118:, at a specific time chosen by the scientist. The scientist can then evaluate the effects of the knocked-out gene and identify the gene's normal function. This is different from having the gene absent starting from conception, whereby inactivation or loss of genes that are essential for the development of the organism may interfere with the normal function of cells and prevent the production of viable offspring.
27:
127:
172:
Translocation events occur when the loxP sites flank genes on two different DNA molecules in a unidirectional orientation. Cre recombinase is then used to generate a translocation between the two DNA molecules, exchanging the genetic material from one DNA molecule to the other, forming a simultaneous
87:
The floxing method is essential in the development of scientific model systems as it allows researchers to have spatial and temporal alteration of gene expression. The Cre-Lox system is widely used to manipulate gene expression in model organisms such as mice in order to study human diseases and drug
138:
experiments through precisely removing segments of or even whole genes. Deletion requires floxing of the segment of interest with loxP sites which face the same direction. The Cre recombinase will detect the unidirectional loxP sites and excise the floxed segment of DNA. The successfully edited
184:(heart muscle tissue) have been shown to express a type of Cre recombinase that is highly specific to cardiomyocytes and can be used by researchers to perform highly efficient recombinations. This is achieved by using a type of Cre whose expression is driven by the
111:
The floxing of genes is essential in the development of scientific model systems as it allows spatial and temporal alteration of gene expression. In layman's terms, the gene can be knocked-out (inactivated) in a specific tissue
155:
Inversion events are useful for inactivating a gene or DNA sequence without actually removing it, and thereby maintaining a consistent amount of genetic material. The inverted genes are not often associated with abnormal
164:. By undergoing Cre recombination, the region flanked by the loxP sites will become inverted, i.e. re-inserted in the same position but in reverse orientation; this process is not permanent and can be reversed.
160:, meaning the inverted genes are generally viable. Cre-LoxP recombination that results in inversion requires loxP sites flanking the gene of interest, with the loxP sites oriented towards each other as
285:). Tamoxifen binds to Cre-ER and disrupts its interactions with the chaperones, which allows the Cre-ER fusion protein to enter the nucleus and perform recombination on the floxed gene. Additionally,
130:
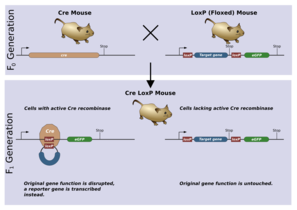
A model experiment in genetics using the Cre-lox system: the premature stop sequence present in floxed mice is removed only from cells that express Cre recombinase when the mice are bred together.
277:. The mutation renders the receptor inactive, which leads to incorrect localization through its interactions with chaperone proteins such as heat shock protein 70 and 90 (
254:
223:
203:
573:"Generation of knock-in mice that express nuclear enhanced green fluorescent protein and tamoxifen-inducible Cre recombinase in the notochord from Foxa2 and T loci"
1071:
431:
360:"Efficient recombination in diverse tissues by a tamoxifen-inducible form of Cre: a tool for temporally regulated gene activation/inactivation in the mouse"
257:-MyHC promoter causes the floxed gene to be inactivated in the heart alone. Further, these knockouts can be made inducible. In several mouse studies,
139:
clones can be selected using a selection marker which can be removed using the same Cre-LoxP system. The same mechanism can be used to create
507:
466:
108:
Floxing a gene allows it to be deleted (knocked out), translocated or inserted (through various mechanisms in Cre-Lox recombination).
324:
307:
1047:
1001:
548:
407:
1089:
Kobayashi K, Kamei Y, Kinoshita M, Czerny T, Tanaka M (January 2013). "A heat-inducible CRE/LOXP gene induction system in medaka".
709:"A cre-transgenic mouse strain for the ubiquitous deletion of loxP-flanked gene segments including deletion in germ cells"
1143:
667:"Adaptation to mutational inactivation of an essential E. coli gene converges to an accessible suboptimal fitness peak"
231:
and allows for the creation of conditional knockouts of the heart, mostly for use as controls. For example, using the
531:
Sakamoto K, Gurumurthy CB, Wagner KU (2014), Singh SR, Coppola V (eds.), "Generation of
Conditional Knockout Mice",
1075:
1025:
435:
30:
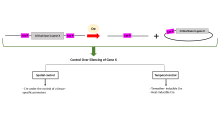
This figure depicts how
Floxing is used in scientific research for spatial and temporal control of gene expression.
140:
65:
19:
This article is about the genetics term. For the nickname for severe reactions to quinolone antibiotics, see
39:
946:"Modification of gene activity in mouse embryos in utero by a tamoxifen-inducible form of Cre recombinase"
274:
144:
73:
69:
52:
20:
957:
227:. These recombinations are capable of disrupting genes in a manner that is specific to heart tissue
61:
1114:
1065:
1019:
780:"A rapid in vitro method to flip back the double-floxed inverted open reading frame in a plasmid"
674:
472:
425:
337:
236:
185:
1106:
1053:
1043:
1007:
997:
975:
926:
860:
831:"Unidirectional Cre-mediated genetic inversion in mice using the mutant loxP pair lox66/lox71"
811:
738:
647:
602:
554:
544:
513:
503:
462:
413:
403:
381:
329:
270:
84:. The term "floxing" is a portmanteau constructed from the phrase "flanking/flanked by LoxP".
239:
208:
188:
1098:
965:
916:
908:
850:
842:
801:
791:
728:
720:
684:
637:
629:
592:
584:
536:
495:
454:
371:
319:
161:
93:
286:
266:
262:
232:
81:
77:
897:"Prolonged Cre expression driven by the α-myosin heavy chain promoter can be cardiotoxic"
961:
921:
896:
806:
779:
642:
621:
597:
572:
181:
135:
970:
945:
878:
Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart WM (2000). "Translocations".
855:
830:
733:
708:
490:
Friedel RH, Wurst W, Wefers B, Kühn R (2011). "Generating
Conditional Knockout Mice".
289:
can be induced by heat when under the control of specific heat shock elements (HSEs).
1132:
57:
1118:
476:
1138:
535:, Methods in Molecular Biology, vol. 1194, Springer New York, pp. 21–35,
341:
126:
88:
development. For example, using the Cre-Lox system, researchers are able to study
26:
633:
499:
912:
540:
879:
757:
895:
Pugach EK, Richmond PA, Azofeifa JG, Dowell RD, Leinwand LA (September 2015).
796:
458:
1057:
944:
Danielian PS, Muccino D, Rowitch DH, Michael SK, McMahon AP (December 1998).
724:
417:
1011:
258:
157:
1110:
930:
864:
815:
651:
606:
558:
517:
385:
376:
359:
333:
979:
742:
846:
89:
325:
10.1002/(SICI)1526-968X(200002)26:2<99::AID-GENE1>3.0.CO;2-B
147:
site which accomplishes the same mechanism but with a different enzyme.
114:
1102:
773:
771:
588:
97:
829:
Oberdoerffer P, Otipoby KL, Maruyama M, Rajewsky K (November 2003).
756:
Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart WM (2000).
402:. Singh, Shree Ram,, Coppola, Vincenzo. New York, NY. 26 July 2014.
689:
679:
666:
571:
Imuta Y, Kiyonari H, Jang CW, Behringer RR, Sasaki H (March 2013).
996:. Clarke, Alan R. (2nd ed.). Totowa, NJ: Humana Press. 2002.
282:
278:
25:
494:. Methods in Molecular Biology. Vol. 693. pp. 205–31.
1040:
Cancer and zebrafish : mechanisms, techniques, and models
43:
308:"Cre recombinase: the universal reagent for genome tailoring"
96:
genes and their role in the development and progression of
994:
Transgenesis techniques : principles and protocols
353:
351:
242:
211:
191:
248:
217:
197:
762:An Introduction to Genetic Analysis. 7th Edition
707:Schwenk F, Baron U, Rajewsky K (December 1995).
620:Hall B, Limaye A, Kulkarni AB (September 2009).
1042:. Langenau, David M. Switzerland. 10 May 2016.
451:Genetically Engineered Mice for Cancer Research
8:
901:Journal of Molecular and Cellular Cardiology
622:"Overview: generation of gene knockout mice"
665:Rodrigues JV, Shakhnovich EI (2019-08-01).
400:Mouse genetics : methods and protocols
1070:: CS1 maint: location missing publisher (
430:: CS1 maint: location missing publisher (
273:(ER) which contains a mutation within its
134:Deletion events are useful for performing
969:
920:
854:
805:
795:
732:
688:
678:
641:
596:
375:
323:
241:
210:
190:
125:
298:
1063:
1017:
492:Transgenic Mouse Methods and Protocols
423:
702:
700:
628:. Chapter 19: Unit 19.12 19.12.1–17.
7:
358:Hayashi S, McMahon AP (April 2002).
261:is used to induce the expression of
173:translocation of both floxed genes.
881:An Introduction to Genetic Analysis
269:is fused to a portion of the mouse
80:between LoxP sites is catalysed by
46:sequence (which is then said to be
56:sequences, creating an artificial
14:
626:Current Protocols in Cell Biology
60:which can then be conditionally
177:Common applications in research
205:-myosin heavy chain promoter (
1:
971:10.1016/s0960-9822(07)00562-3
16:Genetic engineering technique
634:10.1002/0471143030.cb1912s44
500:10.1007/978-1-60761-974-1_12
913:10.1016/j.yjmcc.2015.06.019
778:Xu J, Zhu Y (August 2018).
541:10.1007/978-1-4939-1215-5_2
275:ligand binding domain (LBD)
1160:
168:Mechanism of translocation
18:
797:10.1186/s12896-018-0462-x
459:10.1007/978-0-387-69805-2
449:Green JE, Ried T (2012).
306:Nagy A (February 2000).
34:In genetic engineering,
249:{\displaystyle \alpha }
218:{\displaystyle \alpha }
198:{\displaystyle \alpha }
1024:: CS1 maint: others (
835:Nucleic Acids Research
725:10.1093/nar/23.24.5080
713:Nucleic Acids Research
377:10.1006/dbio.2002.0597
250:
219:
199:
151:Mechanism of inversion
131:
31:
1074:) CS1 maint: others (
434:) CS1 maint: others (
364:Developmental Biology
251:
220:
200:
129:
122:Mechanism of deletion
74:Cre-Lox recombination
29:
21:Quinolone antibiotics
240:
209:
189:
72:in a process called
1144:Genetics techniques
962:1998CBio....8.1323D
141:conditional alleles
847:10.1093/nar/gng140
246:
215:
195:
143:by introducing an
132:
32:
1103:10.1002/dvg.22348
784:BMC Biotechnology
589:10.1002/dvg.22376
509:978-1-60761-973-4
468:978-0-387-69803-8
271:estrogen receptor
100:in mouse models.
1151:
1123:
1122:
1086:
1080:
1079:
1069:
1061:
1036:
1030:
1029:
1023:
1015:
990:
984:
983:
973:
941:
935:
934:
924:
892:
886:
885:
875:
869:
868:
858:
841:(22): 140e–140.
826:
820:
819:
809:
799:
775:
766:
765:
753:
747:
746:
736:
704:
695:
694:
692:
682:
662:
656:
655:
645:
617:
611:
610:
600:
568:
562:
561:
528:
522:
521:
487:
481:
480:
446:
440:
439:
429:
421:
396:
390:
389:
379:
355:
346:
345:
327:
303:
265:. In this case,
255:
253:
252:
247:
224:
222:
221:
216:
204:
202:
201:
196:
162:inverted repeats
104:Uses in research
94:tumor suppressor
1159:
1158:
1154:
1153:
1152:
1150:
1149:
1148:
1129:
1128:
1127:
1126:
1088:
1087:
1083:
1062:
1050:
1038:
1037:
1033:
1016:
1004:
992:
991:
987:
950:Current Biology
943:
942:
938:
894:
893:
889:
884:(7th ed.).
877:
876:
872:
828:
827:
823:
777:
776:
769:
755:
754:
750:
706:
705:
698:
664:
663:
659:
619:
618:
614:
570:
569:
565:
551:
530:
529:
525:
510:
489:
488:
484:
469:
448:
447:
443:
422:
410:
398:
397:
393:
357:
356:
349:
305:
304:
300:
295:
287:Cre recombinase
267:Cre recombinase
263:Cre recombinase
238:
237:
233:Cre recombinase
207:
206:
187:
186:
179:
170:
153:
124:
106:
82:Cre recombinase
64:(knocked out),
24:
17:
12:
11:
5:
1157:
1155:
1147:
1146:
1141:
1131:
1130:
1125:
1124:
1081:
1048:
1031:
1002:
985:
956:(24): 1323–6.
936:
887:
870:
821:
767:
748:
719:(24): 5080–1.
696:
690:10.1101/552240
657:
612:
563:
549:
533:Mouse Genetics
523:
508:
482:
467:
441:
408:
391:
347:
297:
296:
294:
291:
245:
214:
194:
182:Cardiomyocytes
178:
175:
169:
166:
152:
149:
123:
120:
105:
102:
50:) between two
38:refers to the
15:
13:
10:
9:
6:
4:
3:
2:
1156:
1145:
1142:
1140:
1137:
1136:
1134:
1120:
1116:
1112:
1108:
1104:
1100:
1096:
1092:
1085:
1082:
1077:
1073:
1067:
1059:
1055:
1051:
1049:9783319306544
1045:
1041:
1035:
1032:
1027:
1021:
1013:
1009:
1005:
1003:9781592591787
999:
995:
989:
986:
981:
977:
972:
967:
963:
959:
955:
951:
947:
940:
937:
932:
928:
923:
918:
914:
910:
906:
902:
898:
891:
888:
883:
882:
874:
871:
866:
862:
857:
852:
848:
844:
840:
836:
832:
825:
822:
817:
813:
808:
803:
798:
793:
789:
785:
781:
774:
772:
768:
763:
759:
752:
749:
744:
740:
735:
730:
726:
722:
718:
714:
710:
703:
701:
697:
691:
686:
681:
676:
672:
668:
661:
658:
653:
649:
644:
639:
635:
631:
627:
623:
616:
613:
608:
604:
599:
594:
590:
586:
582:
578:
574:
567:
564:
560:
556:
552:
550:9781493912148
546:
542:
538:
534:
527:
524:
519:
515:
511:
505:
501:
497:
493:
486:
483:
478:
474:
470:
464:
460:
456:
452:
445:
442:
437:
433:
427:
419:
415:
411:
409:9781493912155
405:
401:
395:
392:
387:
383:
378:
373:
370:(2): 305–18.
369:
365:
361:
354:
352:
348:
343:
339:
335:
331:
326:
321:
318:(2): 99–109.
317:
313:
309:
302:
299:
292:
290:
288:
284:
280:
276:
272:
268:
264:
260:
256:
243:
234:
230:
226:
212:
192:
183:
176:
174:
167:
165:
163:
159:
150:
148:
146:
142:
137:
128:
121:
119:
117:
116:
109:
103:
101:
99:
95:
91:
85:
83:
79:
78:Recombination
75:
71:
67:
63:
59:
58:gene cassette
55:
54:
49:
45:
41:
37:
28:
22:
1097:(1): 59–67.
1094:
1090:
1084:
1039:
1034:
993:
988:
953:
949:
939:
904:
900:
890:
880:
873:
838:
834:
824:
787:
783:
761:
758:"Inversions"
751:
716:
712:
670:
660:
625:
615:
583:(3): 210–8.
580:
576:
566:
532:
526:
491:
485:
450:
444:
399:
394:
367:
363:
315:
311:
301:
228:
180:
171:
154:
136:gene editing
133:
113:
110:
107:
86:
66:translocated
51:
47:
35:
33:
1133:Categories
680:1902.06630
673:: 552240.
293:References
158:phenotypes
1066:cite book
1058:949668674
1020:cite book
907:: 54–61.
790:(1): 52.
426:cite book
418:885338722
259:tamoxifen
244:α
235:with the
213:α
193:α
90:oncogenes
40:insertion
1119:25211137
1111:23019184
1012:50175106
931:26141530
865:14602933
816:30170595
652:19731224
607:23359409
559:25064096
518:21080282
477:40599715
386:11944939
334:10686599
70:inverted
1091:Genesis
980:9843687
958:Bibcode
922:4558343
807:6119287
743:8559668
671:bioRxiv
643:2782548
598:3632256
577:Genesis
342:2916710
312:Genesis
229:in vivo
145:FRT/Flp
115:in vivo
62:deleted
36:floxing
1117:
1109:
1056:
1046:
1010:
1000:
978:
929:
919:
863:
856:275577
853:
814:
804:
741:
734:307516
731:
650:
640:
605:
595:
557:
547:
516:
506:
475:
465:
416:
406:
384:
340:
332:
225:-MyHC)
98:cancer
48:floxed
1115:S2CID
675:arXiv
473:S2CID
338:S2CID
283:Hsp90
279:Hsp70
68:, or
42:of a
1107:PMID
1076:link
1072:link
1054:OCLC
1044:ISBN
1026:link
1008:OCLC
998:ISBN
976:PMID
927:PMID
861:PMID
812:PMID
739:PMID
648:PMID
603:PMID
555:PMID
545:ISBN
514:PMID
504:ISBN
463:ISBN
436:link
432:link
414:OCLC
404:ISBN
382:PMID
330:PMID
281:and
92:and
53:LoxP
1139:DNA
1099:doi
966:doi
917:PMC
909:doi
851:PMC
843:doi
802:PMC
792:doi
729:PMC
721:doi
685:doi
638:PMC
630:doi
593:PMC
585:doi
537:doi
496:doi
455:doi
372:doi
368:244
320:doi
44:DNA
1135::
1113:.
1105:.
1095:51
1093:.
1068:}}
1064:{{
1052:.
1022:}}
1018:{{
1006:.
974:.
964:.
952:.
948:.
925:.
915:.
905:86
903:.
899:.
859:.
849:.
839:31
837:.
833:.
810:.
800:.
788:18
786:.
782:.
770:^
760:.
737:.
727:.
717:23
715:.
711:.
699:^
683:.
669:.
646:.
636:.
624:.
601:.
591:.
581:51
579:.
575:.
553:,
543:,
512:.
502:.
471:.
461:.
453:.
428:}}
424:{{
412:.
380:.
366:.
362:.
350:^
336:.
328:.
316:26
314:.
310:.
76:.
1121:.
1101::
1078:)
1060:.
1028:)
1014:.
982:.
968::
960::
954:8
933:.
911::
867:.
845::
818:.
794::
764:.
745:.
723::
693:.
687::
677::
654:.
632::
609:.
587::
539::
520:.
498::
479:.
457::
438:)
420:.
388:.
374::
344:.
322::
23:.
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.