:
3347:
3343:
3339:
3335:
3331:
3327:
3323:
3319:
3312:
3309:
3304:
3300:
3296:
3292:
3288:
3284:
3280:
3276:
3272:
3265:
3262:
3257:
3253:
3248:
3243:
3239:
3235:
3231:
3227:
3223:
3216:
3213:
3208:
3204:
3200:
3196:
3192:
3188:
3184:
3180:
3173:
3171:
3167:
3162:
3158:
3153:
3148:
3144:
3140:
3136:
3132:
3128:
3121:
3118:
3113:
3109:
3105:
3101:
3097:
3093:
3089:
3085:
3077:
3074:
3069:
3065:
3061:
3057:
3053:
3049:
3045:
3041:
3034:
3032:
3028:
3023:
3019:
3014:
3009:
3005:
3001:
2997:
2990:
2987:
2982:
2978:
2974:
2970:
2966:
2962:
2955:
2952:
2947:
2943:
2939:
2935:
2931:
2927:
2920:
2917:
2912:
2908:
2904:
2900:
2896:
2892:
2885:
2882:
2877:
2873:
2869:
2865:
2861:
2857:
2853:
2849:
2841:
2839:
2837:
2833:
2828:
2824:
2820:
2816:
2812:
2808:
2801:
2798:
2792:
2787:
2783:
2779:
2775:
2768:
2765:
2760:
2756:
2752:
2748:
2741:
2738:
2733:
2729:
2724:
2719:
2715:
2711:
2707:
2703:
2699:
2692:
2689:
2684:
2680:
2675:
2670:
2666:
2662:
2658:
2651:
2648:
2643:
2639:
2635:
2631:
2627:
2623:
2619:
2615:
2611:
2604:
2601:
2596:
2592:
2588:
2584:
2580:
2576:
2572:
2568:
2564:
2560:
2556:
2549:
2546:
2541:
2537:
2532:
2527:
2523:
2519:
2515:
2511:
2510:Physics Today
2507:
2505:
2496:
2493:
2488:
2484:
2479:
2474:
2470:
2466:
2462:
2458:
2450:
2448:
2444:
2439:
2435:
2431:
2427:
2422:
2417:
2413:
2409:
2405:
2398:
2395:
2390:
2386:
2381:
2376:
2372:
2368:
2364:
2360:
2356:
2349:
2346:
2341:
2337:
2333:
2329:
2325:
2321:
2314:
2312:
2308:
2303:
2299:
2294:
2289:
2285:
2281:
2277:
2273:
2269:
2262:
2259:
2254:
2250:
2246:
2242:
2238:
2234:
2230:
2226:
2222:
2218:
2214:
2207:
2204:
2199:
2195:
2191:
2187:
2183:
2179:
2175:
2171:
2164:
2157:
2155:
2151:
2146:
2142:
2138:
2134:
2130:
2126:
2121:
2116:
2112:
2108:
2104:
2097:
2095:
2093:
2089:
2084:
2082:9780582382107
2078:
2074:
2067:
2065:
2063:
2059:
2054:
2050:
2046:
2042:
2038:
2034:
2027:
2025:
2021:
2016:
2012:
2008:
2004:
2000:
1996:
1992:
1988:
1984:
1980:
1973:
1971:
1967:
1962:
1958:
1954:
1948:
1944:
1937:
1934:
1928:
1923:
1920:
1918:
1915:
1913:
1910:
1908:
1905:
1903:
1900:
1898:
1895:
1893:
1890:
1888:
1885:
1883:
1880:
1878:
1875:
1874:
1869:
1864:
1860:
1856:
1852:
1849:
1847:
1842:
1841:
1837:
1832:
1829:
1826:
1823:
1822:
1818:
1816:
1814:
1810:
1806:
1800:
1792:
1787:
1784:
1780:
1777:
1774:
1770:
1767:
1763:
1759:
1756:
1752:
1751:
1747:
1741:
1737:
1734:
1730:
1727:
1723:
1720:
1716:
1712:
1708:
1704:
1703:
1699:
1693:
1689:
1686:
1683:
1679:
1675:
1674:
1670:
1664:
1660:
1656:
1653:
1650:
1647:
1644:
1641:
1638:
1635:
1632:
1627:
1624:
1621:
1617:
1614:
1611:
1607:
1603:
1599:
1596:
1595:
1594:
1592:
1588:
1580:
1574:
1571:
1568:
1564:
1561:
1557:
1553:
1549:
1546:
1542:
1539:
1534:
1531:
1528:
1525:
1520:
1517:
1513:
1510:
1507:
1504:
1503:Arieh Warshel
1500:
1497:
1494:
1493:
1489:
1487:
1483:
1479:
1475:
1467:
1465:
1463:
1459:
1455:
1452:of molecular
1451:
1448:
1444:
1440:
1436:
1432:
1427:
1423:
1419:
1415:
1411:
1407:
1403:
1399:
1395:
1391:
1386:
1384:
1380:
1376:
1372:
1367:
1363:
1359:
1355:
1349:
1347:
1343:
1339:
1335:
1330:
1326:
1322:
1317:
1313:
1309:
1306:All types of
1304:
1302:
1298:
1294:
1289:
1284:
1282:
1278:
1274:
1266:
1264:
1261:
1259:
1254:
1246:
1244:
1242:
1238:
1234:
1231:example, the
1225:
1223:
1221:
1217:
1212:
1208:
1206:
1202:
1198:
1194:
1190:
1186:
1185:atomic charge
1182:
1178:
1175:
1171:
1167:
1162:
1160:
1156:
1152:
1148:
1144:
1138:
1136:
1132:
1128:
1124:
1115:
1113:
1111:
1107:
1103:
1094:
1092:
1088:
1070:
1050:
1028:
1025:
1021:
1000:
993:
990:
986:
979:
975:
969:
965:
953:
949:
945:
942:
938:
933:
924:
915:
911:
910:ionic bonding
907:
903:
885:
881:
873:
865:
863:
861:
855:
853:
849:
831:
828:
824:
815:
811:
807:
789:
786:
782:
772:
756:
753:
750:
747:
743:
722:
702:
680:
677:
674:
671:
667:
644:
641:
637:
614:
611:
607:
586:
581:
571:
568:
565:
562:
558:
554:
549:
546:
542:
533:
528:
525:
521:
515:
506:
497:
489:
487:
485:
481:
480:Coulomb's law
477:
476:Mie potential
473:
468:
466:
462:
458:
454:
448:
434:van der Waals
429:
425:
421:electrostatic
416:
412:
403:
377:
373:
364:
360:
351:
347:
338:
311:
307:
298:
294:
285:
276:
272:
271:electrostatic
268:
264:
260:
251:
244:
242:
240:
239:nucleic acids
236:
232:
228:
224:
220:
219:methyl groups
216:
212:
208:
204:
200:
199:methyl groups
194:
192:
188:
184:
180:
176:
171:
169:
165:
161:
157:
153:
149:
145:
141:
137:
133:
129:
128:computational
125:
121:
117:
113:
109:
100:
94:
80:
77:
69:
59:
54:
50:
46:
45:
38:
29:
28:
19:
7876:Applications
7827:size scaling
7736:
7552:
7533:
7514:
7469:
7465:
7459:
7442:
7438:
7432:
7415:
7411:
7405:
7370:
7366:
7356:
7321:
7317:
7307:
7264:
7260:
7254:
7211:
7207:
7201:
7150:
7146:
7136:
7085:
7081:
7075:
7048:
7044:
7034:
7007:
7003:
6993:
6950:
6946:
6940:
6905:
6901:
6891:
6872:
6868:
6858:
6813:
6809:
6799:
6785:
6766:
6725:
6721:
6715:
6674:
6670:
6664:
6623:
6619:
6613:
6602:. Retrieved
6595:the original
6564:
6558:
6541:
6527:
6502:
6498:
6488:
6455:
6451:
6445:
6408:
6404:
6394:
6369:
6365:
6359:
6316:
6312:
6302:
6259:
6255:
6245:
6210:
6206:
6196:
6187:
6178:
6153:
6149:
6143:
6118:
6114:
6108:
6083:
6079:
6073:
6038:
6034:
6024:
6007:
6003:
5997:
5972:
5968:
5962:
5937:
5933:
5927:
5902:
5898:
5892:
5867:
5863:
5857:
5822:
5818:
5808:
5783:
5779:
5773:
5738:
5734:
5724:
5689:
5685:
5675:
5640:
5636:
5626:
5617:
5608:
5583:
5579:
5573:
5540:
5536:
5497:(2): 69–85.
5494:
5490:
5484:
5443:
5439:
5433:
5400:
5396:
5390:
5365:
5361:
5355:
5330:
5326:
5320:
5308:
5291:
5287:
5281:
5264:
5260:
5254:
5237:
5233:
5227:
5210:
5206:
5200:
5192:the original
5182:
5165:
5161:
5155:
5112:
5108:
5078:
5074:
5049:
5040:
5015:
5011:
4993:
4984:
4959:
4955:
4949:
4924:
4920:
4914:
4897:
4893:
4886:
4877:
4868:
4843:
4839:
4833:
4816:
4812:
4806:
4761:
4757:
4747:
4704:
4700:
4690:
4681:
4677:
4671:
4638:
4634:
4628:
4603:
4599:
4593:
4558:
4554:
4544:
4509:
4503:
4481:(4): 441–6.
4478:
4474:
4468:
4443:
4439:
4433:
4398:
4394:
4384:
4351:
4347:
4341:
4316:
4312:
4306:
4281:
4277:
4271:
4246:
4242:
4236:
4201:
4197:
4187:
4152:
4148:
4138:
4105:
4101:
4095:
4070:
4066:
4060:
4036:10.1038/5794
4027:
4023:
4017:
3984:
3980:
3973:
3940:
3936:
3930:
3918:|title=
3838:
3834:
3828:
3803:
3799:
3755:
3751:
3711:
3707:
3647:
3643:
3601:
3597:
3591:
3566:
3562:
3556:
3511:
3507:
3497:
3454:
3450:
3443:
3418:
3414:
3401:
3376:
3372:
3366:
3325:
3321:
3311:
3278:
3274:
3264:
3229:
3225:
3215:
3182:
3178:
3134:
3130:
3120:
3087:
3083:
3076:
3043:
3039:
3003:
2999:
2989:
2964:
2960:
2954:
2929:
2925:
2919:
2894:
2890:
2884:
2854:(1): 31–47.
2851:
2847:
2810:
2806:
2800:
2781:
2777:
2767:
2750:
2746:
2740:
2705:
2701:
2691:
2664:
2660:
2650:
2617:
2613:
2603:
2562:
2558:
2548:
2516:(1): 61–65.
2513:
2509:
2503:
2495:
2460:
2456:
2411:
2407:
2397:
2362:
2358:
2348:
2323:
2319:
2275:
2271:
2261:
2220:
2216:
2206:
2173:
2169:
2110:
2106:
2072:
2036:
2032:
1982:
1978:
1942:
1936:
1802:
1718:observables.
1584:
1556:hydrocarbons
1485:
1461:
1409:
1387:
1357:
1350:
1345:
1328:
1324:
1320:
1316:Fritz London
1305:
1296:
1293:polarization
1285:
1276:
1270:
1262:
1250:
1229:
1213:
1209:
1193:bond lengths
1163:
1139:
1119:
1098:
1089:
869:
856:
773:
493:
469:
449:
266:
256:
226:
210:
202:
195:
172:
160:force fields
123:
105:
72:
66:January 2024
63:
56:Please help
52:
41:
7867:von Neumann
7737:force field
7729:percolation
7418:: 745–752.
6968:10454/16776
5543:(1): 1–15.
4956:Biopolymers
4635:Biopolymers
4284:(1): 62–6.
2223:: 105–113.
1805:water model
1799:Water model
1610:Benoit Roux
1581:Polarizable
1526:and FANTOM.
1447:sublimation
1439:alpha helix
1394:sublimation
1288:anisotropic
1267:Limitations
1211:procedure.
1197:bond angles
1181:atomic mass
1131:sublimation
1110:Drude model
1091:solutions.
914:Coulomb law
496:Hooke's law
267:noncovalent
211:united-atom
152:Monte Carlo
124:force field
93:Force field
60:if you can.
7937:Categories
7724:Heisenberg
7487:1956/10456
7221:1902.08408
7160:1712.06113
7095:1503.04987
6882:2206.07697
6604:2015-08-29
6548:Goddard WA
5115:(1): 716.
4684:: 559–567.
3657:1505.05938
2478:2117/24209
2278:: 113876.
2120:1904.05206
1929:References
852:wavenumber
808:spectrum,
7847:Boltzmann
7770:H-theorem
7648:Ensembles
7274:0809.2811
7143:Müller KR
6750:0295-5075
6707:250913111
6699:0295-5075
6656:250796817
6648:0295-5075
6577:CiteSeerX
6519:1573-4951
6480:238230196
5519:0010-4655
5476:250752417
5468:0953-8984
5425:0031-899X
5382:0897-4756
5347:0002-7863
5081:: 68–89.
4171:0739-1102
4009:208700020
4001:1616-301X
3909:cite book
3626:0021-9606
3489:2469-9950
3435:0897-4756
3358:265103133
3342:1549-9596
3295:0892-7022
3256:1047-4838
3232:(7): 17.
3207:202745539
3104:0892-7022
3068:1520-6106
2981:0020-7608
2827:0002-7863
2683:0920-2307
2642:0163-1829
2587:0163-1829
2540:0031-9228
2340:0002-7863
2302:0378-3812
2253:266440296
2237:0039-3681
2145:119199372
2137:0892-7022
1961:254835355
1781:SchNet a
1739:packages.
1490:Classical
1422:solvation
1277:empirical
1168:atoms in
950:ε
946:π
555:−
498:formula:
408:nonbonded
316:nonbonded
217:atoms in
140:parameter
108:chemistry
7857:Tsallis
7496:25970002
7397:24932669
7348:24489522
7299:20782587
7291:18956896
7246:85543050
7238:31042390
7185:29960322
7128:28672393
7120:26574412
7067:29211469
7026:27224739
6985:54524604
6977:30507180
6932:28507695
6850:19717427
6774:Archived
6758:14385201
6550:(2001).
6472:34586817
6437:32343883
6386:19637900
6351:17115732
6294:16542062
6237:18978934
6170:26592288
6100:26641692
6065:18958290
5989:11749341
5954:96616243
5849:20300554
5800:26641126
5765:16741310
5757:15224394
5716:31136175
5667:31865706
5600:16805629
5565:39320318
5557:14634989
5147:29459638
5032:23276161
5012:Langmuir
4990:"GROMOS"
4976:94519023
4941:16526746
4860:91705326
4663:26981215
4585:16731967
4536:15063647
4495:12163065
4460:10449371
4425:19768677
4395:Proteins
4376:17613729
4333:10753811
4298:10679345
4263:12203463
4228:33186021
4220:14579324
4198:Proteins
4179:98441607
4130:43575627
4087:12069625
4044:10048917
3957:11771120
3855:11484218
3835:Proteins
3820:17049320
3782:12142453
3728:29179536
3708:ACS Nano
3682:26390325
3583:26588145
3548:22540809
3393:26750724
3350:37947503
3303:95716947
3199:31549507
3161:26273869
3112:36710284
3022:26583247
2946:26605626
2911:26633217
2848:Proteins
2732:26815602
2487:24828263
2438:18734898
2430:15116359
2389:23832629
2245:38128443
2198:13874240
2190:17569554
2053:26596756
2007:16392929
1870:See also
1671:Reactive
1601:AMOEBA+.
1454:crystals
1450:enthalpy
1410:freezing
1390:enthalpy
1174:carbonyl
1147:proteins
1145:such as
1127:enthalpy
902:covalent
806:infrared
457:aromatic
382:dihedral
235:proteins
233:such as
209:, while
207:hydrogen
187:minerals
183:polymers
168:gradient
42:require
7852:Shannon
7839:Entropy
7388:4277759
7339:3904396
7193:4897444
7165:Bibcode
7100:Bibcode
6923:5414547
6841:2734882
6818:Bibcode
6730:Bibcode
6679:Bibcode
6628:Bibcode
6569:Bibcode
6428:7267649
6342:2080839
6321:Bibcode
6285:2080832
6264:Bibcode
6228:2367138
6123:Bibcode
6056:2572772
5919:5227353
5840:2838399
5707:6615954
5658:7384396
5499:Bibcode
5448:Bibcode
5405:Bibcode
5138:5818522
5117:Bibcode
4874:"ECEPP"
4798:2011594
4766:Bibcode
4739:8889177
4730:1233669
4709:Bibcode
4655:2597723
4620:1660931
4576:2242528
4416:2922016
4356:Bibcode
4122:9794034
4052:3162636
3965:8401242
3863:9912122
3773:2373680
3690:5455024
3662:Bibcode
3606:Bibcode
3516:Bibcode
3459:Bibcode
3234:Bibcode
3152:9649520
3048:Bibcode
2876:2845395
2868:3054871
2755:Bibcode
2753:: 451.
2723:4865892
2622:Bibcode
2595:9943969
2567:Bibcode
2518:Bibcode
2380:3800559
2280:Bibcode
2015:9757894
1987:Bibcode
1851:VALBOND
1762:Kriging
1754:system.
1722:MARTINI
1682:enzymes
1431:H-bonds
929:Coulomb
474:or the
203:ll-atom
44:cleanup
7701:Models
7609:Theory
7559:
7540:
7521:
7494:
7395:
7385:
7346:
7336:
7297:
7289:
7244:
7236:
7191:
7183:
7126:
7118:
7065:
7024:
6983:
6975:
6930:
6920:
6848:
6838:
6791:"SAFT"
6756:
6748:
6705:
6697:
6654:
6646:
6579:
6517:
6478:
6470:
6435:
6425:
6384:
6349:
6339:
6292:
6282:
6235:
6225:
6168:
6098:
6063:
6053:
5987:
5952:
5917:
5884:985660
5882:
5847:
5837:
5798:
5763:
5755:
5714:
5704:
5665:
5655:
5598:
5563:
5555:
5517:
5474:
5466:
5423:
5380:
5345:
5145:
5135:
5030:
4974:
4939:
4858:
4796:
4786:
4737:
4727:
4661:
4653:
4618:
4583:
4573:
4534:
4524:
4493:
4458:
4423:
4413:
4374:
4331:
4296:
4261:
4226:
4218:
4177:
4169:
4128:
4120:
4085:
4050:
4042:
4007:
3999:
3963:
3955:
3897:
3861:
3853:
3818:
3780:
3770:
3726:
3688:
3680:
3624:
3581:
3546:
3487:
3433:
3391:
3356:
3348:
3340:
3301:
3293:
3254:
3205:
3197:
3159:
3149:
3110:
3102:
3066:
3020:
2979:
2944:
2909:
2874:
2866:
2825:
2730:
2720:
2681:
2640:
2593:
2585:
2538:
2485:
2436:
2428:
2387:
2377:
2338:
2300:
2251:
2243:
2235:
2196:
2188:
2143:
2135:
2079:
2051:
2013:
2005:
1959:
1949:
1861:, and
1846:Tinker
1688:ReaxFF
1543:MMFF (
1530:GROMOS
1509:CHARMM
1480:, and
1334:helium
1220:openMD
1216:openMM
1199:, and
1166:oxygen
1153:, and
1013:where
599:where
343:bonded
303:bonded
215:carbon
191:metals
189:, and
118:, and
7910:chaos
7862:Rényi
7719:Potts
7714:Ising
7295:S2CID
7269:arXiv
7242:S2CID
7216:arXiv
7189:S2CID
7155:arXiv
7124:S2CID
7090:arXiv
6981:S2CID
6877:arXiv
6754:S2CID
6703:S2CID
6652:S2CID
6598:(PDF)
6555:(PDF)
6476:S2CID
5950:S2CID
5915:S2CID
5761:S2CID
5561:S2CID
5472:S2CID
5071:(PDF)
4972:S2CID
4856:S2CID
4789:51338
4659:S2CID
4372:S2CID
4224:S2CID
4175:S2CID
4126:S2CID
4048:S2CID
4005:S2CID
3961:S2CID
3859:S2CID
3686:S2CID
3652:arXiv
3411:(PDF)
3354:S2CID
3299:S2CID
3203:S2CID
3108:S2CID
2872:S2CID
2434:S2CID
2249:S2CID
2194:S2CID
2166:(PDF)
2141:S2CID
2115:arXiv
2011:S2CID
1838:Other
1793:Water
1715:CECAM
1705:DPD (
1676:EVB (
1559:MM4).
1496:AMBER
1170:water
810:Raman
369:angle
290:total
126:is a
7792:and
7557:ISBN
7538:ISBN
7519:ISBN
7492:PMID
7393:PMID
7344:PMID
7287:PMID
7234:PMID
7181:PMID
7116:PMID
7063:PMID
7022:PMID
6973:PMID
6928:PMID
6846:PMID
6746:ISSN
6695:ISSN
6644:ISSN
6515:ISSN
6468:PMID
6433:PMID
6382:PMID
6347:PMID
6290:PMID
6233:PMID
6166:PMID
6096:PMID
6061:PMID
5985:PMID
5880:PMID
5845:PMID
5796:PMID
5753:PMID
5712:PMID
5663:PMID
5596:PMID
5553:PMID
5515:ISSN
5464:ISSN
5421:ISSN
5378:ISSN
5343:ISSN
5314:247.
5143:PMID
5028:PMID
4937:PMID
4794:PMID
4735:PMID
4651:PMID
4616:PMID
4581:PMID
4532:PMID
4522:ISBN
4491:PMID
4456:PMID
4421:PMID
4329:PMID
4294:PMID
4259:PMID
4216:PMID
4167:ISSN
4118:PMID
4083:PMID
4040:PMID
3997:ISSN
3953:PMID
3922:help
3895:ISBN
3851:PMID
3816:PMID
3804:1764
3778:PMID
3724:PMID
3678:PMID
3622:ISSN
3579:PMID
3544:PMID
3485:ISSN
3431:ISSN
3389:PMID
3346:PMID
3338:ISSN
3291:ISSN
3252:ISSN
3195:PMID
3157:PMID
3100:ISSN
3064:ISSN
3018:PMID
2977:ISSN
2942:PMID
2907:PMID
2864:PMID
2823:ISSN
2728:PMID
2679:ISSN
2638:ISSN
2591:PMID
2583:ISSN
2536:ISSN
2483:PMID
2426:PMID
2385:PMID
2336:ISSN
2298:ISSN
2241:PMID
2233:ISSN
2186:PMID
2133:ISSN
2077:ISBN
2049:PMID
2003:PMID
1957:OCLC
1947:ISBN
1563:OPLS
1443:coil
1400:and
1364:and
1271:All
1218:and
1063:and
908:and
715:and
511:bond
356:bond
273:and
221:and
179:ions
138:and
122:, a
7482:hdl
7474:doi
7447:doi
7443:212
7420:doi
7383:PMC
7375:doi
7334:PMC
7326:doi
7279:doi
7265:113
7226:doi
7173:doi
7151:148
7108:doi
7053:doi
7012:doi
6963:hdl
6955:doi
6918:PMC
6910:doi
6836:PMC
6826:doi
6814:106
6738:doi
6687:doi
6636:doi
6587:doi
6565:105
6507:doi
6460:doi
6456:125
6423:PMC
6413:doi
6374:doi
6370:113
6337:PMC
6329:doi
6317:125
6280:PMC
6272:doi
6260:124
6223:PMC
6215:doi
6158:doi
6131:doi
6119:117
6088:doi
6051:PMC
6043:doi
6012:doi
5977:doi
5973:100
5942:doi
5907:doi
5872:doi
5868:103
5835:PMC
5827:doi
5788:doi
5743:doi
5702:PMC
5694:doi
5653:PMC
5645:doi
5588:doi
5584:110
5545:doi
5507:doi
5495:172
5456:doi
5413:doi
5401:112
5370:doi
5335:doi
5331:114
5296:doi
5269:doi
5265:111
5242:doi
5238:111
5215:doi
5211:111
5170:doi
5133:PMC
5125:doi
5083:doi
5079:102
5020:doi
4964:doi
4929:doi
4925:110
4902:doi
4848:doi
4821:doi
4784:PMC
4774:doi
4725:PMC
4717:doi
4682:126
4643:doi
4608:doi
4604:222
4571:PMC
4563:doi
4514:doi
4483:doi
4448:doi
4411:PMC
4403:doi
4364:doi
4321:doi
4286:doi
4251:doi
4206:doi
4157:doi
4110:doi
4075:doi
4032:doi
3989:doi
3945:doi
3887:doi
3843:doi
3808:doi
3768:PMC
3760:doi
3716:doi
3670:doi
3614:doi
3602:112
3571:doi
3534:hdl
3524:doi
3512:108
3475:hdl
3467:doi
3423:doi
3381:doi
3330:doi
3283:doi
3242:doi
3226:JOM
3187:doi
3147:PMC
3139:doi
3092:doi
3056:doi
3044:108
3008:doi
2969:doi
2934:doi
2899:doi
2856:doi
2815:doi
2811:118
2786:doi
2782:306
2718:PMC
2710:doi
2706:116
2669:doi
2630:doi
2575:doi
2526:doi
2473:hdl
2465:doi
2416:doi
2375:PMC
2367:doi
2328:doi
2324:116
2288:doi
2276:573
2225:doi
2221:103
2178:doi
2174:111
2125:doi
2041:doi
1995:doi
1983:123
1441:to
1437:or
1392:of
1155:RNA
1151:DNA
1129:of
904:to
162:in
150:or
7939::
7665::
7490:.
7480:.
7470:55
7468:.
7441:.
7416:71
7414:.
7391:.
7381:.
7369:.
7365:.
7342:.
7332:.
7320:.
7316:.
7293:.
7285:.
7277:.
7263:.
7240:.
7232:.
7224:.
7212:15
7210:.
7187:.
7179:.
7171:.
7163:.
7149:.
7122:.
7114:.
7106:.
7098:.
7086:11
7084:.
7061:.
7049:14
7047:.
7043:.
7020:.
7008:12
7006:.
7002:.
6979:.
6971:.
6961:.
6951:15
6949:.
6926:.
6916:.
6904:.
6900:.
6873:35
6871:.
6867:.
6844:.
6834:.
6824:.
6812:.
6808:.
6752:.
6744:.
6736:.
6726:30
6724:.
6701:.
6693:.
6685:.
6675:21
6673:.
6650:.
6642:.
6634:.
6624:19
6622:.
6585:.
6575:.
6563:.
6557:.
6513:.
6501:.
6497:.
6474:.
6466:.
6454:.
6431:.
6421:.
6409:59
6407:.
6403:.
6380:.
6368:.
6345:.
6335:.
6327:.
6315:.
6311:.
6288:.
6278:.
6270:.
6258:.
6254:.
6231:.
6221:.
6209:.
6205:.
6186:.
6164:.
6152:.
6129:.
6117:.
6094:.
6082:.
6059:.
6049:.
6037:.
6033:.
6008:99
6006:.
5983:.
5971:.
5948:.
5938:32
5936:.
5913:.
5903:15
5901:.
5878:.
5866:.
5843:.
5833:.
5821:.
5817:.
5794:.
5782:.
5759:.
5751:.
5739:25
5737:.
5733:.
5710:.
5700:.
5690:15
5688:.
5684:.
5661:.
5651:.
5641:11
5639:.
5635:.
5616:.
5594:.
5582:.
5559:.
5551:.
5541:25
5539:.
5527:^
5513:.
5505:.
5493:.
5470:.
5462:.
5454:.
5442:.
5419:.
5411:.
5399:.
5376:.
5366:17
5364:.
5341:.
5329:.
5292:11
5290:.
5263:.
5236:.
5209:.
5166:99
5164:.
5141:.
5131:.
5123:.
5111:.
5107:.
5095:^
5077:.
5073:.
5058:^
5048:.
5026:.
5016:29
5014:.
5002:^
4992:.
4970:.
4960:29
4958:.
4935:.
4923:.
4898:79
4896:.
4876:.
4854:.
4842:.
4817:98
4815:.
4792:.
4782:.
4772:.
4762:88
4760:.
4756:.
4733:.
4723:.
4715:.
4705:71
4703:.
4699:.
4680:.
4657:.
4649:.
4639:28
4637:.
4614:.
4602:.
4579:.
4569:.
4559:15
4557:.
4553:.
4530:.
4520:.
4489:.
4479:12
4477:.
4454:.
4442:.
4419:.
4409:.
4399:77
4397:.
4393:.
4370:.
4362:.
4352:14
4350:.
4327:.
4317:10
4315:.
4292:.
4282:11
4280:.
4257:.
4247:41
4245:.
4222:.
4214:.
4202:53
4200:.
4196:.
4173:.
4165:.
4153:31
4151:.
4147:.
4124:.
4116:.
4106:31
4104:.
4081:.
4071:35
4069:.
4046:.
4038:.
4026:.
4003:.
3995:.
3985:29
3983:.
3959:.
3951:.
3941:34
3939:.
3913::
3911:}}
3907:{{
3893:.
3871:^
3857:.
3849:.
3839:44
3837:.
3814:.
3802:.
3790:^
3776:.
3766:.
3756:11
3754:.
3750:.
3736:^
3722:.
3712:11
3710:.
3698:^
3684:.
3676:.
3668:.
3660:.
3648:14
3646:.
3634:^
3620:.
3612:.
3600:.
3577:.
3567:10
3565:.
3542:.
3532:.
3522:.
3510:.
3506:.
3483:.
3473:.
3465:.
3455:93
3453:.
3429:.
3419:26
3417:.
3413:.
3387:.
3377:45
3375:.
3352:.
3344:.
3336:.
3326:63
3324:.
3320:.
3297:.
3289:.
3279:40
3277:.
3273:.
3250:.
3240:.
3230:63
3228:.
3224:.
3201:.
3193:.
3183:59
3181:.
3169:^
3155:.
3145:.
3133:.
3129:.
3106:.
3098:.
3088:43
3086:.
3062:.
3054:.
3042:.
3030:^
3016:.
3004:10
3002:.
2998:.
2975:.
2965:90
2963:.
2940:.
2928:.
2905:.
2893:.
2870:.
2862:.
2850:.
2835:^
2821:.
2809:.
2780:.
2776:.
2749:.
2726:.
2716:.
2704:.
2700:.
2677:.
2663:.
2659:.
2636:.
2628:.
2618:29
2616:.
2612:.
2589:.
2581:.
2573:.
2563:37
2561:.
2557:.
2534:.
2524:.
2514:30
2512:.
2508:.
2481:.
2471:.
2461:43
2459:.
2446:^
2432:.
2424:.
2412:25
2410:.
2406:.
2383:.
2373:.
2363:34
2361:.
2357:.
2334:.
2322:.
2310:^
2296:.
2286:.
2274:.
2270:.
2247:.
2239:.
2231:.
2219:.
2215:.
2192:.
2184:.
2172:.
2168:.
2153:^
2139:.
2131:.
2123:.
2111:45
2109:.
2105:.
2091:^
2061:^
2047:.
2035:.
2023:^
2009:.
2001:.
1993:.
1981:.
1969:^
1955:.
1815:.
1476:,
1377:,
1195:,
1187:,
1183:,
1149:,
1133:,
1125:,
1087:.
916::
467:.
237:,
185:,
181:,
177:,
114:,
110:,
7684:G
7679:F
7674:H
7669:U
7594:e
7587:t
7580:v
7565:.
7546:.
7527:.
7498:.
7484::
7476::
7453:.
7449::
7426:.
7422::
7399:.
7377::
7371:3
7350:.
7328::
7322:9
7301:.
7281::
7271::
7248:.
7228::
7218::
7195:.
7175::
7167::
7157::
7130:.
7110::
7102::
7092::
7069:.
7055::
7028:.
7014::
6987:.
6965::
6957::
6934:.
6912::
6906:8
6885:.
6879::
6852:.
6828::
6820::
6793:.
6760:.
6740::
6732::
6709:.
6689::
6681::
6658:.
6638::
6630::
6607:.
6589::
6571::
6535:.
6521:.
6509::
6503:8
6482:.
6462::
6439:.
6415::
6388:.
6376::
6353:.
6331::
6323::
6296:.
6274::
6266::
6239:.
6217::
6211:3
6190:.
6172:.
6160::
6154:9
6137:.
6133::
6125::
6102:.
6090::
6084:1
6067:.
6045::
6039:3
6018:.
6014::
5991:.
5979::
5956:.
5944::
5921:.
5909::
5886:.
5874::
5851:.
5829::
5823:6
5802:.
5790::
5784:1
5767:.
5745::
5718:.
5696::
5669:.
5647::
5620:.
5602:.
5590::
5567:.
5547::
5521:.
5509::
5501::
5478:.
5458::
5450::
5444:5
5427:.
5415::
5407::
5384:.
5372::
5349:.
5337::
5302:.
5298::
5275:.
5271::
5248:.
5244::
5221:.
5217::
5176:.
5172::
5149:.
5127::
5119::
5113:9
5089:.
5085::
5034:.
5022::
4996:.
4978:.
4966::
4943:.
4931::
4908:.
4904::
4880:.
4862:.
4850::
4844:7
4827:.
4823::
4800:.
4776::
4768::
4741:.
4719::
4711::
4665:.
4645::
4622:.
4610::
4587:.
4565::
4538:.
4516::
4497:.
4485::
4462:.
4450::
4444:9
4427:.
4405::
4378:.
4366::
4358::
4335:.
4323::
4300:.
4288::
4265:.
4253::
4230:.
4208::
4181:.
4159::
4132:.
4112::
4089:.
4077::
4054:.
4034::
4028:6
4011:.
3991::
3967:.
3947::
3924:)
3920:(
3903:.
3889::
3865:.
3845::
3822:.
3810::
3784:.
3762::
3730:.
3718::
3692:.
3672::
3664::
3654::
3628:.
3616::
3608::
3585:.
3573::
3550:.
3536::
3526::
3518::
3491:.
3477::
3469::
3461::
3437:.
3425::
3395:.
3383::
3360:.
3332::
3305:.
3285::
3258:.
3244::
3236::
3209:.
3189::
3163:.
3141::
3135:5
3114:.
3094::
3070:.
3058::
3050::
3024:.
3010::
2983:.
2971::
2948:.
2936::
2930:8
2913:.
2901::
2895:3
2878:.
2858::
2852:4
2829:.
2817::
2794:.
2788::
2761:.
2757::
2751:7
2734:.
2712::
2685:.
2671::
2665:9
2644:.
2632::
2624::
2597:.
2577::
2569::
2542:.
2528::
2520::
2506:"
2502:"
2489:.
2475::
2467::
2440:.
2418::
2391:.
2369::
2342:.
2330::
2304:.
2290::
2282::
2255:.
2227::
2200:.
2180::
2147:.
2127::
2117::
2085:.
2055:.
2043::
2037:8
2017:.
1997::
1989::
1963:.
1684:.
1071:j
1051:i
1029:j
1026:i
1022:r
1001:,
994:j
991:i
987:r
980:j
976:q
970:i
966:q
954:0
943:4
939:1
934:=
925:E
886:i
882:q
832:j
829:i
825:k
790:j
787:i
783:k
757:j
754:i
751:,
748:0
744:l
723:j
703:i
681:j
678:i
675:,
672:0
668:l
645:j
642:i
638:l
615:j
612:i
608:k
587:,
582:2
578:)
572:j
569:i
566:,
563:0
559:l
550:j
547:i
543:l
539:(
534:2
529:j
526:i
522:k
516:=
507:E
430:E
426:+
417:E
413:=
404:E
378:E
374:+
365:E
361:+
352:E
348:=
339:E
312:E
308:+
299:E
295:=
286:E
95:.
79:)
73:(
68:)
64:(
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.