55:
212:
transfer genetic material through horizontal gene transfer, phylogenies and genome histories are difficult to define. Studies mostly focus on the analysis of the 16S rRNA gene to define relations and build phylogenic trees. However, to confirm the connections made from 16S rRNA analysis studies have started to analyze the house keeping genes within
207:
circular and plasmids are currently not found on the chromosome. The chromosome CR2-3X is the major chromosome that has been focused on the most. There are 27 genes located on that chromosome, 23 of those genes are involved in chromosome partitioning, DNA replication, transcription, and translation. The small genomes of
33:
171:, which are small bacteria that all share a common cell wall-less feature. They are characterized by helical and spherical morphology, they actually have the ability to be spherical or helical depending on the circumstances. The cells movement is bound by a membrane. The cell size ranges from 0.15 to 0.20 micrometers.
211:
is close to the minimal complement necessary for life and pathogenesis. Their ability to survive in specific niches gives in sight to unique ability of microorganisms to be able to evolve pathogenesis while experiencing severe genome reductions. Due to the nature of bacteria and their abilities to
206:
was established according to biochemical and phenotypic characteristics. These methods provided a general phylogeny, but large gaps in knowledge were made apparent when considering the possibility of horizontal gene transfer. The genome has some unique identities, the chromosomal structure if
361:
are take very seriously, due to it being a vector borne pathogen, it is able to be transferred between plants while only requiring one infected insect. Studies are on going to find proper resistance to this pathogen and will be progressing along with the disease.
379:
349:. It is considered a significant economic risk. Corn stunt disease results in smaller corn husks and loose or missing kernels. The corn industry is a billion-dollar industry that is already being threatened by global warming and the introduction of
256:
is an insect pathogen, it is transferred to other organisms by insects and it uses the host insect vector to multiply, but it is also important to note that some of the hosts are within the plants. Although they are transferred by insects,
186:
allows for the bacterium to be motile through flexional and rotational motility. The cell sizes are approximately 0.15-0.2 μm in diameter and 2.0 15 μm in length. The elongated shape of
971:
194:
can change into a coccoid shape. The ends of the helical shape have a tapered ends and blunt or rounded tips. The tip structure exposes adhesins that are used to attach to cells.
190:
aids in nutrient import. The helical shape is thought to be the result of the cytoskeletal protein fibril. Though it has been observed that with environmental changes
932:
984:
493:"Phylogenetic positions of 'Candidatus Phytoplasma asteris' and Spiroplasma kunkelii as inferred from multiple sets of concatenated core housekeeping proteins"
919:
696:"Use of spectral vegetation indices derived from airborne hyperspectral imagery for detection of European corn borer infestation in Iowa corn plots"
958:
1030:
322:
963:
695:
945:
54:
294:
have simple metabolism. They tend to grow well between 35-38 C and around the normal temperature of the human body.
989:
534:"Complete Genome Sequence of Spiroplasma kunkelii Strain CR2-3x, Causal Agent of Corn Stunt Disease in Zea mays L"
305:
in detail. From the genetic relations many of the characteristics of their metabolism have been inferred from the
1025:
712:
694:
Carroll MW, Glaser JA, Hellmich RL, Hunt TE, Sappington TW, Calvin D, Copenhaver K, Fridgen J (October 2008).
314:
237:
231:
835:
223:
genome have been done and the complete results have been posted to the GenBank. Studies have shown that
139:
893:
787:
661:
Pollack JD, McElwain MC, DeSantis D, Manolukas JT, Tully JG, Chang CJ, et al. (October 1989).
756:
643:
346:
49:
937:
737:"An Economic Analysis of Corn Yield, Corn Profitability, and Risk at the Edge of the Corn Belt"
997:
880:
815:
748:
717:
635:
604:
563:
514:
473:
225:
1002:
805:
795:
707:
674:
594:
553:
545:
504:
463:
453:
329:
lack cytidine, dCMP deaminases, transaldolase and deoxyribose 5-phosphate aldolase enzymes.
106:
96:
774:
Oleszczuk JD, Catalano MI, Dalaisón L, Di Rienzo JA, Giménez Pecci MP, Carpane P (2020).
532:
Davis RE, Shao J, Dally EL, Zhao Y, Gasparich GE, Gaynor BJ, et al. (October 2015).
791:
810:
775:
599:
583:"A genome sequence survey of the mollicute corn stunt spiroplasma Spiroplasma kunkelii"
582:
558:
533:
468:
437:
76:
44:
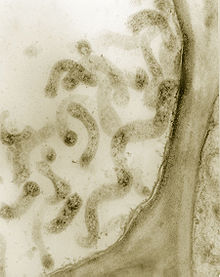
in phloem cells. Thick section (0.4 micrometers) observed in a TEM. Magnified 75,000X.
1019:
976:
885:
279:
273:
800:
298:
uses the host insect vector for multiplication, but it uses plants for survival.
357:
is spreading and has been found in south eastern United states. Though cases of
116:
858:
679:
662:
458:
86:
752:
639:
549:
265:
have a mutualistic relationship between arthropods and plants. Hosts include
182:
is a helical prokaryote that does not have a cell wall. The helical shape of
906:
168:
950:
819:
721:
608:
567:
518:
477:
410:
509:
492:
317:
due to their consumption of organic carbon and their parasitic lifestyle.
852:
267:
66:
760:
736:
647:
924:
872:
867:
829:
776:"Characterization of components of resistance to Corn Stunt disease"
911:
667:
International
Journal of Systematic and Evolutionary Microbiology
497:
International
Journal of Systematic and Evolutionary Microbiology
32:
898:
833:
417:. Centre for Agriculture and Bioscience International (CABI)
713:
10.1603/0022-0493(2008)101[1614:uosvid]2.0.co;2
622:
Renaudin J (2006). "Sugar metabolism and pathogenicity of
301:
Little research has been done regarding the metabolism of
735:
Chavas JP, Kim K, Lauer JG, Klemme RM, Bland WL (2001).
842:
663:"Metabolism of members of the Spiroplasmataceae"
411:"Spiroplasma kunkelii (corn stunt spiroplasma)"
741:Journal of Agricultural and Resource Economics
353:increases the loss possible in the industry.
8:
491:Zhao Y, Davis RE, Lee IM (September 2005).
436:Harne S, Gayathri P, Béven L (2020-10-21).
830:
261:need plants in order to survive and grow.
31:
20:
809:
799:
711:
678:
598:
557:
508:
467:
457:
371:
442:Biology: Opportunities and Challenges"
405:
403:
401:
380:"Spiroplasma--discovery and phylogeny"
241:genomes bare the closest relations to
7:
384:Agricultural Research Service (ARS)
600:10.1111/j.1574-6968.2002.tb11153.x
581:Bai X, Hogenhout SA (April 2002).
14:
325:pathway. Studies have found that
386:. U.S. Department of Agriculture
309:family and their close relative
53:
700:Journal of Economic Entomology
1:
801:10.1371/journal.pone.0234454
415:Invasive Species Compendium
1047:
1031:Bacteria described in 1986
628:Journal of Plant Pathology
345:, as it is known to cause
680:10.1099/00207713-39-4-406
587:FEMS Microbiology Letters
459:10.3389/fmicb.2020.589279
446:Frontiers in Microbiology
145:
138:
50:Scientific classification
48:
39:
30:
23:
550:10.1128/genomeA.01216-15
315:chemoorganoheterotrophs
343:corn stunt spiroplasma
323:Embden-Meyerhof-Parnas
238:Spiroplasma melliferum
510:10.1099/ijs.0.63655-0
232:Spiroplasma phoencium
219:Full analysis of the
202:Initial phylogeny of
155:Whitcomb, Chen et al.
977:spiroplasma-kunkelii
844:Spiroplasma kunkelii
538:Genome Announcements
339:Spiroplasma kunkelii
296:Spiroplasma kunkelii
292:Spiroplasma kunkelii
263:Spiroplasma kunkelii
259:Spiroplasma kunkelii
254:Spiroplasma kunkelii
198:Genome and phylogeny
180:Spiroplasma kunkelii
164:Spiroplasma kunkelii
149:Spiroplasma kunkelii
25:Spiroplasma kunkelii
792:2020PLoSO..1534454O
503:(Pt 5): 2131–2141.
337:The common name of
16:Species of bacteria
347:corn stunt disease
333:Corn stunt disease
313:. Spiroplasma are
1013:
1012:
998:Open Tree of Life
836:Taxon identifiers
624:Spiroplasma citri
226:Spiroplasma citri
160:
159:
1038:
1026:Mycoplasmataceae
1006:
1005:
993:
992:
980:
979:
967:
966:
954:
953:
941:
940:
928:
927:
915:
914:
902:
901:
889:
888:
876:
875:
863:
862:
861:
831:
824:
823:
813:
803:
786:(10): e0234454.
771:
765:
764:
732:
726:
725:
715:
691:
685:
684:
682:
658:
652:
651:
619:
613:
612:
602:
578:
572:
571:
561:
544:(5): e01216–15.
529:
523:
522:
512:
488:
482:
481:
471:
461:
433:
427:
426:
424:
422:
407:
396:
395:
393:
391:
376:
167:is a species of
151:
131:S. kunkelii
107:Mycoplasmataceae
58:
57:
35:
21:
1046:
1045:
1041:
1040:
1039:
1037:
1036:
1035:
1016:
1015:
1014:
1009:
1001:
996:
988:
983:
975:
970:
962:
957:
949:
944:
936:
931:
923:
918:
910:
905:
897:
892:
884:
879:
871:
866:
857:
856:
851:
838:
828:
827:
773:
772:
768:
734:
733:
729:
693:
692:
688:
660:
659:
655:
621:
620:
616:
580:
579:
575:
531:
530:
526:
490:
489:
485:
435:
434:
430:
420:
418:
409:
408:
399:
389:
387:
378:
377:
373:
368:
335:
289:
251:
200:
177:
156:
153:
147:
134:
97:Mycoplasmatales
52:
17:
12:
11:
5:
1044:
1042:
1034:
1033:
1028:
1018:
1017:
1011:
1010:
1008:
1007:
994:
981:
968:
955:
942:
929:
916:
903:
890:
877:
864:
848:
846:
840:
839:
834:
826:
825:
766:
747:(1): 230–247.
727:
706:(5): 1614–23.
686:
653:
634:(2): 129–139.
614:
573:
524:
483:
428:
397:
370:
369:
367:
364:
334:
331:
288:
285:
250:
247:
199:
196:
176:
173:
158:
157:
154:
143:
142:
136:
135:
128:
126:
122:
121:
114:
110:
109:
104:
100:
99:
94:
90:
89:
84:
80:
79:
77:Mycoplasmatota
74:
70:
69:
64:
60:
59:
46:
45:
37:
36:
28:
27:
15:
13:
10:
9:
6:
4:
3:
2:
1043:
1032:
1029:
1027:
1024:
1023:
1021:
1004:
999:
995:
991:
986:
982:
978:
973:
969:
965:
960:
956:
952:
947:
943:
939:
934:
930:
926:
921:
917:
913:
908:
904:
900:
895:
891:
887:
882:
878:
874:
869:
865:
860:
854:
850:
849:
847:
845:
841:
837:
832:
821:
817:
812:
807:
802:
797:
793:
789:
785:
781:
777:
770:
767:
762:
758:
754:
750:
746:
742:
738:
731:
728:
723:
719:
714:
709:
705:
701:
697:
690:
687:
681:
676:
673:(4): 406–12.
672:
668:
664:
657:
654:
649:
645:
641:
637:
633:
629:
625:
618:
615:
610:
606:
601:
596:
592:
588:
584:
577:
574:
569:
565:
560:
555:
551:
547:
543:
539:
535:
528:
525:
520:
516:
511:
506:
502:
498:
494:
487:
484:
479:
475:
470:
465:
460:
455:
451:
447:
443:
441:
432:
429:
416:
412:
406:
404:
402:
398:
385:
381:
375:
372:
365:
363:
360:
356:
352:
348:
344:
340:
332:
330:
328:
324:
321:utilizes the
320:
316:
312:
308:
304:
299:
297:
293:
286:
284:
282:
281:
276:
275:
270:
269:
264:
260:
255:
248:
246:
244:
240:
239:
234:
233:
228:
227:
222:
217:
215:
210:
205:
197:
195:
193:
189:
185:
181:
174:
172:
170:
166:
165:
152:
150:
144:
141:
140:Binomial name
137:
133:
132:
127:
124:
123:
120:
119:
115:
112:
111:
108:
105:
102:
101:
98:
95:
92:
91:
88:
85:
82:
81:
78:
75:
72:
71:
68:
65:
62:
61:
56:
51:
47:
43:
38:
34:
29:
26:
22:
19:
843:
783:
779:
769:
744:
740:
730:
703:
699:
689:
670:
666:
656:
631:
627:
623:
617:
590:
586:
576:
541:
537:
527:
500:
496:
486:
449:
445:
439:
431:
419:. Retrieved
414:
388:. Retrieved
383:
374:
358:
354:
350:
342:
338:
336:
326:
318:
310:
306:
302:
300:
295:
291:
290:
280:Zea perennis
278:
274:Zea mexicana
272:
266:
262:
258:
253:
252:
242:
236:
230:
224:
220:
218:
213:
209:Spiroplasmas
208:
203:
201:
191:
187:
183:
179:
178:
163:
162:
161:
148:
146:
130:
129:
117:
41:
24:
18:
593:(1): 7–17.
440:Spiroplasma
438:"Exploring
359:S. kunkelii
355:S. kunkelii
351:S. kunkelii
327:S. kunkelii
319:S. kunkelii
307:Spiroplasma
303:S. kunkelii
243:S. kunkelii
221:S. kunkelii
214:S. kunkelii
204:S. kunkelii
192:S. kunkelii
188:S. kunkelii
184:S. kunkelii
118:Spiroplasma
42:Spiroplasma
40:Corn stunt
1020:Categories
452:: 589279.
421:5 December
390:5 December
366:References
287:Metabolism
175:Morphology
169:Mollicutes
87:Mollicutes
753:1068-5502
640:1125-4653
249:Pathology
125:Species:
938:10034269
859:Q3966876
853:Wikidata
820:33075073
780:PLOS ONE
761:40987105
722:18950044
648:41998303
609:12023071
568:26494665
519:16166721
478:33193251
311:S. citri
268:Zea mays
103:Family:
73:Phylum:
67:Bacteria
63:Domain:
925:3226167
868:BacDive
811:7571704
788:Bibcode
559:4616174
469:7609405
113:Genus:
93:Order:
83:Class:
1003:753213
964:966261
912:SPIRKU
899:977408
818:
808:
759:
751:
720:
646:
638:
607:
566:
556:
517:
476:
466:
990:47834
951:50978
933:IRMNG
886:4Z8BD
873:14356
757:JSTOR
644:JSTOR
985:NCBI
972:LPSN
959:ITIS
920:GBIF
907:EPPO
816:PMID
749:ISSN
718:PMID
636:ISSN
605:PMID
564:PMID
515:PMID
474:PMID
423:2021
392:2021
277:and
235:and
946:ISC
894:EoL
881:CoL
806:PMC
796:doi
708:doi
704:101
675:doi
626:".
595:doi
591:210
554:PMC
546:doi
505:doi
464:PMC
454:doi
341:is
1022::
1000::
987::
974::
961::
948::
935::
922::
909::
896::
883::
870::
855::
814:.
804:.
794:.
784:15
782:.
778:.
755:.
745:26
743:.
739:.
716:.
702:.
698:.
671:39
669:.
665:.
642:.
632:88
630:.
603:.
589:.
585:.
562:.
552:.
540:.
536:.
513:.
501:55
499:.
495:.
472:.
462:.
450:11
448:.
444:.
413:.
400:^
382:.
283:.
271:,
245:.
229:,
216:.
822:.
798::
790::
763:.
724:.
710::
683:.
677::
650:.
611:.
597::
570:.
548::
542:3
521:.
507::
480:.
456::
425:.
394:.
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.