113:) about 6 nm in diameter and 900 nm long. Several thousand copies of a small (50 amino-acid residues) elongated alpha-helical major coat protein subunit (the product of gene 8, or p8) in an overlapping shingle-like array form a hollow cylinder enclosing the circular single-stranded DNA genome. Each p8 subunit has a collection of basic residues near the C-terminus of the elongated protein and acidic residues near the N-terminus; these two regions are separated by about 20 hydrophobic (non-polar) residues. The shingle-like arrangement places the acidic residues of p8 near the outside surface of the cylinder, where they cause the virus particle to be negatively-charged; non-polar regions near non-polar regions of neighbouring p8 subunits, where non-polar interactions contribute to a notable physical stability of the virus particle; and basic residues near the centre of the cylinder, where they interact with the negatively-charged DNA phosphates at the core of the virion. Longer (or shorter) DNA molecules can be packaged, since more (or fewer) p8 subunits can be added during assembly as required to protect the DNA, making the phage useful for genetic studies. (This effect should not be confused with
102:
366:
Such modified phage are correspondingly longer that wild-type filamentous fd, because the longer DNA is coated with correspondingly more gene 8 coat proteins, but the phage life-cycle is not otherwise disrupted. The traditional “tadpole” or isometric shaped-phage, on the other hand, which have a limited-sized capsid, cannot be so easily used to encapsidate a larger DNA molecule. The modified phage can be selected by infecting kanamycin-sensitive bacteria with modified phage to introduce resistance to kanamycin, and growing the infected bacteria in media containing an otherwise lethal concentration of kanamycin.
31:
180:
305:, and the pilus retracts into the cell. This retraction may involve depolymerization of the pilus subunit assembly into the cell membrane at the base of the pilus by a reversal of the pilus growth and polymerization process. As the tip of the pilus bearing p3 approaches the cell wall, the N1 domain of p3 interacts with the bacterial TolQRA protein to complete infection and release the genome into the cytoplasm of the host.
140:
85:(virus particle) is a flexible filament measuring about 6 by 900 nm, comprising a cylindrical protein tube protecting a single-stranded circular DNA molecule at its core. The phage codes for only 11 gene products, and is one of the simplest viruses known. It has been widely used to study fundamental aspects of molecular biology. George Smith and Greg Winter used f1 and fd for their work on
314:
The host DNA polymerase III then uses this primer to synthesize the full complementary strand of DNA, yielding a double-stranded circle, sometimes called the replicative form (RF) DNA. The complementary strand of the RF is the transcription template for phage coded proteins, especially p2 and p10, which are necessary for further DNA replication.
340:
extrusion process picks up the p7 and p9 proteins which form the outer tip of the progeny phage. As the p5 is stripped off the DNA, the progeny DNA is extruded across the membrane and wrapped in a helical casing of p8, to which p3 and p6 are added at the end of assembly. The p4 protein may form an extrusion pore in the outer membrane.
334:
Infection does not kill the host bacteria, in contrast to most other families of phage. Progeny phage are assembled as they extrude through the membrane of growing bacteria, probably at adhesion sites joining inner and outer membranes. The five phage proteins that form the coat of the completed phage
160:
Five gene products are part of the virion: the major coat protein (p8) and the minor proteins capping the two ends, p3 and p6 at one end, and p7 and p9 at the other end. Three gene products (p2, p5, and p10) are cytoplasmic proteins needed for DNA synthesis and the rest are membrane proteins involved
339:
that are not present in the phage, p1, p11, and p4, are also involved in assembly. Replication of RF DNA is converted to production of phage ssDNA by coating of the DNA with p5 to form an elongated p5/DNA replication/assembly complex, which then interacts with the membrane-bound phage proteins. The
369:
This result was extended by inserting foreign DNA expressing a foreign peptide into fd phage gene 3, rather than into the intergenic sequence, so that the foreign peptide appears on the surface of the phage as a part of the gene 3 adsorption protein. Phage carrying the foreign peptide can then be
317:
The p2 protein cleaves the viral strand of the RF DNA, and host DNA polymerase III synthesizes a new viral strand. The old viral strand is displaced as the new one is synthesized. When a circle is complete, the covalently linked p2 cuts the displaced viral strand at the junction between the old and
313:
After the single-stranded viral DNA enters the cytoplasm, it serves as a template for the synthesis of a complementary DNA strand. This synthesis is initiated in the intergenic region of the DNA sequence by host RNA polymerase, which synthesizes a short RNA primer on the infecting DNA as template.
373:
These techniques have been extended over the years in many ways, for instance by inserting foreign DNA into the genes coding for phage coat proteins other than gene 3, and/or duplicating the gene of interest to modify only some of the corresponding gene products. Phage display technology has been
365:
Ff phages have been engineered for applications in biological and medical sciences. Many applications build on experiments showing that the DNA sequence determining resistance to the antibiotic kanamycin can be inserted in a functional form into the non-coding intergenic sequence of fd phage DNA.
351:
Intermediate assemblies of p8 can be generated by treating the phage with chloroform. The helical content of p8 in these intermediate forms is similar to that in the phage, suggesting that the structural change during assembly may involve just a sliding of the shingled p8 subunits with respect to
347:
which are essential for phage assembly, suggesting that p1 is a molecular motor involved in phage assembly. The p1 protein has a membrane-spanning hydrophobic domain with the N-terminal portion in the cytoplasm and the C-terminal portion in the periplasm (the reverse of the orientation of p8).
156:
producing 11 proteins, since two genes, gene 2 and gene 1, have internal in-frame translation starts, generating two additional proteins, p10 and p11. The genome also contains a short non-coding intergenic sequence. M13 and f1 sequences are slightly different from fd. They both have only 6407
325:
When the concentration of phage proteins has increased, new viral strands are coated by the replication/assembly protein p5 rather than by the complementary DNA strands. The p5 also inhibits translation of p2, so that progeny viral ssDNA production and packaging are in synchrony.
300:
The p3 protein is anchored to one end of the virion by the C-terminal domain of p3. Infection of host bacteria involves interaction of two different N-terminal regions of p3 with two different sites of the host bacteria. First, the N2 domain of p3 attaches to the outer tip of the
157:
nucleotides; f1 differs from fd in 180 positions (only 10 of these changes are reflected in amino-acid changes in gene products) and M13 has only 59 nucleotide differences from f1. For many purposes the phages in the Ff group can be considered as interchangeable.
2035:
Dorval
Courchesne NM, Klug MT, Huang KJ, Weidman MC, Cantú VJ, Chen PY, et al. (2015). "Constructing Multifunctional Virus-Templated Nanoporous Composites for Thin Film Solar Cells: Contributions of Morphology and Optics to Photocurrent Generation".
348:
Adjacent to the cytoplasmic side of the membrane-spanning domain is a 13- residue sequence of p1 having a pattern of basic residues closely matching the pattern of basic residues near the C terminus of p8, but inverted with respect to that sequence.
286:
The gene encoding p1 has been used as a conserved marker gene, along with three other features specific for inovirus genomes, in an automatic machine-learning approach to identify over 10000 inovirus-like sequences from microbial genomes.
335:
enter the inner membrane; for p8 and p3, N-terminal leader sequences (later removed) help the proteins to enter the bacterial membrane, with their N-termini directed away from the cytoplasm towards the periplasm. Three other phage
1724:
Webster, R.E., 2001. Filamentous phage biology. In: Barbas III, C.F., Burton, D.R., Scott, J.K., Silverman, G.J. (Eds.), Phage
Display: A Laboratory Manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, pp.
2212:
Li L, Belcher AM, Loke DK (December 2020). "Simulating selective binding of a biological template to a nanoscale architecture: a core concept of a clamp-based binding-pocket-favored N-terminal-domain assembly".
93:. Early experiments on Ff phages used M13 to identify gene functions, and M13 was also developed as a cloning vehicle, so the name M13 is sometimes used as an informal synonym for the whole group of Ff phages.
136:
data collection and computational analysis. Structures of the p3 capsid protein and the p5 replication/assembly protein have also been determined from X-ray crystallography and deposited in the PDB.
2256:
Brogan AP, Heldman N, Hallett JP, Belcher AM (September 2019). "Thermally robust solvent-free biofluids of M13 bacteriophage engineered for high compatibility with anhydrous ionic liquids".
739:
Pratt D, Tzagoloff H, Erdahl WS (November 1966). "Conditional lethal mutants of the small filamentous coliphage M13. I. Isolation, complementation, cell killing, time of cistron action".
847:
Herrmann R, Neugebauer K, Zentgraf H, Schaller H (February 1978). "Transposition of a DNA sequence determining kanamycin resistance into the single-stranded genome of bacteriophage fd".
343:
Interaction of the double-stranded packaging DNA signal with the p1-thioredoxin complex at the host inner membrane triggers the formation of a pore. The p1 protein contains
370:
detected using appropriate antibodies. The reverse of this approach is to insert DNA coding for antibodies into gene 3 and detect their presence by appropriate antigens.
2392:
101:
1506:
Stopar D, Spruijt RB, Wolfs CJ, Hemminga MA (July 1998). "Mimicking initial interactions of bacteriophage M13 coat protein disassembly in model membrane systems".
2124:
Casey JP, Barbero RJ, Heldman N, Belcher AM (November 2014). "Versatile de novo enzyme activity in capsid proteins from an engineered M13 bacteriophage library".
382:
Ff phages have been engineered for applications such as remediation, electrochemical, photovoltaic, catalytic, sensing and digital memory devices, especially by
2405:
1428:
Griffith J, Manning M, Dunn K (March 1981). "Filamentous bacteriophage contract into hollow spherical particles upon exposure to a chloroform-water interface".
1835:"Beyond phage display: non-traditional applications of the filamentous bacteriophage as a vaccine carrier, therapeutic biologic, and bioconjugation scaffold"
518:
1272:
Hoffmann-Thoms S, Jakob RP, Schmid FX (April 2014). "Energetic communication between functional sites of the gene-3-protein during infection by phage fd".
774:
Pratt D, Tzagoloff H, Beaudoin J (September 1969). "Conditional lethal mutants of the small filamentous coliphage M13. II. Two genes for coat proteins".
647:
Rakonjac J, Bennett NJ, Spagnuolo J, Gagic D, Russel M (2011). "Filamentous bacteriophage: biology, phage display and nanotechnology applications".
1237:
Bennett NJ, Rakonjac J (February 2006). "Unlocking of the filamentous bacteriophage virion during infection is mediated by the C domain of pIII".
1393:
Rapoza MP, Webster RE (May 1995). "The products of gene I and the overlapping in-frame gene XI are required for filamentous phage assembly".
961:
691:
538:
117:, which can package several separate and distinct DNA molecules). About 5 copies each of four minor proteins cap the two ends of the virion.
2463:
132:. In particular, the series of fd and Pf1 virion structures deposited in the PDB over decades illustrate the improvements in methods for
1344:"The Transmembrane Morphogenesis Protein gp1 of Filamentous Phages Contains Walker A and Walker B Motifs Essential for Phage Assembly"
1541:
Roberts LM, Dunker AK (October 1993). "Structural changes accompanying chloroform-induced contraction of the filamentous phage fd".
1576:
Xue, Bin; Blocquel, David; Habchi, Johnny; Uversky, Alexey V.; Kurgan, Lukasz; Uversky, Vladimir N.; Longhi, Sonia (2014).
1307:
Hoffmann
Berling H, Maze R (March 1964). "Release of male-specific bacteriophages from surviving host bacteria BACTERIA".
678:. Advances in Experimental Medicine and Biology. Vol. 1053. Cham: Springer International Publishing. pp. 1–20.
674:
Rakonjac J, Russel M, Khanum S, Brooke SJ, Rajič M (2017). "Filamentous Phage: Structure and
Biology". In Lim TS (ed.).
2159:
Zhang G, Wei S, Belcher AM (2018). "Biotemplated Zinc
Sulfide Nanofibers as Anode Materials for Sodium-Ion Batteries".
1034:
Beck E, Zink B (December 1981). "Nucleotide sequence and genome organisation of filamentous bacteriophages fl and fd".
2458:
78:
2448:
2410:
2453:
395:
179:
58:
1884:
Sioud M (April 2019). "Phage
Display Libraries: From Binders to Targeted Drug Delivery and Human Therapeutics".
2370:
1069:
Russel M, Linderoth NA, Sali A (June 1997). "Filamentous phage assembly: variation on a protein export theme".
90:
2443:
2438:
1194:
Craig L, Forest KT, Maier B (July 2019). "Type IV pili: dynamics, biophysics and functional consequences".
2332:
74:
1980:"Biologically enhanced cathode design for improved capacity and cycle life for lithium-oxygen batteries"
468:"'Big things in small packages: the genetics of filamentous phage and effects on fitness of their host'"
1736:
1625:"Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface"
1471:
Manning M, Griffith J (January 1985). "Association of M13 I-forms and spheroids with lipid vesicles".
2090:
2081:
Lee SW, Belcher AM (2004). "Virus-Based
Fabrication of Micro- and Nanofibers Using Electrospinning".
1991:
1636:
174:
2433:
892:"Ff-nano, short functionalized nanorods derived from Ff (f1, fd, or M13) filamentous bacteriophage"
556:"Similarities and differences within members of the Ff family of filamentous bacteriophage viruses"
2291:
2238:
2194:
1909:
1453:
1219:
872:
153:
2321:
2316:
152:
The DNA sequence of the fd genome has 6408 nucleotide comprising 9 genes, but the genome has 11
2397:
30:
2283:
2230:
2186:
2141:
2106:
2063:
2017:
1960:
1901:
1866:
1815:
1764:
1756:
1707:
1699:
1660:
1652:
1605:
1597:
1558:
1523:
1488:
1445:
1410:
1375:
1324:
1289:
1254:
1211:
1176:
1135:
1086:
1051:
1016:
967:
957:
923:
864:
826:
791:
756:
697:
687:
656:
624:
575:
534:
499:
448:
336:
133:
129:
125:
555:
2273:
2265:
2222:
2176:
2168:
2133:
2098:
2053:
2045:
2007:
1999:
1950:
1940:
1893:
1856:
1846:
1805:
1795:
1748:
1691:
1644:
1589:
1550:
1515:
1480:
1437:
1402:
1365:
1355:
1316:
1281:
1246:
1203:
1166:
1125:
1117:
1078:
1043:
1006:
998:
949:
913:
903:
856:
818:
783:
748:
715:
679:
614:
606:
567:
524:
489:
479:
438:
430:
17:
2311:
1104:
Roux S, Krupovic M, Daly RA, Borges AL, Nayfach S, Schulz F, et al. (November 2019).
985:
Beck E, Sommer R, Auerswald EA, Kurz C, Zink B, Osterburg G, et al. (December 1978).
318:
newly synthesized DNA and re-ligates the two ends and liberates p2. RF replicates by this
110:
1680:"Antibody-selectable filamentous fd phage vectors: affinity purification of target genes"
2094:
1995:
1752:
1640:
1106:"Cryptic inoviruses revealed as pervasive in bacteria and archaea across Earth's biomes"
2012:
1979:
1955:
1928:
1861:
1834:
1810:
1783:
1370:
1343:
1130:
1105:
918:
891:
619:
594:
383:
319:
1735:
Winter, Greg; Griffiths, Andrew D.; Hawkins, Robert E.; Hoogenboom, Hennie R. (1994).
1171:
1154:
1082:
1011:
986:
890:
Sattar S, Bennett NJ, Wen WX, Guthrie JM, Blackwell LF, Conway JF, Rakonjac J (2015).
443:
418:
2427:
2295:
2242:
2198:
1695:
1484:
1441:
1320:
1223:
1047:
822:
787:
752:
344:
252:
240:
228:
216:
86:
1913:
1457:
876:
520:
Filamentous
Bacteriophage in Bio/Nano/Technology, Bacterial Pathogenesis and Ecology
466:
Mai-Prochnow A, Hui JG, Kjelleberg S, Rakonjac J, McDougald D, Rice SA (July 2015).
204:
1679:
809:
Messing J (April 1991). "Cloning in M13 phage or how to use biology at its best".
953:
434:
2364:
683:
139:
2355:
1897:
1285:
1250:
1207:
1121:
529:
264:
2190:
2110:
2067:
2049:
1945:
1851:
1760:
1703:
1656:
1624:
1601:
1577:
908:
2379:
1648:
1002:
944:
Straus SK, Bo HE (2018). "Filamentous
Bacteriophage Proteins and Assembly".
610:
484:
467:
114:
70:
2287:
2234:
2172:
2145:
2021:
1964:
1905:
1870:
1819:
1784:"Filamentous bacteriophage fd as an antigen delivery system in vaccination"
1609:
1406:
1379:
1328:
1293:
1258:
1215:
1180:
1139:
971:
927:
701:
660:
628:
579:
503:
1768:
1711:
1664:
1562:
1527:
1492:
1449:
1414:
1090:
1055:
830:
795:
760:
452:
2349:
2278:
2181:
2058:
1800:
1020:
868:
66:
1554:
2269:
2226:
2003:
860:
494:
2137:
2102:
1593:
1519:
571:
1360:
121:
2326:
2384:
124:(the assembly of p8 subunit proteins) has been determined by X-ray
302:
191:
138:
100:
82:
29:
2330:
1929:"Filamentous Phages As a Model System in Soft Matter Physics"
1978:
Oh D, Qi J, Lu YC, Zhang Y, Shao-Horn Y, Belcher AM (2013).
948:. Subcellular Biochemistry. Vol. 88. pp. 261–279.
595:"Filamentous phages: masters of a microbial sharing economy"
1342:
Loh B, Haase M, Mueller L, Kuhn A, Leptihn S (April 2017).
1155:"F factor conjugation is a true type IV secretion system"
1153:
Lawley TD, Klimke WA, Gubbins MJ, Frost LS (July 2003).
419:"Ff coliphages: structural and functional relationships"
143:
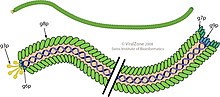
Schematic views showing minor proteins at the two ends
2339:
128:, and structural models have been deposited in the
322:mechanism to generate dozens of copies of the RF.
554:Morag O, Abramov G, Goldbourt A (December 2011).
523:. Frontiers Research Topics. Frontiers Media SA.
89:for which they were awarded a share of the 2018
1833:Henry KA, Arbabi-Ghahroudi M, Scott JK (2015).
1737:"Making Antibodies by Phage Display Technology"
34:Shadowed electron micrograph of unaligned phage
1678:Parmley, Stephen F.; Smith, George P. (1988).
676:Recombinant Antibodies for Infectious Diseases
987:"Nucleotide sequence of bacteriophage fd DNA"
412:
410:
8:
1788:International Journal of Molecular Sciences
105:Assembled major coat protein, exploded view
2327:
163:
2277:
2180:
2057:
2011:
1954:
1944:
1860:
1850:
1809:
1799:
1369:
1359:
1170:
1129:
1010:
946:Virus Protein and Nucleoprotein Complexes
917:
907:
618:
528:
517:Rakonjac J, Bas B, Derda R, eds. (2017).
493:
483:
442:
2126:Journal of the American Chemical Society
1578:"Structural Disorder in Viral Proteins"
1473:Archives of Biochemistry and Biophysics
406:
120:The molecular structure of the virion
939:
937:
842:
840:
417:Rasched I, Oberer E (December 1986).
7:
642:
640:
638:
378:Material sciences and nanotechnology
2038:The Journal of Physical Chemistry C
1753:10.1146/annurev.iy.12.040194.002245
716:"The Nobel Prize in Chemistry 2018"
649:Current Issues in Molecular Biology
560:The Journal of Physical Chemistry B
109:The virion is a flexible filament (
1782:Prisco A, De Berardinis P (2012).
352:their neighbours in the assembly.
25:
57:) is a group of almost identical
849:Molecular & General Genetics
374:widely used for many purposes.
178:
593:Hay ID, Lithgow T (June 2019).
1:
1172:10.1016/S0378-1097(03)00430-0
1083:10.1016/s0378-1119(96)00801-3
1696:10.1016/0378-1119(88)90495-7
1485:10.1016/0003-9861(85)90629-0
1442:10.1016/0092-8674(81)90438-4
1395:Journal of Molecular Biology
1321:10.1016/0042-6822(64)90021-2
1274:Journal of Molecular Biology
1239:Journal of Molecular Biology
1196:Nature Reviews. Microbiology
1048:10.1016/0378-1119(81)90059-7
954:10.1007/978-981-10-8456-0_12
823:10.1016/0378-1119(91)90344-b
788:10.1016/0042-6822(69)90346-8
753:10.1016/0042-6822(66)90118-8
435:10.1128/MR.50.4.401-427.1986
18:Filamentous bacteriophage fd
2464:Single-stranded DNA viruses
1741:Annual Review of Immunology
684:10.1007/978-3-319-72077-7_1
161:in assembly of the virion.
2480:
2161:ACS Applied Nano Materials
1898:10.1007/s12033-019-00156-8
361:Life sciences and medicine
1933:Frontiers in Microbiology
1839:Frontiers in Microbiology
1286:10.1016/j.jmb.2014.01.002
1251:10.1016/j.jmb.2005.11.069
1208:10.1038/s41579-019-0195-4
1159:FEMS Microbiology Letters
1122:10.1038/s41564-019-0510-x
896:Frontiers in Microbiology
530:10.3389/978-2-88945-095-4
472:FEMS Microbiology Reviews
396:Filamentous bacteriophage
173:
166:
2050:10.1021/acs.jpcc.5b00295
1946:10.3389/fmicb.2016.01013
1852:10.3389/fmicb.2015.00755
909:10.3389/fmicb.2015.00316
91:Nobel Prize in Chemistry
2341:Enterobacteria phage f1
2258:Chemical Communications
1886:Molecular Biotechnology
1649:10.1126/science.4001944
611:10.15252/embr.201847427
423:Microbiological Reviews
73:and ZJ/2, which infect
2173:10.1021/acsanm.8b01254
1407:10.1006/jmbi.1995.0247
991:Nucleic Acids Research
330:Assembly and extrusion
144:
106:
35:
1984:Nature Communications
1003:10.1093/nar/5.12.4495
485:10.1093/femsre/fuu007
142:
104:
33:
2371:Escherichia virus f1
1801:10.3390/ijms13045179
175:Virus classification
2264:(72): 10752–10755.
2221:(47): 24214–24227.
2095:2004NanoL...4..387L
2044:(25): 13987–14000.
1996:2013NatCo...4.2756O
1641:1985Sci...228.1315S
1635:(4705): 1315–1317.
1555:10.1021/bi00090a026
1110:Nature Microbiology
154:open reading frames
65:) including phages
2459:Structural biology
2270:10.1039/C9CC04909F
2227:10.1039/D0NR07320B
2004:10.1038/ncomms3756
1623:Smith, G. (1985).
861:10.1007/BF00270890
145:
107:
79:F fertility factor
36:
2449:Membrane proteins
2421:
2420:
2333:Taxon identifiers
2167:(10): 5631–5639.
2138:10.1021/ja506346f
2103:10.1021/nl034911t
1594:10.1021/cr4005692
1588:(13): 6880–6911.
1520:10.1021/bi9718144
1116:(11): 1895–1906.
963:978-981-10-8455-3
693:978-3-319-72076-0
572:10.1021/jp2079742
540:978-2-88945-095-4
337:membrane proteins
284:
283:
134:fiber diffraction
130:Protein Data Bank
126:fiber diffraction
59:filamentous phage
16:(Redirected from
2471:
2454:Membrane biology
2414:
2413:
2401:
2400:
2388:
2387:
2375:
2374:
2373:
2360:
2359:
2358:
2328:
2300:
2299:
2281:
2253:
2247:
2246:
2209:
2203:
2202:
2184:
2156:
2150:
2149:
2132:(47): 16508–14.
2121:
2115:
2114:
2078:
2072:
2071:
2061:
2032:
2026:
2025:
2015:
1975:
1969:
1968:
1958:
1948:
1927:Dogic Z (2016).
1924:
1918:
1917:
1881:
1875:
1874:
1864:
1854:
1830:
1824:
1823:
1813:
1803:
1779:
1773:
1772:
1732:
1726:
1722:
1716:
1715:
1675:
1669:
1668:
1620:
1614:
1613:
1582:Chemical Reviews
1573:
1567:
1566:
1549:(39): 10479–88.
1538:
1532:
1531:
1503:
1497:
1496:
1468:
1462:
1461:
1425:
1419:
1418:
1390:
1384:
1383:
1373:
1363:
1361:10.3390/v9040073
1339:
1333:
1332:
1304:
1298:
1297:
1269:
1263:
1262:
1234:
1228:
1227:
1191:
1185:
1184:
1174:
1150:
1144:
1143:
1133:
1101:
1095:
1094:
1066:
1060:
1059:
1031:
1025:
1024:
1014:
997:(12): 4495–503.
982:
976:
975:
941:
932:
931:
921:
911:
887:
881:
880:
844:
835:
834:
806:
800:
799:
771:
765:
764:
736:
730:
729:
727:
726:
712:
706:
705:
671:
665:
664:
644:
633:
632:
622:
590:
584:
583:
551:
545:
544:
532:
514:
508:
507:
497:
487:
463:
457:
456:
446:
414:
386:and colleagues.
183:
182:
164:
27:Group of viruses
21:
2479:
2478:
2474:
2473:
2472:
2470:
2469:
2468:
2424:
2423:
2422:
2417:
2409:
2404:
2396:
2391:
2383:
2378:
2369:
2368:
2363:
2354:
2353:
2348:
2335:
2308:
2303:
2255:
2254:
2250:
2211:
2210:
2206:
2158:
2157:
2153:
2123:
2122:
2118:
2080:
2079:
2075:
2034:
2033:
2029:
1977:
1976:
1972:
1926:
1925:
1921:
1883:
1882:
1878:
1832:
1831:
1827:
1781:
1780:
1776:
1734:
1733:
1729:
1723:
1719:
1677:
1676:
1672:
1622:
1621:
1617:
1575:
1574:
1570:
1540:
1539:
1535:
1514:(28): 10181–7.
1505:
1504:
1500:
1470:
1469:
1465:
1427:
1426:
1422:
1392:
1391:
1387:
1341:
1340:
1336:
1306:
1305:
1301:
1271:
1270:
1266:
1236:
1235:
1231:
1193:
1192:
1188:
1152:
1151:
1147:
1103:
1102:
1098:
1068:
1067:
1063:
1033:
1032:
1028:
984:
983:
979:
964:
943:
942:
935:
889:
888:
884:
846:
845:
838:
808:
807:
803:
773:
772:
768:
738:
737:
733:
724:
722:
714:
713:
709:
694:
673:
672:
668:
646:
645:
636:
592:
591:
587:
566:(51): 15370–9.
553:
552:
548:
541:
516:
515:
511:
465:
464:
460:
416:
415:
408:
404:
392:
380:
363:
358:
332:
311:
298:
293:
177:
150:
111:worm-like chain
99:
28:
23:
22:
15:
12:
11:
5:
2477:
2475:
2467:
2466:
2461:
2456:
2451:
2446:
2444:Phage proteins
2441:
2439:Bacteriophages
2436:
2426:
2425:
2419:
2418:
2416:
2415:
2402:
2389:
2376:
2361:
2345:
2343:
2337:
2336:
2331:
2325:
2324:
2319:
2314:
2307:
2306:External links
2304:
2302:
2301:
2248:
2204:
2151:
2116:
2089:(3): 387–390.
2073:
2027:
1970:
1919:
1892:(4): 286–303.
1876:
1825:
1794:(4): 5179–94.
1774:
1747:(1): 433–455.
1727:
1717:
1690:(2): 305–318.
1670:
1615:
1568:
1533:
1498:
1479:(1): 297–303.
1463:
1420:
1385:
1334:
1299:
1280:(8): 1711–22.
1264:
1229:
1202:(7): 429–440.
1186:
1145:
1096:
1061:
1042:(1–3): 35–58.
1026:
977:
962:
933:
882:
836:
801:
766:
747:(3): 397–410.
731:
720:NobelPrize.org
707:
692:
666:
634:
585:
546:
539:
509:
458:
405:
403:
400:
399:
398:
391:
388:
384:Angela Belcher
379:
376:
362:
359:
357:
354:
331:
328:
320:rolling circle
310:
307:
297:
294:
292:
289:
282:
281:
274:
270:
269:
262:
258:
257:
250:
246:
245:
242:Faserviricetes
238:
234:
233:
230:Hofneiviricota
226:
222:
221:
214:
210:
209:
202:
195:
194:
189:
185:
184:
171:
170:
149:
146:
98:
95:
26:
24:
14:
13:
10:
9:
6:
4:
3:
2:
2476:
2465:
2462:
2460:
2457:
2455:
2452:
2450:
2447:
2445:
2442:
2440:
2437:
2435:
2432:
2431:
2429:
2412:
2407:
2403:
2399:
2394:
2390:
2386:
2381:
2377:
2372:
2366:
2362:
2357:
2351:
2347:
2346:
2344:
2342:
2338:
2334:
2329:
2323:
2320:
2318:
2315:
2313:
2310:
2309:
2305:
2297:
2293:
2289:
2285:
2280:
2279:1721.1/125988
2275:
2271:
2267:
2263:
2259:
2252:
2249:
2244:
2240:
2236:
2232:
2228:
2224:
2220:
2216:
2208:
2205:
2200:
2196:
2192:
2188:
2183:
2182:1721.1/126086
2178:
2174:
2170:
2166:
2162:
2155:
2152:
2147:
2143:
2139:
2135:
2131:
2127:
2120:
2117:
2112:
2108:
2104:
2100:
2096:
2092:
2088:
2084:
2077:
2074:
2069:
2065:
2060:
2059:1721.1/102981
2055:
2051:
2047:
2043:
2039:
2031:
2028:
2023:
2019:
2014:
2009:
2005:
2001:
1997:
1993:
1989:
1985:
1981:
1974:
1971:
1966:
1962:
1957:
1952:
1947:
1942:
1938:
1934:
1930:
1923:
1920:
1915:
1911:
1907:
1903:
1899:
1895:
1891:
1887:
1880:
1877:
1872:
1868:
1863:
1858:
1853:
1848:
1844:
1840:
1836:
1829:
1826:
1821:
1817:
1812:
1807:
1802:
1797:
1793:
1789:
1785:
1778:
1775:
1770:
1766:
1762:
1758:
1754:
1750:
1746:
1742:
1738:
1731:
1728:
1721:
1718:
1713:
1709:
1705:
1701:
1697:
1693:
1689:
1685:
1681:
1674:
1671:
1666:
1662:
1658:
1654:
1650:
1646:
1642:
1638:
1634:
1630:
1626:
1619:
1616:
1611:
1607:
1603:
1599:
1595:
1591:
1587:
1583:
1579:
1572:
1569:
1564:
1560:
1556:
1552:
1548:
1544:
1537:
1534:
1529:
1525:
1521:
1517:
1513:
1509:
1502:
1499:
1494:
1490:
1486:
1482:
1478:
1474:
1467:
1464:
1459:
1455:
1451:
1447:
1443:
1439:
1436:(3): 747–53.
1435:
1431:
1424:
1421:
1416:
1412:
1408:
1404:
1401:(3): 627–38.
1400:
1396:
1389:
1386:
1381:
1377:
1372:
1367:
1362:
1357:
1353:
1349:
1345:
1338:
1335:
1330:
1326:
1322:
1318:
1315:(3): 305–13.
1314:
1310:
1303:
1300:
1295:
1291:
1287:
1283:
1279:
1275:
1268:
1265:
1260:
1256:
1252:
1248:
1245:(2): 266–73.
1244:
1240:
1233:
1230:
1225:
1221:
1217:
1213:
1209:
1205:
1201:
1197:
1190:
1187:
1182:
1178:
1173:
1168:
1164:
1160:
1156:
1149:
1146:
1141:
1137:
1132:
1127:
1123:
1119:
1115:
1111:
1107:
1100:
1097:
1092:
1088:
1084:
1080:
1076:
1072:
1065:
1062:
1057:
1053:
1049:
1045:
1041:
1037:
1030:
1027:
1022:
1018:
1013:
1008:
1004:
1000:
996:
992:
988:
981:
978:
973:
969:
965:
959:
955:
951:
947:
940:
938:
934:
929:
925:
920:
915:
910:
905:
901:
897:
893:
886:
883:
878:
874:
870:
866:
862:
858:
854:
850:
843:
841:
837:
832:
828:
824:
820:
816:
812:
805:
802:
797:
793:
789:
785:
781:
777:
770:
767:
762:
758:
754:
750:
746:
742:
735:
732:
721:
717:
711:
708:
703:
699:
695:
689:
685:
681:
677:
670:
667:
662:
658:
654:
650:
643:
641:
639:
635:
630:
626:
621:
616:
612:
608:
604:
600:
596:
589:
586:
581:
577:
573:
569:
565:
561:
557:
550:
547:
542:
536:
531:
526:
522:
521:
513:
510:
505:
501:
496:
491:
486:
481:
478:(4): 465–87.
477:
473:
469:
462:
459:
454:
450:
445:
440:
436:
432:
429:(4): 401–27.
428:
424:
420:
413:
411:
407:
401:
397:
394:
393:
389:
387:
385:
377:
375:
371:
367:
360:
355:
353:
349:
346:
345:Walker motifs
341:
338:
329:
327:
323:
321:
315:
308:
306:
304:
295:
290:
288:
280:
279:
275:
272:
271:
268:
267:
263:
260:
259:
256:
255:
254:Tubulavirales
251:
248:
247:
244:
243:
239:
236:
235:
232:
231:
227:
224:
223:
220:
219:
215:
212:
211:
208:
207:
203:
200:
197:
196:
193:
190:
187:
186:
181:
176:
172:
169:
165:
162:
158:
155:
147:
141:
137:
135:
131:
127:
123:
118:
116:
112:
103:
96:
94:
92:
88:
87:phage display
84:
80:
76:
72:
68:
64:
60:
56:
52:
51:
46:
45:
40:
32:
19:
2340:
2261:
2257:
2251:
2218:
2214:
2207:
2164:
2160:
2154:
2129:
2125:
2119:
2086:
2083:Nano Letters
2082:
2076:
2041:
2037:
2030:
1987:
1983:
1973:
1936:
1932:
1922:
1889:
1885:
1879:
1842:
1838:
1828:
1791:
1787:
1777:
1744:
1740:
1730:
1720:
1687:
1683:
1673:
1632:
1628:
1618:
1585:
1581:
1571:
1546:
1543:Biochemistry
1542:
1536:
1511:
1508:Biochemistry
1507:
1501:
1476:
1472:
1466:
1433:
1429:
1423:
1398:
1394:
1388:
1351:
1347:
1337:
1312:
1308:
1302:
1277:
1273:
1267:
1242:
1238:
1232:
1199:
1195:
1189:
1162:
1158:
1148:
1113:
1109:
1099:
1077:(1): 23–32.
1074:
1070:
1064:
1039:
1035:
1029:
994:
990:
980:
945:
899:
895:
885:
855:(2): 171–8.
852:
848:
814:
810:
804:
782:(1): 42–53.
779:
775:
769:
744:
740:
734:
723:. Retrieved
719:
710:
675:
669:
655:(2): 51–76.
652:
648:
602:
599:EMBO Reports
598:
588:
563:
559:
549:
519:
512:
475:
471:
461:
426:
422:
381:
372:
368:
364:
356:Applications
350:
342:
333:
324:
316:
312:
299:
285:
277:
276:
265:
253:
241:
229:
217:
206:Monodnaviria
205:
198:
188:(unranked):
167:
159:
151:
119:
108:
77:bearing the
62:
54:
49:
48:
43:
42:
38:
37:
2365:Wikispecies
1990:(1): 2756.
1165:(1): 1–15.
495:10453/65260
309:Replication
53:ilamentous
2434:Inoviridae
2428:Categories
725:2021-04-10
402:References
291:Life cycle
266:Inoviridae
2312:ViralZone
2296:201115233
2243:227950477
2215:Nanoscale
2199:104742577
2191:2574-0970
2111:1530-6984
2068:1932-7447
1761:0732-0582
1725:1.1-1.37.
1704:0378-1119
1657:0036-8075
1602:0009-2665
1354:(4): 73.
1224:115153017
296:Infection
218:Loebvirae
213:Kingdom:
115:polyphage
97:Structure
47:specific
39:Ff phages
2398:11459734
2356:Q5424175
2350:Wikidata
2322:ATCC M13
2288:31432818
2235:33289758
2146:25343220
2022:24220635
1965:27446051
1939:: 1013.
1914:73434013
1906:30729435
1871:26300850
1820:22606037
1610:24823319
1458:46531024
1380:28397779
1329:14127828
1309:Virology
1294:24440124
1259:16373072
1216:30988511
1181:12855161
1140:31332386
972:29900501
928:25941520
877:22923713
817:: 3–12.
776:Virology
741:Virology
702:29549632
661:21502666
629:30952693
580:22085310
504:25670735
390:See also
278:Inovirus
261:Family:
225:Phylum:
168:Inovirus
148:Genetics
75:bacteria
63:Inovirus
2317:ATCC fd
2091:Bibcode
2013:3930201
1992:Bibcode
1956:4927585
1862:4523942
1845:: 755.
1811:3344273
1769:8011287
1712:3149606
1665:4001944
1637:Bibcode
1629:Science
1563:8399194
1528:9665724
1493:3966795
1450:7226228
1415:7752229
1371:5408679
1348:Viruses
1131:6813254
1091:9224870
1056:6282703
919:4403547
902:: 316.
831:2055478
796:5807970
761:5921643
620:6549030
453:3540571
273:Genus:
249:Order:
237:Class:
81:. The
61:(genus
2385:BPHAF1
2294:
2286:
2241:
2233:
2197:
2189:
2144:
2109:
2066:
2020:
2010:
1963:
1953:
1912:
1904:
1869:
1859:
1818:
1808:
1767:
1759:
1710:
1702:
1663:
1655:
1608:
1600:
1561:
1526:
1491:
1456:
1448:
1413:
1378:
1368:
1327:
1292:
1257:
1222:
1214:
1179:
1138:
1128:
1089:
1054:
1021:745987
1019:
1012:342768
1009:
970:
960:
926:
916:
875:
869:345091
867:
829:
794:
759:
700:
690:
659:
627:
617:
578:
537:
502:
451:
444:373080
441:
122:capsid
83:virion
69:, fd,
55:phages
2411:10863
2393:IRMNG
2292:S2CID
2239:S2CID
2195:S2CID
1910:S2CID
1454:S2CID
1220:S2CID
873:S2CID
605:(6).
303:pilus
199:Realm
192:Virus
41:(for
2406:NCBI
2380:EPPO
2284:PMID
2231:PMID
2187:ISSN
2142:PMID
2107:ISSN
2064:ISSN
2018:PMID
1961:PMID
1902:PMID
1867:PMID
1816:PMID
1765:PMID
1757:ISSN
1708:PMID
1700:ISSN
1684:Gene
1661:PMID
1653:ISSN
1606:PMID
1598:ISSN
1559:PMID
1524:PMID
1489:PMID
1446:PMID
1430:Cell
1411:PMID
1376:PMID
1325:PMID
1290:PMID
1255:PMID
1212:PMID
1177:PMID
1136:PMID
1087:PMID
1071:Gene
1052:PMID
1036:Gene
1017:PMID
968:PMID
958:ISBN
924:PMID
865:PMID
827:PMID
811:Gene
792:PMID
757:PMID
698:PMID
688:ISBN
657:PMID
625:PMID
576:PMID
535:ISBN
500:PMID
449:PMID
2274:hdl
2266:doi
2223:doi
2177:hdl
2169:doi
2134:doi
2130:136
2099:doi
2054:hdl
2046:doi
2042:119
2008:PMC
2000:doi
1951:PMC
1941:doi
1894:doi
1857:PMC
1847:doi
1806:PMC
1796:doi
1749:doi
1692:doi
1645:doi
1633:228
1590:doi
1586:114
1551:doi
1516:doi
1481:doi
1477:236
1438:doi
1403:doi
1399:248
1366:PMC
1356:doi
1317:doi
1282:doi
1278:426
1247:doi
1243:356
1204:doi
1167:doi
1163:224
1126:PMC
1118:doi
1079:doi
1075:192
1044:doi
1007:PMC
999:doi
950:doi
914:PMC
904:doi
857:doi
853:159
819:doi
815:100
784:doi
749:doi
680:doi
615:PMC
607:doi
568:doi
564:115
525:doi
490:hdl
480:doi
439:PMC
431:doi
71:M13
2430::
2408::
2395::
2382::
2367::
2352::
2290:.
2282:.
2272:.
2262:55
2260:.
2237:.
2229:.
2219:12
2217:.
2193:.
2185:.
2175:.
2163:.
2140:.
2128:.
2105:.
2097:.
2085:.
2062:.
2052:.
2040:.
2016:.
2006:.
1998:.
1986:.
1982:.
1959:.
1949:.
1935:.
1931:.
1908:.
1900:.
1890:61
1888:.
1865:.
1855:.
1841:.
1837:.
1814:.
1804:.
1792:13
1790:.
1786:.
1763:.
1755:.
1745:12
1743:.
1739:.
1706:.
1698:.
1688:73
1686:.
1682:.
1659:.
1651:.
1643:.
1631:.
1627:.
1604:.
1596:.
1584:.
1580:.
1557:.
1547:32
1545:.
1522:.
1512:37
1510:.
1487:.
1475:.
1452:.
1444:.
1434:23
1432:.
1409:.
1397:.
1374:.
1364:.
1350:.
1346:.
1323:.
1313:22
1311:.
1288:.
1276:.
1253:.
1241:.
1218:.
1210:.
1200:17
1198:.
1175:.
1161:.
1157:.
1134:.
1124:.
1112:.
1108:.
1085:.
1073:.
1050:.
1040:16
1038:.
1015:.
1005:.
993:.
989:.
966:.
956:.
936:^
922:.
912:.
898:.
894:.
871:.
863:.
851:.
839:^
825:.
813:.
790:.
780:39
778:.
755:.
745:30
743:.
718:.
696:.
686:.
653:13
651:.
637:^
623:.
613:.
603:20
601:.
597:.
574:.
562:.
558:.
533:.
498:.
488:.
476:39
474:.
470:.
447:.
437:.
427:50
425:.
421:.
409:^
301:F-
201::
67:f1
2298:.
2276::
2268::
2245:.
2225::
2201:.
2179::
2171::
2165:1
2148:.
2136::
2113:.
2101::
2093::
2087:4
2070:.
2056::
2048::
2024:.
2002::
1994::
1988:4
1967:.
1943::
1937:7
1916:.
1896::
1873:.
1849::
1843:6
1822:.
1798::
1771:.
1751::
1714:.
1694::
1667:.
1647::
1639::
1612:.
1592::
1565:.
1553::
1530:.
1518::
1495:.
1483::
1460:.
1440::
1417:.
1405::
1382:.
1358::
1352:9
1331:.
1319::
1296:.
1284::
1261:.
1249::
1226:.
1206::
1183:.
1169::
1142:.
1120::
1114:4
1093:.
1081::
1058:.
1046::
1023:.
1001::
995:5
974:.
952::
930:.
906::
900:6
879:.
859::
833:.
821::
798:.
786::
763:.
751::
728:.
704:.
682::
663:.
631:.
609::
582:.
570::
543:.
527::
506:.
492::
482::
455:.
433::
50:f
44:F
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.